Abstract
Marine actinomycetes are less investigated compared to terrestrial strains as potential sources of natural products. To date, few investigations have been performed on culturable actinomycetes associated with South China Sea sediments. In the present study, twenty-eight actinomycetes were recovered from South China Sea sediments after dereplication by traditional culture-dependent method. The 16S rRNA gene sequences analyses revealed that these strains related to five families and seven genera. Twelve representative strains possessed at least one of the biosynthetic genes coding for polyketide synthase I, II, and nonribosomal peptide synthetase. Four strains had anti-Mycobacterium phlei activities and five strains had activities against methicillin-resistant Staphylococcus aureus. 10 L-scale fermentation of strains Salinispora sp. NHF45, Nocardiopsis sp. NHF48, and Streptomyces sp. NHF86 were carried out for novel and bioactive compounds discovery. Finally, we obtained a novel α-pyrone compound from marine Nocardiopsis sp. NHF48, an analogue of paulomenol from marine Streptomyces sp. NHF86 and a new source of rifamycin B, produced by Salinispora sp. NHF45. The present study concluded that marine actinomycetes, which we isolated from South China Sea sediments, will be a suitable source for the development of novel and bioactive compounds.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The class Actinobacteria has considered to be an important source of high value pharmaceutically reagents such as antibiotics, enzymes, and recombinant products [4, 14]. As the search for natural products from Streptomycetes continues [25], the rate of discovery of new compounds from Streptomycetes has decreased. On the other hand, the emergence of drug-resistant bacterial pathogens has caused a resurgence of interest in discovering new biologically active compounds. Nowadays, research shows that rare actinomycete species represent a unique source of novel biologically active compounds [3, 14]. As we know, actinomycetes isolated from terrestrial environments also increase the frequency of rediscovering known natural products [29]. Thus, actinomycetes from deep sea of different regions are mostly relatively novel and untapped resources [25].
Unique biochemical metabolic and physiological capabilities were developed by deep-sea actinomycetes, which ensure both survival and provide potential for the biosynthesis of diverse novel metabolites [18]. Up to date, lots of novel actinomycete species possessing novel biological activities were isolated between 2006 and 2016 from deep-sea environment, especially at depths of abyssal zone [7, 12, 13]. The South China Sea is a typical marginal sea, in which the sediments may contain abundant actinomycetes with diverse taxa. Actinomycetes belonging to seven suborders, 13 families, and 28 genera including 14 new species with diverse metabolites have been obtained by Chinese researchers [27]. For instance, Micromonospora rosaria isolated from sediment sample could synthesis fluostatins with good antimicrobial activities against Staphylococcus aureus [32]; marfuraquinocins from Streptomyces niveus were found to inhibit a NCI-H460 cancer cell line [24]. Thus, the extensive area of South China Sea is a promising biome to explore marine actinomycetes.
Materials and Methods
Marine Sediment Samples Collection and Actinomycetes Isolation
Sediment samples were collected using the mud sampler in the South China Sea from depths 98 to 2974 m (Supplementary Fig. S1). The water contents and pH values of samples were tested using dryer and pH meter. Samples were diluted in sterial seawater and spreaded over the surface of isolation plates (Supplementary Table 1). Then the plates were incubated at 28 °C for 1 month. Actinomycetes were selected based on the presence of filamentous hyphae or spores and purified. The total number of pure actinomycetes was counted and preserved on ISP2 slants.
Detection of 16S rRNA, PKS, and NRPS Biosynthetic Genes
The genes encoding 16S rRNA, polyketide synthases I, II (PKS I and PKS II), and nonribosomal peptide synthetases (NRPS) were amplified and sequenced using the primer pairs shown in Supplementary Table S2 as recommended by Yang and Sun and Qin et al. [20, 31]. The obtained 16S rRNA gene sequences were deposited in GenBank (Accession Numbers: MF467904, MF467905, FJ830632, FJ830634, FJ830630, MF467906, KU500358-KU500370, KU312336-KU312339, KU529470-KU529472, KU550963, JQ911670) and identified using the BLAST program (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Sequences were aligned with most closely related 16S rRNA gene sequences from the GenBank and the phylogenetic trees were constructed using Neighbor-Joining algorithm [22] by MEGA software version 6.0 [26]. The topology of the phylogenetic trees was evaluated by bootstrap re-sampling method with 1000 replicates [6].
