Abstract
Fluorescent pseudomonads isolated from wheat leaves were characterized regarding their antagonistic potential and taxonomy in relation to protect crop plants from infestation by Fusarium and Alternaria fungi causing diseases in wheat. Using a dual culture assay, inhibition of fungal growth was found for 40 isolates of 175 fluorescent pseudomonads. Twenty-two of the antagonists were able to suppress strains of Fusarium as well as Alternaria. By means of real-time qPCR, the phlD gene encoding the antibiotic 2,4-diacetylphloroglucinol was detected in 20 isolates. On the basis of matrix-assisted laser desorption/ionization time-of-flight mass spectrometry spectral patterns, the isolates with antagonistic activity were assigned to the phylogenetic subgroup Pseudomonas fluorescens and the closely related Pseudomonas gessardii subgroup. The results of the study suggest that pseudomonads in the phyllosphere of crop plants may possibly contribute to natural plant protection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The yield and quality of wheat (Triticum spp.)—a staple food consumed by humans in many countries—are threatened by one of the most devastating fungal diseases worldwide: Fusarium head blight (FHB) [14]. The main causal agents in the temperate climates of Central Europe are Fusarium graminearum [teleomorph Gibberella zeae (Schwein) Petch] and Fusarium culmorum [(W.G. Smith) Sacc (teleomorph unknown)] [1]. So-called “black point” is another fungal disease in wheat, which is caused by Alternaria alternata (Fr.) Keissler and Alternaria tenuissima [(Kunze ex Nees et T. Nees: Fries) Wiltshire]. These fungi occur ubiquitously on wheat leaves and ears, and the disease is distributed worldwide in both humid and semi-arid regions [31]. In addition to yield losses and reduced grain quality, these Fusarium as well as Alternaria species are known to produce mycotoxins, which are harmful to both humans and livestock [13, 14].
Bacteria of the genus Pseudomonas are commonly found among the predominant genera in the phyllosphere, independent of the host plant species, and are also found on and in the above-ground parts of wheat plants [12, 33]. The relationship between these bacteria and their host plant can be saprophytic, pathogenic or plant-growth promoting [14, 27], the last of which is attributed to a group of non-phytopathogenic, non-necrogenic species of pseudomonads secreting a yellow-green fluorescent pigment: the siderophore pyoverdine [16]. The genus Pseudomonas comprises mostly fluorescent as well as a few non-fluorescent species in two lineages of intrageneric groups: P. aeruginosa and P. fluorescens. The latter is divided into six groups, with that of P. fluorescens being the most complex and includes nine subgroups [17]. For rapid and efficient species identification, the matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) is also suitable to the genus Pseudomonas [7].
The ecology and function of plant-associated fluorescent pseudomonads have been most extensively studied in the rhizosphere. High densities of these bacteria in field soils can achieve effective natural protection of crop plants against invading pathogens [11, 32]. The suppression of phytopathogens is mainly due to the production of antimicrobial secondary metabolites. Six classes of antibiotic compounds experimentally proven to inhibit fungal and partly bacterial growth are produced by fluorescent pseudomonads: phenazines, phloroglucinols, pyoluteorin, pyrrolnitrin, cyclic lipopeptides, and hydrogen cyanide [6]. The phenolic derivative 2,4-diacetylphloroglucinol (DAPG) has antifungal, antibacterial and antiviral properties, and is therefore, of special relevance in agricultural systems [15].
The antagonistic activity of fluorescent pseudomonads has also been investigated in a few studies in the phyllosphere, e.g. of apple and pears [10, 29]. The potential of fluorescent pseudomonads to control phytopathogenic fusaria is known per se [8, 32, 33]. However, Naik et al. [21] showed that spray inoculation of Pseudomonas fluorescens could also significantly reduce the extent of Alternaria blight of sesame in India. This capability deserves to be investigated thoroughly also under the local conditions of Central Europe, where winter wheat is exposed to both Fusarium and Alternaria fungi and may be contaminated with their mycotoxins [1, 18].
The aim of this study was to examine the potential of a natural population of fluorescent pseudomonads in the phyllosphere of winter wheat to protect crop plants from infestation by Fusarium and Alternaria fungi causing Fusarium head blight and black point diseases. To this end, we tested a representative number of isolates from the phyllosphere of wheat using an in vitro antagonistic dual culture assay and characterized the antagonistic isolates taxonomically by means of MALDI-TOF MS and in part genetically.
