Abstract
Expression and secretion of recombinant proteins in the endotoxin-free bacterium, Bacillus subtilis, has been thoroughly studied, but overexpression in the cytoplasm has been limited to only a few proteins. Here, we used the robust IPTG-inducible promoter, Pgrac212, to overexpress human rhinovirus 3C protease (HRV3C) in the cytoplasm of B. subtilis cells. A novel solubility tag, the N-terminal domain of the lysS gene of B. subtilis coding for a lysyl-tRNA synthetase was placed at the N terminus with a cleavage site for the endoprotease HRV3C, followed by His-HRV3C or His-GST-HRV3C. The recombinant protease was purified by using a Ni–NTA column. In this study, the His-HRV3C and His-GST-HRV3C proteases were overexpressed in the cytoplasm of B. subtilis at 11% and 16% of the total cellular proteins, respectively. The specific protease activities were 8065 U/mg for His-HRV3C and 3623 U/mg for His-GST-HRV3C. The purified enzymes were used to cleave two different substrates followed by purification of the two different protein targets, the green fluorescent protein and the beta-galactosidase. In conclusion, the combination of an inducible promoter Pgrac212 and a solubility tag allowed the overexpression of the HRV3C protease in the cytoplasm of B. subtilis. The resulting fusion protein was purified using a nickel column and was active in cleaving target proteins to remove the fusion tags. This study offers an effective method for producing recombinant proteins in the cytoplasm of endotoxin-free bacteria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacillus subtilis is currently the best-studied Gram-positive bacterium in the scientific world. B. subtilis strain 168 and its derivatives are nonpathogenic and free of exo- and endotoxins. Its ease of genetic manipulation and genetically well-characterized expression systems enable this species to be used as an ideal workhorse for industrial and pharmaceutical purposes including the production of recombinant microbial enzymes and chemicals [1, 2]. Owing to its FDA status as GRAS (generally regarded as safe), its capacity for protein secretion [3,4,5], the absence of significant codon bias [6], the amenability for genetic engineering and abundant toolboxes [7], and extensive information on its fermentation processes [8, 9], B. subtilis has been chosen as an alternative host for overexpression and secretion of heterologous proteins [9,10,11].
In contrast to B. subtilis, the widely used Gram-negative bacterium Escherichia coli produces recombinant proteins mainly in the cytoplasm. Since E. coli contains the potent immunostimulatory endotoxin lipopolysaccharide, scientists sought to use an endotoxin-free platform for the synthesis of desired products [12]. Similar to other expression hosts, in certain cases, E. coli produces little or no recombinant protein, often because of inhibitory effects caused by the heterologous proteins [13]. Different states of recombinant enzymes can also confer low or high activities of the same protein expressed in each cell [14]. Therefore, an alternative intracellular expression system such as that offered by the endotoxin-free and GRAS B. subtilis is needed to produce proteins in the cytoplasm. Only a few proteins have been expressed at levels of more than 10% of the total cellular protein: GFP [15,16,17,18], β-galactosidase [17, 19,20,21], GUS [15], B. subtilis HtpG [22], B. subtilis PbpB4 [22], the B subunit of E. coli heat-labile toxin (LTB) [23], listeriolysin O [24], HIV P24 antigen [25], 2,3-dioxygenase (C23O) (about 25% of total cellular protein) [26], CAT protein from Staphylococcus aureus [27], and two membrane proteins from E. coli, OmpA and OmpF [28]. Recent studies indicate that pMTBs72 backbones [29] such as the pHCMC and pHT01 plasmids use the theta replicating mechanism conferring structural stability of the plasmids. In contrast, most of the B. subtilis overexpression studies published before used the rolling-circle mechanism resulting in structural instability [30, 31]. In this study, a structurally stable plasmid carrying the Pgrac212 promoter was used for overexpression of recombinant proteins in B. subtilis. Pgrac212 is a strong promoter belonging to the Pgrac family [32], but differing from Pgrac01. It contains the mRNA-controllable stabilizing element (CoSE) conferring an extremely stable mRNA [33] that allows beta-galactosidase (BgaB) expression equal to that reported for Pgrac100 [34].
Solubility enhancement tags and affinity purification sequences are essential tools that keep recombinant proteins soluble and simplify purification [35, 36]. However, some tags can alter the structural or functional integrity of a recombinant protein and these have to be removed after purification by specific endoproteases such as thrombin, enterokinase, the tobacco etch virus (TEV) protease or the human rhinovirus 3C protease (HRV3C) [37,38,39]. HRV3C cleaves between glutamine and glycine residues of the canonical site LEVLFQ/GF [35, 38, 40]. The HRV3C endoprotease has up to 10 × higher activity at 4 °C than 37 °C [41] and does not cleave at undesired cryptic sites as reported for other proteases [36].
