Abstract
Macrophomina phaseolina, is a pathogenic soil-borne fungus that affects more than 500 plant species, causing various types of disease to several crops, among which is the crown and root rot disease in strawberry. Its wide variability has been characterized reiteratively in the literature, but little is known about its virulence mechanisms. Morphological, physiological, genetic and phytopathogenic parameters were evaluated among 32 isolates of Macrophomina from different hosts occurring in Argentina and Spain. Colony characteristics, average size of microsclerotia, chlorate phenotype and mycelial growth at different temperatures (5º–40 °C), and pH (3.0–8.0) were recorded. The morphological and physiological traits were heterogeneous and did not show any association with the genetic structure nor with their pathogenicity. Most of the isolates (71.9%) exhibited chlorate-sensitive phenotype. The optimal growth temperature range was between 25 °C and 35 °C, and the optimal pH varied between 4.0 and 6.0. The genetic structure analyzed with four DNA markers (EF-1α, ITS, CAL and TUB) showed little diversity among isolates of M. phaseolina, with no clear association with the country of origin, but a significant association with the host. Based on their phylogenetic affinity, one isolate was reclassified as M. pseudophaseolina and another one as M. tecta. It is the first report of M. pseudophaseolina causing charcoal rot on beans, in Argentina, and the first report of M. tecta outside Australia. Pathogenicity tests on strawberry plants revealed marked host specialization, being the isolates obtained from strawberry more virulent than those from other hosts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The soil-borne pathogenic fungus Macrophomina phaseolina (Tassi) Goidanich (Botryosphaeriales) is widely distributed across the world, causing severe damage to more than 500 plant species worldwide [1], with their respective economic losses [2, 3]. Until 2013, it was considered a monospecific genus, but currently five species are described: M. phaseolina, M. pseudophaseolina [4], M. euphorbiicola [5], M. vaccinia [6] and M. tecta [7]. Isolates are easily identified on potato dextrose agar (PDA), from a characteristic dark mycelium and black microsclerotia. Macrophomina phaseolina shows large unstructured variation in physiology, pathogenicity, chlorate phenotypes and growing conditions which hinders the classification of the different isolates into races [8,9,10,11,12,13,14], despite initial attempts to do so [15]. Isolates of M. phaseolina can also vary significantly in pathogenicity, even between those obtained from a single host species or within the same plant [11, 16]. Its high phenotypic variation and genetic diversity could be important to explain its prevalence across a wide geographical and biological spectra [17].
The severe economic impact of M. phaseolina on crops together with its high variability has encouraged researchers to explore the association of virulence with phenotypic traits and possible host specificity and/or host specialization. While host specificity refers to the interactions of the pathogen on a set of host species- i.e., if the isolates can infect a species or not-; host specialization relates to the aggressiveness of infection between host and pathogen-i.e., if isolates from one host are more virulent/aggressive on that host than isolates from other hosts [18]. Across the literature, the association of morphological and physiological variation to pathogenicity is inconsistent [17, 19, 20]. Pathogenic assays of M. phaseolina isolates on different hosts report mixed results, from both host-specificity and host-specialization specialization [10, 11, 21] to non-specificity [11, 22]. A deeper knowledge of the specificity and specialization of this fungi across species of hosts is important to set the basis for future research on the molecular mechanisms involved in its pathogenicity and the implementation of agricultural practice, for instance, crop rotation [23] that minimize economical losses.
Strawberry (Fragaria x ananassa Duch.) is one of the crops most severely impacted by M. phaseolina and has served as a host model to describe much of the variability and biology from this fungus [21, 22, 24,25,26]. Crown and root rot symptoms in strawberry plants include wilting of the foliage, drying and death of older leaves, and the collapse and death of plants [27]. It is considered an emerging disease with big impact on strawberry crops in Spain and Argentina [28, 29], where it has important consequences given the vast agricultural lands dedicated to strawberry: Argentina is the fifth country in production in South America, with ~ 1300 cultivated ha producing ~ 45,500 t annually [30], while Spain ranks sixth worldwide, with 7260 cultivated ha producing more than 351,900 t (FAO, 2019). However, the control of this disease in strawberry plantations continues to be a challenge in agricultural research [31, 32].
The aim of this study was to investigate the diversity and variability of Macrophomina isolates obtained from multiple crops (strawberry, sunflower, soybean, bean, blueberry, corn, and hemp) occurring in Argentina and Spain, by morphological, physiological, molecular and pathogenic characterization. We also tested its possible host specialization in strawberry. This knowledge could be useful to design mitigation strategies for M. phaseolina diseases.
Materials and Methods
Origin and Isolation of Fungal Isolates
We characterized 32 Macrophomina isolates from Argentina (21) and Spain (11), obtained from opportunistic sampling of plants from experimental plantations with typical symptoms of the charcoal rot disease, or the soil where these plants were grown. The 21 Argentine isolates were obtained from strawberry, sunflower (Helianthus annuus L.), soybeans (Glycine max L.), bean (Phaseolus vulgaris L.) and blueberry (Vaccinum corymbosum L.) grown in the provinces of Tucumán, Salta and Santa Fe. The 11 Spanish isolates came from strawberry, hemp (Cannabis sativa L.), corn (Zea mays L.) and blueberry grown in the provinces of Huelva and Seville (Table 1). Plants were brought to the laboratory and rinsed thoroughly with tap water to remove soil. Affected tissue was cut into small pieces (5‒10 mm), surface sterilized with 2% sodium hypochlorite for 30 s and then rinsed three times with sterilized distilled water. The pieces were drained on sterile absorbent paper for a few minutes and placed on PDA medium (potato, 200 g; dextrose, 20 g; agarose, 17 g; distilled water, 1 L) on 90-mm Petri plates and incubated in the dark for five days at the optimal growth temperature of 30 °C [9, 33, 34]. A small portion of the fastest growing mycelia were taken and placed onto Petri plates containing PDA and incubated in darkness at 30 °C for five days.