Materials and Methods
Strains
Fusarium culmorum 13 (Fc13), Fusarium graminearum 23 (Fg23), Alternaria alternata 219 (Aa219) and Alternaria tenuissima 220 (At220) originating from wheat plants [19] were used as indicators in the dual culture assay. They are stored in the culture collection of microorganisms of the Institute of Landscape Biogeochemistry at the Leibniz-Centre for Agricultural Landscape Research Müncheberg/Germany. The determination of necrotic lesions on leaves, seedling growth inhibition and in vitro mycotoxin production indicated the highly aggressive potential of all four fungal strains [9, 19]. Root-colonizing Pseudomonas protegens CHA0 [25] (formerly Pseudomonas fluorescens CHA0) produces several antagonistic secondary metabolites [22]. It was used as a reference strain for the production of DAPG and its detection in the present study.
Sampling
Leaves of winter wheat (cultivar Meister) were sampled in a field in the Northeast Lowlands of Germany, the Uckermark region, in 2013. At several points in the field, the third leaf on the main tiller of 20 neighbouring plants were removed aseptically and transported in sterile bags on ice. These bulked batches were stored at −20 °C until processed as one sample.
Isolation of Bacteria
Leaves (5 g) were immersed in 100 mL of 1× phosphate-buffered saline, treated for 7 min in an ultrasonic cleaning bath, and subsequently shaken using a linear motion shaker for 20 min at 350 motions per min. The suspensions were diluted in quarter-strength Ringer’s solution and spread-plated on King’s B agar (ROTH, Karlsruhe/Germany). After incubating at 25 °C for 5 days, colonies fluorescing at 366 nm were selected from the agar surface; the isolates were purified twice by streaking on the same agar and then stored in standard I nutrient broth (MERCK, Darmstadt/Germany) and glycerine (1:2, v/v) at −20 °C.
Taxonomic Characterization of Pseudomonas Isolates by MALDI-TOF MS
The isolates were cultured on NAII (SIFIN, Berlin/Germany) for 24 h in preparation for a protein analysis using MALDI-TOF MS according the method of Toth et al. [30]. Briefly, approximately 10 mg of cell mass was suspended in 300 µL water and vortexed to generate a homogeneous suspension. The suspension was mixed with 900 µL ethanol and centrifuged. The pellet was resuspended in 50 µL 70 % formic acid and subsequently carefully mixed with 50 µL acetonitrile. After centrifugation, 1-µL aliquots of the supernatant were placed immediately on spots of a MALDI target. Each spot was allowed to dry and subsequently overlaid with 1 µL of matrix (α-cyano-4-hydroxycinnamic acid in 50 % aqueous acetonitrile containing 2.5 % trifluoroacetic acid). Mass spectra were acquired using a MALDI-TOF MS spectrometer in a linear positive mode (microflexTM LT, Bruker Daltonics, Bremen/Germany) in a mass range of 2–20 kDa. A bacterial test standard (BTS, Bruker Daltonics, Bremen/Germany) was used for instrument calibration. The raw spectra were imported into MALDI Biotyper™ (3.1) software and then processed and analysed using standard pattern matching against the spectra in the MALDI Biotyper™ reference database (version 3.0, Bruker Daltonics, Bremen/Germany). A calculated matching score (LogScore value) provides a measure of the probability of a correct classification.
In Vitro Antagonism Tests
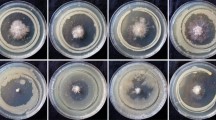
The antagonistic effect of fluorescent pseudomonads against Fusarium and Alternaria indicator strains were tested by dual culturing of the bacteria and fungi on agar as described by Fakhouri and Buchenauer [2]. Tests with the indicators Fc13, Fg23 and At220 were performed using S1 agar [4], and those with Aa219 were performed using Potato Dextrose Agar (ROTH, Karlsruhe/Germany) because this strain is not able to grow on S1 medium. The criterion for antagonistic activity was a distance from the edge of the fungal colony to the bacterial streak (the inhibition zone) of more than 5 mm. The tests were repeated twice.
Screening for the Gene Encoding 2,4-diacetylphloroglucinol (DAPG)
All isolates with antagonistic activity detected in the dual culture assays were tested for the phlD gene, which encodes the antibiotic DAPG. A 1-mL sample of bacteria grown in standard I nutrient broth (MERCK, Darmstadt/Germany) on a rotary shaker for 24 h was centrifuged (8000×g for 20 min). The supernatant was discarded, and DNA was extracted from the bacterial pellet using UltraCleanTM Microbial DNA Isolation Kit (MO BIO Laboratories, Inc., Carlsbad, CA/USA). The presence of the phlD gene was assessed using real-time qPCR with the primers B2BF and BPR4 described by Shizard et al. [28]. The PCR product size and purity were checked by a melting profile and agarose gel electrophoresis.