Overexpression of useful proteins in the cytoplasm of the endotoxin-free bacterium B. subtilis using structurally stable plasmids in combination with the robust IPTG-inducible promoter Pgrac212 has not been reported so far. In this work, we overexpressed the reporter, HRV3C in B. subtilis, purified the protease, and tested its ability to cleave target proteins with subsequent purification.
Materials and Methods
Materials
The strains, plasmids, proteins, and primers used in this study are listed in Table 1. The E. coli strain OmniMAX (Invitrogen) was used for all cloning experiments, and B. subtilis strain 1012 (MoBiTec) was used to express the two proteases fused with six histidines, and in one case with 6×His plus glutathione S-transferase: His-HRV3C and His-GST-HRV3C. Cells were routinely grown in Luria broth (LB) at 37 °C under aeration and shaking at 200 rpm, with additional conditions as indicated. E. coli was grown in the presence of 100 µg/mL ampicillin and B. subtilis with 10 µg/mL chloramphenicol.
Construction of Recombinant Plasmids
The bgaB gene of pHT212 was replaced by the N-terminal part of the B. subtilis lysS gene (lysSN), fused to the HRV3C cleavage site (CS), a 6×His tag and a multi-cloning site (LysSN-HRV3C/CS-His-MCS). Therefore, plasmid pHT399 contained a DNA sequence encoding LysSN-HRV3C/CS-His-MCS with a Strep-tag under control of the Pgrac212 promoter. The BsLysSN enhances the solubility and folding of target proteins [42]. The protein sequence of LysSN-HRV3C/CS-His-MCS with the Strep-tag is shown in Supplement 1. DNA sequences coding for HRV3C (631 bp) and GST-HRV3C (1322 bp) were obtained by PCR reactions with appropriate primer pairs (Table 1) using pGEX 4T-HRV3C as a template (Genbank ID, MN103550). These amplicons were inserted into pHT399 (derived from pHT212 [33]) by the InFusion method to generate pHT399B (His-HRV3C) and pHT399C (His-GST-HRV3C), respectively, and both plasmids were transformed into B. subtilis 1012.
The resulting plasmids were confirmed by restriction enzyme analysis and DNA sequencing as containing the fusion genes encoding BsLysSN-HRV3C/CS-His-HRV3C (pHT399B, for His-HRV3C) and BsLysSN-HRV3C/CS-His-GST-HRV3C-Strep-tag (pHT399C, for His-GST-HRV3C). The protein sequence of BsLysSN-HRV3C/CS-His-HRV3C and His-HRV3C are shown in Supplement 2, while BsLysSN-HRV3C/CS-His-GST-HRV3C-Strep-tag and His-GST-HRV3C with the Strep-tag are shown in Supplement 3. Map of plasmid pHT399B and expression cassette of pHT399B and pHT399C are shown in Supplement 4.
Expression of the Recombinant Proteins
The plasmids obtained from E. coli OmniMAX were inserted into B. subtilis 1012 by natural transformation [20]. To identify an appropriate temperature for the overexpression of the recombinant proteins, B. subtilis 1012 cells harboring pHT399B or pHT399C were grown at 23 °C, 30 °C, and 37 °C till the optical density at 600 nm (OD600nm) reached 0.8. Then, the synthesis of proteins was induced by the addition of 0.5 mM IPTG. Cells were harvested by centrifugation at 6000×g for 10 min. The number of B. subtilis cells was equivalent to those present in 1 ml of culture with an OD600nm of 6. The cells were suspended in 500 µl of lysis buffer (20 mM Tris–HCl, pH 7.2 and 15% (w/v) sucrose) containing 200 µg/ml lysozyme and incubated at 37 °C for 10 min. Then, the samples were sonicated and centrifuged at 2000×g for 5 min. The supernatants representing total proteins were centrifuged at 13,000×g for 5 min, and these supernatants were collected as soluble proteins. The pellets were resuspended at the original volume and represented the insoluble proteins. Each protein sample was analyzed by SDS-PAGE to determine the temperature at which the amount of soluble recombinant protein was highest. The optimal IPTG concentration (0, 0.01, 0.05, 0.1, 0.5, 1 mM) and induction time (0, 2, 4, 6, 8, 10, and 12 h) were also determined.
