Abstract
Lung cancer is the leading cause of cancer death worldwide. Survival is largely dependent on the stage of diagnosis: the localized disease has a 5-year survival greater than 55%, whereas, for spread tumors, this rate is only 4%. Therefore, the early detection of lung cancer is key for improving prognosis. In this study, we present an innovative, non-invasive, cancer detection approach based on measurements of the metabolic activity profiles of immune system cells. For each Liquid ImmunoBiopsy test, a 384 multi-well plate is loaded with freshly separated PBMCs, and each well contains 1 of the 16 selected stimulants in several increasing concentrations. The extracellular acidity is measured in both air-open and hermetically-sealed states, using a commercial fluorescence plate reader, for approximately 1.5 h. Both states enable the measurement of real-time accumulation of ‘soluble’ versus ‘volatile’ metabolic products, thereby differentiating between oxidative phosphorylation and aerobic glycolysis. The metabolic activity profiles are analyzed for cancer diagnosis by machine-learning tools. We present a diagnostic accuracy study, using a multivariable prediction model to differentiate between lung cancer and control blood samples. The model was developed and tested using a cohort of 200 subjects (100 lung cancer and 100 control subjects), yielding 91% sensitivity and 80% specificity in a 20-fold cross-validation. Our results clearly indicate that the proposed clinical model is suitable for non-invasive early lung cancer diagnosis, and is indifferent to lung cancer stage and histological type.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is the third most common cancer diagnosed, but has a higher mortality rate than breast, prostate, and colon cancer combined [1]. Furthermore, since more than half of patients are diagnosed with locally advanced or metastatic disease, and despite advances in treatment, the long-term survival from lung cancer currently remains low [2]. Therefore, significant efforts are being made to make screening and early diagnosis of lung cancer possible, to allow the early treatment and to improve survival [3]. The current recommended method for lung cancer screening and early diagnosis by the US Preventive Services Task Force (USPSTF) is chest low-dose computed tomography (LDCT) in a high-risk population. This recommendation was based on The National Lung Screening Trial (NLST), which demonstrated that scanning with LDCT led to a 20% reduction in mortality rate in this high-risk population [4]. However, LDCT screening has many limitations, including radiation exposure, high false positive rates, and overdiagnosis [5, 6]. In addition, the target population of the USPSTF’s recommendation represents only about 11% of the 94 million former and current smokers in the US [7]. These recommendations also do not take into consideration other high-risk populations, such as patients with chronic obstructive pulmonary disease (COPD), of which about 2.2% develop lung cancer per year [8]. Indeed, there is an urgent need for other non-invasive diagnostic methods, or biomarkers with high accuracy, which might promote the earlier detection of lung cancer, resulting in more efficacious therapeutic interventions, and a higher likelihood of cure.
Liquid biopsy is a new strategy for the non-invasive detection of cancer using body fluids, mainly blood samples [9,10,11,12,13]. Several studies include a quantitative analysis to investigate the role of circulating cell-free tumor DNA, or non-coding RNA, in lung cancer diagnosis [14]. Some methods utilize machine learning to assist the diagnostic process, training a classifier on the covariates obtained from liquid biopsy [15,16,17]. Many challenges remain in this approach, including low frequency of secreted tumor components in blood, their short half-life, cell/DNA fragmentation, high variation in tumor cell mutation, and the incapability to determine tumor origin [18, 19]. Importantly, the current methods detect malignancies mostly in their advanced stages, in which treatment is considerably less effective [18, 20]. In contrast, the liquid immunobiopsy diagnostic platform presented in this study enables a ‘functional’ test that determines the different metabolic activity profiles (MAPs) of PBMCs in response to various antigenic and mitogenic stimulants, which enables us to detect lung cancer at its early stages.
Cancer cells, in contrast to normal differentiated cells, primarily rely on aerobic glycolysis (glycolysis in the presence of oxygen), rather than on mitochondrial oxidative phosphorylation (OXPHOS), a phenomenon discovered almost a century ago and termed “the Warburg effect” [21,22,23]. About 60 years later, a similar paradigm was also found in the immune system. While resting, naive T cells utilize mostly OXPHOS processes for cell maintenance, activated T cells (effector T cells) undergo a metabolic switch from OXPHOS to aerobic glycolysis, characterized by overexpression of glucose transporters [24,25,26,27]. This metabolic switch is essential not only for proliferation of activated T cells, but also for proper acquisition of effector functions [28]. A similar switch from respiration to aerobic glycolysis was observed in both monocytes and B cells during activation, characterized by decreased oxygen consumption [29, 30]. One important finding was that the metabolism of T cells directly influences the expression of PD-1, a negative regulator of T cell function [28]. Moreover, T cell metabolism has a significant implication in immunotherapy, which relies on metabolic profile modification to improve effector function. For example, T cells that receive signals via T cell-inhibitory receptor PD-1 undergo diminished glycolysis and enhanced fatty acid β-oxidation, which dampens their effector function while promoting longevity [31]. The previous studies have shown a link between certain diseases and changes in immunometabolism, which were observable in-vitro in immune cells [32, 33]. Furthermore, studies show that various diseases and physiological conditions result in specific changes in the metabolic profiles of immune cells for each disease [34].
