Abstract
Flagella occur on many prokaryotes, which primarily propel cells to move from detrimental to favorable environments. A variety of species-specific flagellation patterns have been identified. Although it is presumed that for each of these flagellated microorganisms, an evolutionarily fixed flagellation pattern is favored under the normal living conditions, direct evidence is lacking. Here, we use Shewanella oneidensis, a rod-shaped Gram-negative bacterium with a monotrichous polar flagellum (MR-1, the wild-type), as a research model. The investigation has been enabled by multiple mutants with diverse flagellation patterns that had been generated by removing FlhF and FlhG proteins that control flagellar location and number, respectively. Growth assays, as a measure of fitness, revealed that the wild-type strain predominated in spreading on swim plates and in pellicles which form at the air–liquid interface. However, under the pellicles where oxygen is limited, both aflagellated and monotrichous lateral strains showed similar increase in fitness, whereas strains with multiple flagella were less competitive. Moreover, under shaking culturing conditions, the aflagellated strain outcompeted all other strains, including the wild-type, suggesting that cells devoid of flagella would be more likely enriched upon agitation. Overall, these data support the presumption that the monotrichous polar flagellum, as evolutionarily fixed in the wild-type strain, is optimal for the growth fitness of S. oneidensis over any other mutants under most test conditions. However, upon specific changes of environmental conditions, another form could come to predominate. These findings provide insight into the impacts of flagellation patterns and function on bacterial adaptation to differing environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
For many prokaryotes, the ability to actively move from detrimental to favorable niches confers an important advantage for survival and fitness in their varied habitats [1, 2]. The surface appendages that prokaryotic cells have evolved for locomotion are highly varied and include flagella, pili, and mycoplasma “legs” [3]. Among them, flagella/flagellum, semi-rigid filamentous structures which extrude through the cell wall, are probably the most effective for bacteria. By rotation, they are able to propel cells in liquid medium as fast as 60 μm/s for some species [4, 5]. Many bacterial species are motile by means of flagella. Flagella can also play other critical roles relating to adhesion, biofilm (cell community) formation, and host invasion [6, 7].
In bacteria, the flagellation pattern is usually species-specific, whilst many interspecies variations in flagellar location and number occur [8]. For each isolate, the flagellation pattern appears to be fairly fixed and alternatives to the dominant pattern are rare occurrences in the wild-type [9]. The major flagellation patterns that have been characterized are monotrichous (a single flagellum, often polar; e.g., Vibrio cholerae, Pseudomonas aeruginosa); amphitrichous (a single or multiple flagellum at both ends; e.g., Campylobacter jejuni); lophotrichous (a tuft of flagella at one end; e.g., Helicobacter pylori); and peritrichous (flagella distributed along the length of a rod-shaped cell; e.g., Escherichia coli) [10,11,12]. In addition, some bacteria with multiple flagella occurring near the midpoint of the cell body are known as lateral (e.g., Selenomonas ruminantium) [13].
As a complex molecular machine, a typical bacterial flagellum is composed of over 20 different structural proteins assembled to form a basal body (including MS ring, P ring, L ring, and export apparatus), a motor, a switch, a hook, and a filament [14, 15]. The energy cost for flagellar synthesis is enormous, consuming up to 2% of a cell’s metabolic resources [16]. Among flagellar proteins, the flagellin accounts for most of the energy consumption as it requires the synthesis of up to 20,000 units for filament construction [15]. In addition to the cost of synthesis, flagellar rotation spends on ion motive forces, ~ 104–105 protons per second in the case of E. coli [17]. Because of the high-energy investment on motility, flagellar synthesis, assembly and rotation are tightly regulated at multiple levels [18].
Shewanella spp. comprise a highly diverse group of γ-proteobacteria that are widely distributed in marine and freshwater environments [19]. Shewanella spp. are most usually motile by means of a polar flagellum. The species considered as representative of the genus, S. oneidensis, is typical in this regard. However, some atypical Shewanella strains have an accessory peritrichous lateral flagellar system that is conditionally synthesized [9, 20]. Most of the current understanding of the flagellar assembly and regulation of Shewanella strains have been derived from studies on S. oneidensis [21,22,23,24,25,26,27,28,29,30].
S. oneidensis carries a set of genes for a single polar flagellum [23]. The flagellar assembly and the regulatory hierarchy for the expression of flagellar genes in S. oneidensis has been shown to be similar to those of other well studied polar flagellates, such as V. cholerae [24, 30]. In this way, S. oneidensis has been accepted as a primary model species for polar flagella related studies. Despite this, a variety of mutant strains of S. oneidensis have also been previously created displaying all of the other main flagellation patterns (except amphitrichous) [23, 27, 28]. These variations have been achieved by deleting the flhF and flhG genes which, as a cognate pair, are responsible for the spatial and numerical control of flagella. FlhF and FlhG, encoded by genes in a flagellar cluster, have been generally found in polar flagellates as well as occasionally in peritrichous bacteria such as Bacillus subtilis [12]. FlhF, a signal-recognition particle-like (SRP-like) GTPase, binds GTP (not necessarily hydrolysis) to localize the basal body protein FliF to the membrane at the cell pole [28, 31, 32]. In polar flagellates, FlhG, resembling the ParA/MinD superfamily of ATPases, regulates the flagellar number by cycling between two distinct states: a membrane-associated, ATP-bound homodimer and an ADP-bound monomer soluble in the cytoplasm [33]. Loss of FlhG in V. cholerae and P. aeruginosa consistently results in a multi-flagella phenotype, leading to substantially reduced motility [34, 35].