Crude Extract Preparation and Antimicrobial Activity Test
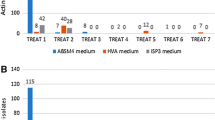
All strains were inoculated into ISP2 medium. Then, the seed culture was incubated at 28 °C for 7–10 days while shaking at 220 rpm. 1 mL of the seed culture was transferred to 40/250 mL flask fermentation medium (Supplementary Table S3). The cultures were grown at 28 °C for 7 days at 220 rpm. An equal volume of ethyl acetate was added to the culture for metabolites extraction. Then, the supernatant was evaporated and dissolved in DMSO used for antimicrobial activity. The antimicrobial activity test of crude extract (100 μg) was carried out using the standard disk diffusion assay [21] against pathogens Mycobacterium phlei and methicillin-resistant Staphylococcus aureus (MRSA) [5]. The antimicrobial potential was assessed as dimeter of the inhibition zone.
Ten-Liter Scale Fermentation, Compound Isolation, and Identification
10 mL seed culture of each isolate was transferred to 200 mL/1000 mL flask fermentation medium and incubated at 28 °C for 7 days at 220 rpm. The fermentation broth was extracted with ethyl acetate three times and subjected to further compound separation. For strain Nocardiopsis sp. NHF48, crude extract was applied on a Sephadex LH-20 column (elution reagent, dichloromethane: methanol = 2:1 (v/v)) and fraction 5 with anti-MRSA activity was subjected to HPLC (Agilent 1200 series) equipped with an Eclipse XDB-C18, 5 μm column (9.4 × 250 mm2, Agilent), with acetonitrile in water (linear gradient elution from 10 to 100% acetonitrile) as mobile phase. Fraction 5 gave pure compound 1. For strain Streptomyces sp. NHF86, the crude extract was dissolved with n-hexane, dichloromethane, and methanol sequentially. The dichloromethane fraction with anti-M. phlei activity was further prepared using HPLC and gave compound 2.
Results
Sample Characteristics and Culturable Actinomycetes
Samples were collected from six different sites of South China Sea with different depths. Results showed that all samples had neutral pH values (7.12–7.48). The water contents of samples collected from different depths exhibited great differences, from 18.6 to 56.4% (Supplementary Table S4).
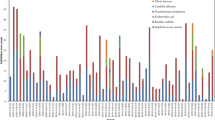
On the basis of the ability to form aerial hyphae and spores, 28 actinobacterial strains were finally purified. The actinomycetes isolated were grouped into five families and seven genera-Streptomyces, Salinispora, Nocardiopsis, Verrucosispora, Micromonospora, Prauserella, and Promicromonospora. The most abundant genus was Streptomyces, which is consistent with previous reports from marine sediments [19]. The percentages of 16S rRNA gene sequence similarities of these isolates to the closest strains were listed in Table 1.
Detection of PKS, NRPS Biosynthetic Genes, and Antimicrobial Strains
The presence of marker genes, such as PKS and NRPS, may indicate production of certain secondary metabolites [15, 16]. Twelve isolates with known species information were screened for the presence of PKS I, PKS II, and NRPS genes, to predict their biosynthetic potential for active or novel compounds. As shown in Table 1, strain NHF86 belonging to genus Streptomyces possessed all three targeted genes. Compared to strain NHF86, other strains possessed one or two target genes. The analysis results confirmed that they indeed encode parts of the expected biosynthetic enzymes. As we know, the detection of biosynthetic genes is not directly correlated with the detection of antimicrobial compounds. Therefore, these strains were also tested for antimicrobial activities against the pathogens MRSA and M. phlei. Results showed that four isolates exhibited activities against MRSA and five isolates were active against M. phlei (Table 1). Among them, three actinobacterial strains, Salinispora sp. NHF45, Nocardiopsis sp. NHF48, and Streptomyces sp. NHF86 were selected for further chemical analyzation.
