Abstract
In Argentina, periurban agriculture is performed by farmers with inadequate training in the use of pesticides and chemical fertilizers, developing horticulture with serious soil deterioration. The aim of this work was to monitor bacterial diversity of a horticultural soil (S) and a reference soil (R) as quality index for the design of future restoration strategies. As crops changed together with the agrochemical applications, sample collection was before harvest for strawberries, post-harvest for red peppers, pre-harvest broccoli crop and of a resting soil in treatment with poultry litter as a fertilizing amendment. Bacterial diversity was analysed by the use of high throughput sequencing of the V1–V3 region of the 16S rRNA gene. Analysis of R soils seemed relatively constant in time, enriched in Alphaproteobacteria and Acidobacteria consistent with a reference to soil health. The effect of the intensive use of S soils was proved by differences in Chloroflexi, Bacteroidetes and Proteobacteria relative abundances. The main evidence of the alteration of S soils was the increase in Bacteroidetes and Betaproteobacteria. A weak recuperation trend of S soil microbiota was registered during a post-harvest inactive period. A strong influence of the soil use routine—consisting in high crop rotation and short time-rest cycles—on microbial community structure was verified. These results indicate the microbiota perturbation, caused by the intense use of periurban agriculture soils and will contribute for further actions to improve environment quality.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Soil quality and structure are determinant factors for the equilibrium of the biodiversity in terrestrial ecosystems. Any disturbance from both natural and anthropogenic origin may cause changes in microbial communities, and hence, their role in the environment can be seriously altered. In the literature, many studies are focused on the description of the variations in microbial community composition and their correlation to functional roles [1,2,3,4,5]. In fact, there is limited information about the use of microbial diversity as an indicator of soil health, including after amendment applications [6, 7].
As a consequence of agricultural production, pesticides and chemical fertilizers are usually introduced in soils affecting soil quality, microbial activities and populations. Particularly, small-scale agriculture around large cities (periurban agriculture) is performed by farmers who generally count with few economical resources and, as receiving inadequate or no training in the use of agrochemicals, they are continuously exposed to consequent damage to their own health [8]. Even though the main objective of the farmers is to obtain high crop yields that are accomplished at short terms, the productivity and environmental health are not sustainable at long terms. This kind of agriculture presents problems that differ from extensive farming and causes a strong environmental impact not only by the intensive use of the land, but also by the practices applied. Agricultural soil research is mainly directed to the evaluation of the influence on soil quality of the different seeding strategies for soybean, wheat, sunflower or corn crops [9, 10]. Also, there are some reports related to microbial communities and soil quality of horticultural units based on monocultures for fruit production [11] or vegetable rotations restricted to greenhouse conditions [12, 13], but scarce information on periurban practices can be found in the literature.
Regarding Buenos Aires Metropolitan Area, small farm clusters (5 ha each or even smaller) are mostly located in the western districts as Moreno [14,15,16]. In this area, periurban horticulture and floriculture have been developed with high crop rotation together with an intensive and uncontrolled use of both pesticides and chemical fertilizers, evidencing serious soil deterioration [17].
Based on the hypothesis that native microbiota composition represents a soil quality indicator, the detection of microbial community alterations is crucial to contribute to the development of sustainable horticultural practices belonging to periurban areas. Then, the aim of this work was to monitor bacterial diversity of a horticultural soil (S) and a reference soil (R) to recover the information as a quality index for the design of future restoration strategies. This work represents an innovative microbiological assessment of periurban horticulture soils characterized by a long history in their intense use, crop rotation, agrochemical applications, since most of the bibliography compiles the information on agricultural soils belonging to extensive farming, especially in Argentina.
