Abstract
Myxococcus xanthus generates diadenosine tetraphosphates (Ap4A) and diadenosine pentaphosphates (Ap5A) under various stress conditions. M. xanthus lysyl-tRNA synthetase (LysS) efficiently synthesizes Ap4A from ATP, Ap5A from ATP and adenosine tetraphosphate (Ap4), and Ap4 from ATP and triphosphate. To identify other M. xanthus enzymes that can catalyze Ap4A and Ap4 synthesis, 15 M. xanthus aminoacyl-tRNA synthetases (aaRSs), four acyl-CoA synthetases (Acys), three acetyl-CoA synthetases (Aces), phosphoglycerate kinase (Pgk), and adenylate kinase (Adk) were expressed in Escherichia coli and examined for Ap4A or Ap4 synthetase activity using ATP or ATP and triphosphate as substrates. Among the tested enzymes, LysS had the highest Ap4A synthetase activity. AlaRS, SerRS, and LeuRS1 showed high ADP synthetase activity with ATP as a substrate in the presence of pyrophosphatase, and also demonstrated the ability to produce Ap4 from ATP and triphosphate in the absence of pyrophosphatase. Ap4 formation by AlaRS, SerRS, and LeuRS1 was approximately 4- to 13-fold higher compared with that of Ap4A, suggesting that these enzymes prefer triphosphate over ATP as a substrate in the second reaction. Some of the recombinant M. xanthus Acys and Aces also synthesized Ap4 from ATP and triphosphate. However, Pgk was capable of catalyzing the production of Ap4 from ATP and 3-phosphoglycerate in the presence of Mg2+ and did not require triphosphate, suggesting that this enzyme is mainly responsible for Ap4 synthesis in M. xanthus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diadenosine polyphosphates (ApnA: n = 3–6) are a diverse group of nucleotide derivatives found in both prokaryotes and eukaryotes; among them, diadenosine tetraphosphate (Ap4A) is the most studied. The intracellular concentration of Ap4A in Escherichia coli, Synechococcus spp., and Salmonella typhimurium is increased by exposure to heat, heavy metals, or oxidative stress [16, 21], and it has been suggested that Ap4A biosynthesis is involved in the regulation of stress response, pathogenesis, and biofilm formation in bacteria [11, 12, 16, 18].
Myxobacteria are a group of soil bacteria that exhibit complex multicellular life cycles and social behavior [26]. Among them, Myxococcus xanthus is used as a model organism to study the formation of the surrounding biofilms, known as fruiting bodies, under starvation conditions; this bacterium also exhibits increased intracellular Ap4A and diadenosine pentaphosphate (Ap5A) levels following exposure to high temperatures, oxidative and osmotic stresses, and nutrient limitation [14]. However, high levels of intracellular Ap4A and/or Ap5A in developing M. xanthus cells under starvation hinder multicellular development [14], suggesting that Ap4A and/or Ap5A may act as signaling molecules during sporulation.
Ap4A is thought to be produced in vivo via a side reaction during aminoacyl-tRNA synthesis, which is catalyzed by aminoacyl-tRNA synthetases (aaRSs); these enzymes that attach an appropriate amino acid to its tRNA. In the absence of tRNA, aaRSs react with ATP and the amino acid to generate aminoacyl-AMP and pyrophosphate; then, Ap4A is synthesized from aminoacyl-AMP and ATP. AaRSs are classified into two distinct classes (I and II) based on consensus motifs in the catalytic domains [6]. Thus, E. coli expresses alanyl-, lysyl-, phenylalanyl-, and prolyl-RSs that belong to class II and synthesize Ap4A in the presence of Mg2+ and Zn2+ [3].
Adenosine 5′-tetraphosphate (Ap4) is a ubiquitous metabolite thought to be involved in cell signaling in mammals [1]. Furthermore, the yeast species Saccharomyces cerevisiae synthesizes Ap4 during sporulation to a maximum concentration reaching 2% of that of ATP [13]. We have previously shown that M. xanthus reacts to various stress conditions by increasing Ap5A synthesis [14], and M. xanthus lysyl-tRNA synthetase (LysS) catalyzes the formation of Ap5A from Ap4 [20]. Ap4 has also been shown to inhibit the activity of the Ap4A-degrading enzyme asymmetrical dinucleoside tetraphosphatase, thus upregulating the intracellular concentration of Ap4A [17]. Overall, these data indicate that Ap4 biosynthesis forms part of a stress response pathway in microorganisms.