As a consequence, flagellation patterns may affect bacterial environmental fitness. This aspect has been little studied, presumably due to the lack of strains with different flagellation patterns that can be derived from a single bacterial species/isolate. In the present study, the wild-type and mutant strains, the latter displaying diverse flagellation patterns, were used to test growth fitness in three different conditions: swim plates, shaking, and static liquid cultures. Results showed that the wild-type strain held great advantages over the other mutants in swim plates and static liquid conditions where motility remains critical. However, in agitated liquid conditions, the aflagellated mutant strain outcompeted all other tested strains, including the wild-type. This study not only provides insight into the mechanism through which the polar flagellum is advantageous over other flagellation patterns in most environments, and is thus evolutionarily conserved in Shewanella, but also demonstrates how another pattern could come to predominate given specific changes in environmental conditions.
Materials and Methods
Bacterial Strains and Culture Conditions
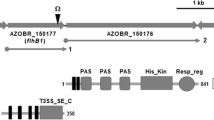
Shewanella oneidensis MR-1 strain (the wild-type, ATCC 700550) and the ΔflaAΔflaB, ΔflhF, ΔflhG, and ΔflhFΔflhG derivatives [23, 27, 28] were used in this study. Their relevant features are shown in Fig. 1 including flagellation diagrams created according to transmission electron microscope photos [27, 28], where the quantities of flagellin subunits were also shown in these references, and motility. The aflagellated strain was ∆flaA∆flaB, in which both flagellin genes flaA and flaB had been removed [23, 27]. The two strains with multiple flagella were ∆flhG and ∆flhF∆flhG, which were lophotrichous-like and peritrichous-like, respectively [28]. Additionally, the strain devoid of the flhF gene carried a single flagellum randomly located anywhere around the periphery of the cell [28].
Flagellar and motile characteristics of the five strains used in this study. a The diagram depicting the flagellation patterns of the indicated strains is drawn according to the published reports [27, 28]. b Identification of the five strains by their motility and PCR. The wild-type and the ΔflhF strains were identified due to their colony sizes on swim plates, which were larger than 1.5 cm and 0.5 cm in diameter, respectively. Scale bar = 0.5 cm. However, the ΔflaAΔflaB, ΔflhG and ΔflhFΔflhG strains formed colonies all smaller than 0.5 cm in diameter. As for strain identification, the ΔflaAΔflaB, ΔflhG and ΔflhFΔflhG strains were identified using PCR analysis (bottom). The products of amplified DNA measuring approximately 2.5 Kb, 1.8 Kb and 0.8 Kb in length indicate the ΔflaAΔflaB, ΔflhG and ΔflhFΔflhG strains, respectively. The lengths of the amplified DNA fragments are shown on the left. Experiments were performed at least three times, and similar results were obtained
With respect to motility, the ΔflaAΔflaB strain had been previously noted as non-motile on swim plates, whereas the ΔflhF, ΔflhFΔflhG, and ΔflhG strains retained approximately 30%, 10%, and < 5% of the motility relative to that of the wild-type, respectively [27, 28]. Multiple lines of evidence suggest that all of the flagellated strains had assembled normal flagella [27, 28, 30]. This was as expected as the changes in genes and in their levels of expression had been restricted to the proteins where their only known function relates to the location and numeric control of flagella. Thus, any motility differences among them could be clearly attributable to the altered flagellation patterns.
All strains were grown in lysogeny broth (LB) medium at 30 °C under aerobic conditions. A single colony on overnight solid LB plates for each of the five strains under testing was used to inoculate 3 mL LB. After an overnight growth period, these cultures were used to inoculate 3 mL fresh LB with a 100-fold dilution. Growth of S. oneidensis strains in liquid LB medium was then measured by recording optical densities at 600 nm (OD600) under aerobic conditions, and converted to colony-forming units per milliliter (CFU/mL). When cultures grew to approximately 1.4 × 108 CFU/mL, medium of each strain were collected and any difference in cell densities among the strains was eliminated with fresh LB, thus providing the startup cultures for use in the competition assay.