Purification of Recombinant Proteins by His-Trap (Ni–NTA) Column
B. subtilis 1012 cells carrying either pHT399B or pHT399C were cultured in one liter of LB medium at 37 °C and when the OD600 reached 0.8, IPTG was added to 1 mM, the temperature was reduced to 23 °C, and the two cultures were further incubated with shaking for 12 h to allow expression of the HRV3C fusion proteins. Cells were harvested by centrifugation at 6000×g for 10 min. Next, the cells were resuspended in binding buffer (30 mM Tris–HCl, pH 8.0; 500 mM NaCl; 5% glycerol) containing lysozyme (20 µg/ml) and incubated at 37 °C for 10 min. DNase I was added to a final concentration of 20 µg/ml and PMSF at 0.5 mM before sonication, and soluble proteins were obtained in the supernatants after centrifugation at 19,000×g at 4 °C for 30 min. The protein solutions were clarified by membrane filtration (pore size 0.22 μm). The clarified samples were pumped through 5 ml His-trap columns at a flow rate of 2 ml/min and recombinant proteins fused to the His-tag bound to the Ni–NTA in the columns. All other proteins without a His-tag were removed from the columns by washing with 30 ml of binding buffer containing 5 mM imidazole. The recombinant proteins were eluted from the columns with buffer containing 10, 20, 40, 60, 80, 100, 120, 160, 250, or 500 mM imidazole. 2-ml fractions were collected from each elution and the fourth fraction containing the eluted proteins was run on SDS-PAGE.
Determination of HRV3C Protease Activity and Purification of Cleaved Proteins
The activity of the eluted proteases (initial concentration, 14 µM) was tested on the substrates, BsLysSN-His-HRV3C/CS-GFP and BsLysSN-His-HRV3C/CS-BgaB. These substrate proteins were first dialyzed against cleavage buffer (50 mM Tris–HCl, 150 mM NaCl, 1 mM EDTA, 1 mM DTT, pH 7.0 at 25 °C) and then 100 µg of each substrate was incubated with 100 µl of protease at dilutions of 5- to 1000-fold for 16 h at 4 °C. The SDS-PAGE and AlphaEaseFC software were used to determine the percentage of intact vs. cleaved substrates, and the activity of HRV3C proteases was calculated. After that, the samples were applied to a His-trap column to confirm the loss of the His-tag.
SDS-PAGE
Bacillus subtilis 1012 carrying either the plasmid pHT399B or pHT399C was grown in LB in the presence of chloramphenicol (Fig. 1). Cells were sedimented by centrifugation and resuspended in lysis buffer. After lysis by sonication and clarification by centrifugation at 2000×g for 5 min, supernatants were sampled for measuring total protein(T) and the remainder was subjected to a second centrifugation step at 13,000 g for 10 min. Further protein samples were taken from the supernatant (S) and from the pellets (P) after resuspension. Aliquots of 100 µl of each protein samples were mixed with 25 µl of 5 × SDS-PAGE sample buffer, heated at 95 °C for 5 min, and then centrifuged at 13,000×g for 5 min. The supernatants were loaded onto SDS-PAGE gels, and electrophoresis was carried out at 25 mA.
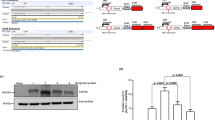
Influence of temperature and IPTG concentration on the production of His-HRV3C and His-GST-HRV3C. B. subtilis containing the plasmids pHT399B (His-HRV3C) or pHT399C (His-GST-HRV3C) were incubated in LB medium at different temperatures (a, b), or induced with different IPTG concentrations (c, d) for up to 12 h (e, f). –, empty vector; T, the total amount of proteins; P, proteins present in the pellets and S, the supernatant after centrifugation. The locations of the recombinant proteins are indicated by the black dot
Results
Construction of pHT399B (His-HRV3C) and pHT399C (His-GST-HRV3C)
Two different expression vectors constructed contain the HRV3C endoprotease gene and additional sequences. Both vectors were based on pHT212, an E. coli–B. subtilis shuttle vector containing the beta-galactosidase (bgaB) gene fused to the IPTG-inducible Pgrac212 promoter [33] derived from the Pgrac promoter [22]. The only difference between these two promoters is that Pgrac212 carries the mRNA stabilizing element CoSE, which results in higher recombinant protein production. It has been shown that RNA molecules can also function as chaperones and are extremely effective in helping the folding of a variety of proteins [42, 43]. In a recent publication, a small tRNA-binding N-terminal domain of lysyl-tRNA synthetase of E. coli has been shown to assist the de novo folding and trimeric assembly of the RID-HA1 hemagglutinin [44]. In the present publication, we evaluated the expression of the recombinant proteins His-HRV3C and His-GST-HRV3C fused to the N-terminal domain of the lysS gene of B. subtilis coding for a lysyl-tRNA synthetase (BsLysSN) to facilitate protein expression.