Extraction of 2,4-diacetylphloroglucinol (DAPG) from Bacterial Cultures and Detection by HPLC
Pseudomonas isolates containing the phlD gene were tested for the production of DAPG on solid medium. The isolates were streaked on S1 agar as described in “In vitro antagonism tests”, but without co-inoculation of a fungal indicator. After incubating at 25 °C for 3 days, the agar was cut into pieces of approximately 5 × 5 mm, packed into 50-mL centrifugation tubes and extracted with 18 mL ethyl acetate (acidified with HCl to pH 1.2) on a linear motion shaker (KS-10, EB Echingen/Germany) for 2.5 h at 350 motions per min. The agar pieces that failed to dissolve were removed from the extract by centrifugation at 7000×g for 12 min. To remove the solvent, the extracts were subjected to a vacuum rotary evaporation at 35 °C. A JASCO HPLC system (Gross-Umstadt/Germany) with a UV-2075 detector, pump PU-1580, autosampler AS 1555-10, degasser and quaternary gradient mixer was used for the qualitative detection of DAPG. Dried extracts were resolved in methanol: water (65:35, v/v). DAPG was separated on a LiChrospher 100 RP 18 column (150 × 4 mm i.d., 5 µm particle size, VDS OptiLab, Berlin/Germany) by isocratic elution using methanol: 0.43 % phosphoric acid (70:30, v/v) at a flow rate of 1 mL min−1 and 25 °C. The DAPG peak was detected by measuring the UV absorbance at 270 nm; the retention time was 6.7 min.
Results
Antagonistic Activity of Pseudomonas Isolates
A total of 40 isolates (23 %) of 175 fluorescent pseudomonads produced sizable inhibition zones against at least one of four fungal phytopathogens (Table 1). Growth inhibition was more pronounced against Alternaria (21 % of 175 bacterial isolates) than against Fusarium (15 %). However, 58 % of the 40 antagonistic isolates were able to suppress at least one strain of both Fusarium and Alternaria. The isolates are listed in Table 1 according to their clusters resulting from the MALDI-TOF MS analyses. There is a notable relationship between these clusters and the patterns of antagonism in the dual culture tests: the isolates of clusters A and B mainly inhibited the growth of the fungal indicator Aa219, whereas most isolates of cluster E and all isolates of cluster F were mainly active against Fc13, Fg23 and At220. The antagonistic activity of the isolates in cluster C was directed against all fungal indicators, whereas that in cluster E was variable.
Detection of the phlD Gene in Antagonistic Isolates and of DAPG in Their Cultures
The gene encoding the antibiotic DAPG was detected in 20 isolates (Table 1), i.e. in 50 % of all antagonists and in 11 % of the total fluorescent Pseudomonas isolates in this study. However, relationships between the patterns of antagonism and the existence of the phlD gene were not evident: isolates 97 and 98 contained the gene and inhibited the growth of all four fungal indicators. In contrast, isolates 8 and 9 did not harbour this gene but were nonetheless antagonistically active. Additionally, several isolates (10, 25, 26, 49, 85, 138, 149, 174, and 179) were found to carry the DAPG-encoding gene, but they showed antagonism only towards the indicator Aa219. Furthermore, the gene was present as well as absent in all the MALDI-TOF MS clusters, yet the HPLC detection of DAPG in the cultures of the isolates carrying the phlD gene was successful for only four of the isolates.
Taxonomic Characterization of Antagonistic Pseudomonas Isolates
A comparison of MALDI-TOF MS spectra obtained for the isolates with reference spectra in the database demonstrated that all isolates belong to the genus Pseudomonas. A proportion of 40 % of the total 40 isolates and one reference strain analysed reached LogScore values between 2.0 and 2.3 for matches with reference spectra, which could be considered as a probable classification at the species level.
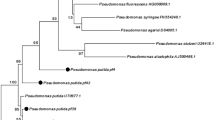
The cluster analysis of the spectral patterns of the investigated isolates revealed grouping into several clusters, which clearly indicated taxonomic differences among the isolates (Fig. 1). Reference spectra of species displaying the closest matches are shown in the dendrogram in Fig. 1. The strains of clusters E and F revealed the highest similarity to one species, whereas the strains of clusters A to C are most likely taxonomically more heterogeneous, as different species were assigned.