A new α-Pyrone Compound from Marine Nocardiopsis sp. NHF48
Strain NHF48 was isolated from a sediment sample No. 32 with anti-MRSA activity (MIC value: 12.5 μg/mL) under medium MPG. The 16S rRNA gene sequence analysis suggested that strain NHF48 had the highest similarity with Nocardiopsis flavescents CGMCC 4.5723T (99.8%) and formed a stable cluster with Nocardiopsis flavescents supported by a high bootstrap value of 99 (Fig. 1a). Basing on these two results, strain NHF48 was a member of the genus Nocardiopsis. Compound 1 was obtained as white amorphous powder. The molecular formula of C16H26O2 was established by the analysis of HRESIMS at m/z 251.1989 [M + H]+ and 13C NMR spectrum which revealed signals of 16 carbons. The final structure of compound 1 was assigned on the basis of 2D NMR data (Supplementary Fig. S2), particularly the combination of HMBC experiment which allowed all protons and carbons to be assigned. The 1H and 13C NMR spectra in combination with 1H–13C HSQC NMR data of 1 indicated signals of five methyl groups, four methylenes, and two methines. Analysis of 1H–1H COSY and HMBC contributed to establish connectivities of most carbons, which was followed by comparison of its spectral data with those from elijopyrone A [28]. In compound 1, the pyrone nucleus (δ C 164.1; δ C 125.1; δ C 144.6, δ H 6.87; δ C 109.4; δ C 162.0) was clearly the same with elijopyrone A. The differences observed between compound 1 and elijopyrone A were the isoamyl substitutions at C-2 (δ C 37.7; δ C 33.1; δ C 29.4; δ C 11.4; δ C 18.9) and C-5 (δ C 34.5; δ C 36.5; δ C 20.7; δ C 14.0; δ C 18.4). The HMBC correlations observed from H-3 (δ H 6.87) to C-12 (δ C 37.7) and H-10 (δ H 1.20, d, 6.8) to C-3 (δ C 144.6), supported the correct locations of two isoamyl groups. The structure of 1 was shown in Fig. 1b and the NMR assignments were shown in Table 2. Compound 1 had activity on mouse melanoma cell line B16 at a GI50 value of 61.7 μg/mL and was a novel compound derived from marine Nocardiopsis sp. NHF48.
An Analogue of Paulomentol from Marine Streptomyces sp. NHF86
Strain NHF86 had anti-M. phlei activity fermented by ISP2 medium. Phylogenetic analysis (Fig. 2a) based on 16S rRNA gene sequence revealed that strain NHF86 was closely associated with the genus Streptomyces. The molecular formular of C29H41NO15 was established at m/z 644.2508 [M + H]+. Compound 2 (Fig. 2b) was determined to an analogue of paulomenol according to the 1D-NMR (1H and 13C NMR) and 2D-NMR (1H-1H COSY, HMBC and HSQC, Supplementary Fig. S3) and by comparison with the data reported in the literatures [1, 2, 30]. Detailed analysis of the 1D and 2D NMR (acetone −d 6) data for compound 2 (Table 3) revealed signals of 29 carbons. Five signals have disappeared from the 13C NMR compared to paulomycin A, indicated the loss of C5H4NOS (corresponding acid, paulic acid). In compound 2, C-5 (δ C 130.6) and C-6 (δ C 150.7) were presented as quinine peaks, and olefinic resonances (C-5, δ C 48.01; C-6, δ C 78.20) were detected in compound paulomenol A.