Expression Levels of His-HRV3C and His-GST-HRV3C Under Different Growth and Induction Conditions
Bacillus subtilis 1012 carrying pHT399B or pTH399C was incubated at three different temperatures, and the levels of recombinant proteins produced under these conditions were quantified by SDS-PAGE. In the case of pHT399B (His-HRV3C), all the recombinant protein was present in the soluble fraction if growth was done at 23 °C or 30 °C, while at 37 °C, about 50% of the protein appeared in the insoluble fraction (Fig. 1a). In the case of pHT399C (His-GST-HRV3C), the recombinant protein remained soluble at all three temperatures (Fig. 1b). To determine the optimal IPTG concentration for inducing protein expression, bacteria carrying pHT399B were grown at 30 °C and those containing pHT399C at 37 °C in the presence of increasing concentrations of IPTG (Fig. 1c, d). A concentration of 1 mM IPTG resulted in the highest expression levels of the HRV3C proteases, and this concentration was chosen for inducing expression in subsequent experiments. The B. subtilis strain carrying pHT399B was grown at 30 °C and that containing pHT399C at 37 °C with 1 mM IPTG and samples were taken every two hours up to 12 h. The amount of His-HRV3C protein expressed from pHT399B was almost identical between 4 and 12 h (11% of total protein), while the His-GST-HRV3C encoded by pHT399C was highest at 12 h (16% of total protein) (Fig. 1e, f).
Purification of His-HRV3C and His-GST-HRV3C
The soluble isolates of the recombinant proteins were purified on a nickel column which allows binding of proteins with a histidine tag. Protein isolates of His-HRV3C showed a band at 22 kDa before loading on the column (BC) which became even more prominent after binding and elution from the column (AC) (Fig. 2a). The His-HRV3C protein was recovered at an imidazole concentration of 80–160 mM. When the same experiment was carried out with bacteria producing the His-GST-HRV3C protein, this protein also bound to the Ni–NTA column and was eluted after addition of 60–160 mM imidazole (Fig. 2b).
Purification of HRV3C using Ni–NTA columns. BC, protein isolates before loading on the Ni–NTA column; AC, flow-through samples collected after protease binding to the Ni–NTA column. The fusion proteins binding to the Ni–NTA column were eluted using a buffer with the indicated concentrations of imidazole (10 to 500 mM). The protein samples were analyzed by SDS-PAGE
The Protease Activity of His-HRV3C and His-GST-HRV3C
The activity of the purified proteases was determined with two substrates, BsLysSN-His-HRV3C/CS-GFP and BsLysSN-His-HRV3C/CS-BgaB. When BsLysSN-His-HRV3C/CS-GFP was used for cleavage, the lowest concentrations of His-HRV3C and His-GST-HRV3C corresponding to 1 unit of HRV3C protease able to cleave > 95% of 100 μg of target protein after 16 h at 4 °C were similar at a 250-fold dilution (Fig. 3a, b). This means that the fusion tag (6×His-GST) did not alter the HRV3C protease activity compared to 6×His alone. The specific protease activity of His-HRV3C was measured at 8065 U/mg and that of His-GST-HRV3C at 3623 U/mg. When the BsLysSN-His-HRV3C/CS-BgaB substrate was used for cleavage, the two proteases cleaved 95% at fivefold dilution (Fig. 3c, d). With the substrate protein BsLysSN-His-HRV3C/CS-BgaB, the specific protease activity of His-HRV3C was 161 U/mg and of His-GST-HRV3C was 72 U/mg. The reason for this difference could be that the folding of this substrate led to partial sequestering of the HRV3C cleavage site from the protease.
Analysis of protease activities of His-HRV3C and His-GST-HRV3C using different substrates. Proteases (P) His-HRV3C (a, c) and His-GST-HRV3C (b, d) were tested by digesting 100 µg aliquots of the two substrates (Sb): BsLysSN-His6-HRV3C/CS-GFP (a, b) and BsLysSN-His-HRV3C/CS-BgaB (c, d). The stocks of protease at an initial concentration of 14 µM diluted from 5 to 1000 times
Application of the HRV3C Proteases for the Purification of GFP and BgaB
The two purified HRV3C endoproteases, His-HRV3C and His-GST-HRV3C, were used to cleave the substrate proteins BsLysSN-GFP (BsLysSN-HisTag-HRV3C/CS-GFP) and BsLysSN-BgaB (BsLysSN-HisTag-HRV3C/CS-BgaB). The BsLysSN-His6-HRV3C/CS-GFP (Fig. 4a, b) target was cleaved to more than 95% by both proteases, while the BsLysSN-His6-HRV3C/CS-BgaB (Fig. 4c, d) was cleaved only to about 95%. The recombinant proteins GFP and BgaB in the cleaved reactions were isolated from the flow-through after loading the mixtures consisting of the purification tag and the recombinant proteins on the His-trap column. The target proteins which had been cut by the HRV3C protease lost their His-tag so they could not bind to the Ni–NTA column and were eluted in the flow-through solution. All the proteins having the 6×His affinity tag were retained on the Ni–NTA column. Finally, pure GFP (Fig. 4a, b, lands F) and BgaB (Fig. 4c, d, lands F) could be collected in the flow-through. These results proved that the purified HRV3C proteases could be applied to release GFP and BgaB from fusion forms containing the HRV3C cleavage site.