Besides protecting the host from virus-induced tumors and the elimination of pathogens for the effective resolution of inflammation and prevention of potential tumorigenesis, the immune system plays a critical role in specifically identifying and eliminating cancer cells, based on their expression of tumor-specific antigens or alternatively molecules excreted by cellular stress of healthy tissues, before they cause harm to the host, a process usually referred as tumor immune surveillance. The idea that the immune system has the capability to specifically recognize and destroy tumor cells was underlined more than a century ago and extensively reviewed recently [35, 36]. In spite of the immune surveillance mechanisms, tumors happen to develop even in the presence of a functioning immune system, and thus, the recently adapted concept of tumor ‘immunoediting’ better describes the role of the immune system during tumor development, which consists of three phases: elimination, equilibrium, and escape [37]. The elimination phase can be complete when all tumor cells are eradicated or incomplete when tumor cells are partially eliminated. In the latter case, a state of equilibrium develops between the immune system and the developing tumor, which remains dormant or continues to accumulate further DNA mutations. The action of the immune system during this phase may be sufficient to control tumor progression, but if it fails to completely eliminate the tumor, the process leads to resistant tumor cell variants leading to the escape phase, in which the immune system is not able to control the tumor growth. During all phases of the immunoediting process, there is a deep and continuous involvement of the immune system, thus differing from the healthy non-cancerous state activity.
Based on these key findings, we initially hypothesized that changes in the body’s physiological function, due to the presence of cancer, may be eventually reflected in different MAPs of PBMCs in response to various antigenic and mitogenic stimulants. Measurement of the MAPs is performed by monitoring and extracting the accumulated acidification by proxy of fluorescently measured pH changes in the extracellular environment of the cells. The resulting detected differences in the acidification rate between lung cancer and control (non-lung-cancer) samples can ultimately be attributed to the differences in PBMCs subpopulations and prevalence [38, 39]. In this work, we present the results of the first diagnostic accuracy study of lung cancer detection using liquid immunobiopsy.
Materials and methods
Study design, demographics, and protocols
Subjects were enrolled between June 2014 and December 2016 in three medical centers. In all cases, the study received Helsinki approval, and subjects read and signed a dedicated consent form. Inclusion criteria included 18 ≤ age ≤ 90 years, no pregnancy and no treatment for lung cancer prior to blood withdrawal. Exclusion criteria included treatment for any type of malignancy in prior 5 years, clinically determined active infection or inflammation, treatment with medication that can affect the immune system, lactation or ongoing fertility treatment, or any of the following conditions: HIV positive, hepatitis B/C, autoimmune disease, hypersensitivity, and/or allergy that cannot be avoided. Lung cancer and control (non-lung-cancer) subjects were enrolled in parallel, the reference standard for lung cancer being biopsy or surgery. Once the number of lung cancer subjects reached 100, they were matched with control subjects via an automated process to obtain a balanced 1:1 cohort, with an optimal matching of age, gender, and COPD distributions, resulting in a total sample size of 200, as described in the Cohort Construction subsection of the results.
Collection, separation, and counting of PBMCs
Blood samples were collected in 9 ml Vacutubes with EDTA (Greiner Bio-One 455,036). For a high viability rate of the blood cells, the samples were transported in thermo-stated containers set to 18 °C until PBMCs separation. Fresh PBMCs were isolated by Lymphoprep™ kit, according to the manufacturer’s instructions (Axis-Shield). Total cell numbers were counted using a hemocytometer. Median cell viability was 99.5%.
