Abstract
The aim of the study is the research and identification of a Streptomyces strain as a new producer of spectinabilin, undecylprodigiosin and metacycloprodigiosin. Among 54 actinomycete isolates isolated from El-Ogbane forest soils in Algeria, only one isolate, designated V002, was selected for its ability to produce prodigiosins. The selected strain was analysed for its ability to produce three different secondary metabolites as well as their biological activities. V002 belongs to the Streptomyces genus and has significant antimicrobial and antioxidant activities. The taxonomic position of V002 by 16S rRNA sequence analysis showed a similarity of 99.93% with Streptomyces lasiicapitis DSM 103124T and 98.96% with Streptomyces spectabilis DSM 40512T. Fractionation of crude secondary metabolites produced by the strain using HPLC–MS revealed the presence of spectinabilin, undecylprodigiosin and metacycloprodigiosin, which demonstrated significant activity. Strain V002 is considered a new producer of spectinabilin, undecylprodigiosin and metacycloprodigiosin with significant antimicrobial and antioxidant activity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The identification of antibiotics was a wondrous innovation and discovery that humanity recognized upon its appearance, but the flame of this miracle was quickly extinguished just after the appearance of antimicrobial resistance in pathogens [1]; moreover, the phenomenon of bio-resistance has not ceased to develop and spread in microorganisms [2]. In confronting this serious problem, cancer constitutes another challenge that humanity has tried to conquer, although its pathology is perhaps more serious following the failure of its eradication because of cytotoxicity of chemicals used during treatment against normal cells [3]. These disturbing situations have pushed researchers to leave classical research platforms to pursue other new strategies based on the detection of new microbial strains, in particular Actinobacteria strains producing new bioactive compounds [4]. New actinobacterial strains can be isolated from marine [5], desertic [6] and glacier soil ecosystems [7], or volcanic fumaroles [8]. Following this strategy, several new bioactive compounds have been discovered in recent years, citing for example α-pyrone [9], the polyhydroxyl macrolide lactones PM100117 and PM100118 [10], novonestmycins A and B [11] and venturicidin C [12].
Other previously discovered substances, in particular spectinabilin, undecylprodigiosin and metacycloprodigiosin, are well known for their biological activities. Spectinabilin is a rare polyketide metabolite, and its chemical structure is substituted by a nitrophenyl; spectinabilin was isolated for the first time from a crude streptovaricin complex produced by Streptomyces spectabilis [13]. Undecylprodigiosin and metacycloprodigiosin are red pigments, belonging to the prodigiosins; these compounds are produced by certain species of the Streptomyces genus, such as S. coelicolor A3 and S. longisporus ruber M-3 [14, 15]. Given the importance and the attractive activities of these three compounds, some researchers tried to develop techniques using plug-and-play scaffolding in order to reactivate the silencing gene cluster for spectinabilin in Streptomyces orinoci [16].
Algeria is the largest country in Africa, and the Mediterranean Sea is the limit of the country in the north, while the Sahara Desert is the limit in the south. Between these two borders, the climate is complex and ranges from wet to arid. This study aims to isolate and identify strains belonging to the Streptomyces genus as new producers of spectinabilin, undecylprodigiosin and metacycloprodigiosin from forest soils located in a semi-arid zone in western Algeria.
Materials and Methods
Sampling Location
Soil samples were taken from El-Ogbane forest (34° 49′ 04.9″ N 0° 09′ 26.5″ E) located in the west of the country in Saida city. The samples were taken from sedimentary lands on the banks of the forest river. After removing stones, tree and root debris, the samples were dried in the open air, then crushed with a mortar and kept in plastic containers.
Isolation and Purification of Actinomycetes
Isolation of actinobacteria was performed by the standard dilution method [17]. First, one gram of soil sample was aseptically diluted in 9 ml of sterile distilled water (10–1), followed by agitation of the liquid for a few minutes, and a series of decimal dilutions (10–2, 10–3 and 10–4) was then generated. An aliquot of 100 μl was spread on the surface of starch casein nitrate agar medium (SCNA supplemented with nalidixic acid (20 mg l−1) and nystatin (50 mg l−1)) at the rate of three plates for each dilution [18, 19]. The plates were finally incubated at 30 °C for 5 to 10 days. According to the morphological and cultural characteristics, typical colonies were selected and then sub-cultured on GYM agar medium and incubated at 30 °C for 5 to 10 days [20]. Once the aerial mycelium was formed, the isolates were labelled and stored at − 80 °C in 50% glycerin.