Marine Salinispora sp. NHF45, a Rifamycin B-Producing Strain
Strain NHF45 isolated from sample No. 37 has the highest similarity with Salinispora arenicola (100%). The HPLC chromatograms showed that crude extract of strain NHF45 contained peak with retention time similar to that of the standard compound rifamycin B (Fig. 3a). The electrospray negative-ion spectrum showed a molecular ion peak of [M−H]− m/z 754.2, confirming this peak was rifamycin B (Fig. 3b). The rediscovery of rifamycin-producing strain from South China Sea sediment supplies another useful source for screening high-yield rifamycin-synthesizing strains.
Discussion
In this study, we isolated twenty-eight marine actinomycetes from the South China Sea sediments. Their phylogenetic properties and potential to synthesize secondary metabolites were investigated. These strains belong to seven genera with 16S rRNA gene sequences similarities of 99.0-100%. Studies of marine actinomycetes as a source of novel bioactive compounds have been acknowledged by many researchers. In this study, in order to explore the active compound producing potential of these isolated strains, a prescreening based on PCR amplification was pursued for the detection of biosynthetic genes including PKS and NRPS [8]. All the selected strains possessed one or more biosynthetic genes, showing that secondary metabolic pathways are widely distributed among actinomycetes isolated from South China Sea. Some corresponding metabolites may not be detected partially due to ‘cryptic’ gene clusters or metabolites with no antimicrobial activities [11]. In recent years, whole-genome sequencing has been widely involved in new gene cluster discovery and kinds of strategies were implicated to stimulate expression of ‘cryptic’ gene clusters [9, 10, 17]. Further efforts can be done to mine active compounds from these strains.
On the other hand, we detected the activities of crude extracts prepared by five fermentation media. Notably, rare actinomycetes showed great activities against MRSA and M. phlei. In detail, Micromonospora, Verrucosispora, Nocardiopsis, and Prauserella showed anti-MRSA activities, Salinispora and Micromonospora showed anti-M. phlei activities. We choose one Streptomyces and two rare actinomycetes for the further chemical analyzation. In the end, compound 1 was obtained from strain Nocardiopsis sp. NHF48. According to literature and database of SciFinder, compound 1 was a new α-pyrone compound with different groups at C2 and C5. Novel α-pyrone compounds have been produced by marine Nocardiopsis with activities against tumor cell lines [33, 34]. Also, α-pyrone compounds can be found in all three kingdoms of life and the biological activities of α-pyrones are very diverse (antimicrobial, anti-tumor, antiviral activities and other biological functions). For example, α-pyrones have been shown to be HIV protease and selective COX-2 inhibitors; α-pyrones can act as signaling molecules in the cell–cell communication system of the bacterium Photorhabdus luminescens [23]. Our finding added another α-pyrone compound produced by Nocardiopsis with anti-tumor (mouse melanoma cell line) activity (a GI50 value of 61.7 μg/mL). To our knowledge, the compound 1 was first reported produced by Nocardiopsis isolated from marine sediments. In addition, for Streptomyces, an analogue of paulomenol was isolated from strain Streptomyces sp. NHF86 with anti-M. phlei activity. It has been reported that paulomycin A can be changed to paulomenol A in the solution of 0.01 N (CH3)3N [30]. We purified compound 2 without (CH3)3N from crude extract of NHF86, which further proved that strain NHF86 could produce paulomenol-related compound. Strain Salinispora sp. NHF45 could be another source for screening high-yield rifamycin-synthesizing strains. This study uncovered the rich source of actinomycetes which provides the basis for the bioprospection of South China Sea.
