Abstract
Purpose
Whether age and inter-individual variability of pharmacogenetics are risk factors for paclitaxel-induced peripheral neuropathy (PIPN) is inconclusive. This study was conducted to evaluate the influence of previously investigated single nucleotide polymorphisms (SNPs) and age, using genotype data from a prospective study of paclitaxel-related toxicity in Japanese patients with breast cancer.
Methods
Peripheral blood mononuclear cells from 127 Japanese women with breast cancer who received weekly adjuvant paclitaxel were used to genotypes SLCO1B3 T334G (rs4149117), CYP2C8 A1196G (rs10509681), ABCB1 C1236T (rs1128503), ABCB1 G2677T/A (rs2032582), and ABCB1 C3435T (rs1045642). Genotypic and clinical factors were investigated for associations with PIPN.
Results
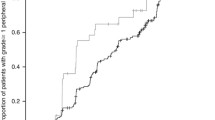
Of the five SNPs evaluated, no SNPs were significantly associated with grade 2 or higher PIPN. However, ABCB1 1236 TT showed a trend to associate with grade 2 or higher PIPN compared to ABCB1 CT/CC (odds ratio 2.1, 95% CI 0.991–4.548, p = 0.051). In subgroup analysis, patients ≥60 years old with an ABCB1 1236 TT had a higher incidence of ≥grade 2 PIPN compared to patients with CT or CC genotype (p = 0.027). On multivariable analysis, age ≥60 years and the ABCB1 1236 TT showed a significant association with ≥grade 2 PIPN (p = 0.005 and p = 0.034, respectively).
Conclusions
ABCB1 1236 TT genotype and older age might be a predictor of PIPN, which diminishes quality of life of cancer survivors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
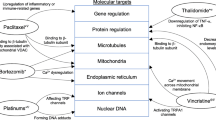
Paclitaxel is a key chemotherapeutic agent in the treatment of early-stage and metastatic breast cancer [1, 2]. Paclitaxel-induced peripheral neuropathy (PIPN) predominantly presents with sensory symptoms such as numbness, tingling, and burning pain in a stocking and glove distribution, without motor symptoms. PIPN may limit the number of paclitaxel doses and may significantly diminish quality of life because it may persist and intensify after completion of chemotherapy. Approximately 50% of patients who developed PIPN recover within 4–6 months, although severe neuropathy may persist for years [3, 4]. The incidence of PIPN depends on several factors: dosage per cycle, treatment schedule, duration of infusion, cumulative dosage, concomitant platinum administration [5], and co-morbidities such as diabetes [6,7,8,9]. In particular, drug exposure is a primary driver of toxicity [10]. Paclitaxel is transported into hepatocytes by solute carrier organic anion transporter family member 1B3 (SLCO1B3) [11], an influx transporter, and is metabolized by cytochrome P450 2C8 (CYP2C8) and cytochrome P450 3A4 (CYP3A4) in the liver [12, 13]. ATP-binding cassette subfamily B member 1 (ABCB1) [14], an efflux transporter, ultimately disposes the metabolite into the bile canaliculi (Fig. 1). Therefore, whether single nucleotide polymorphisms (SNPs) in the genes of these proteins involved in the transport and metabolism of paclitaxel could explain the variability in toxicity has been investigated [15,16,17,18,19,20].