MAP test preparation and measurement
Each well in a black non-binding, low-volume 384 multi-well plate (Greiner Bio-One) was loaded with 10 µl of the PBMCs solution and 10 µl of 10 mM PBS containing 8-Hydroxypyrene-1,3,6-trisulfonic acid (HPTS, Sigma-Aldrich Ltd.), and including 1 of the 16 stimulating reagents (stimulants) in increasing concentrations (Table 1). The final concentration of the HPTS probe in each well was 0.5 µM, and the final concentration of the PBMCs was 5⋅106 cells/ml in 10 mM PBS. Buffer capacity was specifically matched to allow for pH changes to occur as a result of PBMCs metabolic activity. Each well was seeded with 5⋅106 cells/ml, to reflect the average PBMC concentration in adult peripheral blood. The samples were loaded in triplicates, first PBMC samples, followed by stimulants, to obtain a final volume of 20 µl in each well. Furthermore, each test included two controls: one containing only the fluorescent HPTS probe, without cells and without stimulants; the other containing the HPTS probe with cells but without stimulants, which represents the ‘basal state’. The acidification process was monitored for approximately 1.5 h at 37 °C by a commercial fluorescence plate reader (TECAN Infinite M200/ F200; application: Tecan i-control 1.10.4.0, 1.11.1.0, 1.12.4.0). First, the reader monitored the acidification process without a plate seal (‘open’ state), and then, the multi-well plate was sealed hermetically (ThermalSeal RT™, Excel Scientific, Inc.) to avoid ventilation of CO2 and NH3 for the second phase of the test (‘closed’ state). Both states enable the measurement of real-time accumulation of ‘soluble’ versus ‘volatile’ metabolic products. The fluorescence intensities were measured at 513 nm under sequential excitation at wavelengths of 455 and 403 nm. Median time between blood withdrawal and start of scan was 4 h (range 2–9 h).
Measurement of pH using HPTS fluorescent probe
The fluorescent probe HPTS is a non-toxic, membrane-impermeant pH indicator, with a pKa of ~ 7.3 in aqueous buffers. HPTS exhibits a pH-dependent absorption shift, allowing ratio-metric pH measurements as a function of the ratio between the fluorescence intensities measured at a wavelength of 513 nm, under excitation at wavelengths of 455 and 403 nm sequentially. This method is essential for our sensitive measurements of minor pH changes in the physiological range around pH 7. The calibration curve used in the MAP test comprised PBS solutions containing 0.5 µM HPTS and titrated with an acid or base to obtain several pH levels, as measured by a pH glass electrode. The pH measurements and the fluorescence measurements of the titrated samples were carried out at 37 °C. A calibration curve was constructed for both ‘open’ and ‘closed’ plate states, allowing pH measurement as a function of the ratio between the two above-mentioned excitation wavelengths. The concentration of H+ (acidity) in nM units is then derived using the formula: 109−pH.
Type and preparation of stimulants
In each test, the metabolic activity profiles of PBMCs were monitored in the basal state (in the absence of stimulant reagents), and under the influence of either a stimulant, a nutrient or an inhibitor (all referred to as ‘stimulants’ in this paper) as detailed in Table 1. Each stimulant was diluted in PBS to obtain several different concentrations. The selection of stimulants was made by their relation to the immune system, lung cancer, or cancers in general.
Data analysis
All demographic and clinical information, as well as the raw MAP test data, were stored in a secure and dedicated PostgreSQL database. Data analysis was performed using Python.
At the end of the biological analysis, each subject was assigned a datasheet containing raw fluorescent readings of plate wells as a function of time for both ‘open’ and ‘closed’ plate states. The fluorescent readings were transformed into pH values using a calibration curve.
Machine learning was preformed using either decision trees or support vector machines (SVMs), implemented by the publically available scikit-learn Python library [40]. For cross-validation, a stratified k-fold was used. Bagging was executed by training an ensemble of simple classifiers (decision trees or SVMs) on 31 random subsets of the cohort, followed by averaging their binary predictions. When combining predictions from multiple stimulants, the separate predictions of all stimulants were averaged, as well. This yielded a prediction score in the range [0, 1], such that a score > 0.5 means that more than half of the simple classifiers predicted a “lung cancer” clinical status. The scores were scaled to the range [− 10, 10] for easier interpretation. A receiver-operating characteristic (ROC) curve was built using these prediction scores.
Confidence intervals were calculated using the normal approximation for the binomial confidence interval. Significance of differences between subpopulations was estimated using the χ2 test.
Results
Cohort construction
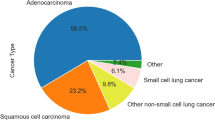
A cohort of 200 subjects was compiled by age- and sex-matching 100 lung cancer subjects with 100 control subjects (Table 2). The control group included both healthy individuals and those diagnosed with COPD, while the lung cancer group included lung cancer patients as well as individuals with both lung cancer and COPD. The prevalence of COPD in both groups was similar by design (17 and 21% respectively), to ensure that the test has no bias towards this condition. As part of the study design, subjects with different stages of lung cancer were included, with emphasis on early stages. Various histological types of lung cancer were included, as well.