The synthesis of Ap4 can be catalyzed by aaRS, acyl-CoA synthetase (Acy), acetyl-CoA synthetase (Ace), adenylate kinase (Adk), and phosphoglycerate kinase (Pgk) [8]. In this study, we investigated the biosynthesis of Ap4A and Ap4 in M. xanthus by cloning and expressing these M. xanthus enzymes in E. coli and assessing their ability to catalyze the production of Ap4A and Ap4.
Methods and Materials
Expression and Purification of aaRSs, Acys, Aces, Pgk, and Adk
Myxococcus xanthus FB (NBRC13542) was used as a reference strain. Nineteen, four, four, one, and one genes encoding aaRSs, Aces, Acys, Pgk, and Adk, respectively, were amplified by PCR using primers listed in Supplementary Table S1 and cloned into the pCold expression vector containing a His tag (Takara Bio.). Expression and purification of recombinant proteins in E. coli was performed as previously described [19].
Enzyme Activity Assay for Ap4A Synthesis
All reactions were performed in a 10-μl volume at 37 °C for 30 min; the composition of the reaction mixtures for each enzyme group is given below.
(1) AaRSs: 50 mM HEPES (pH 8.0), 5 mM MnCl2, 5 mM ATP, 2 mM amino acid, and aaRS (3–8 μg) in the absence or presence of S. cerevisiae pyrophosphatase (0.1 U, Sigma-Aldrich). (2) Aces: 50 mM HEPES (pH 8.0), 5 mM MnCl2, 5 mM ATP, 5 mM potassium acetate, 0.1 mM DTT, and Ace (4–7 µg). (3) Acys: 50 mM HEPES (pH 8.0), 5 mM MgCl2, 5 mM ATP, 0.1 mM DTT, and Acy (1–4 µg).
Enzyme Activity Assay for Ap4 Synthesis
All reactions were performed in a 10-μl volume (unless stated otherwise) at 37 °C; the composition of the reaction mixtures is given below.
(1) AaRSs: 50 mM HEPES (pH 8.0), 5 mM MnCl2, 5 mM ATP, 0.5 mM triphosphate, 2 mM amino acid, and aaRS (3–8 µg) incubated for 30 min. The time course was analyzed in a 30-μl reaction mixture containing 50 mM HEPES (pH 8.0), 5 mM MnCl2, 5 mM ATP, 1 or 5 mM triphosphate, 2 mM amino acid, and aaRS (12–20 μg) in the absence or presence of pyrophosphatase (0.6 U) after 0.5- to 3-h incubation. Then, 4-μl aliquots were removed at the appropriate time intervals and immediately analyzed. (2) Aces: 50 mM HEPES (pH 8.0), 5 mM MnCl2, 0.1 mM DTT, 5 mM ATP, 5 mM potassium acetate, 2 mM triphosphate, and Ace (4–7 µg) incubated for 30 min [10]. (3) Acys: 50 mM HEPES (pH 8.0), 5 mM MgCl2, 4 mM ATP, 0.1 mM DTT, 4 mM triphosphate, and Acy (1–4 μg) incubated for 30 min [7]. (4) Pgk: 50 mM Tris–HCl (pH 7.5), 5 mM MgCl2, 5 mM ATP, 25 mM KCl, 10 mM 3-phosphoglycerate (3PG), and Pgk (10–15 μg) incubated for 20 min [23]. (5) Adk: 50 mM HEPES (pH 8.0), 5 mM MnCl2, 5 mM ATP, 5 mM ADP, and Adk (10 μg) incubated for 180 min [15].
Analysis of Nucleotides by Ion Exchange HPLC
The formation of reaction products was assessed by HPLC using a Resource Q column (GE Healthcare). The nucleotide concentrations were calculated from the peak areas using standard curves derived from known concentrations of the nucleotides. HPLC analysis was performed as previously described [19].