Transport and metabolism of paclitaxel. Paclitaxel is transported into hepatocytes by SLCO1B3, an influx transporter, and is metabolized by CYP2C8 and CYP3A4 in the liver. ABCB1, an efflux transporter, ultimately disposes the metabolite into the bile canaliculi. SLCO1B3 solute carrier organic anion transporter 1B3, CYP2C8 cytochrome P4502C8, CYP3A4 cytochrome P450 3A4, ABCB1 ATP-binding cassette subfamily B1
SLCO1B3 is a polymorphic gene with major SNPs in exon 3 (T334G) and exon 6 (G699A), which are in complete linkage disequilibrium. These SNPs result in a change from serine to alanine at amino acid 112 (S112A) and methionine to isoleucine at amino acid 233 (M233I), respectively [21]. Polymorphic variants in SLCO1B3 exhibit differences in their transport characteristics for several substrates [22], although conflicting data exist regarding the relationship between SLCO1B3 and pharmacokinetics. Smith et al. had reported that SLCO1B3 polymorphisms were not associated with changes in the pharmacokinetics of paclitaxel [23], while van de Steegl et al. had reported that human SLCO1B3 polymorphisms affected the pharmacokinetics of paclitaxel in mice [24]. One of the promising candidates as a risk factor for PIPN is CYP2C8*3. A significant relationship between grade 2 or higher neuropathy and the CYP2C8*3 allele has been reported in both European-American and African-American patient cohorts [15]. The CYP2C8*3 variant (rs11572080 and rs10509681) also leads to decreased paclitaxel metabolism [25] and increased drug exposure [18]. SNPs in ABCB1 (p-glycoprotein, MDR1) have also been associated with neuropathy, since the ABCB1 protein is primarily responsible for excreting agents from the nervous system and back into the systemic circulation. ABCB1 3435 TT variant had significantly higher risk of grade ≥2 neurotoxicity compared to TC and CC genotype [20]. On the other hand, there is a report that ABCB1 3435 TT variant does not explain the substantial inter-individual variability in paclitaxel pharmacokinetics [26]. For the genetic polymorphism in ABCB1 2677G>T/A (rs2032582), a trend of a protective effect for sensory neuropathy was observed in GG wild-type carriers who received paclitaxel and carboplatin [27]. Moreover, ABCB1 1236 polymorphism associated with pharmacokinetics and toxicity in Japanese patients has been reported [28, 29]; however, there are negative reports also regarding associations of ABCB1 1236 and 2677G>T/A with pharmacokinetics and toxicity.
More recently, although genome-wide association studies related to PIPN included variants in TUBB2A, FZD3, FGD4, XKR, EPHA5, and EPHA6 for a possible association with neurotoxicity [30,31,32,33], few results have been replicated in large, independent validation studies. Moreover, pharmacoethnicity in PIPN is reported [34]. However, associations between PIPN and genetic alterations in drug-metabolizing enzymes and transporters have not yet been clearly demonstrated in Asian people. While older age is considered a risk factor for treatment-related toxicity due to age-related changes in drug metabolism, comorbid conditions, and the frequent use of concomitant medications, no definitive associations between PIPN and age have been found [35, 36]. However, PIPN severity was associated with older age in Japanese patients treated with paclitaxel in our previous study [3]. We hypothesized that combination assessment of age and SNPs related to drug exposure would identify the high-risk patients for PIPN. Therefore, we retrospectively evaluated the impact of previously investigated candidate polymorphisms of genes and age on the development of PIPN using genotype data from a prospective study of Japanese breast cancer patients who received paclitaxel as adjuvant therapy.
Materials and methods
Patients
Japanese women with histologically confirmed breast cancer receiving neoadjuvant and/or adjuvant paclitaxel-containing regimens were enrolled in a prospective observational study to evaluate paclitaxel-related toxicity and explore the variants of genes associated with PIPN using the genome-wide association study (GWAS) (UMIN000005294). Eligible patients for this study were required to meet the following criteria: no previous chemotherapy, no history of diabetes, and no preexisting grade 1 or higher neuropathy. All patients provided written informed consent for adjuvant treatment and DNA collection for genetic analysis. The study was conducted in accordance with the Declaration of Helsinki and approved by the local institutional review board (protocol number 2013-199).
Treatment
Patients were pretreated with 4 cycles of cyclophosphamide (600 mg/m2) and doxorubicin (60 mg/m2) (AC) or cyclophosphamide (500 mg/m2) with epirubicin (100 mg/m2) and 5-fluorouracil (500 mg/m2) (CEF) every 3 weeks. Paclitaxel (80 mg/m2) was subsequently administered weekly as a 1-h infusion for 12 weeks. Patients who were positive for human epidermal growth factor receptor 2 (HER2) by immunohistochemistry or gene amplification (in situ hybridization) received trastuzumab concurrently with paclitaxel (weekly or every 3 weeks).
PIPN evaluation
Neuropathy was prospectively evaluated at baseline before paclitaxel treatment, at week 7, within 3 weeks after the final dose, and 1 year after the last dose of paclitaxel, based on the National Cancer Institute Common Terminology Criteria for Adverse Events 4.0 [37]. We decided four assessment points for PIPN assessment, based on our previous study, which showed that median time to onset of PIPN was 28 days. Moreover, 40% of patients showed persistent PIPN at 1 year after completion of treatment [3].