Pellicle Formation Assay
The cultures for each strain were collected and adjusted to 108 CFU/mL with fresh LB. Twenty microliters of the resulting cultures were inoculated into 2 mL LB in 24-well plates as before [36]. After static cultivation, the planktonic biomass was removed from the bottom of the well with a syringe for optical density measurement. This was conducted even before the pellicles were formed. The total bacterial biomass was evaluated using the OD600 reading of the entire biomass in the well (mixing the pellicle cells with cultures underneath by vigorously shaking). The difference between the OD600 readings for planktonic and total biomass was taken as the pellicle biomass.
Growth Competition of S. oneidensis Strains
Two hundred microliters of CFU/mL-adjusted cultures for each strain were transferred to a new tube to create an equally-mixed culture. This was used for the following competition experiments. (i) Motility. Motility competition was performed by spotting 2.5 μL of the mixed cultures on swim LB plates. Samples from 3 different areas (center, radial midpoint and edge of the droplet) were taken after 24 h incubation at 30 °C. (ii) Growth competition under shaking conditions. The mixed culture was inoculated with 10 mL LB by 100-fold dilution and incubated in a flask on a shaker at 250 rpm. Samples were taken and assayed at ~ 2 × 108 CFU/mL. (iii) Growth competition under static conditions. A 50 μL of the mixed culture was inoculated with 2 mL LB in 24-well plates. Samples were taken from pellicles and cultures underneath after incubation for 20 h.
Analysis of Population Composition
To determine the population composition of the competing cultures, the samples from the cultures were collected, properly diluted, and applied onto fresh solid LB plates (agar concentration, 1.5% w/v, on which all test strains were permitted to grow equally) to form separated colonies. From each plate, 36 colonies were chosen randomly for analysis of population composition. All colonies chosen were transferred onto swim LB plates to test their motility (to facilitate comparison, the non-motile ΔflaAΔflaB strain was always included at the center of the same plate). After 18 h incubation at 30 °C, colonies over 1.5 cm in diameter were assigned as the wild-type strain, and colonies between 0.5 and 1.5 cm in diameter were assigned as the ΔflhF strain, in accordance with previous reports [27, 28].
Colonies of less than 0.5 cm in diameter were further tested by polymerase chain reaction (PCR). The cultures were used as templates for amplification of the FlhFG gene locus (all cultures had been preconditioned at 100 °C for 5 min for the disruption of cells and the release of genomic DNA). For PCR, amplification was primed using a pair of oligonucleotides (FlhFG-F: 5′-CCTAATCTCAGAGTGATTTC-3′; and FlhFG-R: 5′-GGTAATCTGGCAAGCAAGTG-3′) derived from the FlhFG sequence. The PCR program consisted of following steps: denaturing at 95 °C for 10 min, 25 cycles of 95 °C for 30 s, 56 °C for 30 s, and 72 °C for 2 min 10 s, then polymerization at 72 °C for 10 min. PCR products were then analyzed using 1% agarose gel electrophoresis. Products of amplified DNA measuring approximately 2.5 Kb, 1.8 Kb and 0.8 Kb in length indicated the ΔflaAΔflaB, ΔflhG and ΔflhFΔflhG strains, respectively.
Statistical Analyses
Data are given as means ± SEM from three independent replicates. In the growth competition experiments of five strains under shaking conditions, analysis of the statistical significance of frequencies of strains against the uniform distribution was performed using χ2 test. In the growth competition experiments between two strains (the wild-type strain verses the aflagellated ΔflaAΔflaB mutant strain), analysis of the statistical significance of frequencies of strains against the uniform distribution was performed using binomial test on untransformed proportions.
Results
Motility Dictates Fitness on Swim Plates
To investigate the influence of flagellation patterns on fitness of S. oneidensis strains, we used competition assays to measure their fitness. All strains used in the study were grown in liquid LB medium and adjusted to ~ 108 CFU/mL with fresh LB. For competition experiments, an evenly mixed culture was deliberately prepared to be composed of the same volume of cultures with identical CFU/mL (~ 108) for each of the five strains under testing (wild-type, ΔflhF, ΔflaAΔflaB, ΔflhG, and ΔflhFΔflhG). A 2.5 µL mixed culture was then dropped on swim plates for evaluating fitness associated with motility.
The diameter of the droplet was about 1 mm. After incubation for 24 h at 30 °C, the droplet grew to approximately 2.5 cm in diameter (Fig. 2a). To examine the strain composition of the droplet, cells were collected from 3 different areas of the droplet: center (blue circle), radial midpoint (green circle), and edge (yellow circle) (Fig. 2a). Collection was conducted using pipette tips (Fig. 2a), with the sample properly diluted (to form 100 to 300 colonies per plate), and applied onto fresh solid LB plates for colony development. In respect to their size, colonies formed on these solid plates appeared indistinguishable from one another (data not shown). This suggests that any differing motility or energy costs for strains with different flagellation patterns had failed to result in any significant differences of colony size on solid LB plates. Subsequently, 36 colonies were randomly chosen from each plate and transferred onto swim LB plates for strain identification according to significant motility differences between the wild-type, ΔflhF, and other strains. Results from samples taken from the droplet center, radial midpoint and edge are shown in Fig. 2b.