Soil samples were taken from the first 20 cm depth using a vertical calibrated drill, in the space between symptomatic plants. The isolations were done as in Papavizas and Klag [35]. Briefly, 10 g of dry soil were mixed with 250 ml of 0.5% NaClO solution. The mix was centrifuged at 2,000 rpm for 30 s and allowed to stand for 3 min, repeating this cycle three times. Then the mixture was sieved, the soil retained in the mesh was washed with sterile distilled water to remove traces of NaClO, and resuspended in 100 ml of sterile distilled water. Aliquots of 1 ml of the final dilution were spread on the surface of Petri plates containing PDA-DOPCNB medium (basal medium containing p-(dimethylamino) benzenediazo sodium sulfonate (DASS, Oxgall) and pentachloronitrobenzene (PCNB) at 50, 2000 and 100 mg l−1, respectively). Inoculated Petri plates were incubated in darkness at 30 °C for 7 days.
Microsclerotia were separated from the mycelia of all grown cultures using a sterile needle, with the help of a magnifying glass, transferred to Petri plates containing PDA and incubated in darkness at 30 °C. Colonies appearing from single sclerotium were transferred to PDA medium in Petri plates and incubated as mentioned above. These isolates were used for morphological, physiological, molecular and pathogenic characterization.
Morphological and Physiological Characterization
Morphological and physiological variability among the 32 isolates of Macrophomina collected from symptomatic samples of Argentina and Spain was studied using the following parameters: type, shape and color of the colonies, chlorate phenotype, growth at different temperatures and pH, and sclerotium size. In each assay, isolates were examined in triplicate using a 5-mm diameter mycelial disc cut from the periphery of 7-day-old cultures grown on PDA at 30 °C in darkness.
Type, Shape and Color of the Colonies
For studying variability in type, shape and color of the colonies, Petri plates (90-mm) with PDA medium were inoculated in the center with a mycelial disc of each isolate. Petri plates were incubated at 30ºC in darkness for 5 days. At that time, the growth type was recorded as “regular”, when the growth of the isolate was even and with a defined edge, or “irregular”, when the edge was irregular. The type of mycelium was classified as “arborescent”, when it presented aerial growth, or “resupinate”, when hyphae remained compact and adhered to the culture medium, without aerial growth.
Chlorate Phenotype
Sensitivity to chlorate was assessed as in Pearson et al. [15]. A mycelial disc of each isolate of the fungus was placed on the center of 90-mm Petri plates containing LBA medium amended with 120 mM potassium chlorate. The Petri plates were then incubated at 30 °C in darkness. Three growth patterns were recorded: dense, feathery and restricted. A dense growth indicated resistance to chlorate, while a feathery or a restricted growth indicated sensitivity to chlorate. Colony phenotypes were recorded after 6 days by comparison with cultures grown on the same medium without chlorate.
Sclerotial Size
The surface of each of the three replicas of Macrophomina isolates grown in PDA at 30 ºC in darkness for 3 days were gently and superficially scraped with a scalpel. The scraping of each replica was placed on a different slide over a drop of sterile distilled water, dispersing well. Ten sclerotia were selected randomly per each of three replicas (n = 30) and size (length) was measured in an optical microscope (20X).
Effect of Temperature and pH on the Growth of Mycelia
For studying the growth at different temperatures, a mycelial disc of each isolate was transferred to the center of 90-mm Petri plates filled with PDA medium. The Petri plates were then placed in incubators with temperatures adjusted to 5, 10, 15, 20, 25, 30, 35 and 40 °C, in the dark.
To assess the effect of pH, PDA media with pH fixed at 3.0, 4.0, 5.0, 6.0, 7.0 and 8.0 were prepared. The pH of the medium was adjusted using 1 N HCl or KOH solutions, checked with a pH-meter, after autoclave sterilization and plated in 90-mm Petri plates. Mycelial discs were placed in the center of the plates and incubated at 30 °C in darkness.
On both assays, total area covered by the colony (A = π*R*r, where: R is the major radius and r is the minor radius) was measured at 2, 3 and 6 days after inoculation.
Molecular Characterization
DNA Extraction
Isolates were grown on Petri plates with PDA for 7 days at 30 ºC in darkness. The surface of the culture was scraped to extract mycelium. Then, samples were frozen in liquid nitrogen and ground to powder with a mortar. For each sample, 0.1 g was digested in 700 µl of lysis buffer (2% PVP, 2% CTAB, 1 M Tris-HCl at pH 8.0, 0,5 M EDTA, 0,5 M NaCl) combined with 2 µl of β-mercaptoethanol, 20 µl of proteinase K (20 mg ml−1) and 10 µl of RNAse (10 mg ml−1). Samples were incubated for 60 min at 65 °C. Then, a phenol–chloroform-isopropanol (25/24/1; v/v/v) DNA extraction was carried out, followed by ethanol precipitation. DNA was quantified in ND-100 NanoDrop spectrophotometer and stored at − 20 °C.
PCR Amplification and DNA Sequencing
We targeted the elongation factor 1 alpha (EF-1α) (~ 350 bp), calmodulin (CAL) (~ 500 bp), tubulin (TUB) (~ 600 bp) and internal transcribed spacer (ITS) regions (~ 650 bp) (Table 2). PCRs were carried out in final volumes of 50 μl containing 1X of PCR buffer, 2 mM ofMgCl2, 20 µM of each dNTP, 0.4 mM of each primer, 1.25U of Taq Polymerase (Thermo Scientific) and 100 ng of template DNA. Each PCR consisted on an initial denaturation of 5 min at 94 °C, followed by 40 cycles of 30 s of denaturation at 94 °C, 30 s of annealing at 55 °C, and 2 min of extension at 72 °C, ended by a final extension at 72 °C. A negative control with no DNA template was included in each set of reactions. The amplification products were run in 1% agarose gels, stained with RedSafe (0.025 µlml−1) and visualized under UV. Amplifications and gel visualization were repeated twice with each primer. After confirming the amplification, the PCR products were sent to StabVida (Caparica, Portugal) for their purification and sequencing.