Genotyping
A 10-mL blood sample was collected from each patient at enrollment. Genotyping was performed for eligible patients who had sufficient DNA in the sample. DNA was extracted, and the concentration was fixed at 10 ng/μL. Blinded genotyping was performed using the i-densy™ genetic testing platform (ARKRAY, Inc., Kyoto, Japan) and the quenching probe system (QP-system) based on the principles of mutant detection. The i-densy™ automatically performs polymerase chain reaction (PCR), and mutant detection using extracted DNA within 60 min as previously described [38]. We used quenching probes (QProbe, Nippon Steel Kankyo Engineering Co., Ltd., Tokyo, Japan).
The SNPs that we analyzed were SLCO1B3 T334G (rs4149117), CYP2C8 A1196G (rs10509681), and ABCB1 C1236T (rs1128503), G2677T/A (rs2032582), and C3435T (rs1045642). The genotypes were categorized as wild type, heterozygous, or homozygous variant. For ABCB1 G2677T/A, each patient was classified as having ABCB1 homozygous (TT) variant, heterozygous (GT or AT), or homozygous (GG) wild type. QP-system cannot distinguish between GT and AT as haplotype of ABCB1 G2677T/A. The primers are shown in Supplementary Table 1. The fluorescent pigments that were used are TAMRA, PACIFIC BLUE, and BODIPY FL.
Genotyping was performed blinded to neuropathy data. For quality control, cutoff value for exclusion was set at 0.95 for call rate by SNP and patient. Hardy–Weinberg equilibrium (HWE) was also assessed to detect possible genotyping issues.
Statistical analysis
The primary objective was to evaluate the association between the grade of PIPN, age, and the variants of SLCO1B3, CYP2C8, and ABCB1. The SLCO1B3, CYP2C8, and ABCB1 genotype frequencies were assessed for concordance with expectations under the HWE with the threshold for significant deviation from theoretical distribution being set at p = 0.05, using the Chi-square test and Fisher’s exact test (degree of freedom = 1). All analyses were performed using the highest grade of treatment-related sensory peripheral neuropathy. Patients in whom paclitaxel was discontinued due to adverse events, except neuropathy, were excluded from the analyses. The univariate association between genotype and severity of PIPN on genotypic or recessive model was assessed using Chi-square test. For the tri-allelic SNP G2677T/A in ABCB1, patients carrying an A allele were analyzed as a T allele because of QP-system treating the A allele as a T allele. Multivariable logistic regression analysis was used to identify the risk factors for grade 2 or higher maximum PIPN at any time after the administration of paclitaxel. Pre-treatment regimen (AC vs. CEF), body surface area (BSA) (<1.5 vs. ≥1.5 m2), and age (<60 vs. ≥60 years) were included as clinical covariates. We pre-planned to dichotomize age at 60 years and compare PIPN, referring to our previous study that showed significant association between age ≥60 years and duration of PIPN [3]. Moreover, we pre-planned to dichotomize BSA by 1.5 and compare PIPN, referring to the administration method of S-1 therapy for Japanese cancer patients [39]. A two-sided P value of <0.05 was considered statistically significant, and all analyses were performed using SAS software (version 9.2; SAS Institute, Cary, NC, USA).
Results
Patient characteristics
One hundred and sixty-two of 197 Japanese women enrolled between February 2011 and October 2013 at two study sites were analyzed. Thirty-five patients were excluded from the analysis including patients treated with docetaxel (n = 20), patients lost to follow-up (n = 9), patients in whom paclitaxel was discontinued due to adverse events other than peripheral neuropathy (n = 3), and patients with a history of prior paclitaxel usage (n = 3). Patient characteristics are listed in Table 1. The median age was 50 years (range 25–75 years), and 33 patients were ≥60 years old (26%).