Dendrogram, generated by the BioTyper program (version 3.1; Bruker Daltonics, Bremen/Germany), showing the similarities of MALDI-TOF mass spectra of investigated strains; species with most similar spectra of their type strains are given for the formed cluster. Strains highlighted in grey revealed scores considering probable classification at species level
Discussion
Regulating the entire plant-associated microbiome for capitalization on beneficial microbial traits to the hosts should be a goal of future research to develop a more sustainable next-generation agriculture [26]. Pseudomonas bacteria appear to play an important role as native biocontrol agents in soils, in the spermosphere and in the rhizosphere [5, 15, 24] with particular relevance of the antifungal compound DAPG in this context [5, 15].
There is, however, an urgent need to improve our knowledge to manage phyllosphere microbiota in terms of plant health [20]. The present paper provides a complementary thought-provoking step in direction of suppressing FHB and black point in wheat.
The potential antagonistic capacity of the Pseudomonas population investigated in this study with 23 % antagonists and 11 % of the total isolates possessing the phlD gene was comparable with results available from investigations of pseudomonads isolated from the rhizosphere of crop plants [21, 23, 28].
Although the taxonomic affiliation of the isolates based on MALDI-TOF MS has been proven to be an effective identification tool, it does have limitations. In particular, the discrimination of closely related species is difficult, as the analysis of spectral patterns is mainly based on ribosomal proteins. Therefore, this method mirrors the restrictions of identification by 16S rRNA gene analyses, which are due to the high sequence similarity between most of the species in the genus Pseudomonas [17]. Hotta et al. [7] demonstrated that the analysis of special biomarkers based on the ribosomal subunit proteins encoded by the S10–spc–alpha operon can improve the discrimination of Pseudomonas species by MALDI-TOF MS. However, the results obtained in our study enabled the classification of the isolates into phylogenetic groups, as determined by a multi-locus sequence analysis on the basis of four housekeeping genes (16S rRNA gene, gyrB, rpoB and rpoD) by Mulet et al. [17]. The species with highly similar spectra belong to the P. fluorescens subgroup and the closely related P. gessardii subgroup, and therefore, these isolates are also related to these phylogenetic groups. Furthermore, cluster analyses of the spectral patterns allowed classification of the isolates into several groups that differed also in their antagonistic activities (Table 1).
A broad spectrum of antagonistic activity against all the fungal indicators used in this study was especially pronounced in the isolates of clusters C and D, which showed the highest spectral similarity to both P. libanensis and P. extremorientalis and to P. gessardii, respectively (Fig. 1). Conversely, the antagonism of the isolates in clusters A and B, which were most similar to P. orientalis and P. synxantha and to P. antarctica, P. fluorescens and P. gessardii, respectively, was found to be limited to a narrow spectrum of fungi. Cluster D also contains the reference strain Pseudomonas protegens CHA0, which was inhibitory against all the fungal indicators tested and caused the widest inhibition zones (Table 1). This strain is characterized by extraordinarily diverse traits responsible for antagonistic activity [6].
In this study, antagonistic activity was demonstrated in isolates containing the phlD gene but also in others that did not possess this gene. Hence, DAPG is obviously not the only antagonistic compound produced by fluorescent pseudomonads isolated from the wheat phyllosphere. This is in accordance with the results of a study by Shirzard et al. [28] on Pseudomonas isolates from the rhizosphere of various crop plants. The authors detected strains possessing the genes encoding phenazine-1-carboxylic acid, pyoluteorin and DAPG.
DAPG could be detected in only four single cultures of 20 isolates containing the phlD gene spread on S1 agar. The reason for this controversial result is possibly the instability of this substance. The DAPG standard spread alone onto S1 agar was decreased by approximately 95 % in only 4 days at 25 °C. Hence, it appears that the detection of the corresponding gene is more reliable than the biochemical analysis of the substance as performed in this study.
Most of the antagonists in the present study were able to inhibit both Fusarium and Alternaria in vitro. The effects of antagonistic compounds and traits of fluorescent pseudomonads perhaps generally address a broad spectrum of fungi as also indicated in a recent study by Funnel-Harries et al. [3].
The results of the present study suggest the antagonistic potential against fungi causing FHB and black point diseases by fluorescent pseudomonads in the phyllosphere. Further studies are necessary to clarify other antibiotics than DAPG involved in the antagonism, to detect these antibiotics in above-ground plant parts, and to check the antagonistic activity of pseudomonads in vivo. Furthermore, the consequences of the co-occurrence of phytopathogenic fungi and fluorescent pseudomonads on above-ground plant parts in wheat fields with regard to plant health and crop quality should be addressed.
