Abstract
Isolates of Vibrio splendidus are ubiquitously presented in various marine environments, and they can infect diverse marine culture animals, leading to high mortality and economic loss. Therefore, a control strategy of the infection caused by V. splendidus is urgently recommended. Tryptanthrin is a naturally extracted bioactive chemical with antimicrobial activity to other bacteria. In this study, the effects of tryptanthrin on the bacterial growth and virulence-related factors of one pathogenic strain V. splendidus AJ01 were determined. Tryptanthrin (10 μg/mL) could completely inhibit the growth of V. splendidus AJ01. The virulence-related factors of V. splendidus AJ01 were affected in the presence of tryptanthrin. Tryptanthrin resulted an increase in biofilm formation, but lead to reduction in the motility and hemolytic activity of V. splendidus cells. In the cells treated with tryptanthrin, two distinctly differentially expressed extracellular proteins, proteases and flagellum, were identified using SDS–PAGE combined with LC–MS. Real-time reverse transcriptase PCR confirmed that the genes involved in the flagellar formation and hemolysin decreased, whereas specific extracellular proteases and the genes involved in the biofilm formation were upregulated. Two previously annotated luxOVs genes were cloned, and their expression levels were analyzed at different cell densities. Molecular docking was performed to predict the interaction between LuxOVs and ATP/tryptanthrin. The two sigma-54-dependent transcriptional regulators showed similar ATP or tryptanthrin binding capacity but with different sites, and the direct competitive binding between ATP and tryptanthrin was present only in their binding to LuxO1. These results indicated that tryptanthrin can be used as a bactericide of V. splendidus by inhibiting the growth, bacterial flagella, and extracellular proteases, but increasing the biofilm. Sigma-54-dependent transcriptional regulator, especially the quorum sensing regulatory protein LuxO1, was determined to be the potential target of tryptanthrin.
Key points
• Tryptanthrin inhibited the growth of V. splendidus in a dose-dependent manner.
• The effect of tryptanthrin on the virulence factors of V. splendidus was characterized.
• LuxO was the potential target for tryptanthrin based on molecular docking.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Vibrio splendidus is an important opportunistic pathogen, and isolates belonging to this species are ubiquitously present in seawater and sediments. They could infect a wide range of cultured animals from fish to invertebrate (Zhang and Li 2021). The infection of V. splendidus could lead to various diseases, resulting in mass mortalities among shellfish (Le Roux et al. 2004) and fish (Jensen et al. 2003). In particular, V. splendidus could infect the highly valued sea cucumber Apostichopus japonicus and result in disease with skin ulcer syndrome. Bacterial swimming, biofilm formation, and hemolysin are the virulence-related factors of V. splendidus (Zhang and Li 2021; Li et al. 2023; Yang et al. 2023).
Antibiotics are the most frequently and commonly used agents to cope with bacterial infection (Han et al. 2016). For example, furan and quinolones have been used to cure sea cucumber A. japonicus suffering from totting edge symptoms at the auricularia stage and the gas bubble–diseased sea cucumber A. japonicus suffering at the stage of auricularia due to Vibrio sp. (Han et al. 2016). However, the remaining antibiotics in the aquatic environment can cause severe environmental problems. The residue of antibiotics can frequently generate and spread antibiotic resistances (Polianciuc et al. 2020). Therefore, new environmentally friendly chemicals to substitute antibiotics have become the promising tools to prevent Vibrio sp. infection (Wang et al. 2017). One representative kind of these chemicals is the quorum sensing inhibitors, and coumarin targeting the quorum sensing regulation has been used for the control of V. splendidus infection (Zhang et al. 2017). Pyoverdine interfering the iron uptake process was determined to be effective in controlling V. splendidus infection (Zhang et al. 2016).
