Abstract
Bacteria of genus Holospora (order Holosporales, class Alphaproteobacteria) are obligate intranuclear symbionts of ciliates Paramecium spp. with strict host species and nuclear (macronucleus or micronucleus) specificity. However, three species under study Holospora undulata, Holospora elegans and ‘Holospora recta' occupy the same ecological niche—micronucleus of Paramecium caudatum and demonstrate some differences in morphology of infectious form. The genetic diversity of holosporas by rrs and rpoB sequence analysis was determined. Phylogenetic and phylogenomic analysis of Holospora spp., as well as some phenotypic features indicate that there is no distinctive difference supporting studied micronuclear endosymbionts as distinct species. Therefore, Holospora elegans and ‘Holospora recta' should be considered subspecies of Holospora undulata (ex Haffkine 1890) Gromov and Ossipov 1981, which was described first. Thus, we confirmed the evolutionary aspects of the development of symbiotic relationships: holosporas have a strict specificity to the host species and the type of nucleus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Unlike most of the eukaryotes the ciliates contain two nucleus types: a macronucleus (Ma), used to control metabolism, and a micronucleus (Mi), which performs reproductive functions. Bacteria of genus Holospora are intranuclear symbionts of ciliates Paramecium spp. with strict host species and nuclear specificity (Mi or Ma). They possess a complex life cycle with alternating infectious (IFs) and reproductive (RFs) that differ morphologically. Moreover, holosporas can be transmitted to the host both vertically and horizontally.
According to the List of Prokaryotic names with Standing in Nomenclature (LPSN) page (https://lpsn.dsmz.de/genus/holospora), the taxon Holospora (ex Haffkine 1890) Gromov and Ossipov 1981 is a homonym of Holospora (ex Haffkine 1890) Preer and Preer 1982 and the type species of the genus Holospora is Holospora undulata (ex Haffkine 1890) Gromov and Ossipov 1981.
Four species, Holospora undulata (ex Haffkine 1890) Gromov and Ossipov 1981, Holospora elegans (ex Haffkine 1890) Preer and Preer 1982, Holospora obtusa (ex Haffkine 1890) Gromov and Ossipov 1981 and Holospora caryophila (ex Preer et al. 1974) Preer and Preer 1982, are validly published; two, Holospora undulata (ex Haffkine 1890) Preer and Preer 1982 and Holospora obtusa (ex Haffkine 1890) Preer and Preer 1982, are illegitimate names.
For seven species, namely Holospora acuminata, Holospora caryophila, Holospora curviuscula, Holospora elegans, Holospora obtusa, Holospora undulata and ‘Candidatus Holospora parva’, the rrs gene sequences are available in GenBank.
Our phylogenetic analysis indicated that most of them belonged to the genus Holospora [1]. Only Holospora caryophila distantly relates to other Holospora species, and it was reclassified into a new genus as Preeria caryophila comb. nov. [2].
The genus Holospora is the nomenclatural type of order Holosporales and it belongs to the family Holosporaceae [1, 3]. This work studies phylogenetic relationships and taxonomic position of Paramecium caudatum Mi symbionts Holospora undulata, Holospora elegans and ‘Holospora recta'.
As we noted earlier, holosporas are specific to host's species and its nucleus type—Ma or Mi [4]. Hence, each of two types of nuclei of host species is infected by strictly specific symbiont species. In spite of that, three different types of holosporas were reported as symbionts for micronucleus of Paramecium caudatum–Holospora undulata (ex Haffkine 1890) Gromov and Ossipov 1981, Holospora elegans (ex Haffkine 1890) Preer and Preer 1982 and “Holospora recta” Fokin 1991. Moreover, description of these species was grounded on morphological traits only, and their taxonomic position was also questioned later [4]. These inconsistencies have triggered our interest for the study of phylogeny and taxonomic position of the three symbionts species.
Materials and Methods
Bacterial Isolation and Culture Conditions
The frozen cells of pure symbionts cultures (the accession number refers to protozoan clones carrying the endosymbionts)—type strains Holospora undulata M1-48, ‘Holospora recta’ k1 and Holospora elegans 02AZ19-20 stored in the former Laboratory of Karyology, St. Petersburg State University were used. We have also studied the clone of Paramecium caudatum 06R3-1 infected by Holospora undulata and clone of Paramecium bursaria AC61-10 with Holospora acuminata. Cultivation and isolation of symbionts were performed as described previously [1].