Results and Discussion
Enzymatic Activity of M. xanthus aaRSs
It has been shown previously that Class II aaRSs, LysRS, PheRS, AlaRS, and ProRS, generate Ap4A [3]. Also, E. coli LysRS has two distinct isoforms (LysS and LysU), and one (LysU) of two isoforms is induced and expressed when the cell is placed under stress conditions; LysU mainly synthesizes Ap4A in E. coli cells [5, 25]. Analysis of the complete M. xanthus genomic sequence indicates that it contains 26 aaRS-encoding genes. Based on the previous findings, we selected 19 aaRS genes coding for class II aaRSs or two distinct isoforms of aaRSs.
Among the 19 enzymes, 15 aaRSs (AlaRS [MXAN_5802], AsnRS [MXAN_2298], GlyRS α and β [MXAN_3100 and _3101], two GluRSs [MXAN_1271 and _2675], HisRS [MXAN_3725], two LeuRSs [MXAN_4051 (LeuRS1) and _4745 (LeuRS2)], two LysRS [MXAN_4731 (LysS) and _5579], PheRS α and β [MXAN_3594 and _3595], ProRS [MXAN_6655], SerRS [MXAN_1941], and two ThrRSs [MXAN_2352 (ThrRS1) and _3591]) were expressed in E. coli as soluble proteins.
After incubation with ATP in the absence of pyrophosphatase, seven aaRSs (AlaRS, two LeuRSs, LysS, ProRS, SerRS, and ThrRS1) showed Ap4A synthetase activity (Table 1), which was significantly induced by Mn2+; in addition, LeuRSs were also activated by Ca2+ (Fig. 1a). Zn2+ is also a potent inducer of Ap4A synthesis by several class II aaRSs (LysRS, PheRS, AlaRS, and ProRS) [1]; however, it inhibits the activity of M. xanthus LysS at 0.15 mM [19]. Here, we observed that Zn2+ at 0.15 and 5 mM stimulated ThrRS1 activity by 1.4- and 2.6-fold, respectively. However, the increase in the Ap4A synthetic activity of ThrRS1 in the presence of 0.15 mM Zn2+ was significantly lower compared with that of E. coli LysU (by 50-180-fold) [2, 22], or AlaRS, PheRS, and ProRS (by 27-188-fold) [3].
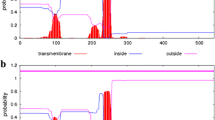
a Effects of bivalent metal cations on Ap4A synthesis by aaRSs. The enzymes were incubated with 5 mM ATP, 2 mM amino acid, and 5 mM of the indicated cations. Also, aaRSs were incubated with 0.15 mM ZnCl2 and 5 mM MnCl2 or MgCl2. The data are presented as the mean ± SE of three independent measurements. b–e Time-dependent changes in the amount of reaction products generated by SerRS (b, d) and LeuRS1 (c, e) from 5 mM ATP and 1 mM triphosphate or 5 mM ATP and 5 mM triphosphate, respectively, in the absence (b, c) or presence (d, e) of pyrophosphatase. (open circle) ATP, (filled circle) Ap4, (filled square) ADP, (filled triangle) Ap4A
In the absence of pyrophosphatase, LysS showed the highest Ap4A synthetic activity, followed by SerRS and ThrRS1 (Table 1) with the Km values for ATP of 1.23 ± 0.06, 2.23 ± 0.19, and 0.79 ± 0.03 mM, respectively (Table 3). In addition, analysis of ThrRS1 kinetics revealed that ATP at concentrations over 2.5 mM inhibited Ap4A synthesis. AlaRS, two LeuRSs, and ProRS also had Ap4A synthetase activity in the presence of Mn2+; however, it was significantly lower than that of LysS (Table 1).
In our previous study, we showed that M. xanthus LysS incubated with lysine and ATP in the absence of pyrophosphatase produced only Ap4A, whereas in the presence of pyrophosphatase it generated Ap4A or ADP from lysyl-AMP with ATP or phosphate, respectively, and then Ap3A from lysyl-AMP with ADP [19]. In the presence of pyrophosphatase, recombinant SerRS, ThrRS1, AlaRS, ProRS, and two LeuRSs mainly produced ADP from aminoacyl-AMP with phosphate, suggesting that these aaRSs exhibited preference for phosphate rather than ATP as a substrate in the second reaction. Because these M. xanthus aaRSs had high ADP synthetase activity (41.7–128.1 nmol/min/mg) in bacterial cells, they may mainly generate ADP from ATP and phosphate in the presence of pyrophosphatase.