Determination of Culture, Physiological, Biochemical and Microscopic Characteristics of Typical Strains
To identify and compare the isolates from different strains, a series of tests was carried out. Culture characteristics such as colony colour, aerial mycelium production and soluble pigment synthesis were revealed on ISP medium (International Streptomyces Project) by flooding GYM medium, yeast extract-malt extract agar (ISP2), oatmeal agar (ISP3), inorganic salt–starch agar (ISP4), glycerol-asparagine agar (ISP5), peptone-yeast extract iron agar (ISP6), tyrosine agar (ISP7), SSM + T and SSM − T medium with two drops of a fresh culture of the selected isolates [20,21,22]. After growth, the ISCC-NBS colour chart was used to determine the colour of the colonies, aerial mycelium and soluble pigment. Growth at different temperatures (5 to 40 °C, with intervals of 5 °C) was resolved in ISP3 agar over 15 days of incubation. Growth at different pH values was revealed in GYM broth (glucose 1%, yeast extract 1%, K2HPO4·3H2O 0.05%, MgSO4·7H2O 0.05%, weight/volume) over a pH range from 4 to 12 (with intervals of 1.0 pH unit) followed by incubation at 20 °C for 15 days in a rotary shaker [23].
The use of carbohydrates (glucose, arabinose, sucrose, xylose, inositol, mannitol, fructose, raffinose, cellulose and rhamnose) as a single carbon source was determined on 5338 medium ((NH4)2SO4, KH2PO4, K2HPO4, MgSO4·7H2O, agar and trace element solution 5342), which is a basic medium. Incubation was carried out at 30° C for 10 to 15 days [20]. Growth on different concentrations of NaCl (0 to 10%, with intervals of 2.5%) was performed on 5339 medium (casein peptone, yeast extract and agar) as a basic medium. Incubation was carried out at 30 °C for 10 to 15 days. Enzymatic characteristics were revealed on Api Zym® and Api Coryne® plates [24,25,26] according to the protocol accompanying the kits. The preparation of the strains selected for scanning electron microscopy was carried out on agar containing a bacterial lawn fixed in glutaraldehyde using cultures after growth for 4 weeks at 30 °C on ISP3 medium [27].
Genomic DNA Extraction, Amplification, Purification and Sequencing of the 16S rRNA Gene
Genomic DNA extraction was performed by using 0.5 ml of liquid culture using a DNA extraction kit (Stratec Molecular, Invisorb Spin plant, mini Kit, Berlin, Germany) according to the supplied protocol. Verification of the DNA extraction efficiency was performed by agarose gel electrophoresis (0.8% of agarose gel, 70 V, 400 mA, 40 min). Once the genomic DNA was extracted, amplification of 16S rRNA was carried out in a thermocycler (Eppendorf, Mastercycler gradient) using two universal primers (27F: 5′-AGTTTGATCCTGGCTCAG-3′ and 1492R: 5′-ACGGCTACCTTGTTACGACTT-3′) [28]. Cleanup was then carried out using a cleanup kit (Macherey–Nagel, Nucleo Spin, gel and PCR clean up, Düran, Germany). The cleanup of PCR products was confirmed by electrophoresis according to the same previous conditions. Sequencing of PCR products was performed using five primers, 27F, 518R, 1100F, 1100R and 1492R, at the DSMZ (Braunschweig. Germany). The obtained DNA sequences were viewed and edited by Geneouis software V7. The 16S rRNA sequence similarities between the strains were calculated by pairwise alignment using the EzTaxon-e server [29]. Phylogenetic trees were produced with the maximum likelihood [30] and neighbour-joining algorithms [31] using Mega software V7 [32]. The stability of the topology of the phylogenetic trees was evaluated by the 1000 repetition bootstrap method [33]. A distance matrix was generated using Kimura's model of two parameters [34]. All positions with gaps and missing data were eliminated.
Extraction of Secondary Metabolites and Screening of Biological Activities
Screening of Antimicrobial Activity of Crude Extracts
Screening of the antimicrobial activity of crude extracts was accomplished for several pathogenic bacteria and fungi, specifically three microscopic fungi, Gram-negative bacteria and Gram-positive bacteria. All selected pathogenic strains were obtained from the DSMZ (German Collection of Microorganisms and Cell Cultures, Braunschweig, Germany) and ATCC (American Type Culture Collection, Manassas, VA 20110, USA).
To prepare the crude extracts, 20 ml of ethyl acetate (Sigma-Aldrich, USA) was mixed with 20 ml of bacterial culture from the previously prepared selected isolate (culture incubated under agitation for 10 days in two different liquid media, namely: 5294 and 5254). The mixtures were agitated for 12 min, and then the media were separated by centrifugation at 9000 rpm for 10 min. The solvent containing the metabolites was subsequently transferred to a rotavapor (Heidolph, Germany) in order to evaporate it completely. Once evaporation was completely achieved, the weight of the extract is calculated and then recovered in 1 ml of methanol.