Materials and Methods
Sampling Area
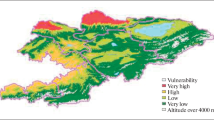
Moreno District belongs to the western Buenos Aires Metropolitan Area of which geographical location is shown in Fig. 1a. Western Buenos Aires Metropolitan Area soils´ classification corresponds to Order Mollisols, Suborder Udolls, Great Group Argiudolls, Subgroup Vertic Argiudolls [17,18,19]. Particularly, the study area is characterized by a continuous production history of at least 20 years in case of S soils and without any activity during the same period in case of R soils. A physicochemical characterization of soil was previously performed and is summarized in Table 1 [20]. Standard soil testing procedures were applied: organic matter by calcination of 5–10 g of soil at 430 °C, 2 h; pH and conductivity in 1:5 in H2O; total N by Kjedhal method and total P by Molybdo-vanado-phosphate method. From the obtained data, total C; C:N and C:N:P ratios were calculated [20].
a Geographic location of sampling site, b Sampling sites belonging to the Cuartel V periurban area soils. I: pre-harvest strawberry crop in spring 2014; II: post-harvest bell pepper in fall 2015; III: pre-harvest broccoli crop in fall 2016; IV: resting soil in treatment with poultry litter as fertilizing amendment (spring 2016); V: reference grassland close to the productive plots; as illustrative, but not sampled VI: poultry litter and VII: pesticide formulation preparation area of the farm
Soil samples were taken in 2014, 2015 and 2016 from different sites of a farm belonging to Cuartel V area, Moreno District (Fig.1a). S (34°34′33.69″ S 58°48′49.71″ W) was a pre-harvest strawberry crop plot performed with mulching techniques in spring 2014, post-harvest bell pepper crop in fall 2015, pre-harvest broccoli crop in fall 2016 and a resting soil in treatment with poultry litter as fertilizing amendment in spring 2016. R, a reference grassland close to the productive plots (34°34′39.31″ S 58°48′44.38″ W), was also sampled as control. Figure 1b illustrates the sampling area for each instance.
Soil Sampling
For each plot, 5 random 10 cm deep soil cores of 5 cm diameter were collected in a sterile recipient. Once in the laboratory, soil cores were independently mortared. At least two independent composite samples were obtained then mixing 500 mg of each core. Each composite sample was homogenized and stored at −80 °C until utilization.
Microbial Community Analysis
DNA Extraction and Sequencing
DNA extraction was performed from each homogenized sample using the MoBio PowerSoil DNA extraction kit (MoBio Laboratories Inc), following manufacturer protocol. DNA content was spectrophotometrically estimated (Nanodrop 2000 Thermo Scientific) and submitted to be sequenced by MR DNA (www.mrdnalab.com, Shallowater, TX, USA) on a MiSeq. Briefly, 16S rRNA gene V1–V3 variable region was amplified using standard primers 27F and -519R and HotStarTaq Plus Master Mix Kit (Qiagen, USA). Amplification conditions were 94 °C for 3 min, 28 cycles of 94 °C for 30 s, 53 °C for 40 s and 72 °C for 1 min, after which a final elongation step at 72 °C for 5 min. Multiple samples were pooled together in equal proportions and were purified using calibrated Ampure XP beads. DNA library was prepared according to Illumina TruSeq DNA library preparation protocol.
Data Processing and Statistical Analysis
Raw sequences (NCBI BioProject ID: PRJNA658339) were processed by demultiplexing (used script: https://github.com/AgustinPardo/demultiplex) and “amplicon sequence variant” (ASV) was obtained using DADA2 package in R. Taxonomic identity was assigned to each ASV according to the SILVA database SSU r132 [21]. In this work, the analysis of the ASV table was adapted to the Bergey’s phylogenetic nomenclature, regarding some considerations. First, Epsilonbacteraeota (a new phylum according to Silva database) was considered as Epsilonbacteria, the Proteobacteria class. Then Betaproteobacteriales, included as a Gammaproteobacteria order in Silva database, was incorporated as Betaproteobacteria, the Proteobacteria class. And last, regarding Rickettsiales class, some of the obtained ASV were labelled as Mitochondria. To analyse if those ASV should be removed from de ASV table, the ten more abundant sequences labelled as Mitochondria were aligned using BLAST® (blastn, nr/nt collection). None of them evidenced similarity with mitochondrial DNA but showed highly scored alignments with uncultured Proteobacteria. To avoid missing information, those sequences were kept in the resulting ASV table.