Unlike most aaRSs, PheRSs exhibit a tetrameric structure (α2β2) formed by the α and β subunits [23]. In E. coli, plants, and animals, PheRSs catalyze Ap4A formation [4, 9]; however, the mixture of recombinant α and β subunits of M. xanthus PheRS did not show Ap4A synthetic activity, indicating that the recombinant subunits did not form an enzymatically active complex.
In the absence of pyrophosphatase, the ability of LysS to produce Ap4A and Ap4 from 5 mM ATP and 5 mM triphosphate dramatically decreased; this was attributed to the presence of both pyrophosphate and triphosphate synergistically blocking the formation of lysyl-AMP or Ap4A through competitive inhibition of the ATP reaction [20]. Therefore, we examined Ap4 synthesis by seven aaRSs capable of generating Ap4A, using 5 mM ATP and 0.5 mM triphosphate as substrates (Table 1). SerRS and LeuRS1 showed high Ap4 synthetase activity, and the Km values for triphosphate of SerRS and LeuRS1 were 0.21 ± 0.01 and 0.82 ± 0.01 mM, respectively, in the absence of inorganic pyrophosphatase (Table 3). When SerRS was incubated with 5 mM ATP and 1 mM triphosphate and LeuRS1 with 5 mM ATP and 5 mM triphosphate, they mainly produced Ap4 (Fig. 1b, c).
In the presence of pyrophosphatase, M. xanthus LysS incubated with 5 mM ATP and 5 mM triphosphate first generated Ap4A from lysyl-AMP with ATP, then Ap4 from lysyl-AMP with triphosphate, and finally ADP from lysyl-AMP with phosphate [20]. SerRS did not generate Ap4A and Ap4; however, it efficiently utilized seryl-AMP and inorganic phosphate cleaved from pyrophosphate by pyrophosphatase to produce ADP in the second reaction (Fig. 1d). At the same time, LeuRS1 generated Ap4 from leucyl-AMP with triphosphate, indicating its preference for triphosphate over ATP in the second reaction, and then produced ADP from leucyl-AMP with phosphate (Fig. 1e). After a 90-min reaction, LeuRS1 generated low amounts of Ap4A or Ap5A from leucyl-AMP with ATP or Ap4, respectively.
Enzymatic Activity of M. xanthus Acys
It was shown that Acy from Pseudomonas fragi synthesizes Ap4 from ATP and triphosphate [7]. M. xanthus has four Acys: Acy1–4 (MXAN_0216, _0225, _6374 (Acy3), and _7148, respectively). When recombinant M. xanthus Acys were incubated with 4 mM ATP and 4 mM triphosphate, they showed Ap4 synthetase activity, which was stimulated by 5 mM Mg2+, Mn2+, or Co2+ (Fig. 2a). Among the four enzymes, Acy3 showed the highest Ap4 synthetase activity (Table 2); the Km for triphosphate was 2.07 ± 0.05 mM, which is similar to that of P. fragi Acy (1.3 mM) [7]. The Vmax for triphosphate was 6.89 ± 0.27 nmol/min/mg (Table 3). The enzyme additionally exhibited weak Ap4A synthetase activity when incubated with ATP (Table 2).
Enzymatic Activity of M. xanthus Aces
Ace from S. cerevisiae catalyzes the synthesis of Ap4 from ATP and triphosphate [10]. M. xanthus possesses four Aces (MXAN_0949, _1573, _2570, and _5856), three of which (MXAN_0949, _2570, and _5856) were expressed in E. coli and designated Ace1–3, respectively. Ace1 was capable only of Ap4 synthesis from ATP and triphosphate, which was stimulated exclusively by 5 mM Mn2+ (Fig. 2a). Its Km for triphosphate was 0.52 ± 0.01 mM, which is approximately ninefold higher than that of S. cerevisiae Ace [10], and its Vmax was 1.05 ± 0.08 nmol/min/mg (Table 3). Ap4 synthesis occurred in the absence of added acetate; however, the enzyme demonstrated a decrease in Ap4 formation by approximately 40%. In addition, Ace1 showed very low Ap4A synthetase activity when ATP was used as a substrate (Table 1).