PCR and Sequencing
DNA of symbionts was extracted by the conventional phenol/chloroform method [5]. PCR amplification targeting the 16S rRNA gene, purification and sequencing of PCR products was carried out as described in [1] with the additional primers for sequencing to yield an expected amplicon of approximately 1500 bp: RFI1 5′-CTCTGTGCCAGCAGCCGCGG-3′ (Escherichia coli positions 460–480) and RRI1 5′-GCGGTATTAGCCAAGGTTTC-3′ (E. coli positions 145–125); RFI2 5′-GATAAATCGGAGGAAGGAGAGG-3′ (E. coli positions 1110–1132) and RRI2 5′-CGTCTAGCACTCATCGTTTAGGG-3′ (E. coli positions 760–737).
The primers CM7 5′-AACCAGTTCCGCGTTGGCCTGG-3′ (E. coli positions 1705–1727) and CM31b 5′-CCTGAACAACACGCTCGGA-3′ (E. coli positions 2436–2455) were used for amplifying and sequencing the partial rpoB gene sequences.
Phylogenetic and Genome Analysis
A BLAST analysis of the obtained sequences was performed against sequences held in GenBank through the NCBI website (http://www.ncbi.nlm.nih.gov/). Phylogenetic and phylogenomic analyses were performed using software package MEGA version X [6].
Analysis of the rrs and rpoB genes was based on sequences received in this study and sequences available in GenBank. Distances for both genes were calculated according to the Kimura's two-parameter model [7] and for the rrs gene phylogenetic constructions the General Time Reversible (GTR) model of nucleotide substitution [8] which often fits the rRNA data better also used. Clustering was based on the neighbour-joining [9] and maximum-likelihood [10] methods. The bootstrap analysis is commonly based on 1000 resamples and was used to evaluate the topology of the phylogenetic tree [11]. Digital analysis and calculations were made by tetranucleotide frequencies and correlation coefficients algorithms described in JSpecies [12], OrthoANI [13] and in silico DDH [14].
Results and Discussion
In the present study sequences of rrs gene of symbionts of Paramecium caudatum Mi—the representative strains Holospora undulata M1-48, ‘Holospora recta’ k1, also Holospora elegans 02AZ19-20 and Holospora undulata 06R3-1 were analysed. The analyses of 16S rRNA gene sequence placed all strains within the genus Holospora (Fig. 1). The 16S rRNA encoding gene sequence of Holospora elegans 02AZ19-20 (1456 bp) showed 100% similarity with respect to that of a strain from Holospora elegans AB297813 (479 bp) held in GenBank. Sequences similarities between the different strains of Holospora undulata, Holospora elegans and ‘Holospora recta’ ranged from 99.45 to 100% and the sequences of Holospora obtusa strains from 99.35 to 100%.
Neighbour-joining phylogenetic tree based on the 16S rRNA gene sequences of Holospora undulata 06R3-1, Holospora undulata M1-48, Holospora elegans 02AZ19-20, Holospora recta k1 and members of related taxa within the family Holosporaceae. Bar, 0.02 substitutions per nucleotide position. The maximum-likelihood tree produced by both Kimura's two-parameter [7] и the General Time Reversible (GTR) models [8] showed essentially the same topology (data not shown)
Gene rpoB encoding DNA-directed RNA polymerase subunit beta is very promising for proteobacteria identification, and the phylogenetic resolution of the rpoB tree was approximately three times higher than that of the rrs tree [15]. The partial rpoB gene sequences of representative strains ‘Holospora recta’ k1 and Holospora acuminata AC61-10 were obtained. The phylogenetic analysis of the rpoB gene sequences confirmed the affiliation of these strains to the genus Holospora, with the three species Holospora undulata, Holospora elegans and ‘Holospora recta’ forming a cluster with high internal similarity values (Fig. 2).
Maximum-likelihood phylogenetic tree based on the rpoB sequences of strains ‘Holospora recta’ k1 and Holospora acuminata AC61-10 and members of related taxa within the order Holosporales. Bar, 0.1 substitutions per nucleotide position. The neighbour-joining tree showed essentially the same topology (data not shown)
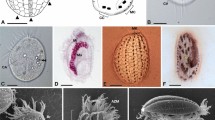
Three morphologically described species Holospora undulata [16], Holospora elegans [17] and ‘Holospora recta’ [18] are symbionts of same ciliates' species Paramecium caudatum and same host nuclear type—micronucleus (Supplementary Table 1). They demonstrate different cell morphology of IF: (1) spiral, ends tapered (Holospora undulata), (2) straight, ends tapered (Holospora elegans) and (3) straight, one-end tapered (‘Holospora recta’). The taxonomical position for ‘Holospora recta’ has been questioned by Ossipov and colleagues, who found a strain of Holospora elegans with a number of individuals exhibiting ‘Holospora recta’-like features [4]. Finally, it was concluded that the question may be resolved after the obtaining of molecular data.