Sequence Editing and Alignment
The chromatograms from Sanger sequencing were converted to FASTAQ files with Seqret tool from EMBOSS 6.6.0 (http://emboss.open-bio.org/). Low-quality terminal sequences were trimmed using cutadapt1.18 [36]. The quality thresholds for cutadapt were empirically determined (-q 40,40) by comparing the trimmed sequences with the original chromatograms in Geneious R8 (https://www.geneious.com). Sequences below 50 nucleotides were discarded. Multiple sequence alignments for each marker were made with MAFFT v7.244 [37] and DECIPHER [38] using defaults parameters.
Phylogenetic Analysis
The information from the four markers was then integrated by concatenating them with AMAS [39] for increased phylogenetic resolution [40]. We included sequences for the same markers from reference strains of all described species in Macrophomina from Poudel et al. [7], and the sequence of the related fungus Lasiodiplodia pseudotheobromae, from the same family of M. phaseolina, was included as an external group. The GenBank accession numbers of the four markers of each isolate is on the Supplementary Material 1. We excluded 59 samples from GenBank which had identical sequences in the concatenated alignment to other samples in the dataset. The phylogenetic reconstruction was performed in a Maximum Likelihood framework with RAxML 7.2.8 [41], using the GTR + G evolution model with one partition for each marker and the “rapid hill-climbing” algorithm. Bootstrap support was calculated over 100 replicates. To further support the evolutionary affinities between isolates, a Bayesian phylogenic tree was inferred on the same dataset in BEAST 2.1.2 [42]. Since BEAST uses a coalescent approach, the outgroup was excluded. PartitionFinder2 [43] was used to determine the best partition scheme out from the four DNA regions considered, proposing a unique partition under a HKY model of evolution. Thus, the four markers were concatenated and HKY was selected as a site model in BEAST. We assumed a coalescent constant population size, since most samples belonged to the same or sister species. Two independent chains were run for 10 million iterations and convergence was checked with Tracer 1.6 [44]. Trees from both chains were merged with LogCombiner 2.4.4 [42] and a maximum clade credibility tree was generated after a 10% burn-in.
Genetic Structure and Diversity
Haplotype networks were performed with the sequences of the four concatenated markers. To maximize the number of positions to be compared, isolates TOR-861 and TOR-862 were excluded since they presented a higher proportion of missing data, and indels were treated as a fifth state. Haplotype networks were built with PopArt [45] using TCS algorithm [46]. This program was also used to make an Analysis of Molecular Variance (AMOVA) and to estimate genetic diversity parameters: nucleotide diversity, polymorphic sites and parsimoniously informative sites. The purpose of AMOVA was to find out if there was an association between the genetic structure and the country (Spain or Argentina) or host (strawberry, sunflower, soybean, bean, blueberry, corn or hemp) of origin. The significance (P ≤ 0.05) of the observed structure (ϕst) was obtained from 1000 permutations.
Genetic Distance
Pairwise Kimura-2 genetic distances were computed with MEGA 7 program [47] using the concatenated sequences.
Pathogenic Characterization
We run two pathogenic assays in strawberry plants to evaluate the virulence of Macrophomina isolates and their possible host specialization. Around half (n = 18) of the isolates used had been isolated from the same species of host, strawberry, while the others (n = 14) came from different host species (Table 1). Macrophomina phaseolina exhibits an ample range of virulence, from very low to very high, depending on the experimental conditions and isolates [8, 48, 49]. To encompass this high range of virulence, we run the two independent assays with different inoculation methods described in the literature to cause different rates of severity. In the first one, we used the toothpick method, described in previous studies to cause higher virulence in shorter times of around one month [21, 50, 51]. In the second one, we induced infection by adding a suspension of microsclerotia. The latter, simulates better a natural infection but requires more time to detect the pathogenic effects of Macrophomina, of around 6 months [25] (details in Supplementary Material 2).
In both assays, a week before inoculation with Macrophomina isolates, plants were transferred to an infection chamber at 28 °C and 16-h photoperiod (100.5 μmolm−2 s−1) and subjected to hydric stress to generate favorable conditions for the infection of the fungus. Plants were watered three days a week with 50 ml of water per pot. The experimental design of each experiment was completely randomized. At the end of the assays, susceptibility to crown and root rot was evaluated as disease severity, using a disease severity rating (DSR) according to Fang et al. [52]: 0 = plant well developed, no disease symptoms; 1 = plant slightly stunted; 2 = plant stunted and yellowing; 3 = plant severely stunted and/or wilting; 4 = majority of leaves of the plant wilted or dead; 5 = plant death. Re-isolations were made from segments of freshly harvested diseased root and/or crown onto PDA to confirm the infection.
Statistical Analyses
Statistical analysis of data from morphological and physiological characterizations were performed in Statistix 9.1 software (http://www.statistix.com/). Means comparison between treatments of a same assay were evaluated using analysis of variance (ANOVA) and Fisher's Least Significant Difference (LSD) tests at α = 0.05.
Host specialization of the isolates was evaluated by comparing DSR in strawberry isolates versus isolates from other hosts. The statistical significance of the medians between both groups in each assay was evaluated with Mann Whitney U tests in R 4.0 (R Core Team 2020).
Results
Morphological and Cultural Characterization
All the isolates exhibited regular growth, except TOR-876 and TOR-855 with irregular growth. Colonies were black on the agar surface, except TOR-854 and TOR-861, which were brown. Regarding the type of the colony, ten isolates had resupinate mycelium and 22 an arborescent mycelium (Table 3).
After six days of incubation, nine isolates were resistant to chlorate (28.1%), whereas 23 were sensitive (71.9%). The bean (Por-Lea) and corn (TOR-860) isolates showed a dense phenotype. The soybean (Soj-Bal) isolate had a feathery phenotype, and blueberry and hemp isolates showed a restricted phenotype. Isolates obtained from strawberry and sunflower presented the three phenotypes (Table 3).
Microsclerotia sizes ranged from 50.3 µm to 114.3 µm. Isolates Fru-Fes (strawberry), TOR-840 (hemp) and TOR-872 (blueberry) had significantly bigger microsclerotia (P ≤ 0.05) to the rest. Isolate Gi-AG4 (sunflower) showed significantly smaller (P ≤ 0.05) microsclerotia than all others, except Por-Lea, Gi-PAN1031 and Fru-CR (Table 3).