The measurement of antimicrobial activity was carried out by microdilution in 96-well microplates. Three liquid culture media were used: Mueller–Hinton (MH), Mycosel (Myc) and Middlebrook (Mid). The choice of medium was determined for each of the pathogenic strains, for which the concentration adjustment was set to 0.01 McFarland units. The dose of extract in the well (A) of the first line corresponded to 1 μl of extract from 14 μl of inoculated culture medium. By successive dilution, the concentration decreased twice in the wells (B–H). The results were obtained by visual observation after 24 and 48 h of incubation at different temperatures [35].
Screening of the Antioxidant Activity of Crude Extracts
ABTS Radical Scavenging Assay
To measure the antioxidant capacity of the crude active extract by ABTS·+ radical cation inhibition, the method described by Surveswaran et al. [36] was applied. ABTS·+ generation was carried out by a chemical reaction between ABTS solution (7 mM) and potassium persulfate solution (2.45 mM) for 16 h in obscurity and at room temperature. The resulting solution (ABTS·+) was diluted in methanol (in a ratio of 1/50) in order to obtain an absorbance of 0.700 ± 0.005 at a wavelength of 734 nm. A concentration of 3.75 µg µl−1 of crude extract was prepared at different volumes (25, 50, 75, 100 µl) and was reacted with 3.9 ml of ABTS·+ solution, and methanol was added to achieve the same volume for all prepared solutions (4 ml). After 10 min of reaction, the absorbance was measured by a spectrophotometer set at 734 nm. Trolox (6-hydroxy-2,5,7,8-tetramethylchroman-2-carboxylic acid) was used as standard antioxidant at a concentration range of 0–500 μM. Neutralization of the ABTS·+ was calculated according to the following formula:
DPPH Radical Scavenging Assay
The antiradical activity of the crude extracts was determined by a DPPH· radical scavenging assay according to the method adopted by Orphanides et al. [37]. A volume of 3.9 ml of DPPH· solution (0.3 mM) was mixed with 3.75 µg µl−1 of crude extract dissolved in methanol at different volumes (25, 50, 75, 100 µl) and incubated for 30 min in obscurity and at room temperature. Methanol volumes were added to achieve the same volume for all prepared solutions (4 ml). The absorbances were then measured by a spectrophotometer set at 517 nm. Trolox (6-hydroxy-2,5,7,8-tetramethylchroman-2-carboxylic acid) was used as standard antioxidant at a concentration range of 0–500 μM. The neutralization of the DPPH· radical was calculated according to the following formula:
Fractionation of Crude Extracts by HPLC and LC–MS
Only the extracts showing good antimicrobial potency were selected for HPLC fractionation. In this study, the HPLC used was an Agilent 1100, with an X-Bridge C18 3.5 µm, 2.1 × 100 mm Column (Waters, Milford, USA) ,and a DAD detector (200–400 nm). Initial pressure was adjusted at 33 bar, and monitoring of the wavelength was located between 210 and 360 nm. The liquid phase was composed of two elution buffers: A1 (950 ml of H2O, 50 ml of C2H3N + 0.05 mM (385 mg l−1) C2H7NO2 + 40 µl of C2H4O2) and B2 (50 ml of H2O, 950 ml of C2H3N, 0.05 mM (385 mg l−1) C2H7NO2 + 40 µl of C2H4O2). After 40 min, the fractions were recovered in a 96-well microplate at a rate of 150 μl per well. Once fractionation was completely achieved, a chromatogram showing the position of the fractions present in the extract was obtained from the HPLC. All obtained fractions were evaporated in a MiniVap (Porvair Sciences, UK) under heated nitrogen (40 °C). After drying the microplates, each well was filled with 150 μl of inoculum containing pathogen adjusted to 0.01 McFarland. After incubation for 24–48 h, the results were determined by visual observation.
To identify the active fractions, HPLC results were compared with those obtained from LC–MS; the instrument used was an LC–MS Agilent 1200 with a DAD detector (200–600 nm) combined with a maXis mass spectrometer (UHR-TOF, Bruker Daltonics, USA). The column was a Waters Acquity UPLC BEH C18 (2.1 × 50 mm, 1.7 μm). The mobile phase was composed of two solvents: A (H2O with 0.1% CH2O2) and B (CH3CN with 0.1% CH2O2) with a flow rate of 0.6 ml min−1. The equilibration time between samples was 5 min [35].
Results
Selection of Isolated Strains
A total of approximately 54 distinct isolates, depending on the colony morphology and pigmentation characteristics of Actinobacteria, were isolated by the dilution method on SCNA medium supplemented with nalidixic acid and nystatin. All isolates were labelled from V000 to V053 and then stored at − 80 °C in 50% glycerol. In the present study, only one isolate (V002) was selected due to its particular cultural characteristics, indicating suspected production of prodigiosins.