Testing of the proteases for purification of GFP and BgaB. The substrates (Sb) BsLysSN-6×His-HRV3C/CS-GFP (a, b) and BsLysSN-6×His-HRV3C/CS-BgaB (c, d) were cleaved by His-HRV3C (a, c) or His-GST-HRV3C (b, d) for 16 h at 4 °C. The mixture after cleavage (C) was loaded on a Ni–NTA column. The flow-through (F) contained the target proteins GFP or BgaB. The dots indicate the location of the isolated proteins
Discussion
This study demonstrates the use of a structurally stable plasmid carrying the robust IPTG-inducible promoter, Pgrac212 [33], to produce recombinant proteins in the cytoplasm of B. subtilis. The human rhinovirus 3C protease (HRV3C) [39] was either fused to a His-tag or a GST-tag, and both the recombinant proteases were overexpressed at 11% and 16%, of the total cellular proteins, respectively, and purified using a nickel column. The specific protease activities were measured to be 8065 U/mg for His-HRV3C and 3623 U/mg for His-GST-HRV3C.
In recent years, HRV3C has been used to develop new purification methods which are more simplified and faster than the traditional approach. For example, in 2014, a new system was developed to produce native proteins in E. coli, allowing for the removal of the fusion tag via intracellular self-cleavage by this protease [45]. In 2019, the simplified method to remove fusion tags named Cell Lysate Purification system based on HRV3C protease (CLP3C) was developed, in which strains expressing HRV3C protease and the other expressing substrates were mixed before (co-fermentation method) or after (post-fermentation method) inducing with IPTG, followed by cell disruption and incubation at 4 °C, overnight for cleavage [46]. However, these methods were only based on naturally lipopolysaccharide (LPS) producing host E. coli, which need the additional steps to remove LPS. In contrast, the HRV3C expressed in the endotoxin-free bacteria B. subtilis could be easily applied for those new purification methods to produce other recombinant proteins in B. subtilis using a similar approach as described for E. coli.
Intracellular expression of recombinant proteins in B. subtilis could be used to produce antigens as an oral vaccine delivery vector. It has been demonstrated that the heat-labile enterotoxin, B subunit (LTB), expressed in the cytoplasm of B. subtilis at deficient levels that could not be seen on SDS-PAGE. The purified protein was able to induce the immune response and confer partial protection to mice using lethal challenges with purified LTB [47]. The high expression system for B. subtilis as described here could be useful to develop B. subtilis as vaccine vehicles.
A large number of strong inducible and constitutive promoters have been identified and used for recombinant protein production, and there are still many attempts to discover novel promoters [48,49,50]. To improve the stability of the recombinant protein and allow simple purification, stabilizing peptides and purification tags can be added [9]. We used the IPTG-inducible Pgrac212 promoter and added three different peptide domains: (i) The small tRNA-binding N-terminal domain of the E. coli lysyl-tRNA synthetase to assist the de novo folding of the recombinant protein [44], (ii) a His-tag or His- and GST-tag to allow a one-step purification of the recombinant proteins, and (iii) the HRV3C sequence recognized by HRV3C protease [35] thereby allowing to cleave between these three tags to release the recombinant protein. As recombinant protein, we used first the HRV3C protease and analyzed the influence of the growth temperature and the IPTG concentration on the production of the protease. Based on these data, we fused the coding sequences of GFP and BgaB to the fusion tags in our vectors to further verify the application of this approach in B. subtilis. We could express GFP and BgaB fusions in B. subtilis (unpublished data) and used the purified HRV3C enzymes to cleave two substrate proteins followed by isolation of the two protein targets, GFP and BgaB. In summary, our data nicely prove the successful application of our vectors for the intracellular production of recombinant proteins in B. subtilis.