Enzymatic Activity of M. xanthus Pgk and Adk
Pgk catalyzes the transfer of the phosphoryl group from ATP to 3-phosphoglycerate, yielding 1,3-bisphosphoglycerate and ADP [24], and can generate Ap4 and 3-phosphoglycerate from 1,3-bisphosphoglycerate and ATP. When M. xanthus Pgk was incubated with 5 mM ATP and 10 mM 3-phosphoglycerate, it catalyzed Ap4 synthesis; this process required bivalent cations, among which 5 mM Mg2+ was the most effective (Fig. 2b). Thus, with 5 mM Co2+, Mn2+, or Ca2+, Pgk activity was 85, 60, and 10%, respectively, of that with Mg2+. The kinetic parameters of M. xanthus Pgk with ATP as a substrate were: Km = 5.22 ± 0.14 mM and Vmax = 26.7 ± 0.49 nmol/min/mg (Table 3).
Adk catalyzes the reversible conversion of ATP and AMP to two ADPs. As rabbit and pig Adks have been shown to catalyze the formation of Ap4 from ATP and ADP [15], we assessed Ap4 synthesis by M. xanthus Adk; however, it demonstrated very low specific activity (0.035 nmol/min/mg).
Triphosphate is generated from dGTP by deoxyguanosine triphosphatase; however, it is generally considered that the intracellular concentration of triphosphate is very low. Although M. xanthus does not utilize glucose or any other sugar as a primary carbon source, Pgk is required for gluconeogenesis to generate polysaccharides, spore coat, and the cell wall. Therefore, in M. xanthus cells, Ap4 may be mainly produced by Pgk, which does not require triphosphate for Ap4 synthesis. The generated Ap4 may be used as a substrate for Ap5A synthesis by LysS. Ap5A binds Adk with nanomolar affinity, and inhibits the Adk reaction. Since Adk plays an important role in cellular viability and cellular energy balance, Ap5A synthesized by LysS from Ap4 may regulate adenine nucleotide homeostasis in starved M. xanthus cells. In addition, Ap4 acts as a strong inhibitor of Nudix Ap4A hydrolase [17], suggesting that this tetraphosphate may play a critical role in the regulation of intracellular Ap4A levels.
References
Amici A, Grolla AA, Del Grosso E, Bellini R, Bianchi M, Travelli C, Garavaglia S, Sorci L, Raffaelli N, Ruggieri S, Genazzani AA, Orsomando G (2017) Synthesis and degradation of adenosine 5′-tetraphosphate by nicotinamide and nicotinate phosphoribosyltransferases. Cell Chem Biol 24:553–564
Blanquet S, Plateau P, Onesti S (2005) Class II lysyl-tRNA synthetases. In Ibba M, Francklyn C, Cusack S (eds) The aminoacyl-tRNA synthetases. Eurekah, Georgetown, pp 227–240
Blanquet S, Plateau P, Brevet A (1983) The role of zinc in 5′,5′-diadenosine tetraphosphate production by aminoacyl-transfer RNA synthetases. Mol Cell Biochem 52:3–11
Brevet A, Plateau P, Cirakoğlu B, Pailliez JP, Blanquet S (1982) Zinc-dependent synthesis of 5′,5′-diadenosine tetraphosphate by sheep liver lysyl- and phenylalanyl-tRNA synthetases. J Biol Chem 257:14613–14615
Brevet A, Chen J, Lévêque F, Blanquet S, Plateau P (1995) Comparison of the enzymatic properties of the two Escherichia coli lysyl-tRNA synthetase species. J Biol Chem 270:14439–14444
Eriani G, Delarue M, Poch O, Gangloff J, Moras D (1990) Partition of tRNA synthetases into two classes based on mutually exclusive sets of sequence motifs. Nature 347:203–206
Fontes R, Sillero MA, Sillero A (1998) Acyl coenzyme A synthetase from Pseudomonas fragi catalyzes the synthesis of adenosine 5′-polyphosphates and dinucleoside polyphosphates. J Bacteriol 130:3152–3158
Fraga H, Fontes R (2011) Enzymatic synthesis of mono and dinucleoside polyphosphates. Biochim Biophys Acta 1810:1195–1204
Goerlich O, Foeckler R, Holler E (1982) Mechanism of synthesis of adenosine (5′) tetraphosphate (5′) adenosine (AppppA) by aminoacyl-tRNA synthetases. Eur J Biochem 126:135–139