Phenotypic Characterization of Isolates
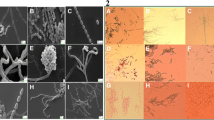
The isolate V002 was observed to be Gram positive, aerobic with abundant mycelium and well developed in the substrate. The aerial mycelium was typical of that of the Streptomyces genus. Scanning electron microscopy allowed visualization of the complex architecture of the isolate, and several morphological characteristics related to the colonies were observed: the aerial mycelia was differentiated by fully matured spores into long chains that were straight and flexible, and the spores were in stick form with a rigorous and non-motile surface. Complete septation of the air chains led to a collapse of mature spores (Fig. 1 in Supplementary Material).
The isolate V002 showed good growth on GYM, ISP3, ISP4, ISP5, ISP7, SSM + T ,and SSM-T medium. Moderate growth was observed on ISP2 and ISP6. The colour of the aerial mass varied between yellow orange and red violet. No diffusible pigment or melanin was observed in the medium. The growth and colour of the aerial and substrate mycelia of V002 on ISP medium are described in Table 1 and Fig. 1 (Supplementary Material). The optimum temperature and pH for mycelial growth were 30 °C and 7.0, respectively. V002 showed significant growth in the absence and presence of 2.5 and 5% NaCl, respectively, and possessed important genetic material that allowed the strain to degrade the whole range of carbohydrates as a single source of carbon (Table 2 and Fig. 1 Supplementary Material).
The enzymatic activity determined by the use of Api Zym® and Api Coryne® showed significant enzymatic potential; V002 exhibited high enzymatic activities in the presence of leucin arylamidase, valine arylamidase, cystine arylamidase, trypsin, chymotrypsin, phosphatase acid, naphtyl-AS-BI-phosphohydrolase, phosphatase alcaline, beta galactosidase, alpha glucosidase, N-acetyl-beta-glucoseamidase and alpha mannosidase. The enzymatic activity of nitrate reductase, pyraziamidase, beta glucuronidase, esculin and urease was almost negative. V002 was unable to ferment sugars; however, it was able to hydrolyse gelatine (Table 3 and Fig. 1 Supplementary Material).
Genotypic Characterization of V002
EzTaxon-e analysis of 16S rRNA gene sequences demonstrated that the bacterial isolate labelled V002 should be classified in the Streptomyces genus. The analysis also showed that the sequence of V002 (1509 bp) had high similarity to Streptomyces lasiicapitis DSM 103124T (99.93%, 1/1494) and Streptomyces spectabilis DSM 40512T (98.96%, 15/1444). A phylogenetic analysis of maximum likelihood confirmed the results (Fig. 1). The 16S rRNA gene sequence of V002 is deposited under accession number MH298058.
Neighbour-joining tree based on 16S rRNA gene sequences showing the relationship between strain V002 and the related taxa. Numbers at nodes are bootstrap values (percentages of 1000 replications); only values > 50% are shown. Asterisks indicate branches that were also recovered in the maximum likelihood tree. Bar, 0.0020 nucleotide substitutions per site
Evaluation of Antibacterial Activity and Determination of Bioactive Compounds
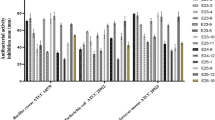
According to the results presented in Table 1, Ext5294.V002 presented low antimicrobial activity (A–B) against C. violaceum, M. smegmatis and P. anomala, minimum inhibitory concentrations (MIC) were 0.125, 0.125 and 0.25 µg µl−1, respectively. An average antimicrobial activity (C–F) was registered against B. subtilis, whereas high activity was registered against S. aureus Newman and M. luteus. Minimum inhibitory concentrations were 0.781 × 10–2, 0.195 × 10–2 and 0.195 × 10–2 µg µl−1, respectively. Ext5254.V002 exhibited moderate activity against B. subtilis. The active extract (Ext5294.V002) was subjected to HPLC fractionation and LC–MS analysis to determine which compounds were active.
The range of the inhibited wells after fractionation of Ext5294.V002 revealed that the fractions exerted strong antimicrobial activity against S. aureus, M. luteus, B. subtilis and C. violaceum For B. subtilis, the correlation test between the peak activity of Ext5294.V002 after HPLC fractionation (Fig. 2 Supplementary Material) and the LC–MS chromatogram (Fig. 3 Supplementary Material) revealed that the active fractions were obtained between 21.5 and 34 min, and the LC–MS data suggested that the peaks correlated with spectinabilin, undecylprodigiosin and metacycloprodigiosin. For M. luteus and S. aureus, peak data from HPLC and LC–MS revealed that active fractions were located between retention time 26.5 and 33.5 min, which suggested that metacycloprodigiosin was the active fraction against these pathogens. For C. violaceum, peak data from HPLC and LC–MS revealed that the active fractions were obtained between 24 and 25 min, which suggested that spectinabilin was the active fraction.