Alpha and beta diversity were studied using QIIME1 platform [22]. Rarefaction was carried out between 5000 and 50,000 subsamples, and alpha diversity indexes (CHAO1, Shannon, Simpson and Dominance) were obtained at the iteration of 25,000.
For statistical analysis, Levene and Shapiro tests were performed in order to check for homoscedasticity and normality in the data. Alpha diversity indexes and taxonomic analysis by relative abundance calculations (%) were compared by Student’s t test. For beta diversity, principal coordinates analysis (PcoA) was created using the weighted version of the UniFrac metric distance. In order to maximize comparability with analysis of beta diversity, Bray-Curtis distances were used to check for dissimilarities by the non-parametric statistical test ANOSIM (Analysis of Similarities).
Results
Sequence Analysis and Microbial Community Structure
Microbial community was evaluated by the analysis of 16S rRNA gene sequences as described in Materials and Methods. Obtained sequences were binned into 20,122 unique ASVs to calculate diversity non-parametric estimators (Fig. 2). Bacterial richness (CHAO1 index) showed a significant decrease (Student’s t test p = 0.00280) when poultry litter was used as soil amendment (S soil in spring 2014–2016). No significant differences in CHAO1 index were observed in the reference soil. Diversity indexes (Shannon’s H index, Simpson) showed no significant differences between S and R soils.
To deepen the study on the microbiota differences between the samples, beta diversity was analysed. Figure 3 shows a PCoA multidimensional diagram where R and S soils presented two different clusters (ANOSIM, R = 0.97 and p < 0.001) despite the season and the year. One cluster matched the R soils with no history of horticultural activities in the last 20 years, and the other cluster included the exploited S soils, confirming that soil use is the main component that affects the autochthonous microbiota and explained 21.35% of the variance along the first axis of the plot.
Bacterial Community Composition
When analysing phylum distribution across the samples (Fig. 4), R soils showed that Acidobacteria relative abundance changed from 44.8% to 26.3% between spring 2014 and fall 2015 (Student’s t test p = 0.023) and remained unchanged in 2016 despite the season. A different effect was observed in Actinobacteria relative abundance revealing non-significant shifts (3.7–10.3% between spring 2014 and fall 2015 and 7.7–5.0% in fall and spring 2016 samples).
Bacteria phylum diversity of R and S soils from each sampling campaign: spring 2014-R soils (R 2014), fall 2015-R soils (R 2015), fall 2016-R soils (R 2016f), spring 2016-R soils (R 2016s), spring 2014-S soils (S 2014), fall 2015-S soils (S 2015), fall 2016-S soils (S 2016f) and spring 2016-S soils (S 2016s). The graphic represents the average of two independent sequencing dataset
In S soils, the main observed difference is the increase in the Bacteroidetes proportion compared with R soil. In fall 2015, bell pepper had already been collected and no use of the fertilizer was declared by farmers resulting in a decrease in Bacteroidetes relative abundance from 29.6% in S spring 2014 to 13.0% in S fall 2015 and again increased to 21.5% in spring 2016.
The disturbance in soils caused by the already mentioned intense use could be also evidenced by an increase in Chloroflexi proportion (Student’s t test p = 0.0003) (Fig. 4) and by Proteobacteria diversity when comparing S to R soils (Student’s t test p = 0.0376) (Figs. 5 and 6). R soils showed a high Alphaproteobacteria proportion (63% in spring 2014, 73% in fall 2015, 66% in fall 2016 and 72% in spring 2016), particularly in the Order Rhizobiales, while in S soils, Alphaproteobacteria relative abundance was 15% in spring 2014. Although Betaproteobacteria was the most abundant class in this sample (55%), in fall 2015, S soil samples showed a significant decrease (Student’s t test p = 0.0312). Interestingly, although a recovery of Alphaproteobacteria proportion was observed in fall 2015 (35%), fall 2016 (32%) and spring 2016 (58%), the Order Rhizobiales still appeared at low percentages while comparing to Rhodospirillales (Fig. 6).