Distribution of the SLCO1B3, CYP2C8, and ABCB1 genotypes
Genotyping call rates by SNP and patient were estimated. It was found that the both call rates (by SNP and patient) were >0.95, and all the SNPs and samples were included in analyses. G2677T/A genotypes were confirmed to be in HWE (χ 2 = 0.475, p = 0.491 for SLCO1B3 T334G, χ 2 = 0.388, p = 0.533 for ABCB1 C1236T, χ 2 = 0.239, p = 0.625 for ABCB1 C3435T, and χ 2 = 0.385, p = 0.535 for ABCB1 G2677T/A). The genotype distributions were all in HWE except CYP2C8 A1196G, for which genotypes of all of the patients were homozygous for the major allele, and thus, statistical analyses were not conducted. Minor allele frequencies (MAF) were 0.244 for SLCO1B3 T334G, 0.413 for ABCB1 C1236T, 0.394 for ABCB1 C3435T, and 0.350 for ABCB1 G2677T/A.
Age and grade of PIPN
Among the 127 patients who were treated with weekly paclitaxel (median cumulative dose 933 mg/m2, range 560–960 mg/m2), 66 (52%) patients experienced grade 2 or higher PIPN. Grade 2 or higher PIPN was significantly associated with an age ≥60 years compared to age <60 years [odds ratio 3.30, 95% confidence interval (CI) 1.39–7.86, p = 0.006].
Association between SNPs and PIPN
None of the four SNPs were significantly associated with grade 2 or higher PIPN in the genotypic model (Table 2). However, ABCB1 1236 TT showed a trend to associate with grade 2 or higher PIPN compared to ABCB1 CT/CC in the recessive model. (odds ratio 2.1, 95% CI 0.991–4.548, p = 0.051) (Supplementary Table 2). For patients who were ≥60 years old, ABCB1 1236 TT was significantly associated with PIPN (p = 0.027). In contrast, no association was observed for ABCB1 1236 TT among patients who were <60 years old (Table 3).
Multivariable analyses
The clinically relevant covariates [SNPs, age, pre-treatment regimen (CEF vs. AC), and body surface area (<1.5 vs. ≥1.5 m2)] were included in logistic regression analysis. The age of 60 years (odds ratio 3.652, 95% CI 1.468–9.085, p = 0.005) and ABCB1 1236 TT (odds ratio 2.404, 95% CI 1.067–5.419, p = 0.034) were identified as independent risk factors associated with grade 2 or higher PIPN. The grade of PIPN in patients with the SLCO1B3 334 TT tended to be higher than in those with SLCO1B3 334 GG or GT (p = 0.052) (Table 4).
Discussion
We retrospectively evaluated the influence of candidate genes with functions in either transport or metabolism of paclitaxel and PIPN and age on the development of PIPN for breast cancer patients treated with paclitaxel, using prospectively collected data. Multivariable analysis revealed that the ABCB1 1236 TT variant and patient age of 60 years were significantly associated with grade 2 or higher PIPN. In addition, there was a trend for SLCO1B3 334 TT to be associated with an increased risk of PIPN. To the best of our knowledge, the present study is the first study for Japanese patients to demonstrate that older age and SNPs are associated with an increased severity of PIPN.
Our findings indicate that age ≥60 years was the most significant risk factor for PIPN. Prior studies have given conflicting results regarding the pharmacokinetics of paclitaxel in elderly patients [40]. Multiple factors may contribute to a decrease in drug clearance in the liver of older individuals [41]. Liver size and hepatic blood flow are 25–35% lower in older individuals compared to younger adults. Older individuals also exhibit decreased bile flow [42]. Moreover, the specific cytochrome P450 (CYP) contents of the liver decrease with aging [43].
In addition to age, candidate genes in the pharmacokinetic and pharmacodynamic pathways for paclitaxel appear to play a role in the development of PIPN. ABCB1 1236 TT and SLCO1B3 TT were identified as risk factors for PIPN in our study. However, ABCB1 variants have been reported to be associated with both increased and decreased risks of PIPN previously [19, 44]. Moreover, there is no report of the association between a SLCO1B3 genotype and PIPN although it is associated with clinical effect and toxicity except for PIPN. Most previous pharmacogenetic studies were conducted in a small number of patients with different tumor types undergoing a variety of treatment regimens, and differences in the paclitaxel dosing schedule may partially explain the conflicting results. CYP2C8*3 variants have been most frequently reported as being positively associated with an increased risk of neurotoxicity [15]. However, no patients had CYP2C8*3 in the present study. The CYP2C8*3 variant occurs primarily in Caucasian individuals with a frequency of approximately 0.13, while most Japanese individuals do not have the CYP2C8*3 variant [45]. Considering these ethnic differences in SNP distribution, it is reasonable to identify the risk factors, except for CYP2C8*3, in Japanese population.