Effect of Temperature on the Mycelial Growth
The optimal growth temperature range of the analyzed isolates varied between 25 °C and 35 °C, although the isolates could still grow relatively well at 20 °C. At 5 °C, no mycelial growth was observed even with the longest incubation times. Growth was very slow at 10 ºC (mean ± SE for the 32 isolates: 5.4 ± 5.1 cm2) and 15 °C (12.8 ± 8.1 cm2). At 20 °C most of the isolates completely covered the 90-mm Petri plates (56.7 cm2) on the 6th day (52.0 ± 13.1 cm2). Between 25 and 35 °C, mycelial growth was maximal, covering completely the plates after the 3rd day in most cases. At 40 °C, a steep decrease on the mycelial growth was observed, with great differences of mycelium growth extension on the 6th day, ranging from 56.7 cm2 for Gi-ACA885 and Gi-A963 (both from sunflower) to 0.6 cm2 for TOR-878 (strawberry) (Fig. 1a; Supplementary Material 3).
Effect of pH on the Growth of Mycelia
The optimal pH range for the development of M. phaseolina isolates was between 4.0 and 6.0. On the 2nd and 3rd days, maximum growth was observed at pH 5.0 (mean ± SE for the 32 isolates: 36.3 ± 7.4 and 54.9 ± 6.1 cm2, respectively). On the 6th day, the average area of mycelium growth was higher than 52.0 cm2 overall all pHs tested (3.0–8.0). A pH 3.0 was the least favorable for the pathogen. At pH 4.0, 5.0, 6.0, 7.0 and 8.0, the mycelia covered completely the Petri-dish area (56.7 cm2) on the 6th day, except the isolates TOR-855 (strawberry) and TOR-888 (hemp) (Fig. 1b; Supplementary Material 4).
Phylogenetic Analysis
The DNA alignment used in the phylogenetic analysis contained the four concatenated DNA markers, with a total of 71 isolates, 2359 positions and 9.8% of missing data. Both, the Maximum Likelihood and Bayesian phylogenetic trees supported well the different species described in the current taxonomy, except for the position of M. vaccini. The latter was a sister species of M. phaseolina in the Maximum Likelihood tree with low bootstrap support of 67%, whereas in the Bayesian tree grouped together with M. tecta with a posterior probability of 0.94 (Fig. 2). All isolates from this study grouped with M. phaseolina except for the isolates Por-Lea that showed higher affinity with M. pseudophaseolina and TOR-860 with M. tecta. A very small genetic variability was observed among the rest of the isolates grouped together, without any apparent association by country or host of origin.
Genetic Structure and Diversity
The concatenated alignment used in the haplotype network represented 30 isolates and 1,313 nucleotide positions. Eleven different haplotypes were identified (Fig. 3; Supplementary Material 5). The bean isolate (Por-Lea) was the most differentiated one, with more than 27 mutation steps with the closest isolate, followed by the corn isolate (TOR-860), with at least 10 mutations steps. This agrees with the phylogenetic tree, where these isolates grouped with reference strains of M. pseudophaseolina and M. tecta, respectively.
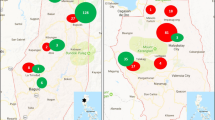
TCS Haplotype networks obtained from the four concatenated DNA markers, represented according to the country (a) and host (b) of origin of the isolates. Isolates that have the same haplotype (same sequence) are grouped in the same circle. Its size varies according to the number of isolates with the same haplotype. Each transversal line represents a mutational step between haplotypes. The small black circles represent missing haplotypes (may exist but were not sampled)
The AMOVA showed that the genetic structure of the isolates was not associated with their country of origin (ϕst = − 0.012, P = 0.543). However, there was some association between the genetic structure of the isolates and their host of origin (ϕst = 0.810, P = 0.001). Nucleotide diversity was 0.0039, and there were 43 polymorphic sites and 13 parsimoniously informative sites.
Genetic Distance
Genetic distance between isolates of M. phaseolina was low, varying from 0 to 1.1% (Supplementary Material 6). The Por-Lea isolate, identified as M. pseudophaseolina, differed the most, with distances between 2.1 and 3.0% from the isolates of M. phaseolina. The TOR-860 isolate, identified as M. tecta, differed with distances between 0.7 and 1.3% from the isolates of M. phaseolina. The greatest pairwise distance corresponded to 3.0% between Por-Lea (bean, Argentina) and TOR-861 (blueberry, Spain), while many isolates shared the same haplotype across the markers studied (distance of 0%; Supplementary Material 6).
Virulence of M. phaseolina Isolates
In both assays, all the isolates induced symptoms of crown and root rot except Por-Lea (M. pseudophaseolina, bean) and Soj-Bal (soybean) on strawberry plants cv. ‘Candonga’. The more virulent isolates were always strawberry isolates: Fru-SWA, Fru-Sab, Fru-Alb, Fru-For and Fru-Fes on cv. ‘Pájaro’; and TOR-854 and TOR-878 on cv. ‘Candonga’. The least virulent isolates on strawberry plants cv. ‘Pájaro’ were Gi-ACA885 and Gi-P65A25, both coming from sunflower (Table 3). According to the Mann Whitney U tests, the virulence of isolates coming from strawberry, as measure in the Fang scale, was 2.44 higher in assay 1 (W = 214, P value < 0.001) and 2.39 higher in assay 2 (W = 269.5, P value < 0.001 (Fig. 4).
Discussion
In this study, we report high morphological and physiological variability, and describe the genetic structure between 32 isolates of Macrophomina from diverse economically important crops in Argentina and Spain. We also found a high host specialization: isolates coming from strawberry were more virulent on strawberry plants than isolates coming from other plant hosts.