Extracting biochemical agents from Chinese traditional plants is an effective method to obtain new antimicrobial agents to inhibit the biomass or the virulence of bacterial pathogens (Pu et al. 2020). Tryptanthrin is a natural alkaloidal compound with the chemical name 6,12-dihydro-6,12-dioxoindolo-(2,1-b)-quinazoline, and it is distributed worldwide (Bigoniya and Rana 2009; Kawakami et al. 2011). The therapeutic effect of tryptanthrin has been confirmed to inhibit the expression of matrix metalloproteinase (MMP)-3 gene in different cell lines (Kirpotina et al. 2020). Tryptanthrin also possesses the antituberculotic activity, antiprotozoal activity, antioxidant, and antimicrobial activities (Kaur et al. 2017). As for the antimicrobial activity, tryptanthrin from the leaves of Strobilanthes cusia can inhibit the growth of Trichophyton mentagrophytes (Honda et al. 1979). Recently, tryptanthrin has also been used to strongly inhibit the biofilm formation activity of Vibrio cholerae (Narendrakumar et al. 2019).
In the present study, the growth and virulence-related factors of V. splendidus AJ01 by tryptanthrin were determined. The biofilm formation, hemolytic activity, extracellular protein secretion, and bacterial motility of V. splendidus AJ01 were characterized. Molecular docking between LuxO and tryptanthrin was performed to detect whether LuxO could be the target of tryptanthrin in V. splendidus. This study offers a new antibacterial chemical to treat V. splendidus infection in the future.
Materials and methods
Bacteria, culture, and regents
V. splendidus AJ01 was stocked in our lab and was preserved with a name Vs in CGMCC with an accession number of 7.242. V. splendidus AJ01 was cultured in 2216E medium, which was prepared with 1-g yeast extract, 5 g tryptone, and 0.01 g FePO4 in 1-L aged seawater. The bacterial culture was incubated in one shaker with speeds of 120–150 rpm/min (Jiangnan Instrument Co., Ningbo, China). Tryptanthrin was bought from Shanghai Yuanye Biotechnology Co. (CAS 13220–57-0, Shanghai, China), and it was dissolved into dimethyl sulfoxide (DMSO) to make a stock solution of 2.5 mg/mL. The other chemicals were bought from Shanghai Sangon Co. (China), unless otherwise stated.
Molecular technology
Through searching the genomic DNA of AJ01, two genes annotated to code LuxO repressor protein were selected. Both were amplified by PCR using two pairs of primers of luxO1-F/luxO1-R and luxO2-F/luxO2-R (Table 1), respectively. The sequences of the luxO1 and luxO2 genes were sequenced by Sangon (Shanghai, China) and verified by BLAST software in the website of the National Centre for Biotechnology Information (http://www.ncbi.nlm.nih.gov/blast). The expert protein analysis system (http://www.expasy.org/) was used to determine the molecular mass, amino acid sequence, and theoretical isoelectric point (pI). SMART was used to analyze the domains (http://smart.embl.de/). The three-dimensional structure of LuxO1 and LuxO2 was constructed using the SWISS-MODEL (https://web.expasy.org/protparam/). Molecular phylogenetic tree was constructed using MEGA7.0 software with neighbor-joining tree method.
Measurement of minimal inhibitory concentration (MIC)
MIC of tryptanthrin was determined as described previously (Narendrakumar et al. 2019). Briefly, 2216E media supplemented with 0.5, 1, 2.5, 5, 10, 25, and 50 μg/mL tryptanthrin were used to culture V. splendidus AJ01. The control sample was the culture of V. splendidus AJ01 grown in medium without tryptanthrin. After being cultured for 24 h, OD600 was recorded using a microplate reader (FlexA-200, Allsheng, China). Each growth was performed and measured in triplicate.
Biofilm measurement
Biofilm was determined as previous description (O’Toole 2011) with minor modification. V. splendidus AJ01 was reinoculated into fresh 2216E medium to make an initial bacterial cell suspension of 1.0 × 106 CFU/mL. The cell suspension was split into a 96-well polystyrene microplate. 2.5 μg/mL tryptanthrin was added as the experimental group, and the culture with the same volume of DMSO was used as the control sample. The plates were statically cultured in an incubator at 28 ℃ for 36 h, and then, the OD600 of the culture was measured. After that, biofilm was stained using crystal violet, and the absorbance at OD590 using a spectrophotometer (FlexA-200, Allsheng, China).