Growth competition of the five strains on swim plate. a A single droplet composed of the evenly mixed cultures of the five strains was applied onto the swim plate and incubated until the droplet grew to approximately 2.5 cm in diameter. Samples from the center (blue), radial midpoint (green) and edge (yellow) were picked up for the determination of population composition. Scale bar = 1 cm. b Motility of 36 colonies from cells collected at the center, radial midpoint and edge as shown in (a), respectively. Colonies larger than 1.5 cm in diameter were identified to be the wild-type strain. The colony (purple arrow) larger than 0.5 cm in diameter was identified to be the ΔflhF strain. Colonies (red and black arrows) smaller than 0.5 cm in diameter were unidentified. To facilitate comparison, the non-motile ΔflaAΔflaB strain was always included at the center of each plate. Scale bar = 1 cm. c Identification of the two colonies indicated by red and black arrows in (b) using PCR analysis. The lengths of the amplified DNA fragments are shown on the left. Asterisk indicates non-specific amplification. d The distribution of each strain in the center, radial midpoint, and edge of the single droplet. Experiments in the center area were performed in triplicate, a–c only show the representative data. Data are shown as means with error bars representing the standard error of the mean (SEM) (Color figure online)
Among the 36 colonies separated from the droplet center, 33 were of the wild-type strain, based on their motility capable of forming colonies larger than 1.5 cm in diameter on swim LB plates (Fig. 2b). One colony (purple arrow) was assigned as ΔflhF, due to its intermedia colony size (larger than 0.5 cm, but smaller than 1.5 cm in diameter). The remaining two colonies with significantly impaired motility showed no obvious visual differences on colony size and had to be determined by PCR. Using primers specific for the mutation regions, we were able to demonstrate that they were ΔflaAΔflaB (red arrow) and ΔflhFΔflhG (black arrow) (Fig. 2b, c). In the case of cells from the radial midpoint and edge areas, all 36 colonies were shown to be of the wild-type, based on their motility on swim LB plates (Fig. 2b).
The strain identification from center areas showed that the highly non-motile mutants were similar in general, though with slight variations (Fig. 2d). Although the wild-type strain predominated in the population at the center of the droplet, the other strains still could be found here. In contrast, all colonies from both the radial midpoint and edge areas of the culture droplet were identified to be of the wild-type. These results collectively indicate that the monotrichous polar flagellum of the wild-type strain offers an overwhelming competitive advantage in situations where motility matters.
Loss of Flagellum Confers S. oneidensis a Fitness Gain in Shaking Liquid Culture
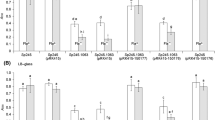
Since the flagellar filament consists of a large number of flagellins, this may impose an energy burden on flagellated cells. In a previous study [28], the flagellin amount in each of the strain used here was tested. The study showed that the strain with the highest flagellin production was ΔflhFΔflhG (in which flagellin protein amount was elevated by 19-fold compared to the wild-type), whereas the strain with the lowest flagellin production was ΔflaAΔflaB (in which no flagellin was detectable). To analyze the impact of different expression levels of flagellin on the growth of these strains, we measured the growth of the ΔflhFΔflhG strain and the ΔflaAΔflaB strain in contrast to the wild-type. Under shaking (250 rpm) liquid LB aerobic conditions, although the aflagellated ΔflaAΔflaB strain grew slightly faster than the wild-type at the early stages of the cultivation, the overall growths of the mutants were similar to those of the wild-type, especially as the cultivation continued (Fig. S1).
As conventional characterization is unable to reveal subtle differences in the influence of the individual mutations on growth, competition experiments were also conducted. A volume of 100 µL of evenly mixed cultures of the five strains under test was inoculated into 10 mL LB in a flask and then grown to ~ 2 × 108 CFU/mL on a shaker at 250 rpm, from which samples were taken. The strain composition of the samples was then determined as described above (Fig. S2a, b). As summarized in Fig. 3a, the share of the ΔflaAΔflaB strain increased from the initial, ~ 20%, to ~ 37%, as compared to ~ 18% of the wild-type, implying that the aflagellated strain has a competitive growth advantage over others, even including the wild-type, in shaking liquid culture. Since motilities were no longer required under such conditions, we presume it is the differences of the expression levels of flagellin, and thus the energy expenditures, that caused their competitive differences. As revealed before [28], the ΔflhF mutant only produced 46% of the flagellin relative to that of the wild-type, whereas the flagellin protein amounts in the ΔflhG and ΔflhFΔflhG mutants were elevated by 14- and 19-fold compared to the wild-type, respectively. In addition, the ΔflaAΔflaB strain, previously noted as the flagellin-free mutant (FFM) [28], could not synthesize flagellin proteins. After the mixed culture, the distribution of the strains showed significant imbalance from the initial status of uniform distribution (P < 0.05), and the average percentage of each tested strain indicated a negative trend with their expression levels of flagellin. It therefore seems that the flagellin protein level, and thus the energy expenditure, is a critical factor affecting growth.