Previous studies also described high variability in morphology and physiology of M. phaseolina. Indeed, type of growth can be very different, from light dense to dense and colony color can vary between grey, dark green, black or brown [14, 19, 53]; microsclerotia can have large variation in size, from 50 to 150 μm [54, 55], and sensitivity to chlorate vary from around 30% [53, 56] to 90% [57]. Despite morphological or phenotypic variations have not been associated to country or host of origin of the isolates [19], for chlorate sensitivity some authors affirm isolates from the same host generally share phenotype [11, 56, 57] and others report mixed results regarding its association with virulence [20, 58]. The optimal growth temperature is in agreement with previous studies [9, 33, 34]. Acidity of the soil affects the survival of M. phaseolina and influence charcoal rot incidences in crops, although there is a high variance in the optimal pH conditions, from 3.6 to 6, between isolates, as we and previous authors have described [33, 59,60,61]. The wide morphological and physiological variability observed among isolates, together with the lack of associations between phenotypes and the country or host of origin, makes it difficult to classify them into subspecies [2, 19].
We observed little genetic variation among isolates of M. phaseolina (Figs. 2 and 3). Some genetic structure was detected between isolates and the host of origin, but not with the country (Fig. 3). These results must be interpreted cautiously as the number of isolates evaluated was low (30) relative to the number of hosts (n = 7). Most studies do not observe associations between isolates of M. phaseolina and their geographical origin or host [4, 12, 62]. However, with highly polymorphic DNA markers, Jana et al. [63] and Purkayastha et al. [56] use RAPDs to correlate DNA polymorphisms with the host of origin, while Jana et al. [64] and Sánchez et al. [34] used SSRs to group M. phaseolina isolates according to their geographic origin.
The bean (Por-Lea) and corn (TOR-860) isolates classified together with reference samples of M. pseudophaseolina [4] and M. tecta [7], respectively, in the phylogenetic tree. Despite being different species, Por-Lea and TOR-860 did not exhibit noticeable differences in phenotype compared to the other isolates of M. phaseolina (Table 3, Supplementary Material 3 and 4). This is the first report of M. pseudophaseolina causing charcoal rot on beans, in Argentina. This recently-described Macrophomina species has been only reported in different host species in Senegal, Brazil, India and Australia [4, 5, 7, 65,66,67]. This is also the first report of M. tecta causing charcoal rot on corn, in Spain. This new species was found on plants of both Sorghum bicolor and Vigna radiata with symptoms of charcoal rot, in Australia [7]. So far, there are no more records of this species anywhere.
In the pathogenicity tests, strawberry isolates scored an average of 2.4 points higher on the Fang scale, suggesting specialization between the isolates and their host (Fig. 4), disregarding their phenotypic background (Table 3). Many studies have attempted, with unclear results, to relate the morphological and molecular characterizations of M. phaseolina isolates with their pathogenicity in diverse hosts [10,11,12, 19, 20, 53, 56, 68]. Das et al. [20] suggested a possible relation between the sensitivity to chlorate of M. phaseolina isolates and their virulence on sorghum. However, we report resistance to chlorate in isolates with large variation in virulence: e.g., Gi-PAN1031 was barely virulent while chlorate-resistant isolates such as TOR-878, TOR-854 and Fru-Trea were highly virulent (Table 3). When looking at host specificity and host specialization in M. phaseolina, previous studies show mixed evidence. Su et al. [11] and Mayek-Pérez et al. [10], reported isolates from corn and beans, respectively, to be more virulent when infecting the same host than those from other hosts. Koike et al. [21] also found host specialization of M. phaseolina in strawberry. However, other authors reported no host specificity and/or host specialization in several host species [22, 69,70,71]. Virulence factors and host-specificity/specialization factors play a fundamental role in pathogenicity [72, 73]. A pioneer study suggested an adaptive mechanism behind host specialization laying on the ability of adapted M. phaseolina isolates to use nitrogenous compounds of a specific host [15]. However, the mechanisms involved in host-isolate specificity/specialization of M. phaseolina have not been studied so far. The lack of a clear genetic structure or phenotypic traits associated to virulence in this study, contrasted with the effect of host specialization. This suggests a heritable non-measured variation responsible for this differential virulence between hosts. The combined effect of fast-evolving adaptive mechanisms of virulence with certain heritability, such as epigenetics marks [74,75,76], could be at the base of the success of this versatile fungi to invade new species of hosts across variable environmental conditions. For instance, mechanisms of DNA methylation, modification of histone proteins and sRNA could be involved [77,78,79]. Further studies on the molecular mechanisms and on the epigenetic variation are required to understand the mechanisms of host specificity and host specialization in Macrophomina.
Conclusion
In this work we described a high diversity of Macrophomina in its morphological, genetic and pathogenic characteristics. We clearly differentiate isolates of M. phaseolina from its sister species M. pseudophaseolina and M. tecta, and found some association between the genetic structures of the isolates and their host of origin. One of the important aspects of this study reside in the finding of host specialization: isolates that had previously infested the same plant species (strawberry) showed greater virulence in strawberry plants. This contributes to the general knowledge of the behavior of this phytopathogen, but the molecular mechanisms leading to this host specialization are still little addressed. These latter considerations raise interesting directions for future research on interactions between pathogenic fungi and crops.
Data Availability
DNA sequences from the nuclear loci have been deposited in GenBank with accession numbers MK447797-MK447922. Data and code have been deposited in a GitHub repository (https://github.com/csmiguel/characterization-macrophomina). A permanent release has been deposited in Zenodo doi: https://doi.org/10.5281/zenodo.6528544.
Code Availability
References
Farr DF, Rossman AY (2022) Fungal databases, U.S. National Fungus Collections, ARS, USDA. https://nt.ars-grin.gov/fungaldatabases/. Accessed 3 Mar 2022.
Kaur S, Dhillon GS, Brar SK et al (2012) Emerging phytopathogen Macrophomina phaseolina: biology, economic importance and current diagnostic trends. Crit Rev Microbiol 38:136–151. https://doi.org/10.3109/1040841X.2011.640977
CABI (2021) Macrophomina phaseolina. Invasive Species Compend.