There are some limitations in this study. First, QP-system is designed not to distinguish between 2677G and 2677A. However, because the heterozygous genotype can be specified and frequency of 2677A is low, its influence is considered small. Moreover, we consider that QP-system is potentially convenient in clinic because of its fast turnaround time. Second, in whole analysis, the current study had 90% power to detect the differences between groups using Chi-squared test at a two-sided α error of 0.05. In subgroup analysis (n = 33; patients aged ≥60 years), it had 67% power at a two-sided α error of 0.2. Therefore, it is underpowered in the subgroup population. Additionally, no adjustments were made for multiplicity of inferences.
In conclusion, PIPN was associated with age ≥60 years and ABCB1 1236 TT in Japanese female breast cancer patients who received paclitaxel as adjuvant treatment for primary breast cancer. Optimal dosing of paclitaxel in elderly patients warrants further investigation to mitigate the risk of PIPN. Although our data are not yet sufficiently conclusive to clinically recommend genetic tests to predict neurotoxicity, we believe that combination assessment of older age and ABCB1 1236 variant may help to identify high-risk patients for PIPN.
References
Mamounas EP, Bryant J, Lembersky B et al (2005) Paclitaxel after doxorubicin plus cyclophosphamide as adjuvant chemotherapy for node-positive breast cancer: results from NSABP B-28. J Clin Oncol 23:3686–3696
De Laurentiis M, Cancello G, D’Agostino D et al (2008) Taxane based combinations as adjuvant chemotherapy of early breast cancer: a meta-analysis of randomized trials. J Clin Oncol 26:44–53
Tanabe Y, Hashimoto K, Shimizu C et al (2013) Paclitaxel-induced peripheral neuropathy in patients receiving adjuvant chemotherapy for breast cancer. Int J Clin Oncol 18:132–138
Hershman DL, Weimer LH, Wang A et al (2011) Association between patient reported outcomes and quantitative sensory tests for measuring long-term neurotoxicity in breast cancer survivors treated with adjuvant paclitaxel chemotherapy. Breast Cancer Res Treat 125:767–774
Muggia FM, Braly PS, Brady MF et al (2000) Phase III randomized study of cisplatin versus paclitaxel versus cisplatin and paclitaxel in patients with suboptimal stage III or IV ovarian cancer: a gynecologic oncology group study. J Clin Oncol 18:106–115
Grisold W, Cavaletti G, Windebank AJ (2012) Peripheral neuropathies from chemotherapeutics and targeted agents: diagnosis, treatment, and prevention. Neurol Oncol 14(Suppl 4):iv45–iv54
Lee JJ, Swain SM (2006) Peripheral neuropathy induced by microtubule-stabilizing agents. J Clin Oncol 24:1633–1642
Nabholtz JM, Gelmon K, Bontenbal M et al (1996) Multicenter, randomized comparative study of two doses of paclitaxel in patients with metastatic breast cancer. J Clin Oncol 14:1858–1867
Seidman AD, Berry D, Cirrincione C et al (2008) Randomized phase III trial of weekly compared with every-3-weeks paclitaxel for metastatic breast cancer, with trastuzumab for all HER-2 overexpressors and random assignment to trastuzumab or not in HER-2 nonoverexpressors: final results of Cancer and Leukemia Group B protocol 9840. J Clin Oncol 26:1642–1649
van Gerven JM, Moll JW, van den Bent MJ et al (1994) Paclitaxel (Taxol) induces cumulative mild neurotoxicity. Eur J Cancer 30:1074–1077
Smith NF, Figg WD, Sparreboom A (2005) Role of the liver-specific transporters OATP1B1 and OATP1B3 in governing drug elimination. Expert Opin Drug Metab Toxicol 1:429–445
Rahman A, Korzekwa KR, Grogan J et al (1994) Selective biotransformation of taxol to 6 alpha-hydroxytaxol by human cytochrome P450 2C8. Cancer Res 54:5543–5546