References
Argoudelis AD, Baczynskyj L, Haak WJ, Knoll WM, Mizsak SA, Shilliday FB (1988) New paulomycins produced by Streptomyces paulus. J Antibiot 41:157–169
Argoudelis AD, Baczynskyj L, Mizsak SA, Shilliday FB (1988) O-demethylpaulomycin-A and O-demethylpaulomycin-B, U-77,802 and U-77,803, paulomenol-A and paulomenol-B, new metabolites produced by Streptomyces paulus. J Antibiot 41:1316–1330
Baltz RH (2006) Marcel Faber Roundtable: is our antibiotic pipeline unproductive because of starvation, constipation or lack of inspiration? J Ind Microbiol Biotechnol 33:507–513. doi:10.1371/journal.pone.0149216
Berdy J (2005) Bioactive microbial metabolites—a personal view. J Antibiot 58:1–26
Chen C, Wang J, Guo H, Hou W, Yang N, Ren B, Liu M, Dai H, Liu X, Song F, Zhang L (2013) Three antimycobacterial metabolites identified from a marine-derived Streptomyces sp. MS100061. Appl Microbiol Biotechnol 97:3885–3892. doi:10.1007/s00253-012-4681-0
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. doi:10.1111/j.1558-5646.1985.tb00420.x
Fenical W, Jensen PR (2006) Developing a new resource for drug discovery: marine actinomycete bacteria. Nat Chem Biol 2:666–673. doi:10.1038/nchembio841
Fischbach MA, Walsh CT (2006) Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: logic, machinery, and mechanisms. Chem Rev 106:3468–3496. doi:10.1007/s10295-013-1348-5
Gomez-Escribano JP, Bibb MJ (2014) Heterologous expression of natural product biosynthetic gene clusters in Streptomyces coelicolor: from genome mining to manipulation of biosynthetic pathways. J Ind Microbiol Biotechnol 41:425–431. doi:10.1007/s10295-013-1348-5
Guo X, Geng P, Bai F, Bai G, Sun T, Li X, Shi L, Zhong Q (2012) Draft genome sequence of Streptomyces coelicoflavus ZG0656 reveals the putative biosynthetic gene cluster of acarviostatin family alpha-amylase inhibitors. Lett Appl Microbiol 55:162–169. doi:10.1111/j.1472-765X.2012.03274.x
Hodges TW, Slattery M, Olson JB (2012) Unique actinomycetes from marine caves and coral reef sediments provide novel PKS and NRPS biosynthetic gene clusters. Mar Biotechnol 14:270–280. doi:10.1007/s10126-011-9410-7 (NY)
Kamjam M, Sivalingam P, Deng Z, Hong K (2017) Deep sea actinomycetes and their secondary metabolites. Front Microbiol 8:760. doi:10.3390/md15030071
Kwon HC, Kauffman CA, Jensen PR, Fenical W (2006) Marinomycins A-D, antitumor-antibiotics of a new structure class from a marine actinomycete of the recently discovered genus “Marinispora”. J Am Chem Soc 128:1622–1632. doi:10.1021/ja0558948
Lazzarini A, Cavaletti L, Toppo G, Marinelli F (2001) Rare genera of actinomycetes as potential producers of new antibiotics. Anton Leeuw Int J G 79:399–405. doi:10.1023/A:1010287600557
Liao L, Chen R, Jiang M, Tian X, Liu H, Yu Y, Fan C, Chen B (2016) Bioprospecting potential of halogenases from Arctic marine actinomycetes. BMC Microbiol 16:34. doi:10.1186/s12866-016-0662-2
Liu W, Ahlert J, Gao Q, Wendt-Pienkowski E, Shen B, Thorson JS (2003) Rapid PCR amplification of minimal enediyne polyketide synthase cassettes leads to a predictive familial classification model. Proc Natl Acad Sci USA 100:11959–11963. doi:10.1073/pnas.2034291100
Moore JM, Bradshaw E, Seipke RF, Hutchings MI, McArthur M (2012) Use and discovery of chemical elicitors that stimulate biosynthetic gene clusters in Streptomyces bacteria. Methods Enzymol 517:367–385. doi:10.1016/B978-0-12-404634-4.00018-8