Sarr MP, Ndiaye MB, Groenewald JZ, Crous PW (2014) Genetic diversity in Macrophomina phaseolina, the causal agent of charcoal rot. Phytopathol Mediterr 53:250–268
Machado AR, Pinho DB, Soares DJ et al (2019) Bayesian analyses of five gene regions reveal a new phylogenetic species of Macrophomina associated with charcoal rot on oilseed crops in Brazil. Eur J Plant Pathol 153:89–100. https://doi.org/10.1007/s10658-018-1545-1
Zhao L, Cai J, He W, Zhang Y (2019) Macrophomina vaccinii sp. nov. causing blueberry stem blight in China. MycoKeys 55:1–14. https://doi.org/10.3897/mycokeys.54.35015
Poudel B, Shivas RG, Adorada DL et al (2021) Hidden diversity of Macrophomina associated with broadacre and horticultural crops in Australia. Eur J Plant Pathol. https://doi.org/10.1007/s10658-021-02300-0
Tok FM, Dervis S, Arslan M (2018) Host selective virulence, temperature response and genetic diversity in Macrophomina phaseolina isolates from sesame and peanut in southern Turkey. Fresenius Environ Bull 27:7374–7380
Manici LM, Caputo F, Cerato C (1995) Temperature responses of isolates of Macrophomina phaseolina from different climatic regions of sunflower production in Italy. Plant Dis 79:834–838
Mayek-Pérez N, Lopez-Castañeda C, González-Chavira M et al (2001) Variability of Mexican isolates of Macrophomina phaseolina based on pathogenesis and AFLP genotype. Physiol Mol Plant Pathol 59:257–264. https://doi.org/10.1006/pmpp.2001.0361
Su G, Suh SO, Schneider RW, Russin JS (2001) Host specialization in the charcoal rot fungus, Macrophomina phaseolina. Phytopathology 91:120–126. https://doi.org/10.1094/PHYTO.2001.91.2.120
Reyes-Franco MC, Hernández-Delgado S, Beas-Fernández R et al (2006) Pathogenic and genetic variability within Macrophomina phaseolina from Mexico and other countries. Phytopathology 154:447–453
Saleh AA, Ahmed HU, Todd TC et al (2010) Relatedness of Macrophomina phaseolina isolates from tallgrass prairie, maize, soybean and sorghum. Mol Ecol 19:79–91. https://doi.org/10.1111/j.1365-294X.2009.04433.x
Mishra PK, Kumari M, Dantre RK (2021) Morpho-cultural and pathogenic variability in Macrophomina phaseolina isolates from soybean. Pharma Innov J 10:777–785
Pearson C, Leslie J, Schwenk F (1986) Variable chlorate resistance in Macrophomina. Phytopathology 76:646–649
Mayek-Pérez N, Lopez Castaneda C, Acosta Gallegos JA (1997) Variation in vitro cultural characteristics of Macrophomina phaseolina isolates and its virulence on common bean. Agrociencia 31:187–195
Ashraf W, Sahi ST, ul Haq I, Ahmed S (2015) Morphological and pathogenic variability among Macrophomina phaseolina isolates associated with maize (Zea mays) in Punjab-Pakistan. Int J Agric Biol 17:1037–1042
Gomez AO (2020) Pathogenicity studies on strawberry to identify sources of resistance against Macrophomina phaseolina. University of Queensland
Beas-Fernández R, De Santiago-De SA, Hernández-Delgado S, Mayek-Pérez N (2006) Characterization of Mexican and non-Mexican isolates of Macrophomina phaseolina based on morphological characteristics, pathogenicity on bean seeds and endoglucanase genes R. J Plant Pathol 88:53–60
Das IK, Fakrudin B, Arora DK (2008) RAPD cluster analysis and chlorate sensitivity of some Indian isolates of Macrophomina phaseolina from sorghum and their relationships with pathogenicity. Microbiol Res 163:215–224. https://doi.org/10.1016/j.micres.2006.05.006
Koike ST, Arias RS, Hogan CS et al (2016) Status of Macrophomina phaseolina on strawberry in california and preliminary characterization of the pathogen. Int J Fruit Sci 16:148–159. https://doi.org/10.1080/15538362.2016.1195313
Zveibil A, Mor N, Gnayem N, Freeman S (2012) Survival, host-pathogen interaction, and management of Macrophomina phaseolina on strawberry in Israel. Plant Dis 96:265–272. https://doi.org/10.1094/PDIS-04-11-0299
Bargués-Ribera M, Gokhale CS (2020) Eco-evolutionary agriculture: host-pathogen dynamics in crop rotations. PLOS Comput Biol 16:e1007546. https://doi.org/10.1371/journal.pcbi.1007546
Viejobueno J, Ramallo AC, Kirschbaum DS et al (2017) Severe outbreaks of strawberry crown and root charcoal rot caused by Macrophomina phaseolina in Tucumán, Argentina. Rev agronómica del noroeste argentino 37:111–114
Gomez AO, De Faveri J, Neal JM et al (2020) Response of strawberry cultivars inoculated with Macrophomina phaseolina in Australia. Int J Fruit Sci 20:164–177. https://doi.org/10.1080/15538362.2019.1709114
Sánchez S, Henríquez JL, Urcola LA et al (2016) Susceptibility of strawberry cultivars to root and crown rot caused by Macrophomina phaseolina. J Berry Res 6:345–354. https://doi.org/10.3233/JBR-150114
Koike ST (2008) Crown rot of strawberry caused by Macrophomina phaseolina in California. Plant Dis 92:1253–1253
Baino OM, Salazar SM, Ramallo AC, Kirschbaum DS (2011) First report of Macrophomina phaseolina causing strawberry crown and root rot in Northwestern Argentina. Plant Dis 95:1477. https://doi.org/10.1094/PDIS-00-00-0000
Avilés M, Castillo S, Bascon J, et al (2008) First report of Macrophomina phaseolina causing crown and root rot of strawberry in Spain. Plant Pathol 57
Kirschbaum DS, Vicente CE, Cano-Torres MA et al (2017) Strawberry in South America: from the Caribbean to Patagonia. Acta Hortic 1156:947–956
Lodha S, Mawar R (2019) Population dynamics of Macrophomina phaseolina in relation to disease management: a review. J Phytopathol 168:1–17. https://doi.org/10.1111/jph.12854
Baggio JS, Cordova LG, Peres NA (2019) Sources of inoculum and survival of Macrophomina phaseolina in florida strawberry fields. Plant Dis 103:2417–2424. https://doi.org/10.1094/PDIS-03-19-0510-RE