Harris JW, Rahman A, Kim BR et al (1994) Metabolism of taxol by human hepatic microsomes and liver slices: participation of cytochrome P450 3A4 and an unknown P450 enzyme. Cancer Res 54:4026–4035
Sparreboom A, van Asperen J, Mayer U et al (1997) Limited oral bioavailability and active epithelial excretion of paclitaxel (Taxol) caused by P-glycoprotein in the intestine. Proc Natl Acad Sci USA 4:2031–2035
Hertz DL, Roy S, Motsinger-Reif AA et al (2013) CYP2C8*3 increases risk of neuropathy in breast cancer patients treated with paclitaxel. Ann Oncol 24:1472–1478
Sissung TM, Mross K, Steinberg SM et al (2006) Association of ABCB1 genotypes with paclitaxel mediated peripheral neuropathy and neutropenia. Eur J Cancer 42:2893–2896
Leskelä S, Jara C, Leandro-García LJ et al (2011) Polymorphisms in cytochromes P450 2C8 and 3A5 are associated with paclitaxel neurotoxicity. Pharmacogenomics J 11:121–129
Bergmann TK, Brasch-Andersen C, Gréen H et al (2011) Impact of CYP2C8*3 on paclitaxel clearance: a population pharmacokinetic and pharmacogenomic study in 93 patients with ovarian cancer. Pharmacogenom J 11:113–120
Abraham JE, Guo Q, Dorling L et al (2014) Replication of genetic polymorphisms reported to be associated with taxane-related sensory neuropathy in patients with early breast cancer treated with Paclitaxel. Clin Cancer Res 20:2466–2475
Kus T, Aktas G, Kalender ME et al (2016) Polymorphism of CYP3A4 and ABCB1 genes increase the risk of neuropathy in breast cancer patients treated with paclitaxel and docetaxel. Onco Targets Ther 9:5073–5080
Tsujimoto M, Hirata S, Dan Y et al (2006) Polymorphisms and linkage disequilibrium of the OATP8 (OATP1B3) gene in Japanese subjects. Drug Metab Pharmacokinet 21:165–169
Letschert K, Keppler D, Konig J (2004) Mutations in the SLCO1B3 gene affecting the substrate specificity of the hepatocellular uptake transporter OATP1B3 (OATP8). Pharmacogenetics 14:441–452
Smith NF, Marsh S, Scott-Horton TJ et al (2007) Variants in the SLCO1B3 gene: interethnic distribution and association with paclitaxel pharmacokinetics. Clin Pharmacol Ther 81:76–82
van de Steeg E, van Esch A, Wagenaar E et al (2013) Influence of human OATP1B1, OATP1B3, and OATP1A2 on the pharmacokinetics of methotrexate and paclitaxel in humanized transgenic mice. Clin Cancer Res 19:821–832
Dai D, Zeldin DC, Blaisdell JA et al (2001) Polymorphisms in human CYP2C8 decrease metabolism of the anticancer drug paclitaxel and arachidonic acid. Pharmacogenetics 11:597–607
Henningsson A, Marsh S, Loos WJ et al (2005) Association of CYP2C8, CYP3A4, CYP3A5, and ABCB1 polymorphisms with the pharmacokinetics of paclitaxel. Clin Cancer Res 11:8097–8104
Gréen H, Söderkvist P, Rosenberg P et al (2009) Pharmacogenetic studies of Paclitaxel in the treatment of ovarian cancer. Basic Clin Pharmacol Toxicol 104:130–137
Fujiwara Y, Hamada A, Mizugaki H et al (2016) Pharmacokinetic profiles of significant adverse events with crizotinib in Japanese patients with ABCB1 polymorphism. Cancer Sci 107:1117–1123
Hamada A, Sasaki J, Saeki S et al (2012) Association of ABCB1 polymorphisms with erlotinib pharmacokinetics and toxicity in Japanese patients with non-small-cell lung cancer. Pharmacogenomics 13:615–624
Baldwin RM, Owzar K, Zembutsu H et al (2012) A genome-wide association study identifies novel loci for paclitaxel-induced sensory peripheral neuropathy in CALGB 40101. Clin Cancer Res 18:5099–5109
Leandro-García LJ, Inglada-Pérez L, Pita G et al (2013) Genome-wide association study identifies ephrin type A receptors implicated in paclitaxel induced peripheral sensory neuropathy. J Med Genet 50:599–605
Njiaju UO, Gamazon ER, Gorsic LK et al (2012) Whole-genome studies identify solute carrier transporters in cellular susceptibility to paclitaxel. Pharmacogenet Genom 22:498–507