Guranowski A, Gunther Sillero MA, Sillero A (1994) Adenosine 5′-tetraphosphate and adenosine 5′-pentaphosphate are synthesized by yeast acetyl coenzyme A synthetase. J Bacteriol 176:2986–2990
Hansen S, Lewis K, Vulic M (2008) Role of global regulators and nucleotide metabolism in antibiotic tolerance in Escherichia coli. Antimicrob Agents Chemother 52:2718–2726
Ismail TM, Hart CA, McLennan AG (2003) Regulation of dinucleoside polyphosphate pools by the YgdP and ApaH hydrolases is essential for the ability of Salmonella enterica serovar Typhimurium to invade cultured mammalian cells. J Biol Chem 278:32602–32607
Jakubowski H (1986) Sporulation of the yeast Saccharomyces cerevisiae is accompanied by synthesis of adenosine 5′-tetraphosphate and adenosine 5′-pentaphosphate. Proc Natl Acad Sci USA 83:2378–2382
Kimura Y, Tanaka C, Sasaki K, Sasaki M (2017) High concentrations of intracellular Ap4A and/or Ap5A in developing Myxococcus xanthus cells inhibit sporulation. Microbiology 163:86–93
Kupriyanov VV, Ferretti JA, Balaban RS (1986) Muscle adenylate kinase catalyzes adenosine 5′-tetraphosphate synthesis from ATP and ADP. Biochim Biophys Acta 869:107–111
Lee PC, Bochner BR, Ames BN (1983) AppppA, heat-shock stress, and cell oxidation. Proc Nat Acad Sci USA 80:7496–7500
Lobatón CD, Vallejo CG, Sillero A, Sillero MAG (1975) Diguanosinetetraphosphatase from rat liver: activity on diadenosine tetraphosphate and inhibition by adenosine tetraphosphate. Eur J Biochem 50:495–501
Monds RD, Newell PD, Wagner JC, Schwartzman JA, Lu W, Rabinowitz JD, O’Toole GA (2010) Di-adenosine tetraphosphate (Ap4A) metabolism impacts biofilm formation by Pseudomonas fluorescens via modulation of c-di-GMP-dependent pathways. J Bacteriol 192:3011–3023
Oka M, Takegawa K, Kimura Y (2015) Enzymatic characterization of a class II lysyl-tRNA synthetase, LysS, from Myxococcus xanthus. Arch Biochem Biophys 579:33–39
Oka M, Takegawa K, Kimura Y (2016) Lysyl-tRNA synthetase from Myxococcus xanthus catalyzes the formation of diadenosine penta- and hexaphosphates from adenosine tetraphosphate. Arch Biochem Biophys 604:152–158
Pàlfi Z, Surànyi G, Borbély G (1991) Alterations in the accumulation of adenylylated nucleotides in heavy-metal-ion-stressed and heat-stressed Synechococcus sp. strain PCC 6301, a cyanobacterium, in light and dark. Biochem J 276:487–491
Plateau P, Blanquet S (1982) Zinc-dependent synthesis of various dinucleoside 5′,5′′′-P1,P3-Tri- or 5″,5′′′-P1,P4-tetraphosphates by Escherichia coli lysyl-tRNA synthetase. Biochemistry 21:5273–5279
Sanni A, Mirande M, Ebel J-P, Boulanger Y, Waller J-P, Fasiolo F (1988) Structure and expression of the genes encoding the α and β subunits of yeast phenylalanyl-tRNA synthetase. J Biol Chem 263:15407–15415
Small GD, Cooper C (1966) Studies on the occurrence and biosynthesis of adenosine tetraphosphate. Biochemistry 5:26–33
VanBogelen RA, Vaughn V, Neidhardt FC (1983) Gene for heatinducible lysyl-tRNA synthetase (lysU) maps near cadA in Escherichia coli. J Bacteriol 153:1066–1068
Whitworth DE (2008) Myxobacteria: multicellularity and differentiation. ASM Press, Washington
Acknowledgements
This study was supported by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan (16K07667).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kimura, Y., Tanaka, C. & Oka, M. Identification of Major Enzymes Involved in the Synthesis of Diadenosine Tetraphosphate and/or Adenosine Tetraphosphate in Myxococcus xanthus. Curr Microbiol 75, 811–817 (2018). https://doi.org/10.1007/s00284-018-1452-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-018-1452-x