Csöndes I, Cseh A, Taller J, Poczai P (2012) Genetic diversity and effect of temperature and pH on the growth of Macrophomina phaseolina isolates from sunflower fields in Hungary. Mol Biol Rep 39:3259–3269
Sánchez S, Chamorro M, Henríquez JL et al (2017) Genetic and biological characterization of Macrophomina phaseolina (Tassi) Goid. causing crown and root rot of strawberry. Chil J Agric Res 77:325–331. https://doi.org/10.4067/S0718-58392017000400325
Papavizas GC, Klag NG (1975) Macrophomina phaseolina from soil. Phytopathology 65:182–187
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnetJournal 17:10–12
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Wright ES (2016) Using DECIPHER v2.0 to analyze big biological sequence data in R”. R J 8:352–359
Borowiec ML (2016) AMAS: a fast tool for alignment manipulation and computing of summary statistics. PeerJ 4:e1660. https://doi.org/10.7717/peerj.1660
Gadagkar SR, Rosenberg MS, Kumar S (2005) Inferring species phylogenies from multiple genes: concatenated sequence tree versus consensus gene tree. J Exp Zool Part B Mol Dev Evol 304B:64–74. https://doi.org/10.1002/jez.b.21026
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Bouckaert R, Heled J, Kuehnert D et al (2014) BEAST 2: A software platform for Bayesian evolutionary analysis. PLoS Comput Biol 10:e1003537
Lanfear R, Frandsen PB, Wright AM et al (2016) PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol Biol Evol 34:772–773. https://doi.org/10.1093/molbev/msw260
Rambaut A, Drummond AJ, Xie D et al (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67:901–904. https://doi.org/10.1093/sysbio/syy032
Leigh JW, Bryant D (2015) Popart: full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116. https://doi.org/10.1111/2041-210X.12410
Clement M, Snell Q, Walker P et al (2002) TCS: estimating gene genealogies. Int Proc Parallel Distrib Process Symp 2:184–190
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Vengadeshkumar L, Kalaiselvi M, Sanjaygandhi S, Jaiganesh V (2019) Survey on the incidence of cowpea root rot disease and assessing the cultural characters and virulence of Macrophomina phaseolina. Plant Arch 17:143–148
Mohanapriya R, Naveenkumar R, Balabaskar P (2017) Survey, virulence and pathogenicity of root rot incidence of cowpea in selected districts of Tamilnadu caused by Macrophomina phaseolina (Tassi.) Goid. Int J Curr Microbiol Appl Sci 6:694–705
Youssef S, Shalaby AA, Pickel B et al (2021) Selecting tolerant strawberry germplasm to the fungus Macrophomina phaseolina. Acta Hortic 1309:751–758
de Assis Melo NJ, de Lima AG, Negreiros AMP et al (2021) Pathogenicity of Macrophomina phaseolina in cultivars and accessions of Cucumis melo. J Plant Pathol. https://doi.org/10.1007/s42161-021-00832-2
Fang X, Phillips D, Li H et al (2011) Comparisons of virulence of pathogens associated with crown and root diseases of strawberry in Western Australia with special reference to the effect of temperature. Sci Hortic (Amsterdam) 131:39–48. https://doi.org/10.1016/j.scienta.2011.09.025
Aboshosha SS, Attaalla SI, El-Korany AE, El-Argawy E (2007) Characterization of Macrophomina phaseolina isolates affecting sunflower growth in El-Behera governorate, Egypt. Int J Agric Biol 9:807–815
Nishad I, Srivastava AK, Saroj A et al (2018) First report of Nepeta cataria root rot caused by Macrophomina phaseolina in India. Plant Dis 102:2380
Wagan KH, Khaskheli MI, Lanjar AG (2018) Isolation and characterization of Macrophomina phaseolina isolates prevailing in Sindh, Pakistan. Pure Appl Biol 7:1309–1315
Purkayastha S, Kaur B, Dilbaghi N, Chaudhury A (2006) Characterization of Macrophomina phaseolina, the charcoal rot pathogen of cluster bean, using conventional techniques and PCR-based molecular markers. Plant Pathol 55:106–116. https://doi.org/10.1111/j.1365-3059.2005.01317.x
Mahdizadeh V, Safaie N, Goltapeh EM (2011) Diversity of Macrophomina phaseolina based on morphological and genotypic characteristics in Iran. Plant Pathol J 27:128–137. https://doi.org/10.5423/PPJ.2011.27.2.128
Mihail JD, TayIor SJ (1995) Interpreting of variability among isolates of Macrophomina phaseolina in patogenicity, pycnidium production, and chlorate utilization. Can Joumal Bot 73:1596–1603. https://doi.org/10.1139/b95-172
Dhingra OD, Sinclair JB (1978) Biology and pathology of Macrophomina phaseolina. Universidade Federal de Viçosa, Viçosa-Mina Gerais- Brasil
Bruton BD, Biles CL, Dunlap JR (1995) Nutrient utilization of Macrophomina phaseolina: a chromogenic isolate from cantaloupe fruit. Subtrop Plant Sci 47:46–52
Nischwitz C, Olsen M, Rasmussen S (2004) Effect of irrigation type on inoculum density of Macrophomina phaseolina in melon fields in Arizona. J Phytopathol 152:133–137
Khan AN, Shair F, Malik K et al (2017) Molecular identification and genetic characterization of Macrophomina phaseolina strains causing pathogenicity on sunflower and chickpea. Front Microbiol 8:1309. https://doi.org/10.3389/fmicb.2017.01309
Jana T, Sharma TR, Prasad RD, Arora DK (2003) Molecular characterization of Macrophomina phaseolina and Fusarium species by a single primer RAPD technique. Microbiol Res 158:249–257. https://doi.org/10.1078/0944-5013-00198
Jana TK, Singh NK, Koundal KR, Sharma TR (2005) Genetic differentiation of charcoal rot pathogen, Macrophomina phaseolina, into specific groups using URP-PCR. Can J Microbiol 51:159–164. https://doi.org/10.1139/w04-122