As the next step, our vector systems carrying different recombinant genes could be analyzed in the genome-reduced B. subtilis 168 cells. Several genomes reduced B. subtilis strains have been constructed so far. The first strain constructed lost 332 nonessential genes [51]; another group deleted 0.99 Mbp [52], and the smallest chromosome constructed so far lost 1.51 Mbp [53, 54]. All these strains allowed an increase production of recombinant proteins as compared with the wild-type strain 168.
References
Westers L, Westers H, Quax WJ (2004) Bacillus subtilis as cell factory for pharmaceutical proteins: a biotechnological approach to optimize the host organism. Biochim Biophys Acta 1694:299–310. https://doi.org/10.1016/j.bbamcr.2004.02.011
Zweers JC, Barák I, Becher D et al (2008) Towards the development of Bacillus subtilis as a cell factory for membrane proteins and protein complexes. Microb Cell Fact 7:10. https://doi.org/10.1186/1475-2859-7-10
Harwood CR, Cranenburgh R (2008) Bacillus protein secretion: an unfolding story. Trends Microbiol 16:73–79. https://doi.org/10.1016/j.tim.2007.12.001
Pohl S, Harwood CR (2010) Heterologous protein secretion by Bacillus species from the cradle to the grave. Adv Appl Microbiol 73:1–25. https://doi.org/10.1016/S0065-2164(10)73001-X
Wong SL (1995) Advances in the use of Bacillus subtilis for the expression and secretion of heterologous proteins. Curr Opin Biotechnol 6:517–522
Shields DC, Sharp PM (1987) Synonymous codon usage in Bacillus subtilis reflects both translational selection and mutational biases. Nucleic Acids Res 15:8023–8040
Kang Z, Yang S, Du G, Chen J (2014) Molecular engineering of secretory machinery components for high-level secretion of proteins in Bacillus species. J Ind Microbiol Biotechnol 41:1599–1607. https://doi.org/10.1007/s10295-014-1506-4
Schallmey M, Singh A, Ward OP (2004) Developments in the use of Bacillus species for industrial production. Can J Microbiol 50:1–17. https://doi.org/10.1139/w03-076
Schumann W (2007) Production of recombinant proteins in Bacillus subtilis. Adv Appl Microbiol 62:137–189. https://doi.org/10.1016/S0065-2164(07)62006-1
van Dijl JM, Hecker M (2013) Bacillus subtilis: from soil bacterium to super-secreting cell factory. Microb Cell Fact 12:3. https://doi.org/10.1186/1475-2859-12-3
Heinrich J, Drewniok C, Neugebauer E et al (2019) The YoaW signal peptide directs efficient secretion of different heterologous proteins fused to a StrepII-SUMO tag in Bacillus subtilis. Microb Cell Fact 18:31. https://doi.org/10.1186/s12934-019-1078-0
Taguchi S, Ooi T, Mizuno K, Matsusaki H (2015) Advances and needs for endotoxin-free production strains. Appl Microbiol Biotechnol 99:9349–9360. https://doi.org/10.1007/s00253-015-6947-9
Rosano GL, Ceccarelli EA (2014) Recombinant protein expression in Escherichia coli: advances and challenges. Front Microbiol 5:172. https://doi.org/10.3389/fmicb.2014.00172
Zhao Y, He W, Liu W-F et al (2012) Two distinct states of Escherichia coli cells that overexpress recombinant heterogeneous β-galactosidase. J Biol Chem 287:9259–9268. https://doi.org/10.1074/jbc.M111.327668
Cui W, Han L, Cheng J et al (2016) Engineering an inducible gene expression system for Bacillus subtilis from a strong constitutive promoter and a theophylline-activated synthetic riboswitch. Microb Cell Fact 15:199. https://doi.org/10.1186/s12934-016-0599-z
Nguyen NH, Phan TTP, Tran TL, Nguyen HD (2014) Investigating the expression of GFP fused with HIS-TAG at N- or C-terminus using plasmid pHT253 and pHT254 in Bacillus subtilis. Sci Technol Dev J 17(4), 5–11. 10.32508/stdj.v17i4.1550.