Metabolic activity profile measurements
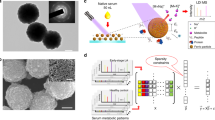
Our Liquid ImmunoBiopsy test, also called the MAP test, is designed to be performed on freshly separated PBMCs, which are isolated through Lymphoprep and gradient centrifugation. For this study, the test was carried out using PBMCs from each of the test subjects. The use of the whole population of PBMCs, as opposed to cells of a certain type, was hypothesized to allow the measurement of the combined response of immune cells to the various stimulations. The possible problems of inter-subject heterogeneity were dealt with using appropriate controls of medium alone, basal PBMC state (no stimulant), and several concentrations of the same stimulant. For each test, a 384-well plate was loaded with PBMCs, and each well contained 1 of the 16 selected stimulants in several increasing concentrations (see “Materials and methods” for details). The extracellular acidity was measured in both air-open (‘open’) and hermetically-sealed (‘closed’) states. Both states enable the measurement of real-time accumulation of ‘soluble’ versus ‘volatile’ metabolic products (lactic acid versus CO2 and NH3), thereby differentiating between OXPHOS, anaerobic glycolysis and aerobic glycolysis. The measurements were performed using a commercial fluorescence plate reader, whose output comprises intensity readings for both ‘open’ and ‘closed’ states across a timespan of approximately 1.5 h. The metabolic activity profiles were calculated and examined for cancer diagnosis by machine learning and data mining tools (see a graphical illustration of the process in Fig. 1).
Diagnostic prediction model construction
Machine-learning methods were utilized to distinguish between lung cancer and control subjects. Before this could be done, meaningful features needed to be extracted from the raw data of the MAP test. The raw data comprise fluorescent readings, which are converted to acidity units using a calibration curve (Fig. 2a). The values represent the acidification levels of the extracellular environment while exposed to varying concentrations of stimulants. We hypothesized that the presence of cancer, associated with changes in the physiological function of the immune system, will be reflected in different metabolic activity profiles of the tested PBMC samples. Thus, the change in acidity as a function of time, defined as the reaction rate (r), was calculated for each concentration of each stimulant (Fig. 2b). The value of r was then observed as a function of stimulant concentration (C) (Supplementary Fig. 1). At many concentrations, a clear difference could be observed between the mean values of lung cancer samples and the mean of control samples (Fig. 2c).
Extraction of features from raw data. a Conversion of raw data to acidity units: in this example, PBMCs from one of the subjects were mixed with D-Glucose (concentration #7) as a stimulant, and the acidity was measured at different times. b Calculation of reaction rates: using the same example subject as in the previous panel, the change in acidity as a function of time is measured for several increasing concentrations of d-Glucose. c Mean reaction rate (r) of lung cancer (n = 100, red) and control (n = 100, green) subjects, as a function of stimulant concentration index (C). There are instances in which a difference was found between the two populations (c1), and instances with no significant difference (c2). The specific stimulant is written in the title of each graph, with the plate state in parentheses. Error bars depict the standard error of the mean for each data point
Several mathematical models were used to describe the relationship between C and r, using a small number of coefficients. Some of the models also take into consideration the inter-dependence of different stimulants. To enhance the difference between the two populations with each stimulant, decision tree classifiers were trained to predict the clinical status of subjects (“lung cancer” or “control”). The best mathematical model and best classifier parameters were selected for each stimulant, maximizing accuracy, and the final prediction model comprised an ensemble of decision trees, taking into consideration predictions from multiple stimulants. This was coupled with bootstrap aggregation (“bagging”) to obtain robust results. For verification, the entire process was repeated using an ensemble of SVM classifiers instead of trees.
Diagnostic prediction of lung cancer and control subjects
The constructed model produced an almost complete separation between the populations of lung cancer and control subjects (Fig. 3a), with an apparent performance sensitivity of 100% and specificity of 98%. As a next step, cross-validation (CV) was utilized to assess the predictive capability of the model (Fig. 3b). Specifically, a stratified 20-fold CV analysis was used, in which ten samples (1/20 of the cohort) are left out for validation, and the rest are used as a training set for the prediction model. The process is repeated iteratively, with a different set of 10 samples each time, until every subject in the cohort is given a prediction. The resulting prediction is a score between − 10 (strong control) and 10 (strong lung cancer). A receiver-operating characteristic (ROC) curve can then be plotted, and a positivity cut-off (or discrimination threshold) can be set to determine sensitivity and specificity. The obtained area under the ROC curve (AUC) was 0.91, with a sensitivity of 91%, specificity of 80%, and precision of 82.0% [95% confidence intervals are (87.7%, 94.3%), (75.3%, 84.7%), and (77.5%, 86.5%), respectively] with the cut-off value set to -0.3. For further assessment of the predictive capabilities of the model, tenfold and fivefold CV analyses were performed as well (leaving out 20 and 40 samples, respectively, each time), resulting in AUCs of 0.91 and 0.86 respectively. The result of the SVM-based model was an AUC of 0.84 using a 20-fold CV analysis.