Antioxidant Activity of Crude Extracts
The antioxidant potential of Ext5294.V002 as revealed by DPPH· and ABTS·+ radical assays is presented in Table 2. The figure clearly shows that the volume range between 25 and 100 µl had significant antioxidant activity, which was visible by a change in the solution colour. The percentage of ABTS·+ radical scavenging ranged from 71.48 ± 1.30% to 97.43 ± 0.87%. The ABTS·+ radical assay showed consistent activity between 50 to 100 µl. The percentage of DPPH· radical scavenging ranged from 7.39 ± 0.41% to 25.45 ± 0.29%.
Discussion
Microorganisms of the genus Streptomyces are considered potential producers of secondary metabolites with interesting biological activities and beneficial effects on human health. The research and exploitation of Streptomyces from unexplored environments is one of the most effective approaches for discovering new bioactive metabolites [28]. Streptomyces sp. V002 was isolated from El-Ogbane forest, located in the semi-arid zone in Algeria; this forest contains plants and trees with muddy soils in winter and high temperatures in summer. The river that divides the forest creates a unique environment that contains a diversity of flora, freshwater sediments and microorganisms.
The search for new strains producing spectinabilin and prodigiosins requires reliable screening and identification. Despite the specificity of the 16S rRNA region, a dendrogram showed that Streptomyces sp. V002 had a high concordance with the grouping and topology of S. lasiicapitis and S. spectabilis; however, the comparison of cultural and biochemical characteristics revealed the dissimilarity of both types of strains. LC–MS results demonstrated that Streptomyces sp. V002 isolated from the sedimentary lands on the banks of the forest river was considered a new producer of spectinabilin, metacycloprodigiosin and undecylprodigiosin; however, S. lasiicapitis identified by Ye produced kanchanamycin [38], and S. spectabilis produced spectinabilin and metacycloprodigiosin [13, 39].
Only a few compounds containing nitro groups are known and among them is spectinabilin which is a rare polyketide metabolite substituted with a nitrophenyl group; also, few studies have unveiled the antimicrobial activity of spectinabilin, metacycloprodigiosin and undecylprodigiosin [40]. Evaluation of the antimicrobial activity of secondary metabolites of crude and fractionated extracts revealed good inhibitory activity. Spectinabilin possesses significant biological activities against P. falciparum K1 [39], and this compound also has significant nematocidal activity against Bursaphelenchus xylophilus, with an LC50 equal to 0.84 µg ml−1 [41]. Spectinabilin isolated from Streptomyces sp. ZQ4BG demonstrated suppression of C. albicans with an MIC of 12.5 μg ml−1 [42]. The antimicrobial activity of undecylprodigiosin was confirmed in the study of Stankovic and his team, in which the compound was able to inhibit the growth of M. luteus and B. subtilis at a concentration of 50 μg ml−1, while for C. albicans ATCC 10231 and C. albicans ATCC 10259, the inhibition concentrations were 100 and 200 μg ml−1, respectively [43]. In the study of Zainal Abidin et al., undecylprodigiosin demonstrated strong antibacterial activities against S. aureus, B. subtilis and C. albicans, with algicidal activity against A. minutum and P. bahamense [44]. Metacycloprodigiosin is known for its anti-malaria activity [39], and it also induces cell death in β-catenin-mutated tumour cells [45]. Metacycloprodigiosin and undecylprodigiosin possess anticancer activity against several human cancer cell lines (P388, HL60, A-549, BEL-7402 et SPCA4) [46].
The complexity and multifunctionality of bioactive compounds in an extract make it difficult to choose a single assay to detect antioxidant activity, although the analyses performed by DPPH· and ABTS·+ radical scavenging assays are robust and simple to perform [28]. In this study, both assays were used for a preliminary screening to determine the antioxidant capacity of the crude extract. The antioxidant reactions were measured by a hydrogen atom transfer or an election transfer on probe molecules [47]. The most active crude extracts against selected microorganisms revealed good antioxidant potential, and these results may be pursued by further research to determine which compound is more active than the others and to develop important products. The radical scavenging activity of the extract was proportionally related to the compositions of secondary metabolites and the concentration of bioactive compounds, which was corroborated by Tan and his team, who said that antioxidant activity correlates with content for bioactive compounds [28].