Phan TTP, Tran LT, Schumann W, Nguyen HD (2015) Development of Pgrac100-based expression vectors allowing high protein production levels in Bacillus subtilis and relatively low basal expression in Escherichia coli. Microb Cell Fact 14:72. https://doi.org/10.1186/s12934-015-0255-z
Wenzel M, Müller A, Siemann-Herzberg M, Altenbuchner J (2011) Self-inducible Bacillus subtilis expression system for reliable and inexpensive protein production by high-cell-density fermentation. Appl Environ Microbiol 77:6419–6425. https://doi.org/10.1128/AEM.05219-11
Phan TTP, Nguyen HD, Schumann W (2012) Development of a strong intracellular expression system for Bacillus subtilis by optimizing promoter elements. J Biotechnol 157:167–172. https://doi.org/10.1016/j.jbiotec.2011.10.006
Phan T, Huynh P, Truong T, Nguyen H (2017) A generic protocol for intracellular expression of recombinant proteins in Bacillus subtilis. Methods Mol Biol 1586:325–334. https://doi.org/10.1007/978-1-4939-6887-9_21
Tran DTM, Phan TTP, Huynh TK et al (2017) Development of inducer-free expression plasmids based on IPTG-inducible promoters for Bacillus subtilis. Microb Cell Fact 16:130. https://doi.org/10.1186/s12934-017-0747-0
Phan TTP, Nguyen HD, Schumann W (2006) Novel plasmid-based expression vectors for intra- and extracellular production of recombinant proteins in Bacillus subtilis. Protein Expr Purif 46:189–195. https://doi.org/10.1016/j.pep.2005.07.005
Phan TTP, Nguyen ALT, Nguyen HD (2013) Cloning and expression of LTB in Escherichia coli and Bacillus subtilis. Nat Sci 16(1), 13–22. https://doi.org/10.32508/stdj.v16i1.1392
Ngo HK, Nguyen ALT, Huynh PTK, et al. (2016) Expression and purification of listeriolysin O from Listeria monocytogenes harbouring E247M and D320K mutations in Bacillus subtilis. Sci Technol Dev J 19:20–31. https://doi.org/10.32508/stdj.v19i3.470
Truong TTT, Phan TTP, Nguyen HD (2017) Purification of P24 protein expressed in Bacillus subtilis and evaluation of its immunogenicity in mice. Sci Technol Dev J 1:69–79. https://doi.org/10.32508/stdjns.v1iT1.436
Zukowski MM, Miller L (1986) Hyperproduction of an intracellular heterologous protein in a sacUh mutant of Bacillus subtilis. Gene 46:247–255
Peschke U, Beuck V, Bujard H et al (1985) Efficient utilization of Escherichia coli transcriptional signals in Bacillus subtilis. J Mol Biol 186:547–555
Puohiniemi R, Butcher S, Tarkka E, Sarvas M (1991) High level production of Escherichia coli outer membrane proteins OmpA and OmpF intracellularly in Bacillus subtilis. FEMS Microbiol Lett 67:29–33. https://doi.org/10.1111/j.1574-6968.1991.tb04383.x
Titok MA, Chapuis J, Selezneva YV et al (2003) Bacillus subtilis soil isolates: plasmid replicon analysis and construction of a new theta-replicating vector. Plasmid 49:53–62
Nguyen HD, Nguyen QA, Ferreira RC et al (2005) Construction of plasmid-based expression vectors for Bacillus subtilis exhibiting full structural stability. Plasmid 54:241–248. https://doi.org/10.1016/j.plasmid.2005.05.001
Nguyen HD, Phan TTP, Schumann W (2007) Expression vectors for the rapid purification of recombinant proteins in Bacillus subtilis. Curr Microbiol 55:89–93. https://doi.org/10.1007/s00284-006-0419-5
Phan TTP, Nguyen HD, Schumann W (2010) Establishment of a simple and rapid method to screen for strong promoters in Bacillus subtilis. Protein Expr Purif 71:174–178. https://doi.org/10.1016/j.pep.2009.11.010
Phan TTP, Nguyen HD, Schumann W (2013) Construction of a 5′-controllable stabilizing element (CoSE) for over-production of heterologous proteins at high levels in Bacillus subtilis. J Biotechnol 168:32–39. https://doi.org/10.1016/j.jbiotec.2013.07.031
Trang PTP (2007) Construction and analysis of novel controllable expression vectors for Bacillus subtilis. Doctoral thesis, University of Bayreuth, Bayreuth
Young CL, Britton ZT, Robinson AS (2012) Recombinant protein expression and purification: a comprehensive review of affinity tags and microbial applications. Biotechnol J 7:620–634. https://doi.org/10.1002/biot.201100155
Waugh DS (2011) An overview of enzymatic reagents for the removal of affinity tags. Protein Expr Purif 80:283–293. https://doi.org/10.1016/j.pep.2011.08.005