Growth competition of the strains under shaking liquid conditions. a Growth competition of the five strains under 250 rpm shaking condition. The culture, composed of an evenly mixed culture of the five strains under test, was grown to ~ 2 × 108 CFU/mL. The strain composition of the culture was then determined according to motility and PCR analysis. Experiments were performed three times, and data are given as means with error bars representing SEM. Analysis of the statistical significance of frequencies of strains against the uniform distribution was performed using Chi-squared test. b Growth competition of wild-type and aflagellated ΔflaAΔflaB strains under 100 and 300 rpm shaking conditions in contrast to static culture. Experiments were performed three times, and data are given as means with error bars representing SEM. Statistical significance of frequencies of strains against the uniform distribution was performed using binomial test on untransformed proportions
The observation that the ΔflaAΔflaB mutant strain appears to have higher fitness than the flagellated wild-type strain under shaking conditions was somewhat as expected. For confirmation, we performed head-to-head growth competition experiments with the ΔflaAΔflaB and the wild-type strains under 100 rpm and 300 rpm agitated conditions in contrast to static condition. When the culture grew to ~ 2 × 108 CFU/mL, samples were taken and their strain composition was determined (Fig. S3). As shown in Fig. 3b, the major portion of the population was made up of the aflagellated ΔflaAΔflaB mutant strain, 66.7% and 77.8% in 100-rpm and 300-rpm cultures, respectively. These results thus confirmed that the aflagellated strain has a significant advantage over the polarly flagellated wild-type when cultured under shaking liquid conditions and that the higher the rotation speed, the greater the advantage for the aflagellated ΔflaAΔflaB strain.
The Wild-Type Strain is Advantageous When Cultivated Under Static Conditions
As bacterial cells grow, they commonly exist in the form of assemblages, which are composed of both living cells and an adhesive matrix secreted by the cells [37]. A typical assemblage that S. oneidensis develops is known as pellicle, a type of biofilm formed at the air–liquid interface [36, 38, 39]. We therefore continued to investigate the roles of motility and the locomotive device per se related to pellicle formation. To this end, the pellicle formation of the five above-mentioned strains was assessed. For each strain, 20 µL cultures were grown to ~ 108 CFU/mL, inoculated into 2 mL liquid LB in 24-well plates and incubated under static conditions.
To quantify the differences of these strains in pellicle formation, we measured the density of planktonic cells and cells in pellicles. Consistent with the previous findings of the wild-type [38], the density of planktonic cells increased with time before the initiation of pellicle formation but remained relatively stable beyond this point (Fig. 4a). However, among mutants, ΔflaAΔflaB grew almost exclusively in the planktonic form. All other mutants retained the ability of pellicle formation to a certain extent, although the times of the initiation of the process differed significantly. ΔflhG, as a flagellated non-motile strain, was slower to form pellicles, and exhibited slower growth rates than did the other strains during the planktonic phase. This implied that motility facilities growth under static conditions.
Growth comparison of the five strains in the pellicle formation under static liquid condition. a Growth dynamics of the five strains under static liquid condition. The values of OD600 were measured in the total biomass including pellicle and planktonic portions (blue, combined) and planktonic only (black) at 0, 4, 8, 12, 16 and 20 h culture. The difference between the OD600 for planktonic and total biomass was taken as the pellicle biomass (orange). Experiments were performed three times, and data are given as means with error bars representing SEM. b Growth competition of the five strains in pellicle and planktonic portions under static liquid condition. The cultures composed of an even mixture of the five strains under test were grown statically for 20 h. Experiments were performed three times, and data are given as means with error bars representing SEM (Color figure online)
To assess the fitness of flagellar patterns upon pellicle formation, 50 µL cultures, consisting of these five strains mixed evenly, were inoculated into 2 mL liquid LB in 24-well plates and incubated under static conditions. Samples were collected in the pellicle and underneath of the medium (planktonic) at 20 h after the inoculation when the wild-type had begun to grow predominantly in the pellicle and the planktonic biomass accumulation had started to slow down. The proportion of each strain was determined using the same method as described above to quantify their fitness. Thirty-six colonies were obtained from each of the pellicle and planktonic population (72 in total) and were subjected to identification. Data are presented in Fig. 4b, with representative results shown in Fig. S4a (pellicle) and Fig. S4b, c (underneath, planktonic). From the planktonic population, 59%, 17%, and 22% were identified to be the wild-type, ΔflaAΔflaB, and ΔflhF, respectively. In contrast, strains with multiple flagella (ΔflhG and ΔflhFΔflhG) were not detected in the population, with only one exception. These data indicate that the impacts of motility on planktonic growth under static conditions are critical. In the pellicles, 89%, 1%, and 10% were identified to be the wild-type, ΔflaAΔflaB, and ΔflhF, respectively, whereas the strains with multiple flagella were absent (Fig. 4b). Our results indicate that the wild-type with a polar flagellum existing in assemblages is advantageous and this flagellation pattern has thus been fixed during evolution in nature.