Proteobacteria class relative abundances in R and S soils from each sampling campaign: spring 2014-R soils (R 2014), fall 2015-R soils (R 2015), fall 2016-R soils (R 2016f), spring 2016-R soils (R 2016s), spring 2014-S soils (S 2014), fall 2015-S soils (S 2015), fall 2016-S soils (S 2016f) and spring 2016-S soils (S 2016s). The graphic represents the average of two independent sequencing dataset
Comparison of Proteobacteria order distribution between S and R soil samples from each campaign: spring 2014-R soils (R 2014), fall 2015-R soils (R 2015), fall 2016-R soils (R 2016f), spring 2016-R soils (R 2016s), spring 2014-S soils (S 2014), fall 2015-S soils (S 2015), fall 2016-S soils (S 2016f) and spring 2016-S soils (S 2016s). Proteobacteria orders: Alphaproteobacteria (blue), Betaproteobacteria (orange), Gammaproteobacteria (green), Deltaproteobacteria (yellow) and Epsilonproteobacteria (red). The graphic represents the average of two independent sequencing data set
Focusing on S soils, Betaproteobacteria population’s deep analysis showed a predominance of the Order Burkholderiales (30% in spring 2014, 8% in fall 2015, 24% in fall 2016 and 25% in spring 2016), in contrast to R soils where Nitrosomonadales was the most abundant order specially in spring 2014 and in fall 2015 (Fig. 6). The fall 2016 pre-harvest broccoli crop soil analysis revealed a new increase in Betaproteobacteria, especially in Burkholderiales, while the poultry litter amendment during spring 2016 positively influenced Alphaproteobacteria relative abundance. However, Sphingomonadales and Rickettsiales proportions were higher than Rhizobiales in this case (Fig. 6).
Another evidence of perturbed soil is the presence of Epsilonproteobacteria in spring 2014-S soils, not registered in other samples (Fig. 6). Finally, no significant changes were detected in both Gamma and Deltaproteobacteria relative abundances that could infer other additional data for soil quality assessment.
Discussion
Different weather conditions, crop rotation, the absence of control in agrochemical applications and the use of amendments of unknown nature are factors that contributed to deeply impact on the microbiota of these periurban horticulture soils. In this case, there was no available information about the types of agrochemicals and fertilizers used, the amounts applied and dates of application of each product, so the evaluation of soil quality mainly relied on the analysis of a combination of biological responses.
Regarding physicochemical characterization of sampled soils previously performed [20], main differences were found in organic matter-C:N values, conductivity and copper concentration. R soils showed low conductivities with the highest organic matter-C:N values. S-2014 soils evidenced high copper concentration related to the utilization of Cu-based agrochemicals as the fungicide Cotacuatro®, which decreased in 2015 samples studies. Both R soils—spring 2014 and fall 2015—showed low conductivities with the highest organic matter-C:N values. During fall 2015, notable amounts of leaf litter were observed in R soils, deriving an almost 20% increase in C:N ratio while compared to spring 2014 samples. Changes in 1 pH unit were detected in S soils, correlated to the respective use. Being S a post-harvest soil in fall 2015, the observed tendency to acidic pH values as in R soils would represent signals to a slight soil restoration [20]. Considering the fluctuations previously found in physicochemical characteristics of both R and S soils, different microbial community responses were registered.