The results of the phylogenetic analysis of the rpoB gene sequences confirmed those of the 16S rRNA suggesting that Holospora undulata, Holospora elegans and ‘Holospora recta’ belong to the same species (Figs. 1, 2).
The genome is the ultimate source of information for taxonomic purposes and its use has been accelerated significantly thanks to advances in high-throughput DNA-sequencing technologies [19]. There are four draft genomes of holosporas deposited in GenBank: Holospora obtusa, Holospora undulata, Holospora elegans [20] and Holospora curviuscula [21]. Their genomic characteristics are presented in Supplementary Table2.
The phylogenomic analysis of Holospora spp., OrthoANI (Supplementary Fig. S1), dDDH (Table 1) confirmed that the studied micronuclear endosymbionts refer to the same species. The threshold value of OrthoANI is 95% and dDDH > 70% [22]. Construction of the classification system was based on the fact that DDH can reveal coherent genomic groups (genospecies) of strains generally sharing DDH values with greater than 70% similarity [23]. The 95%–96% ANI threshold can be readily used as an objective boundary for species circumscription, especially if it is reinforced by high TETRA correlation values [22]. Other algorithms for ANI calculations (basing the calculations on BLAST and MUMmer algorithm) of this species showed similar results (> 96%).
In this regard and as a general observation, TETRA values 0.99 may support the species circumscription based on the ANI range 95%–96%, but both values should agree [22]. Tetranucleotide analysis by pairwise comparison of strains (Table 1) shows value 0.99245.
Currently, bacterial taxonomy relies on a polyphasic approach based on the combination of phenotypic and genotypic characteristics including the DNA–DNA hybridization, the 16S rRNA gene sequence nucleotide similarity, the phylogeny and the DNA G + C content, as well as the genomic criteria (genome characteristics and genomic sequence similarity). Identification and classification of holosporas are generally difficult, since they have not yet been cultured in the laboratory and are contained within the cultures of their hosts. We used methods of phylogenetic analysis and basic genomic characteristics that significantly complement the phenotypic observations. This approach allowed us to clarify the systematic position of previously described species [4]. Studied species showed some phenotypic and genotypic differences. However, the values of most parameters were above the same species threshold. Therefore, Holospora elegans (ex Haffkine 1890) Preer and Preer 1982 and ‘Holospora recta’ Fokin 1991 should be considered as subspecies of Holospora undulata (ex Haffkine 1890) Gromov and Ossipov 1981, which was for the first time described by Haffkine [24]. Diagnostic characteristics of Holospora undulata subsp. elegans, H. undulata subsp. recta and Holospora undulata subsp. undulata are given in Supplementary Table 1. Hence, we confirmed the evolutionary aspects of the development of symbiotic relationships: holosporas have a strict specificity to the host species and the type of nucleus.
Emended Description of Holospora undulata (Ex Haffkine 1890) Gromov and Ossipov 1981
Holospora undulata (un.du.láta. L. fem. adj. undulatus undulated, with waves).
The properties are as given in the previous description [16], with the following amendments. Lives in the micronucleus of Paramecium caudatum, very rarely can be found in the macronucleus. Infectious form long 0.7–1 × 10–25 μm, spiral shaped or straight.
Emended Description of Holospora undulata subsp. undulata (Ex Haffkine 1890) Gromov and Ossipov 1981
Holospora undulata subsp. undulata (un.du.láta. L. fem. adj. undulatus undulated, with waves).
Basonym: Holospora undulata (ex Haffkine 1890) Gromov and Ossipov 1981. The properties are as given in the previous description [16].
Emended Description of Holospora undulata subsp. elegans (Ex Haffkine 1890) Preer and Preer 1982
Holospora undulata subsp. elegans (éle.gans. L. fem. adj. elegans elegant).
Basonym: Holospora elegans (ex Haffkine 1890) Preer and Preer 1982. The properties are as given in the previous description [17].
Emended Description of H. undulata subsp. recta Fokin 1991
H. undulata subsp. recta (rećta. L. fem. adj. rectus straight).
Basonym: H. undulata recta Fokin 1991. The properties are as given in the previous description [18].