Discussion
In nature, bacteria have developed diverse flagellation patterns as a highly conserved, species-specific feature. These seem to have resulted during the process of evolution as a response to their exposure to constantly changing environments. Despite the notion that each flagellated species has an evolutionarily fixed flagellation pattern which is favored under their normal living conditions, direct evidence is lacking. With S. oneidensis as the research model, multiple strains with distinct flagellation patterns have been previously reported [23,24,25, 27, 28]. In this study, we presented data to illustrate that the above notion largely holds.
Throughout the process of evolution, S. oneidensis seemed to have been fine-tuned to possess a set of flagellar genes for a monotrichous polar flagellum [23]. For the facilitation of motility, this wild-type flagellation pattern is overwhelmingly superior to all other possible flagellar arrangements found on S. oneidensis mutants, including aflagellated (ΔflaAΔflaB), monotrichous lateral (ΔflhF), lophotrichous-like (ΔflhG), and peritrichous-like (ΔflhFΔflhG). The differences of these strains in motility can be confidently attributed to the differing aspects of flagellation as it seems that only the location and number of flagella are affected by these mutations [27, 28, 30] with no other off-target effects apparent. When the wild-type is grown on swim plates together with other mutants (whose motility is significantly impaired), the competition assays reveal that the wild-type predominated in the central area of the droplet, where initial inoculation occurred, and was also the only strain to spread to the outer areas. While this observation further strengthens the understanding that strong motility, as provided by the wild-type flagellation, is critical for cells to move to uninhabited habitat under such poorly-mixed conditions, it is possible that the highly mobile wild-type cells quickly use up nutrients in the area into which they spread, thus limiting growth of other strains under the test by this means.
The monotrichous polar flagellum also confers a tremendous fitness gain in pellicles under static culturing conditions. In a medium with fully developed pellicles, oxygen is accessible only to cells in the pellicles whereas those underneath are oxygen-starved. As a result, fast growth can be only achieved at the air–liquid interface [36, 38]. As revealed previously [38], cells need effective locomotive ability to reach the air–liquid interface. The finding that approximately 89% of the cells in pellicles are of the wild-type strain indicates that the polar flagellum is the best means for moving away from the lower, low-oxygen environments to reach the favorable niche of the interface. The importance of motility in this case is also supported by the observation that ΔflhF makes up ~ 10% of the cells in the pellicles. In contrast, motility appears to be dispensable for cells living under the pellicles. This is probably due to the diminished difference in growth rates between the highly motile and non-motile strains because growth rates of all strains are substantially reduced as a consequence of oxygen limitation [38].
When cultivated in the agitated conditions, the aflagellated ΔflaAΔflaB mutant strain was found to have an advantage over the other four strains, including the wild-type. To validate this observation, the head-to-head comparison of the wild-type and ΔflaAΔflaB strains further revealed the enhancement and advantage of the aflagellated strain upon agitation. Clearly, environments become even upon agitation: nutrition and oxygen become well-distributed, and cell densities are consistent throughout the entire medium. In such situations, means of active locomotion no long confer a survival advantage for bacterial strains. As only the motility and the energy expenditure on flagellin synthesis were altered in the mutants, we can therefore confidently attribute the advantage of the aflagellated strain to its lower energy consumption. This result may also predict that flagellated bacterial strains may evolve towards aflagellation in agitated environments. More studies in the natural environment may be required to confirm this hypothesis. With shaking cultivation being the most-used culturing method for laboratory works with microorganisms, the complete loss of flagella or a declined ability to produce flagellar filaments, might also be predicted to occur during long periods of domestication under laboratory conditions. In the case of Bacillus subtilis, while the undomesticated wild strain 3610 swarms across agar surface, propelled by highly flagellated cells at the leading edge, strain 168 (a widely-used and long-domesticated laboratory strain) fails to swarm [40]. The failure to produce hyperflagellated cells in strain 168 could contribute to its swarming incapability. In nature, perhaps the energy spent on the flagellar rotation, rather than on biosynthesis, is the most substantial factor since the flagellar filament may be involved in other physiological processes that affect its growth as it also serves as a sensor for environmental cues such as surface and wetness [41]. This merits further investigation.
Although the lophotrichous-like ΔflhG strain and peritrichous-like ΔflhFΔflhG strain require additional energy and metabolic input for the biosynthesis of their multiple flagella, as compared with a monotrichous flagellum of the wild-type, they do not appear to exhibit corresponding negative aspects of growth or fitness when inter-strain competition is absent. On solid LB plates, all strains under testing could form colonies that were indistinguishable in size. In line with this, when independently measured, the growth of all strains under testing was comparable under agitated conditions. However, strains with multiple flagella were much less competitive when growing as planktonic cells under the pellicles. We speculate that additional energy and metabolic input for biosynthesis of multiple flagella may be responsible for their lowered fitness, at least partially. Given that growth under pellicles is supported by respiration of non-oxygen electron acceptors, which is extremely low in efficacy, any extra energy cost such as in the production of multiple flagella, could make a particular difference in growth fitness in such conditions.