Several studies reported that extracts from Streptomyces provided antioxidant potential; Raghava Rao and Raghava Rao [48] demonstrated that Streptomyces isolated from mangrove soil of the Visakhapatnam region was endowed with antioxidant activity, while another study confirmed our findings and stated that ethyl acetate extract obtained from Streptomyces sp. AM-S1 possessed antioxidant potential against DPPH· and ABTS·+ with IC50 values of 90.2 and 13.2 μl ml−1, respectively [49]. Streptomyces V002 is recognized as a good source of antioxidants. With the results obtained, these antioxidants may prevent the progression of various disorders related to oxidative stress, and they have good potential to avoid cell damage resulting from a redox imbalance following the rise of O2·− products after exceeding the cell defence capacity [28]. Undecylprodigiosin has demonstrated its antioxidant activity against the oxidation of linoleic acid [43], and it can also exert gastroprotective effects and attenuate induced gastric lesions via antioxidant and anti-inflammatory mechanisms by decreasing the levels of inflammatory mediators and apoptotic markers [50].
Adaptation of microorganisms of the El-Ogbane forest to climatic and environmental conditions allows them to develop metabolic capacities and to synthesize prodigious compounds that allow them to survive in this forest ecosystem. In this study, the ability of Streptomyces sp. V002 isolated from sedimentary lands on the banks of the forest river to produce spectinabilin, undecylprodigiosin and metacycloprodigiosin was confirmed. Previously published results strongly suggest that the three substances could be selected as important lead molecules for the development of chemotherapy treatment. The present study has demonstrated the antimicrobial and antioxidant activities of the three molecules, which clarifies their importance with the producing strain. In-depth investigation of the underlying mechanism of the antimicrobial effect of spectinabilin, undecylprodigiosin and metacycloprodigiosin would be valuable in the future.
References
Davies J, Davies D (2010) Origins and evolution of antibiotic resistance. Microbiol Mol 74:417–433
Friedman ND, Temkin E, Carmeli Y (2016) The negative impact of antibiotic resistance. Clin Microbiol Infect 22:416–422. https://doi.org/10.1016/j.cmi.2015.12.002
McLachlan A, Kekre N, McNulty J, Pandey S (2005) Pancratistatin: a natural anti-cancer compound that targets mitochondria specifically in cancer cells to induce apoptosis. Apoptosis 10:619–630. https://doi.org/10.1007/s10495-005-1896-x
Mahajan GB, Balachandran L (2012) Antibacterial agents from actinomycetes—a review. Front Biosci 4:240–253. https://doi.org/10.2741/373
Hassan SS, Anjum K, Abbas SQ, Akhter N, Shagufta BI, Shah SA, Tasneem U (2017) Emerging biopharmaceuticals from marine actinobacteria. Environ Toxicol Pharmacol 49:34–47. https://doi.org/10.1016/j.etap.2016.11.015
Yang ZW, Salam N, Mohany M, Chinnathambi A, Alharbi SA, Xiao M, Hozzein WN, Li WJ (2018) Microbacterium album sp. nov. and Microbacterium deserti sp. nov., two halotolerant actinobacteria isolated from desert soil. Int J Syst Evol Microbiol 68:217–222. https://doi.org/10.1099/ijsem.0.002485
Hauptmann AL, Stibal M, Bælum J, Sicheritz-Pontén T, Brunak S, Bowman JS, Hansen LH, Jacobsen CS, Blom N (2014) Bacterial diversity in snow on North Pole ice floes. Extremophiles 18:945–951. https://doi.org/10.1007/s00792-014-0660-y
Medrano-Santillana M, Souza-Brito EM, Duran R, Gutierrez-Corona F, Reyna-López GE (2014) Bacterial diversity in fumarole environments of the Paricutín volcano, Michoacán (Mexico). Extremophiles 21(3):499–511. https://doi.org/10.1007/s00792-017-0920-8
Yang N, Song F (2018) Bioprospecting of novel and bioactive compounds from marine actinomycetes isolated from south china sea sediments. Curr Microbiol 75:142–149. https://doi.org/10.1007/s00284-017-1358-z
Pérez M, Schleissner C, Fernández R, Rodríguez P, Reyes F, Zuñiga P, de la Calle F, Cuevas C (2016) PM100117 and PM100118, new antitumor macrolides produced by a marine Streptomyces caniferus GUA-06-05-006A. J Antibiot 69:388–394. https://doi.org/10.1038/ja.2015.121