Cancer detection results. a Histogram of prediction scores (a1) and ROC curve (a2) showing a separation between populations of lung cancer and control subjects. For these results, the prediction model was trained and validated on the entire data set. b Histogram of prediction scores (b1) and ROC curve (b2) of the 20-fold cross-validation
The prediction model seems to be equally strong in predicting the late and early stages of lung cancer. When defining stages 1–2 as early and stages 3–4 as late, there is no observable difference in sensitivity between the two groups (p = 0.74). This can be visualized by breaking down the results of the positive group into stages (Fig. 4a).
Prediction model performance for specific subpopulations. P value of the χ2 test is shown in both panels. a Sensitivity of the diagnostic model, broken down into lung cancer stages. b Accuracy of the model, broken down into medical condition groups. c Accuracy of the model, broken down according to subjects’ smoking habits
Identifying lung cancer over other chronic lung diseases
One important challenge, addressed here, is the ability of our diagnostic MAP test to distinguish not only between lung cancer and control subjects, but also between patients with cancer versus those with other diseases that increase immune system activity. To this end, subjects diagnosed with COPD were included in both the lung cancer and the control groups in approximately the same ratio. We observed that the percentage of correct predictions is similar between subjects with and without COPD, in both the control group (p = 0.69) and the lung cancer group (p = 0.92) (Fig. 4b). These results suggest that the MAP test’s ability to identify lung cancer is not affected by the presence of chronic lung comorbidities.
Comparison between smoking and non-smoking subpopulations
Since smoking habits have a major influence on the development of lung cancer, it is important to verify the integrity of the prediction model in regard to this variable. The percentage of correct predictions was compared between subjects labeled as smokers (either former or current) and non-smokers. There was no significant difference in success rates in both the control group (p = 0.23) and the lung cancer group (p = 0.48) (Fig. 4c). These results suggest that the MAP test’s ability to identify lung cancer is not affected by the smoking habits of tested subjects.
Discussion
Liquid ImmunoBiopsy is a novel functional test that measures the relative acidification levels of the PBMCs extracellular environment, revealing the metabolic activity profiles (MAPs) of the immune system cells as an indicator of disease status. Since the immune system is extremely sensitive, it is expected to be inherently suited for the early cancer detection.
In general, the metabolic activity of normal differentiated cells involves predominantly OXPHOS, while cancer cells primarily rely on aerobic glycolysis [21,22,23]. Interestingly, a similar effect was also found in the immune system, where activated (effector) T cells undergo a metabolic switch from OXPHOS to aerobic glycolysis, which is essential to their proper acquisition of effector functions, in addition to their proliferation [24,25,26,27,28]. The immune system plays a critical role in specifically identifying and eliminating cancer cells, a process referred to as tumor immune surveillance, which is based, among other things, on the expression of tumor-specific antigens [36]. More recently, the concept of tumor ‘immunoediting’ was adapted to better describe the role of the immune system during tumor development, consisting of three phases: elimination, equilibrium, and escape [35, 37]. The continuous involvement of the immune system in all phases of this process, in response to tumor development, differs from the non-cancerous state activity. Based on these important discoveries, we conclude that the resulting detected differences between lung cancer and control samples, exploited in this study for the development of a novel diagnostic model, can be attributed to the differences in PBMCs subpopulations [38, 39].
In Liquid ImmunoBiopsy, the raw MAP test data are first analyzed to extract meaningful classifier features, which are then used as input parameters for a machine-learning diagnostic prediction model. We present 20-fold cross-validation (CV) results of the diagnostic model, with an AUC of 0.91, displaying high sensitivity and specificity of 91 and 80% respectively. Further and more stringent examinations, using both 10- and fivefold CV procedures, reveal a slight-to-no decrease in AUC, which indicates robustness of the presented diagnostic model.
The diagnostic model demonstrates a statistically uniform sensitivity across different cancer stages, indicating that the early detection is possible using our liquid immunobiopsy method. This is of great diagnostic importance, since lung cancer survival is largely and directly dependent on the stage of diagnosis [43]. Moreover, the presence of COPD comorbidity in the tested subjects was shown not to affect the diagnostic results, in either sensitivity or specificity, indicating that the model’s results are not influenced by these pathological conditions. COPD increases fivefold the risk for lung carcinogenesis [44], thus, being able to detect lung cancer in this high-risk population can have a major impact on patient survival. Finally, the test was shown to be unaffected by the smoking habits of tested subjects, yielding similar results in smoking and non-smoking subpopulations.