Schneider BP, Li L, Radovich M et al (2015) Genome-wide association studies for taxane-induced peripheral neuropathy in ECOG-5103 and ECOG-1199. Clin Cancer Res 21:5082–5091
Komatsu M, Wheeler HE, Chung S et al (2015) Pharmacoethnicity in paclitaxel-induced sensory peripheral neuropathy. Clin Cancer Res 21:4337–4346
Argyriou AA, Polychronopoulos P, Koutras A et al (2006) Is advanced age associated with increased incidence and severity of chemotherapy-induced peripheral neuropathy? Support Care Cancer 14:223–229
Lichtman SM, Hurria A, Cirrincione CT, Cancer and Leukemia Group B et al (2012) Paclitaxel efficacy and toxicity in older women with metastatic breast cancer: combined analysis of CALGB 9342 and 9840. Ann Oncol 23:632–638
Chen AP, Setser A, Anadkat MJ et al (2012) Grading dermatologic adverse events of cancer treatments: the Common Terminology Criteria for Adverse Events Version 4.0. J Am Acad Dermatol 67:1025–1039
Suzuki S, Komori M, Hirai M et al (2012) Development of a novel, fully-automated genotyping system: principle and applications. Sensors (Basel) 12:16614–16627
Kubota T (2008) The role of S-1 in the treatment of gastric cancer. Br J Cancer 98:1301–1304
Biganzoli L, Licitra S, Moretti E et al (2009) Taxanes in the elderly: can we gain as much and be less toxic? Crit Rev Oncol Hematol 70:262–271
Ginsberg G, Hattis D, Russ A, Sonawane B (2005) Pharmacokinetic and pharmacodynamic factors that can affect sensitivity to neurotoxic sequelae in elderly individuals. Environ Health Perspect 113:1243–1249
Zeeh J, Platt D (2002) The aging liver: structural and functional changes and their consequences for drug treatment in old age. Gerontology 48:121–127
Sotaniemi EA, Arranto AJ, Pelkonen O et al (1997) Age and cytochrome P450-linked drug metabolism in humans: an analysis of 226 subjects with equal histopathologic conditions. Clin Pharmacol Ther 61:331–339
Chang H, Rha SY, Jeung HC et al (2009) Association of the ABCB1 gene polymorphisms 2677G>T/A and 3435C>T with clinical outcomes of paclitaxel monotherapy in metastatic breast cancer patients. Ann Oncol 20:272–277
The International HapMap Consortium (2003) The international hapmap project. Nature 426:789–796
Acknowledgements
This work was supported by a Scientific Research Grant of the Ministry of Health, Labor, and Welfare (H21-021), and the National Cancer Center Research and Development Fund (23-A-30, 26-A-20). We thank Ms. Masayo Kawamura, Ms. Nao Nakamura, Ms. Kiyomi Nonogaki, and Ms. Hitomi Sato for helping with the data collection.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
YF reports grants from Taiho Pharmaceutical Co. Ltd, grants from Takeda Pharmaceutical Company Ltd, grants from Takeda Bio Development Center Ltd, grants and other from Chugai Pharmaceutical Co Ltd, other from Astra Zeneca KK, other from Eisai Co Ltd, other from Daiichi Sankyo Co Ltd, other from Sanofi-Aventis KK, grants and other from Eli Lilly Japan KK, other from Yakult Honsha Co Ltd, other from NEC Corporation, outside the submitted work. CS reports grants from Eli Lilly Japan KK and Pfizer KK. KH is currently an employee at Chugai Pharmaceutical Europe. The other authors have no conflicts of interest to declare.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committee and with the 1964 Declaration of Helsinki.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tanabe, Y., Shimizu, C., Hamada, A. et al. Paclitaxel-induced sensory peripheral neuropathy is associated with an ABCB1 single nucleotide polymorphism and older age in Japanese. Cancer Chemother Pharmacol 79, 1179–1186 (2017). https://doi.org/10.1007/s00280-017-3314-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00280-017-3314-9