Wan Z, Fang W, Shi L, Wang K, Zhang Y, Zhang Z, Wu Z, Yang Z, Gu Y (2015) Novonestmycins A and B, two new 32-membered bioactive macrolides from Streptomyces phytohabitans HBERC-20821. J Antibiot 68:185–190. https://doi.org/10.1038/ja.2014.123
Shaaban KA, Singh S, Elshahawi SI, Wang X, Ponomareva LV, Sunkara M, Copley GC, Hower JC, Morris AJ, Kharel MK, Thorson JS (2014) Venturicidin C, a new 20-membered macrolide produced by Streptomyces sp. TS-2-2. J Antibiot 67:223–230
Kakinuma K, Hanson CA, Rinehart Jr KL (1976) Spectinabilin, a new nitro-containing metabolite isolated from Streptomyces spectabilis. Tetrahedron 32:217–222. https://doi.org/10.1016/0040-4020(76)87004-4
Sevcikova B, Kormanec J (2004) Differential production of two antibiotics of Streptomyces coelicolor A3(2), actinorhodin and undecylprodigiosin, upon salt stress conditions. Arch Microbiol 181:384–389. https://doi.org/10.1007/s00203-004-0669-1
Wasserman HH, Rodgers GC, Keith DD (1969) Metacycloprodigiosin, a tripyrrole pigment from Streptomyces longisporus ruber. J Am Chem Soc 91:1263–1264. https://doi.org/10.1021/ja01033a065
Shao Z, Rao G, Li C, Abil Z, Luo Y, Zhao H (2013) Refactoring the silent spectinabilin gene cluster using a plug-and-play scaffold. ACS Synth Biol 2:662–669
Valanarasu M, Duraipandiyan V, Agastian P, Ignacimuthu S (2009) In vitro antimicrobial activity of Streptomyces from Western Ghats rock soil (India). J Mycol Med 19:22–28. https://doi.org/10.1016/j.mycmed.2008.12.002
Tanasupawat S, Jongrungruangchok S, Kudo T (2010) Micromonospora marina sp. nov., isolated from sea sand. Int J Syst Evol Microbiol 60:648–652. https://doi.org/10.1099/ijs.0.014068-0
Songsumanus A, Tanasupawat S, Thawai C, Suwanborirux K, Kudo T (2011) Micromonospora humi sp. nov., isolated from peat swamp forest soil. Int J Syst Evol Microbiol 61:1176–1181. https://doi.org/10.1099/ijs.0.024281-0
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340. https://doi.org/10.1099/00207713-16-3-313
Kämpfer P, Glaeser SP, Parkes L, van Keulen G, Dyson P (2014) The family Streptomycetaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes, 4th edn. Springer, Berlin, pp 889–1010
Suter MA (1978) Isolierung von Melanin-negativen Mutanten aus Streptomyces glaucescens, thesis no. 6276. Eidgenossische Technische Hochschule, Switzerland
Jia F, Liu C, Wang X, Zhao J, Liu Q, Zhang J, Gao R, Xiang W (2013) Wangella harbinensis gen. nov., sp. nov., a new member of the family Micromonosporaceae. Antonie Van Leeuwenhoek 103:399–408. https://doi.org/10.1007/s10482-012-9820-1
MacFaddin JF (2000) Biochemical tests for the identification of medical bacteria, 3rd edn. Lippincott Williams & Wilkins, Philadelphia
Humble MW, King A, Phillips I (1977) API ZYM: a simple rapid system for the detection of bacterial enzymes. J Clin Pathol 30:275–277. https://doi.org/10.1136/jcp.30.3.275
Kilian M (1978) Rapid identification of Actinomycetaceae and related bacteria. J Clin Microbiol 8:127–133
Wink J (2002) Polyphasic taxonomy and antibiotic formation in some closely related genera of the family Pseudonocardiaceae. Recent Res Dev Antibiot 2:97–140
Tan LT, Chan KG, Khan TM, Bukhari SI, Saokaew S, Duangjai A, Pusparajah P, Lee LH, Goh BH (2017) Streptomyces sp. MUM212 as a source of antioxidants with radical scavenging and metal chelating properties. Front Pharmacol 8:276
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721. https://doi.org/10.1099/ijs.0.038075-0
Kim SB, Brown R, Oldfield C, Gilbert SC, Iliarionov S, Goodfellow M (2000) Gordonia amicalis sp. nov., a novel dibenzothiophene-desulphurizing actinomycete. Int J Syst Evol Microbiol 50:2031–2036. https://doi.org/10.1099/00207713-50-6-2031
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. https://doi.org/10.1007/BF01731581
Charousová I, Medo J, Hleba L, Javoreková S (2018) Streptomyces globosus DK15 and Streptomyces ederensis ST13 as new producers of factumycin and tetrangomycin antibiotics. Braz J Microbiol 49:816–822
Surveswaran S, Cai YZ, Corke H, Sun M (2007) Systemic evaluation of natural phenolic antioxidants from 133 Indian medicinal plants. Food Chem 102:938–953. https://doi.org/10.1016/j.foodchem.2006.06.033