References
Bottalico A, Perrone G (2002) Toxigenic Fusarium species and mycotoxins associated with head blight in small-grain cereals in Europe. Eur J Plant Pathol 108:124–611
Fakhouri W, Buchenauer H (2003) Characteristics of fluorescent pseudomonad isolates towards controlling of tomato wilt caused by Fusarium oxysporum f. sp. lycopersici. J Plant Dis Protect 110:143–156
Funnell-Harris DL, Sattler SE, Pedersen JF (2013) Characterization of fluorescent Pseudomonas spp. associated with roots and soil of two sorghum genotypes. Eur J Plant Pathol 136:469–481
Gould WD, Hagedorn C, Bardinelli TR, Zablotowicz RM (1985) New selective media for enumeration and recovery of fluorescent pseudomonads from various habitats. Appl Environ Microbiol 49:28–32
Haas D, Défago G (2005) Biological control of soil-borne pathogens by fluorescent pseudomonads. Nat Rev Microbiol 3:307–319
Haas D, Keel C (2003) Regulation of antibiotic production in root-colonizing Pseudomonas spp. and relevance for biological control of plant disease. Annu Rev Phytopathol 41:117–153
Hotta Y, Teramoto K, Sato H, Yoshikawa H, Hosoda A, Tamura H (2010) Classification of genus Pseudomonas by MALDI-TOF MS based on ribosomal protein coding in S10–spc–alpha operon at strain level. J Proteome Res 9:6722–6728
Khan MR, Fischer S, Egan D, Doohan FM (2006) Biological control of Fusarium seedling blight disease of wheat and barley. Phytopathol 96:386–394
Korn U, Müller T, Ulrich A, Müller MEH (2011) Impact of aggressiveness of Fusarium graminearum and F. culmorum isolates on yield parameters and mycotoxin production in wheat. Mycotox Res 27:195–206
Kucheryava N, Fiss M, Auling G, Kroppenstedt RM (1999) Isolation and characterization of epiphytic bacteria from the phyllosphere of apple, antagonistic in vitro to Venturia inaequalis, the causal agent of apple scab. Syst Appl Microbiol 22:472–478
Landa BB, Mavrodi OV, Raaijmakers JM, McSpadden Gardener BB, Thomashow LS, Weller DM (2002) Differential ability of genotypes of 2,4-diacetylphloroglucinol-producing Pseudomonas fluorescens strains to colonize the roots of pea plants. Appl Environ Microbiol 68:3226–3237
Legard DE, McQuilken MP, Whipps JM, Fenlon JS, Fermor TR, Thompson IP et al (1994) Studies of seasonal changes in the microbial populations on the phyllosphere of spring wheat as a prelude to the release of a genetically modified microorganism. Agric Ecosyst Environ 50:87–101
Logrieco A, Moretti A, Solfrizzo M (2009) Alternaria toxins and plant diseases: an overview of origin, occurrence and risks. World Mycotoxin J 2:129–140
McMullen M, Bergstrom G, de Wolf E, Dill-Macky R, Hershman D, Shaner G et al (2012) A united effort to fight an enemy of wheat and barley: Fusarium head blight. Plant Dis 96:1712–1728
McSpadden Gardener BB (2007) Diversity and Ecology of biocontrol Pseudomonas spp. in agricultural systems. Phytopathol 97:220–226
Meyer JM, Geoffroy VA, Baida N, Gardan L, Izard D, Lemanceau P et al (2002) Siderophore typing, a powerful tool for the identification of fluorescent and nonfluorescent pseudomonads. Appl Environ Microbiol 68:2745–2753
Mulet M, Lalucat J, García-Valdés E (2010) DNA sequence-based analysis of the Pseudomonas species. Environ Microbiol 12:1513–1530
Müller MEH, Korn U (2013) Alternaria mycotoxins in wheat—a 10 years survey in the Northeast of Germany. Food Control 34:191–197
Müller MEH, Steier I, Köppen R, Siegel D, Proske M, Korn U et al (2012) Cocultivation of phytopathogenic Fusarium and Alternaria strains affects fungal growth and mycotoxin production. J Appl Microbiol 113:874–887
Müller T, Ruppel S (2014) Progress in cultivation-independent phyllosphere microbiology. FEMS Microbiol Ecol 87:2–14