Acidobacteria was described as an oligotrophic phylum, and its relative abundance is strongly related with carbon content [23]. In agreement, the decrease observed in this phylum proportion was consistent with the variation of the physicochemical properties of R soils (Fig. 4), especially C content increment in fall 2015 (from 21.26% to 42.93%, Table 1 [20]). On the other hand, regarding Fierer et al. [23], Actinobacteria relative abundance variation did not correlate with C content and any trend of change in this phylum would not be related to this physicochemical characteristic. In this case, Actinobacteria distribution in R soils seemed relatively constant in time, as well as observed with an enriched Alphaproteobacteria proportion within Proteobacteria (Fig. 5), which provides a good reference of soil health.
The relative abundance of Bacteroidetes could be a consequence of the addition of poultry litter as fertilizing amendment in S spring 2014 and spring 2016 soils (Fig. 4), since this phylum represents one of the most abundant of the phyla usually found in its microbiota [24, 25]. The observed community shifts are probable related to the intensive and uncontrolled treatments applied by farmers concerned with constant productivity improvement according to their economic needs.
The enrichment in Betaproteobacteria population detected in spring 2014—S soils (Fig. 5) could be also related to the metabolic diversity and the selection pressure imposed: C:N:P was clearly unbalanced in spring 2014 (106:2:1, Table 1 [20]). This selection pressure increased Burkholderiales relative abundance while compared with the other orders of this class (Fig. 6). In this same direction, Ancion et al. [26] detected an increase in Betaproteobacteria population in stream biofilms when they were exposed to synthetic urban runoff moderately contaminated with Cu, Zn and Pb. By contrast, a higher N level and a better C:N:P ratio (80:3:1) were detected in fall 2015 (Table 1, [20]) probably associated with Alphaproteobacteria—specially Rhizobiales—relative abundance with a consequently decrease in Betaproteobacteria proportion (Fig. 6). These results are in agreement to those obtained by Hartman et al. [27] describing that Betaproteobacteria are highly abundant in agricultural soils, but their relative abundance decreases when the soil quality is restored. In fact, fall 2015-S soil samples showed a decrease of Betaproteobacteria (Student’s t test p = 0.0312) associated with their condition of being collected during an inactive period accompanied by an increase in autochthonous vegetation, which also contributed to acid pH restoration (7.6–6.3 Table 1, [20]). This apparent restoration was rapidly interrupted, consequence of intensive crop rotation as in fall 2016 with broccoli production. Betaproteobacteria again increased enriched in Burkholderiales during this period, shifting to an Alphaproteobacteria with Sphingomonadales and Rickettsiales predominance in spring 2016 (Fig. 6). This shift also proved the perturbation level caused by the use declared (poultry litter) and undeclared amendments as Rhizobiales relative abundance remained unchanged.
Interestingly, Epsilonproteobacteria was detected in spring 2014-S soils (Fig. 6), class usually associated with waste treatment plant sludges and related to gut microbiomes [28]. Horticultural practices usually applying unknown origin amendments and using for irrigation untreated both surface and ground waters from suburban sources with high contamination levels could support this detection [29,30,31,32].
In this work, the evaluation of soils through microbiological approaches gave a wide insight of the environmental perturbations that periurban agriculture could generate, not usually explored. Therefore, this study contributed to the knowledge of the problematic proving that microbial community composition operated as an indicator of soil health, becoming an excellent tool to plan further actions for the environment quality improvement.
Conclusions
In this work, the effect of intensive use of the horticultural soil was observed in a variation of the microbial diversity compared with a reference healthy soil. The detected changes seemed to be also related to the soil use routine—crop development period, harvest and soil rest times—since alterations on microbial community structure were verified. This knowledge will contribute to improve periurban soils usage leading to environmentally friendly and sustainable practices combined with high productivity.