Orphanides A, Goulas V, Gekas V (2013) Effect of drying method on the phenolic content and antioxidant capacity of spearmint. Czech J Food Sci 31:509–513
Ye L, Zhao S, Li Y, Jiang S, Zhao Y, Li J, Yan K, Wang X, Xiang W, Liu C (2017) Streptomyces lasiicapitis sp. nov., an actinomycete that produces kanchanamycin, isolated from the head of an ant (Lasius fuliginosus L.). Int J Syst Evol Microbiol 67:1529–1534. https://doi.org/10.1099/ijsem.0.001756
Isaka M, Jaturapat A, Kramyu J, Tanticharoen M, Thebtaranonth Y (2002) Potent in vitro antimalarial activity of metacycloprodigiosin isolated from Streptomyces spectabilis BCC 4785. Antimicrob Agents Chemother 46:1112–1113
Choi YS, Johannes TW, Simurdiak M, Shao Z, Lu H, Zhao H (2010) Cloning and heterologous expression of the spectinabilin biosynthetic gene cluster from Streptomyces spectabilis. Mol Biosyst 6:336–338. https://doi.org/10.1039/b923177c
Liu MJ, Hwang BS, Jin CZ, Li WJ, Park DJ, Seo ST, Kim CJ (2019) Screening, isolation and evaluation of a nematicidal compound from actinomycetes against the pine wood nematode, Bursaphelenchus xylophilus. Pest Manag Sci 75:1585–1593. https://doi.org/10.1002/ps.5272
Wang W, Song T, Chai W, Chen L, Chen L, Lian XY, Zhang Z (2017) Rare polyene-polyol macrolides from mangrove-derived Streptomyces sp. ZQ4BG. Sci Rep 7:1703
Stankovic N, Radulovic V, Petkovic M, Vuckovic I, Jadranin M, Vasiljevic B, Nikodinovic-Runic J (2012) Streptomyces sp. JS520 produces exceptionally high quantities of undecylprodigiosin with antibacterial, antioxidative, and UV-protective properties. Appl Microbiol Biotechnol 96:1217–1231. https://doi.org/10.1007/s00253-012-4237-3
Zainal Abidin ZA, Ahmad A, Latip J, Usup G (2016) Marine Streptomyces sp. UKMCC_PT15 producing undecylprodigiosin with algicidal activity. J Teknologi 78:55–60
Ikeda H, Shikata Y, Watanapokasin R, Tashiro E, Imoto M (2017) Metacycloprodigiosin induced cell death selectively in β-catenin-mutated tumor cells. J Antibiot 70:109–112. https://doi.org/10.1038/ja.2016.75
Liu R, Cui CB, Duan L, Gu QQ, Zhu WM (2005) Potent in vitro anticancer activity of metacycloprodigiosin and undecylprodigiosin from a sponge-derived actinomycete Saccharopolyspora sp. nov. Arch Pharm Res 28:1341–1344. https://doi.org/10.1007/BF02977899
Apak R, Özyürek M, Güçlü K, Çapanoğlu E (2016) Antioxidant activity/capacity measurement. 2. Hydrogen atom transfer (HAT)-based, mixed-mode (electron transfer (ET)/HAT), and lipid peroxidation assays. J Agric Food Chem 64:1028–1045. https://doi.org/10.1021/acs.jafc.5b04743
Raghava Rao KV, Raghava Rao TJ (2013) Molecular characterization and its antioxidant activity of a newly isolated Streptomyces coelicoflavus BC 01 from mangrove soil. J Young Pharm 5(4):121–126
Sowndhararajan K, Kang SC (2013) Evaluation of in vitro free radical scavenging potential of Streptomyces sp. AM-S1 culture filtrate. Saudi J Biol Sci 20:227–233. https://doi.org/10.1016/j.sjbs.2012.12.003
Abdelfattah MS, Elmallah MIY, Ebrahim HY, Almeer RS, Eltanany RMA, Abdel Moneim AE (2019) Prodigiosins from a marine sponge-associated actinomycete attenuate HCl/ethanol-induced gastric lesion via antioxidant and anti-inflammatory mechanisms. PLoS ONE 14:e0216737
Acknowledgements
We are thankful to the researchers and technician from microbial strain collection department, Helmholtz Centre for Infection Research for their support during this work.
Author information
Authors and Affiliations
Contributions
MAG carried out experimental work. MAG and BB co-wrote the manuscript. AOHK and JW conceived and designed the study. All authors contributed to interpretation of results, read and approved the final draft.
Corresponding author
Ethics declarations
Conflict of interest
We declare that we have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gacem, M.A., Ould-El-Hadj-Khelil, A., Boudjemaa, B. et al. Antimicrobial and Antioxidant Effects of a Forest Actinobacterium V002 as New Producer of Spectinabilin, Undecylprodigiosin and Metacycloprodigiosin. Curr Microbiol 77, 2575–2583 (2020). https://doi.org/10.1007/s00284-020-02007-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02007-1