One present limitation of the current Liquid ImmunoBiopsy test is the inability to detect the type of lung cancer under test. Not having an independent test group is an apparent disadvantage of this study, and a prospective validation study is being planned to further validate the diagnostic prediction model.
Our groundbreaking novel liquid immunobiopsy diagnostic method is initially aimed at assisting in cancer diagnosis, following the detection of nodules in the lungs. It could also potentially be used as a screening test for high-risk populations, such as COPD patients. We see benefits of our approach as a diagnostic tool, in which a negative result will lead to surveillance imaging and the avoidance of unnecessary invasive procedures, while a positive result could lead to an early, life-saving intervention. We hypothesize that liquid immunobiopsy methods may, in the future, contribute to monitoring immuno-responsiveness to immunotherapy procedures, because the MAPs should reflect the expected enhanced immune response.
We believe that liquid immunobiopsy will be capable of detecting diseases other than lung cancer. Preliminary results, using a breast-specific MAP test design, showed a good differentiation between breast cancer and control subjects, while observing a clear difference between cancer and other benign breast diseases such as fibro-adenoma (data not shown). The use of multiple stimulants, varying in accordance with the disease in question, combined with sophisticated machine-learning methods to infer MAPs, harbors great potential for the detection of various diseases. In this current work, we demonstrated the capability of an accurate, non-invasive diagnostic tool for the early detection of lung cancer.
Abbreviations
- AUC:
-
Area under the curve
- COPD:
-
Chronic obstructive pulmonary disease
- CV:
-
Cross-validation
- LDCT:
-
Low-dose computed tomography
- MAP:
-
Metabolic activity profile
- OXPHOS:
-
Oxidative phosphorylation
- ROC:
-
Receiver-operating characteristic
- SCLC:
-
Small cell lung cancer
- SVM:
-
Support vector machine
- USPSTF:
-
US Preventive Services Task Force
References
Dela Cruz CS, Tanoue LT, Matthay RA (2011) Lung cancer: epidemiology, etiology, and prevention. Clin Chest Med 32:605–644. https://doi.org/10.1016/j.ccm.2011.09.001
Howlader N, Noone A, Krapcho M et al (2016) SEER Cancer Statistics Review 1975–2013. National Cancer Institute; Bethesda, MD. https://seer.cancer.gov/csr/1975_2013/. Accessed 4 May 2017
Smith RA, Cokkinides V, Brooks D et al (2011) Cancer screening in the United States, 2011. CA Cancer J Clin 61:8–30. https://doi.org/10.3322/caac.20096
Moyer VA (2014) Screening for lung cancer: U.S. preventive services task force recommendation statement. Ann Intern Med 160:330–338. https://doi.org/10.7326/M13-2771
Rampinelli C, De Marco P, Origgi D et al (2017) Exposure to low dose computed tomography for lung cancer screening and risk of cancer: secondary analysis of trial data and risk-benefit analysis. BMJ 356:j347. https://doi.org/10.1136/bmj.j347
Marshall HM, Bowman RV, Yang IA et al (2013) Screening for lung cancer with low-dose computed tomography: a review of current status. J Thorac Dis 5:S524–S539. https://doi.org/10.3978/j.issn.2072-1439.2013.09.06
Pinsky PF, Kramer BS (2015) Lung cancer risk and demographic characteristics of current 20–29 pack-year smokers: implications for screening. J Natl Cancer Inst 107:djv226. https://doi.org/10.1093/jnci/djv226
Rodriguez-Roisin R, Soriano JB (2008) Chronic obstructive pulmonary disease with lung cancer and/or cardiovascular disease. Proc Am Thorac Soc 5:842–847. https://doi.org/10.1513/pats.200807-075TH
Siravegna G, Marsoni S, Siena S, Bardelli A (2017) Integrating liquid biopsies into the management of cancer. Nat Rev Clin Oncol 14:531–548. https://doi.org/10.1038/nrclinonc.2017.14
Aravanis AM, Lee M, Klausner RD (2017) Next-generation sequencing of circulating tumor DNA for early cancer detection. Cell 168:571–574. https://doi.org/10.1016/j.cell.2017.01.030
Fujii T, Barzi A, Sartore-Bianchi A et al (2017) Mutation-enrichment next-generation sequencing for quantitative detection of KRAS mutations in urine cell-free DNA from patients with advanced cancers. Clin Cancer Res 23:3657–3666. https://doi.org/10.1158/1078-0432.CCR-16-2592