Walker PA, Leong LE, Ng PW et al (1994) Efficient and rapid affinity purification of proteins using recombinant fusion proteases. Biotechnology (NY) 12:601–605
Vergis JM, Wiener MC (2011) The variable detergent sensitivity of proteases that are utilized for recombinant protein affinity tag removal. Protein Expr Purif 78:139–142. https://doi.org/10.1016/j.pep.2011.04.011
Antoniou G, Papakyriacou I, Papaneophytou C (2017) Optimization of soluble expression and purification of recombinant human rhinovirus type-14 3C protease using statistically designed experiments: isolation and characterization of the enzyme. Mol Biotechnol 59:407–424. https://doi.org/10.1007/s12033-017-0032-9
Cordingley MG, Callahan PL, Sardana VV et al (1990) Substrate requirements of human rhinovirus 3C protease for peptide cleavage in vitro. J Biol Chem 265:9062–9065
Raran-Kurussi S, Tözsér J, Cherry S et al (2013) Differential temperature dependence of tobacco etch virus and rhinovirus 3C proteases. Anal Biochem 436:142–144. https://doi.org/10.1016/j.ab.2013.01.031
Choi SI, Han KS, Kim CW et al (2008) Protein solubility and folding enhancement by interaction with RNA. PLoS ONE 3:e2677. https://doi.org/10.1371/journal.pone.0002677
Horowitz S, Bardwell JCA (2016) RNAs as chaperones. RNA Biol 13:1228–1231. https://doi.org/10.1080/15476286.2016.1247147
Yang SW, Jang YH, Kwon SB et al (2018) Harnessing an RNA-mediated chaperone for the assembly of influenza hemagglutinin in an immunologically relevant conformation. FASEB J 32:2658–2675. https://doi.org/10.1096/fj.201700747RR
Feng Y, Xu Q, Yang T et al (2014) A novel self-cleavage system for production of soluble recombinant protein in Escherichia coli. Protein Expr Purif 99:64–69. https://doi.org/10.1016/j.pep.2014.04.001
Xu H, Wang Q, Zhang Z et al (2019) A simplified method to remove fusion tags from a xylanase of Bacillus sp. HBP8 with HRV 3C protease. Enzyme Microb Technol 123:15–20. https://doi.org/10.1016/j.enzmictec.2019.01.004
Paccez JD, Nguyen HD, Luiz WB et al (2007) Evaluation of different promoter sequences and antigen sorting signals on the immunogenicity of Bacillus subtilis vaccine vehicles. Vaccine 25:4671–4680. https://doi.org/10.1016/j.vaccine.2007.04.021
Cui W, Han L, Suo F et al (2018) Exploitation of Bacillus subtilis as a robust workhorse for production of heterologous proteins and beyond. World J Microbiol Biotechnol 34:145. https://doi.org/10.1007/s11274-018-2531-7
Öztürk S, Ergün BG, Çalık P (2017) Double promoter expression systems for recombinant protein production by industrial microorganisms. Appl Microbiol Biotechnol 101:7459–7475. https://doi.org/10.1007/s00253-017-8487-y
Song Y, Nikoloff JM, Zhang D (2015) Improving protein production on the level of regulation of both expression and secretion pathways in Bacillus subtilis. J Microbiol Biotechnol 25:963–977. https://doi.org/10.4014/jmb.1501.01028
Westers H, Dorenbos R, van Dijl JM et al (2003) Genome engineering reveals large dispensable regions in Bacillus subtilis. Mol Biol Evol 20:2076–2090. https://doi.org/10.1093/molbev/msg219
Ara K, Ozaki K, Nakamura K et al (2007) Bacillus minimum genome factory: effective utilization of microbial genome information. Biotechnol Appl Biochem 46:169–178. https://doi.org/10.1042/BA20060111
Aguilar Suárez R, Stülke J, van Dijl JM (2019) Less is more: toward a genome-reduced Bacillus cell factory for “difficult proteins”. ACS Synth Biol 8:99–108. https://doi.org/10.1021/acssynbio.8b00342
Saito H, Shibata T, Ando T (1979) Mapping of genes determining nonpermissiveness and host-specific restriction to bacteriophages in Bacillus subtilis Marburg. Mol Gen Genet 170:117–122
Funding
This research was funded by the Department of Science and Technology of Ho Chi Minh City, Vietnam under Grant Number 1022/QĐ-SKHCN.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Research Involving Human and Animal Rights
This study did not use animals or samples from human for experiments.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Le, V.D., Phan, T.T.P., Nguyen, T.M. et al. Using the IPTG-Inducible Pgrac212 Promoter for Overexpression of Human Rhinovirus 3C Protease Fusions in the Cytoplasm of Bacillus subtilis Cells. Curr Microbiol 76, 1477–1486 (2019). https://doi.org/10.1007/s00284-019-01783-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01783-9