Pathom-aree W, Stach JEM, Ward AC, Horikoshi K, Bull AT, Goodfellow M (2006) Diversity of actinomycetes isolated from Challenger Deep sediment (10,898 m) from the Mariana Trench. Extremophiles 10:181–189. doi:10.1007/s00792-005-0482-z
Prieto-Davó A, Dias T, Gomes SE, Rodrigues S, Parera-Valadezl Y, Borralho PM, Pereira F, Rodrigues CMP, Santos-Sanches I, Gaudencio SP (2016) The madeira archipelago as a significant source of marine-derived actinomycete diversity with anticancer and antimicrobial potential. Front Microbiol. doi:10.3389/fmicb.2016.01594
Qin S, Li J, Chen HH, Zhao GZ, Zhu WY, Jiang CL, Xu LH, Li WJ (2009) Isolation, diversity, and antimicrobial activity of rare actinobacteria from medicinal plants of tropical rain forests in Xishuangbanna, China. Appl Environ Microbiol 75:6176–6186. doi:10.1128/AEM.01034-09
Raahave D (1974) Paper disk-agar diffusion assay of penicillin in the presence of streptomycin. Antimicrob Agents Chemother 6:603–605
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schäberle TF (2016) Biosynthesis of alpha-pyrones. Beilstein J Org Chem 12:571–588. doi:10.3762/bjoc.12.56
Song Y, Huang H, Chen Y, Ding J, Zhang Y, Sun A, Zhang W, Ju J (2013) Cytotoxic and antibacterial marfuraquinocins from the deep South China Sea-derived Streptomyces niveus SCSIO 3406. J Nat Prod 76:2263–2268. doi:10.1021/np4006025
Subramani R, Aalbersberg W (2012) Marine actinomycetes: an ongoing source of novel bioactive metabolites. Microbiol Res 167:571–580. doi:10.1016/j.micres.2012.06.005
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. doi:10.1093/molbev/mst197
Tian XP, Zhang S, Li W (2011) Advance in marine actinobacterial research—a review. Wei sheng wu xue bao= Acta microbiologica Sinica 51:161–169
Toske SG, Jensen PR, Kauffman CA, Fenical W (1995) Elijopyrones A-D: new α-pyrones from a marine actinomycete. Nat Prod Lett 6:303–308. doi:10.1080/10575639508043175
Wawrik B, Kutliev D, Abdivasievna UA, Kukor JJ, Zylstra GJ, Kerkhof L (2007) Biogeography of actinomycete communities and type II polyketide synthase genes in soils collected in New Jersey and Central Asia. Appl Environ Microbiol 73:2982–2989. doi:10.1128/AEM.02611-06
Wiley PF, Mizsak SA, Baczynskyj L, Argoudelis AD (1984) The structure of paulomycin. J Antibiot 37:1273–1275
Yang N, Sun C (2016) The inhibition and resistance mechanisms of actinonin, isolated from marine Streptomyces sp. NHF165, against Vibrio anguillarum. Front Microbiol 7:1467. doi:10.3389/fmicb.2016.01467
Zhang W, Liu Z, Li S, Lu Y, Chen Y, Zhang H, Zhang G, Zhu Y, Zhang G, Zhang W, Liu J, Zhang C (2012) Fluostatins I-K from the South China Sea-derived Micromonospora rosaria SCSIO N160. J Nat Prod 75:1937–1943. doi:10.1021/np300505y
Zhang XM, Sun MW, Shi H, Lu CH (2017) α-Pyrone derivatives from a marine actinomycete Nocardiopsis sp. YIM M13066. Nat Prod Res. doi:10.1080/14786419.2017.1299730
Zou G, Liao XJ, Peng Q, Chen GD, Wei FY, Xu ZX, Zhao BX, Xu SH (2017) A new alpha-pyrone from the deep-sea actinomycete Nocardiopsis dassonvillei subsp. dassonvillei DSM 43111(T). J Asian Nat Prod Res. doi:10.1080/10286020.2017.1307186
Acknowledgements
This work was supported by grants from Shandong Provincial Natural Science Foundation, China (ZR2016CB13) and National Natural Science Foundation of China (31600136).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, N., Song, F. Bioprospecting of Novel and Bioactive Compounds from Marine Actinomycetes Isolated from South China Sea Sediments. Curr Microbiol 75, 142–149 (2018). https://doi.org/10.1007/s00284-017-1358-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-017-1358-z