Li X, Hayward C, Fong P-Y et al (2013) A blood-based proteomic classifier for the molecular characterization of pulmonary nodules. Sci Transl Med 5:207ra142. https://doi.org/10.1126/scitranslmed.3007013
Lanman RB, Mortimer SA, Zill OA et al (2015) Analytical and clinical validation of a digital sequencing panel for quantitative, highly accurate evaluation of cell-free circulating tumor DNA. PLoS One 10:e0140712. https://doi.org/10.1371/journal.pone.0140712
Brock G, Castellanos-Rizaldos E, Hu L et al (2015) Liquid biopsy for cancer screening, patient stratification and monitoring. Transl Cancer Res 4:280–290. https://doi.org/10.3978/j.issn.2218-676X.2015.06.05
Vachani A, Hammoud Z, Springmeyer S et al (2015) Clinical utility of a plasma protein classifier for indeterminate lung nodules. Lung 193:1023–1027. https://doi.org/10.1007/s00408-015-9800-0
Whitney DH, Elashoff MR, Porta-Smith K et al (2015) Derivation of a bronchial genomic classifier for lung cancer in a prospective study of patients undergoing diagnostic bronchoscopy. BMC Med Genom 8:18. https://doi.org/10.1186/s12920-015-0091-3
Anderson D, Najafzadeh M, Gopalan R et al (2014) Sensitivity and specificity of the empirical lymphocyte genome sensitivity (LGS) assay: implications for improving cancer diagnostics. FASEB J 28:4563–4570. https://doi.org/10.1096/fj.14-254748
Pantel K, Alix-Panabières C (2013) Real-time liquid biopsy in cancer patients: Fact or fiction? Cancer Res 73:6384–6388. https://doi.org/10.1158/0008-5472.CAN-13-2030
Hiley CT, Le Quesne J, Santis G et al (2016) Challenges in molecular testing in non-small-cell lung cancer patients with advanced disease. Lancet 388:1002–1011. https://doi.org/10.1016/S0140-6736(16)31340-X
Bettegowda C, Sausen M, Leary R (2014) Detection of circulating tumor DNA in early-and late-stage human malignancies. Sci Transl 6:224ra24. https://doi.org/10.1126/scitranslmed.3007094
Vander Heiden MG, Cantley LC, Thompson CB (2009) Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324:1029–1033. https://doi.org/10.1126/science.1160809
Kaelin WG Jr, Thompson CB (2010) Q&A: cancer: clues from cell metabolism. Nature 465:562–564. https://doi.org/10.1038/465562a
DeBerardinis RJ, Lum JJ, Hatzivassiliou G, Thompson CB (2008) The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab 7:11–20. https://doi.org/10.1016/j.cmet.2007.10.002
MacIver NJ, Jacobs SR, Wieman HL et al (2008) Glucose metabolism in lymphocytes is a regulated process with significant effects on immune cell function and survival. J Leukoc Biol 84:949–957. https://doi.org/10.1189/jlb.0108024
Fox C, Hammerman P, Thompson C (2005) Fuel feeds function: energy metabolism and the T-cell response. Nat Rev Immunol 5:844–852. https://doi.org/10.1038/nri1710
Michalek RD, Rathmell JC (2010) The metabolic life and times of a T-cell. Immunol Rev 236:190–202. https://doi.org/10.1111/j.1600-065X.2010.00911.x
Pearce E (2010) Metabolism in T cell activation and differentiation. Curr Opin Immunol 22:314–320. https://doi.org/10.1016/j.coi.2010.01.018
Chang CH, Curtis JD, Maggi LB Jr et al (2013) Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell 153:1239–1251. https://doi.org/10.1016/j.cell.2013.05.016
Dietl K, Renner K, Dettmer K et al (2010) Lactic acid and acidification inhibit TNF secretion and glycolysis of human monocytes. J Immunol 184:1200–1209. https://doi.org/10.4049/jimmunol.0902584
Jellusova J, Cato MH, Apgar JR et al (2017) Gsk3 is a metabolic checkpoint regulator in B cells. Nat Immunol 18:303–312. https://doi.org/10.1038/ni.3664
Patsoukis N, Bardhan K, Chatterjee P et al (2015) PD-1 alters T-cell metabolic reprogramming by inhibiting glycolysis and promoting lipolysis and fatty acid oxidation. Nat Commun 6:6692. https://doi.org/10.1038/ncomms7692
Yang Z, Matteson EL, Goronzy JJ, Weyand CM (2015) T-cell metabolism in autoimmune disease. Arthritis Res Ther 17:29. https://doi.org/10.1186/s13075-015-0542-4
Beezhold K, Byersdorfer CA (2018) Targeting immuno-metabolism to improve anti-cancer therapies. Cancer Lett 414:127–135. https://doi.org/10.1016/J.CANLET.2017.11.005
Chimenti MS, Triggianese P, Conigliaro P et al (2015) The interplay between inflammation and metabolism in rheumatoid arthritis. Cell Death Dis 6:e1887. https://doi.org/10.1038/cddis.2015.246