References
Fierer N, Ladau J, Clemente JC, Leff JW, Owens SM, Pollard KS, Knight R, Gilbert JA, McCulley RL (2013) Reconstructing the microbial diversity and function of pre-agricultural tallgrass prairie soils in the United States. Science 342:621–624
Fierer N, Lauber CL, Ramirez KS, Zaneveld J, Bradford MA, Knight R (2012) Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J 6:1007–1017
Griffiths RI, Thomson BC, James P, Bell T, Bailey M, Whiteley AS (2011) The bacterial biogeography of British soils. Environ Microbiol 13:1642–1654
Mendes LW, Tsai SM, Navarrete AA, de Hollander M, van Veen JA, Kuramae EE (2015) Soil-borne microbiome: linking diversity to function. Microb Ecol 70:255–265
Wu T, Chellemi DO, Graham JH, Martin KJ, Rosskopf EN (2008) Comparison of soil bacterial communities under diverse agricultural land management and crop production practices. Microb Ecol 55:293–310
Hermans SM, Buckley HL, Case BS, Curran-Cournane F, Taylor M, Lear G (2017) Bacteria as emerging indicators of soil condition. Appl Environ Microbiol 83:e02826–e02816
De Corato U (2020) Agricultural waste recycling in horticultural intensive farming systems by on-farm composting and compost-based tea application improves soil quality and plant health: a review under the perspective of a circular economy. Sci Total Environ 738:139840. https://doi.org/10.1016/j.scitotenv.2020.139840
UNDP, United Nations (1997) The Second International Colloquium of Mayors on Governance for Sustainable Growth and Equity, New York City. http://www.fao.org/unfao/bodies/coag/coag15/x0076e.htm. Accessed 15 May 2017
Carbonetto B, Rascovan N, Álvarez R, Mentaberry A, Vázquez MP (2014) Structure, composition and metagenomic profile of soil microbiomes associated to agricultural land use and tillage systems in Argentine Pampas. PLoS One 9(6):e99949. https://doi.org/10.1371/journal.pone.0099949
Figuerola ELM, Guerrero LD, Rosa SM, Simonetti L, Duval ME, Galantini JA, Bedano JC, Wall LG, Erijman L (2012) Bacterial indicator of agricultural management for soil under no till crop production. PLoS One 7(11):e51075. https://doi.org/10.1371/journal.pone.0051075
Liang B, Ma C, Fan L, Wang Y, Yuan Y (2018) Soil amendment alters soil physicochemical properties and bacterial community structure of a replanted apple orchard. Microbiol Res 216:1–11. https://doi.org/10.1016/j.micres.2018.07.010
Li N, Gao D, Zhou X, Chen S, Li C, Wu F (2020) Intercropping with potato-onion enhanced the soil microbial diversity of tomato. Microorganisms 8(6):834. https://doi.org/10.3390/microorganisms8060834
Lyu J, Jin L, Jin N, Xie J, Xiao X, Hu L, Tang Z, Wu Y, Niu L, Yu J (2020) Effects of different vegetable rotations on fungal community structure in continuous tomato cropping matrix in greenhouse. Front Microbiol 11:829. https://doi.org/10.3389/fmicb.2020.00829
Berenstein GA, Hughes EA, March H, Rojic G, Zalts A, Montserrat JM (2014) Pesticide exposure during the manipulation of concentrated mixtures at small horticultural and floricultural production units in Argentina: the formulation effect. Sci Total Environ 472:509–516
Craig E, Falco L, Sabaté L (2002) Municipal strategies for the primary sector of the district of Moreno, Buenos Aires. UA Mag 7:7–9
Hughes EA, Flores AP, Ramos LM, Zalts A, Glass CR, Montserrat JM (2008) Potential dermal exposure to deltamethrin and risk assessment for manual sprayers: influence of crop type. Sci Total Environ 391:34–40
INTA, Instituto Nacional de Tecnologia Agropecuaria (2014). http://anterior.inta.gov.ar/suelos/cartas/index.htm. Accessed 13 September 2017
Querejeta GA, Ramos LM, Hughes EA, Vullo DL, Zalts A, Montserrat JM (2014) Environmental fate of trifluralin, procymidone and chlorpyrifos in small horticultural production units in Argentina. Water Air Soil Pollut 225:1952