Abbreviations
- Holospora , Paramecium :
-
Intranuclear Symbiosis
- DDH:
-
DNA–DNA hybridization
- ANI:
-
Average nucleotide identity
- TETRA:
-
Tetranucleotide frequency correlation coefficients, rrs and rpoB genes
References
Rautian M, Wackerow-Kouzova ND (2013) Phylogenetic placement of two previously described intranuclear bacteria from the ciliate Paramecium bursaria (Protozoa, Ciliophora): ‘Holospora acuminata′ and ‘Holospora curviuscula. Int J Syst Evol Microbiol 32:140–141
Potekhin A, Schweikert M, Nekrasova I, Vitali V, Schwarzer S, Anikina A, Kaltz O, Petroni G, Schrallhammer M (2018) Complex life cycle, broad host range and adaptation strategy of the intranuclear Paramecium symbiont Preeria caryophila comb. nov. FEMS Microbiol Ecol. https://doi.org/10.1093/femsec/fiy076
Amann R, Springer N, Ludwig W, Görtz HD, Schleifer KH (1991) Identification in situ and phylogeny of uncultured bacterial endosymbionts. Nature 351:161–164
Görtz HD, Schmidt HJ (2005) Holosporaceae. In: Garrity GM (ed) Bergey′s Manual of Systematic Bacteriology. Springer, New York, pp 146–160
Grimont F, Grimont PAD (1995) Determination of rRNA gene restriction patterns. In: Howard J, Whitcombe DM (eds) Methods in Molecular Biology. Humana Press, New York, pp 181–200
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol Biol Evol 35:1547–1549
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Waddell P, Steel M (1997) General time-reversible distances with unequal rates across sites: mixing gamma and inverse Gaussian distributions with invariant sites. Mol Phylogenet Evol 8:398–414
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Kishino H, Hasegawa M (1989) Evaluation of the maximum likelihood estimate of the evolutionary tree topologies from DNA sequence data, and the branching order in Hominoidea. J Mol Evol 29:170–179
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Richter M, Rosselló-Móra R, Glöckner FO, Peplies J (2016) JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32:929–931
Lee I, Ouk Kim Y, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103. https://doi.org/10.1099/ijsem.0.000760
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics 14:60
Ait Tayeb L, Ageron E, Grimont F, Grimont PA (2005) Molecular phylogeny of the genus Pseudomonas based on rpoB sequences and application for the identification of isolates. Res Microbiol 156:763–773
Gromov BV, Ossipov DV (1981) Holospora (Ex Haffkine 1890) nom. rev., a genus of bacteria inhabiting the nuclei of paramecia. Int J Syst Bacteriol 31:348–352
Görtz HD, Dieckmann J (1980) Life cycle and infectivity of Holospora elegans Haffkine, a micronucleus-specific symbiont of Paramecium caudatum (Ehrenberg). Protistologica 16:591–603
Fokin SI (1991) Holospora recta sp. nov. - a micronucleus-specific endobiont of the ciliate Paramecium caudatum. Tsitologiya 33:135–141
Chun J, Rainey FA (2014) Integrating genomics into the taxonomy and systematics of the Bacteria and Archaea. Int J Syst Evol Microbiol 64:316–324
Dohra H, Tanaka K, Suzuki T, Fujishima M, Suzuki H (2014) Draft genome sequences of three Holospora species (Holospora obtusa, Holospora undulata, and Holospora elegans), endonuclear symbiotic bacteria of the ciliate Paramecium caudatum. FEMS Microbiol Lett 359:16–18
Garushyants SK, Beliavskaia AY, Malko DB, Logacheva MD, Rautian MS, Gelfand MS (2018) Comparative Genomic Analysis of Holospora spp., Intranuclear Symbionts of Paramecia. Front Microbiol. https://doi.org/10.3389/fmicb.2018.00738
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 45:19126–21931. https://doi.org/10.1073/pnas.0906412106
Goebel BM, Stackebrandt E (1994) Cultural and phylogenetic analysis of mixed microbial populations found in natural and commercial bioleaching environments. Appl Environ Microbiol 60:1614–1621. https://doi.org/10.1128/AEM.60.5.1614-1621
Haffkine MW (1890) Maladies infectieuses des Paramécies. Ann Inst Pasteur 4:148–162
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or nonprofit sectors.
Author information
Authors and Affiliations
Contributions
NDW-K performed the molecular, phylogenetic and taxonomic analysis and wrote the manuscript. DVM carried out genome analysis and read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
All sequences obtained in this study are deposited in GenBank under accession numbers: for rrs gene—M1-48-MZ934711, 06R3-1-MZ934710, k1-MZ934709 and 02AZ19-20-MZ934708; for rpoB gene—k1–OK150102 and AC61-10-OK169910. OrthoANI values diagram, diagnostic table of Holospora undulata group and table with main features of the four Holospora endosymbiont genomes are available as Supplementary materials in Current Microbiology Online.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wackerow-Kouzova, N.D., Myagkov, D.V. Clarification of the Taxonomic Position of Paramecium caudatum Micronucleus Symbionts. Curr Microbiol 78, 4098–4102 (2021). https://doi.org/10.1007/s00284-021-02667-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-021-02667-7