Naik MK, Rajalaxmi K, Rangeshwaran R, Amaresh YS, Reddy MS (2013) Search for 2,4 DAPG positive gene in fluorescent Pseudomonas and their exploitation for sustainable disease management. In: Reddy MS, Ilao RI, Faylon PS, Dar WD, Sayed R, Sudini H et al (eds) Recent advances in biofertilizers and biofungicides (PGPR) for sustainable agriculture. Proceedings of 3rd Asian conference on plant growth-promoting rhizobacteria (PGPR) and other microbials. Manila, Philipines, 21–24 Apr, 2013, pp 152–163
Natsch A, Keel C, Pfirter HA, Haas D, Défago G (1994) Contribution of the global regulator gene gacA to persistence and dissemination of Pseudomonas fluorescens biocontrol strain CHA0 introduced into soil microcosms. Appl Environ Microbiol 60:2553–2560
Picard C, Di Cello F, Ventura M, Fani R, Guckert A (2000) Frequency and biodiversity of 2,4-diacetylphloroglucinol-producing bacteria isolated from the maize rhizosphere at different stages of plant growth. Appl Environ Microbiol 66:948–955
Raaijmakers JM, Vlami M, de Souza JT (2002) Antibiotic production by bacterial biocontrol agents. Antonie Van Leeuwenhoek 81:537–547
Ramettea A, Frapolli M, Fischer-Le Saux M, Gruffaz C, Meyer J-M, Défago G et al (2011) Pseudomonas protegens sp. nov., widespread plant-protecting bacteria producing the biocontrol compounds 2,4-diacetylphloroglucinol and pyoluteorin. Syst Appl Microbiol 34:180–188
Schlaeppi K, Bulgarelli D (2015) The plant microbiome at work. Mol Plant Microbe Interact 28:212–217
Schroth MN, Hildebrand DC, Panopoulos N (1992) Phytopathogenic pseudomonads and related plant-associated pseudomonads. In: Balows A, Trüper HG, Dworkin M, Harder W, Schleifer K-H (eds) The procaryotes, 2nd edn. Springer, New York, pp 3104–3131
Shirzad A, Fallahzadeh-Mamaghani V, Pazhouhandeh M (2012) Antagonistic potential of fluorescent pseudomonads and control of crown and root rot of cucumber caused by Phythophtora drechsleri. Plant Pathol J 28:1–9
Stockwell VO, Jonhson KB, Loper JE (2002) Biological control of fire blight: understanding interactions among introduced and indigenous microbial communities. In: Lindow SE, Hecht-Poinar EI, Elliot VJ (eds) Phyllosphere microbiology. APS, St. Paul, pp 225–239
Tóth EM, Schumann P, Borsodi AK, Kéki Z, Kovács AL, Márialigeti K (2008) Wohlfahrtiimonas chitiniclastica gen. nov., sp. nov., a new γ-proteobacterium isolated from Wohlfahrtia magnifica (Diptera: Sarcophagidae). Int J Syst Evol Microbiol 58:976–998
Vučković JN, Brkljača JS, Bodroža-Solarov MI, Bagi FF, Stojšin VB, Ćulafić JN et al (2012) Alternaria spp. on small grains. Food Feed Res 39:79–88
Weller DM, Landa BB, Mavrodi OV, Schroeder KL, De La Fuente L, Blouin Bankhead S et al (2007) Role of 2,4-diacetylphloroglucinol-producing fluorescent Pseudomonasspp. in the defense of plant roots. Plant Biol 9:4–20
Yoshida S, Ohba A, Liang Y-M, Koitabashi M, Tsushima S (2012) Specificity of Pseudomonas isolates on healthy and Fusarium head blight-infected spikelets of wheat heads. Microb Ecol 64:214–225
Acknowledgments
The authors thank Anne Winding (Aarhus University) for providing Pseudomonas protegens CHA0 and Birgit Wernitz as well as Beate Selch for excellent technical assistance. This work was supported by the Bundesministerium für Verbraucherschutz, Ernährung und Landwirtschaft and the Ministerium für Wissenschaft, Forschung und Kultur des Landes Brandenburg.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Müller, T., Behrendt, U., Ruppel, S. et al. Fluorescent Pseudomonads in the Phyllosphere of Wheat: Potential Antagonists Against Fungal Phytopathogens. Curr Microbiol 72, 383–389 (2016). https://doi.org/10.1007/s00284-015-0966-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-015-0966-8