In summary, the data presented here illustrate the intriguing and previously underappreciated impact of flagellation on ecophysiological fitness in bacteria. In nature, bacteria evolve the best strategies to survive and to proliferate in their respective niches. Apparently, multiple flagella, which are commonly found on bacteria associated with solid surfaces such as soil and intestinal tracts, do not provide fitness gain for S. oneidensis. Instead, S. oneidensis, which lives in water bodies and sediments, adopts monotrichous polar flagellation. This flagellation pattern confers a selective advantage because motility and energy cost are elegantly balanced. The findings of a recent study coincide with our conclusion [42]. In B. subtilis, the mutant with lower flagellar number (9 flagella) than the wild-type (26 flagella) was noted as more efficient in long-distance transport and spread faster. On the contrary, having more flagella (41 flagella) slows spreading, and thus become beneficial for the formation of biofilm. The flagellar number found on the wild-type cells is moderate, which is optimal for efficient searching and exploring. Finally, the data suggest that the monotrichous polar flagellation of S. oneidensis may evolve towards aflagellation in a constantly agitated environment such as vents, jet streams and laboratory domestication, due to lower energy cost and protein synthesis requirement that is no longer required for the production of useless flagella.
References
Armitage JP (1999) Bacterial tactic responses. Adv Microb Physiol 41:229–289
Johnson CN (2013) Fitness factors in Vibrios: a mini-review. Microb Ecol 65:826–851
Jarrell KF, McBride MJ (2008) The surprisingly diverse ways that prokaryotes move. Nat Rev Microbiol 6:466–476
Atsumi T, McCarter L, Imae Y (1992) Polar and lateral flagellar motors of marine Vibrio are driven by different ion-motive forces. Nature 355:182–184
Vaituzis Z, Doetsch RN (1969) Motility tracks: technique for quantitative study of bacterial movement. Appl Microbiol 17:584–588
Moens S, Vanderleyden J (1996) Functions of bacterial flagella. Crit Rev Microbiol 22:67–100
Kirov SM (2003) Bacteria that express lateral flagella enable dissection of the multifunctional roles of flagella in pathogenesis. FEMS Microbiol Lett 224:151–159
Fujii M, Shibata S, Aizawa S (2008) Polar, peritrichous, and lateral flagella belong to three distinguishable flagellar families. J Mol Biol 379:273–283
Wang F, Wang J, Jian H, Zhang B, Li S, Wang F et al (2008) Environmental adaptation: genomic analysis of the piezotolerant and psychrotolerant deep-sea iron reducing bacterium Shewanella piezotolerans WP3. PLoS ONE 3:e1937
Chevance FF, Hughes KT (2008) Coordinating assembly of a bacterial macromolecular machine. Nat Rev Microbiol 6:455–465
Schuhmacher JS, Thormann KM, Bange G (2015) How bacteria maintain location and number of flagella? FEMS Microbiol Rev 39:812–822
Guttenplan SB, Shaw S, Kearns DB (2013) The cell biology of peritrichous flagella in Bacillus subtilis. Mol Microbiol 87:211–229
Haya S, Tokumaru Y, Abe N, Kaneko J, Aizawa S (2011) Characterization of lateral flagella of Selenomonas ruminantium. Appl Environ Microbiol 77:2799–2802
Liu R, Ochman H (2007) Stepwise formation of the bacterial flagellar system. Proc Natl Acad Sci USA 104:7116–7121
Macnab RM (2003) How bacteria assemble flagella. Annu Rev Microbiol 57:77–100
Smith DR, Chapman MR (2010) Economical evolution: microbes reduce the synthetic cost of extracellular proteins. mBio 1:e00131–e00210
Berg HC (2003) The rotary motor of bacterial flagella. Annu Rev Biochem 72:19–54
Guttenplan SB, Kearns DB (2013) Regulation of flagellar motility during biofilm formation. FEMS Microbiol Rev 37:849–871
Fredrickson JK, Romine MF, Beliaev AS, Auchtung JM, Driscoll ME, Gardner TS et al (2008) Towards environmental systems biology of Shewanella. Nat Rev Microbiol 6:592–603
Bubendorfer S, Held S, Windel N, Paulick A, Klingl A, Thormann KM (2012) Specificity of motor components in the dual flagellar system of Shewanella putrefaciens CN-32. Mol Microbiol 83:335–350
Koerdt A, Paulick A, Mock M, Jost K, Thormann KM (2009) MotX and MotY are required for flagellar rotation in Shewanella oneidensis MR-1. J Bacteriol 191:5085–5093