Dunn GP, Bruce AT, Ikeda H et al (2002) Cancer immunoediting: from immunosurveillance to tumor escape. Nat Immunol 3:991–998. https://doi.org/10.1038/ni1102-991
Swann JB, Smyth MJ (2007) Immune surveillance of tumors. J Clin Invest 117:1137–1146. https://doi.org/10.1172/JCI31405
Dunn GP, Old LJ, Schreiber RD (2004) The three Es of cancer immunoediting. Annu Rev Immunol 22:329–360. https://doi.org/10.1146/annurev.immunol.22.012703.104803
Michalek RD, Gerriets VA, Jacobs SR et al (2011) Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J Immunol 186:3299–3303. https://doi.org/10.4049/jimmunol.1003613
Pearce E, Poffenberger M, Chang C (2013) Fueling immunity: insights into metabolism and lymphocyte function. Science 342:1242454. https://doi.org/10.1126/science.1242454
Pedregosa F, Varoquaux G, Gramfort A et al (2011) Scikit-learn: machine learning in Python. J Mach Learn Res 12:2825–2830
Weissferdt A, Moran CA (2014) Reclassification of early stage pulmonary adenocarcinoma and its consequences. J Thorac Dis 6:S581–S588. https://doi.org/10.3978/j.issn.2072-1439.2014.07.41
Goldstraw P, Ball D, Jett JR et al (2011) Non-small-cell lung cancer. Lancet 378:1727–1740. https://doi.org/10.1016/S0140-6736(10)62101-0
Siegel RL, Miller KD, Jemal A (2015) Cancer statistics, 2015. CA Cancer J Clin 65:5–29. https://doi.org/10.3322/caac.21254
Sekine Y, Katsura H, Koh E et al (2012) Early detection of COPD is important for lung cancer surveillance. Eur Respir J 39:1230–1240. https://doi.org/10.1183/09031936.00126011
Acknowledgements
The authors wish to thank the principal investigators, Prof. Fernando Patolsky, Prof. Nadir Arber, Dr. Mirjana Wollner, and Dr. Ran Kremer; the coordinators of this study, including multi-site coordinator Janna Bronstein, for patient recruitment, collection of specimens, and clinical data; Dr. Sara Bar Yehuda for regulatory support; Dr. Uri Itay and Chaim Singal for their help in the statistical analysis of the data; and furthermore, all the donor patients for their contribution to this clinical study.
Funding
This work was supported by Savicell Diagnostics and by Grants from the Israel Science Foundation (ISF) through the Legacy Program.
Author information
Authors and Affiliations
Contributions
FP conceived and designed the experiments and assisted in manuscript preparation. HPS and RT designed and performed preliminary pre-clinical studies and helped analyzing preliminary data. ST, ED, GD, and IA synchronized the study, helped analyzing data, and assisted in manuscript preparation. SA, CB, TS, and AL performed clinical testing, assisted in data analysis, and assisted in manuscript preparation. ST and ED developed classification model and performed statistical analysis. YA, SS, and MS assisted in clinical sample collection, clinical study coordination, and manuscript preparation.
Corresponding author
Ethics declarations
Conflict of interest
Fernando Patolsky received consulting fees from Savicell Diagnostics. Shoval Tirman, Aviv Lutaty, Tali Scheinmann, Eyal Davidovits, Irit Arbel, and Giora Davidovits are employed by Savicell Diagnostics. Shirley Abramovitch and Cynthia Botbol were employed by Savicell Diagnostics. Eyal Davidovits, Irit Arbel, and Giora Davidovits are officers in Savicell Diagnostics. Eyal Davidovits and Giora Davidovits are on the board of directors of Savicell Diagnostics. Yochai Adir, Shoval Tirman, Shirley Abramovitch, Cynthia Botbol, Aviv Lutaty, Tali Scheinmann, Eyal Davidovits, Irit Arbel, Giora Davidovits, and Fernando Patolsky own stock and/or options in Savicell Diagnostics’ parent company. The other authors declare that they have no conflict of interest.
Ethical approval and ethical standards
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committees and with the 1964 Helsinki declaration and its later amendments. Institutional review board approval numbers are: 0105-13-CMC for Carmel Medical Center, Haifa; 0274-15-RMB for Rambam Medical Center, Haifa; and 0009-13-TLV for Sourasky Medical Center, Tel Aviv.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Adir, Y., Tirman, S., Abramovitch, S. et al. Novel non-invasive early detection of lung cancer using liquid immunobiopsy metabolic activity profiles. Cancer Immunol Immunother 67, 1135–1146 (2018). https://doi.org/10.1007/s00262-018-2173-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-018-2173-5