de Queiroz Brito AC, Ferreira de Mello J, Michereff SJ et al (2019) First report of Macrophomina pseudophaseolina causing stem dry rot in cassava in Brazil. J Plant Pathol 101:1245–1245
Negreiros AMP, Sales Júnior R, León M et al (2019) Identification and pathogenicity of Macrophomina species collected from weeds in melon fields in Northeastern Brazil. J Phytopathol 167:326–337. https://doi.org/10.1111/jph.12801
Mastan A, Bharadwaj R, Kushwaha RK, Vivek Babu CS (2019) Functional fungal endophytes in Coleus forskohlii regulate labdane diterpene biosynthesis for elevated forskolin accumulation in roots. Microb Ecol 78:914–926. https://doi.org/10.1007/s00248-019-01376-w
Berbegal M, Ortega A, Jiménez-Gasco MM et al (2010) Genetic diversity and host range of Verticillium dahliae isolates from artichoke and other vegetable crops in Spain. Plant Dis 94:396–404. https://doi.org/10.1094/PDIS-94-4-0396
Almeida ÁMR, Sosa-Gomez DR, Binneck E et al (2008) Effect of crop rotation on specialization and genetic diversity of Macrophomina phaseolina. Trop Plant Pathol 33:257–264. https://doi.org/10.1590/S1982-56762008000400001
Fuhlbohm MJ, Ryley MJ, Aitken EAB (2012) New weed hosts of Macrophomina phaseolina in Australia. Australas Plant Dis Notes 7:193–195. https://doi.org/10.1007/s13314-012-0082-6
Mihail JD, Taylor SJ (1995) Interpreting variability among isolates of Macrophomina phaseolina in pathogenicity pycnidium production, and chlorate utilization. Can J Bot 73:1596–1603
Borah N, Albarouki E, Schirawski J (2018) Comparative methods for molecular determination of host-specificity factors in plant-pathogenic fungi. Int J Mol Sci 19:863. https://doi.org/10.3390/ijms19030863
van der Does HC, Rep M (2007) Virulence genes and the evolution of host specificity in plant-pathogenic fungi. Mol Plant-Microbe Interact 20:1175–1182. https://doi.org/10.1094/MPMI-20-10-1175
Lind MI, Spagopoulou F (2018) Evolutionary consequences of epigenetic inheritance Heredity (Edinb) 121:205–209. https://doi.org/10.1038/s41437-018-0113-y
Stajic D, Jansen LET (2021) Empirical evidence for epigenetic inheritance driving evolutionary adaptation. Philos Trans R Soc B Biol Sci 376:20200121. https://doi.org/10.1098/rstb.2020.0121
Kasuga T, Gijzen M (2013) Epigenetics and the evolution of virulence. Trends Microbiol 21:575–582. https://doi.org/10.1016/j.tim.2013.09.003
He C, Zhang Z, Li B, Tian S (2020) The Pattern and function of DNA methylation in fungal plant pathogens. Microorganisms 8:227. https://doi.org/10.3390/microorganisms8020227
Tini F, Beccari G, Marconi G et al (2021) Identification of putative virulence genes by DNA methylation studies in the cereal pathogen Fusarium graminearum. Cells 10:1192. https://doi.org/10.3390/cells10051192
Dubey A, Jeon J (2017) Epigenetic regulation of development and pathogenesis in fungal plant pathogens. Mol Plant Pathol 18:887–898. https://doi.org/10.1111/mpp.12499
Carbone I, Kohn LM (1999) A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 91:553–556. https://doi.org/10.1080/00275514.1999.12061051
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR protocols: a guide to methods and applications. Academic Press Inc, New York, pp 315–322
O´Donnell K, Cigelnik E (1997) Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol Phylogenet Evol 7:103–116
Acknowledgements
This study is part of a joint project between Instituto Nacional de Tecnología Agropecuaria-EEA Famaillá and Instituto Andaluz de Investigación y Formación Agraria, Pesquera y Alimentaria y de la Producción Ecológica. It was partially supported with grants of the Universidad Nacional de Tucumán (Program A621), Agencia Nacional de Promoción Científica y Tecnológica (PICT 2017-0653) and INTA (PNHFA 1106073, 2019-PD-E4-I069-001, 2019-PD-E4-I085-001) and by Instituto Andaluz de Investigación y Formación Agraria, Pesquera y Alimentaria y de la Producción Ecológica and 80 % Fondo Europeo de Desarrollo Regional (FEDER) (PP.AVA 2016.010). Authors are grateful to Strawberry Active Germplasm Bank (BGA) from Universidad Nacional de Tucumán (UNT). We thank María del Carmen and Olga Gómez Saavedra, and Miguel Ángel Fernández Ayala for helping in the laboratory. JV is CONICET fellowship.
Funding
Partial financial support was received from Universidad Nacional de Tucumán (Program A621), Agencia Nacional de Promoción Científica y Tecnológica (PICT 2017-0653), INTA (PNHFA 1106073, 2019-PD-E4-I069-001, 2019-PD-E4-I085-001), Instituto de Investigación y Formación Agraria y Pesquera de Andalucía (IFAPA) and 80% Fondo Europeo de Desarrollo Regional (FEDER) (PP.AVA 2016.010).
Author information
Authors and Affiliations
Contributions
Conceptualization: JV, BS, SMS; Methodology: JV, BS, AA, SMS; Formal analysis and investigation: JV, BS, AA, MCS; Writing—original draft preparation: JV; Writing—review and editing: JV, BS, MCS, MC, SMS; Funding acquisition: BS, MC, SMS; Resources: BS, AA, MC, SMS; Supervision: BS, AA, MC, SMS.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Consent to Participate
All authors consent to participate in this study.
Consent for Publication
All authors consent to publish this work in Current Microbiology.
Ethical Approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Viejobueno, J., de los Santos, B., Camacho-Sanchez, M. et al. Phenotypic Variability and Genetic Diversity of the Pathogenic Fungus Macrophomina phaseolina from Several Hosts and Host Specialization in Strawberry. Curr Microbiol 79, 189 (2022). https://doi.org/10.1007/s00284-022-02883-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-022-02883-9