Paulick A, Koerdt A, Lassak J, Huntley S, Wilms I, Narberhaus F et al (2009) Two different stator systems drive a single polar flagellum in Shewanella oneidensis MR-1. Mol Microbiol 71:836–850
Wu L, Wang J, Tang P, Chen H, Gao H (2011) Genetic and molecular characterization of flagellar assembly in Shewanella oneidensis. PLoS ONE 6:e21479
Shi M, Wu L, Xia Y, Chen H, Luo Q, Sun L et al (2013) Exoprotein production correlates with morphotype changes of nonmotile Shewanella oneidensis mutants. J Bacteriol 195:1463–1474
Sun L, Jin M, Ding W, Yuan J, Kelly J, Gao H (2013) Posttranslational modification of flagellin FlaB in Shewanella oneidensis. J Bacteriol 195:2550–2561
Sun L, Dong Y, Shi M, Jin M, Zhou Q, Luo Z et al (2014) Two residues predominantly dictate functional difference in motility between Shewanella oneidensis flagellins FlaA and FlaB. J Biol Chem 289:14547–14559
Shi M, Gao T, Ju L, Yao Y, Gao H (2014) Effects of FlrBC on flagellar biosynthesis of Shewanella oneidensis. Mol Microbiol 93:1269–1283
Gao T, Shi M, Ju L, Gao H (2015) Investigation into FlhFG reveals distinct features of FlhF in regulating flagellum polarity in Shewanella oneidensis. Mol Microbiol 98:571–585
Gao T, Meng Q, Gao H (2017) Thioesterase YbgC affects motility by modulating c-di-GMP levels in Shewanella oneidensis. Sci Rep 7:3932
Gao T, Shi M, Gao H (2018) Partially Reciprocal Replacement of FlrA and FlrC in Regulation of Shewanella oneidensis Flagellar Biosynthesis. J Bacteriol 200:e00796–e00817
Green JC, Kahramanoglou C, Rahman A, Pender AM, Charbonnel N, Fraser GM (2009) Recruitment of the earliest component of the bacterial flagellum to the old cell division pole by a membrane-associated signal recognition particle family GTP-binding protein. J Mol Biol 391:679–690
Kusumoto A, Nishioka N, Kojima S, Homma M (2009) Mutational analysis of the GTP-binding motif of FlhF which regulates the number and placement of the polar flagellum in Vibrio alginolyticus. J Biochem 146:643–650
Schuhmacher JS, Rossmann F, Dempwolff F, Knauer C, Altegoer F, Steinchen W et al (2015) MinD-like ATPase FlhG effects location and number of bacterial flagella during C-ring assembly. Proc Natl Acad Sci USA 112:3092–3097
Dasgupta N, Arora SK, Ramphal R (2000) fleN, a gene that regulates flagellar number in Pseudomonas aeruginosa. J Bacteriol 182:357–364
Correa NE, Peng F, Klose KE (2005) Roles of the regulatory proteins FlhF and FlhG in the Vibrio cholerae flagellar transcription hierarchy. J Bacteriol 187:6324–6332
Yuan J, Chen Y, Zhou G, Chen H, Gao H (2013) Investigation of roles of divalent cations in Shewanella oneidensis pellicle formation reveals unique impacts of insoluble iron. Biochim Biophys Acta 1830:5248–5257
Teschler JK, Zamorano-Sánchez D, Utada AS, Warner CJ, Wong GC, Linington RG et al (2015) Living in the matrix: assembly and control of Vibrio cholerae biofilms. Nat Rev Microbiol 13:255–268
Liang Y, Gao H, Chen J, Dong Y, Wu L, He Z et al (2010) Pellicle formation in Shewanella oneidensis. BMC Microbiol 10:291
Zhou G, Yuan J, Gao H (2015) Regulation of biofilm formation by BpfA, BpfD, and BpfG in Shewanella oneidensis. Front Microbiol 6:790
Kearns DB, Losick R (2003) Swarming motility in undomesticated Bacillus subtilis. Mol Microbiol 49:581–590
Wang Q, Suzuki A, Mariconda S, Porwollik S, Harshey RM (2005) Sensing wetness: a new role for the bacterial flagellum. EMBO J 24:2034–2042
Najafi J, Shaebani MR, John T, Altegoer F, Bange G, Wagner C (2018) Flagellar number governs bacterial spreading and transport efficiency. Sci Adv 4:eaar6425
Acknowledgements
This work was supported by grants from the National Natural Science Foundation Project (No. 81874355) and the Zhejiang Provincial Natural Science Foundation (No. LY18H280008). The authors would like to express sincere gratitude to professor Haichun Gao for the gift of the strains used in this study. The authors are also grateful to Mr. Chris Wood for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
RY and YC conceived and designed the experiments. RY performed the experiments and analyzed the data. RY and YC wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, RS., Chen, YT. Flagellation of Shewanella oneidensis Impacts Bacterial Fitness in Different Environments. Curr Microbiol 77, 1790–1799 (2020). https://doi.org/10.1007/s00284-020-01999-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-01999-0