Soil Survey Staff (2014) Keys to soil taxonomy, 12th edn. USDA-Natural Resources Conservation Service, Washington, DC
Di Schiena J, Berenstein G, Cáceres Wenzel M, Oneto ML, Fuchs J, Montserrat JM, Casabé N, Basack S (2015) Biomarcadores en Eisenia andrei expuestas a suelos frutihortícolas tratados con clorpirifós, oxicloruro de cobre y miclobutanil. Acta Toxicológica Argent 23:50
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2012) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41(D1):D590–D596
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Gonzalez Pena A, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88(6):1354–1364
Huber DH, Chavarria-Palma JE, Malkaram SA, Montenegro-Garcia NA, Lhilhi Noundou V, Ugwuanyi IR, Espinosa-Solares T (2018) Metagenome sequences of a thermophilic anaerobic digester adapted to a low C/N ratio, high-ammonia feedstock (poultry litter). Genome Announc 6:e00598-18. https://doi.org/10.1128/genomeA.00598-18
Weidhaas J, Mantha S, Hair E, Nayak B, Harwood VJ (2015) Evidence for extraintestinal growth of Bacteroidales originating from poultry litter. Appl Environ Microbiol 81:196–202. https://doi.org/10.1128/AEM.02354-14
Ancion PY, Lear G, Lewis GD (2010) Three common metal contaminants of urban runoff (Zn, Cu & Pb) accumulate in freshwater biofilm and modify embedded bacterial communities. Environ Pollut 158:2738–2745
Hartman WH, Richardson CJ, Vilgalys R, Bruland GL (2008) Environmental and anthropogenic controls over bacterial communities in wetland soils. Proc Natl Acad Sci U S A 105(46):17842–17847
Sidhu C, Vikram S, Pinnaka AK (2017) Unraveling the microbial interactions and metabolic potentials in pre-and post-treated sludge from a wastewater treatment plant using metagenomic studies. Front Microbiol 8:1382
Feijoó CS, Giorgi A, García ME, Momo F (1999) Temporal and spatial variability in streams of a pampean basin. Hydrobiologia 394:41–52. https://doi.org/10.1023/A:1003583418401
Massone HE, Martinez DE, Cionchi JL, Bocanegra E (1998) Suburban areas in developing countries and their relationship to groundwater pollution: a case study of Mar del Plata, Argentina. Environ Manag 22(2):245–254. https://doi.org/10.1007/s002679900100
Topalian ML, Rovedatti MG, Castañe PM, Salibian A (1999) Pollution in a lowland river system. A case study: the Reconquista river (Buenos Aires, Argentina). Water Air Soil Pollut 114:287–302. https://doi.org/10.1023/A:1005180714913
Vullo DL, Ceretti HM, Hughes EA, Ramirez SA, Zalts A (2005) Indigenous heavy metal multiresistant microbiota of Las Catonas stream. Environ Monit Assess 105:81–97. https://doi.org/10.1007/s10661-005-3157-4
Acknowledgements
This work was supported by the Universidad Nacional de General Sarmiento, Universidad de Buenos Aires, Consejo Nacional de Investigaciones Científicas y Tecnológicas (CONICET) and grant PIO CONICET FYPF2014-2015 N°13320130100204CO. The sampling sites are located in non-protected areas. Sampling permissions were obtained from the farm owner Mr. Severino C., from IMDEL (Instituto Municipal de Desarrollo Económico Local) and Moreno District municipality. We are grateful to Miss Leticia Rossi for the English language revision and to Mr. Agustin Pardo for the bioinformatic platform assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Raiger Iustman, L.J., Almasqué, F.J. & Vullo, D.L. Microbiota Diversity Change as Quality Indicator of Soils Exposed to Intensive Periurban Agriculture. Curr Microbiol 78, 338–346 (2021). https://doi.org/10.1007/s00284-020-02298-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02298-4