Swimming test
Swimming motility of V. splendidus AJ01 was performed according to the previous method (Wang et al. 2023). V. splendidus AJ01 was grown in medium supplemented with 2.5 μg/mL tryptanthrin for 24 h; then, 10-μL bacterial culture was plugged into the swimming agar, i.e., 2216E medium containing 2.5 μg/mL of tryptanthrin and 0.3% agar. The cells grown without tryptanthrin in both liquid medium and swimming agar were used as control. Diameter of the swimming circle was measured after inoculated for 24 h. Each experiment was carried out in triplicate.
Hemolytic activity assay
Defatted sheep blood was used to detect the hemolytic activity of V. splendidus AJ01 according to the description by De et al. (1954). Briefly, V. splendidus AJ01 was grown with 2.5 μg/mL tryptanthrin for 24 h; then, 10-μL bacterial culture was separately dropped onto the solid culture medium simultaneously containing 5% sheep blood cells and 2.5 μg/mL tryptanthrin. The cells grown without tryptanthrin in both liquid medium and agar were used as control. Then, all these plates were left statically at 28 ℃ for 24 h, and the diameter of hemolytic circle with or without tryptanthrin was measured.
Extraction of extracellular proteins
Overnight culture of V. splendidus AJ01 was reinoculated into 2216E medium at a volume of 1%. 2.5 μg/mL tryptanthrin was added and equal volume of DMSO was added as a control. The cultures were grown to a OD600 value of approximately 0.4, and then, the supernatants were collected after centrifugation, followed by filtration through a 0.22 μM membrane (Millipore, the USA) to obtain the cell-free supernatant. Then, 1/6 volume of 10% TCA was added. The mixture was left at 4 ℃ for 30 min, followed by centrifugation at 10,000 × g for 10 min to pellet the extracellular proteins. The precipitate was finally dissolved in buffer B (100 mM NaH2PO4 and 8 M urea in 10 mM Tris–Cl).
SDS–PAGE and LC–MS
Twenty microliters of the sample was taken and mixed with 5 μL SDS–PAGE loading buffer, and the samples were boiled at 100 ℃ for 5 min. Then, the differentially expressed proteins (DEPs) in the extracellular proteins collected from the cultures grown with and without tryptanthrin were detected using SDS–PAGE. The bands with DEPs were cut off and smashed into about 1-mm3 small pieces. Finally, the proteins in the gels were analyzed using LC–MS by Sangon Co. (Shanghai, China).
Scanning electron microscopy (SEM)
Bacterial cells were obtained by centrifugation of the cell culture at 3000 rpm at 4 ℃ for 2 min, and then, the cells were washed in triplicate in 100 mM PBS (pH 7.2–7.4). The cells were mixed with glutaraldehyde (2.5%) for 12 h, followed by washes with PBS, twice with 10 min for each time, and another twice washes with pure water. The sample was subsequently dehydrated with ethanol solution at gradients ranging from 30 to 90% for 15 min and finally dehydrated twice with pure ethanol, 15 min for each time. The samples were then resuspended in the mixture of tert-butanol and ethanol (1:1, v/v) for 15 min, followed by suspension in tert-butanol for another 15 min for twice. The samples were placed on 5 × 5 mm cover slips, frozen at − 80 °C, then put into a freeze dryer, and finally were observed under Hitachi SU8100 SEM (Japan).
Real-time reverse transcriptase PCR (RT–PCR)
Real-time RT–PCR was used to determine the mRNA level of specific genes in the cells grown with tryptanthrin, when compared to their expression in the cells grown without tryptanthrin. One biofilm formation–related gene, six genes involved in the flagellar biosynthesis, and three hemolysins and luxO genes were chosen. To collect samples for quantitative real-time RT–PCR, overnight culture of V. splendidus AJ01 was inoculated into media with or without 2.5 μg/mL tryptanthrin, respectively. The cultures were grown to OD600 value of approximately 0.4; then, the cultures were centrifuged to collect the cell pellet. TRIzol was used to extract RNA, and ABclonal reverse transcription kit (ABclonal Technology Co., Wuhan, China) was used to synthesize cDNA. The primers in the real-time RT–PCR for each gene are listed in Table 1. Real-time RT–PCR experiment was performed in the ABI 7500 real-time system (Applied Biosystems, the USA) based on the SYBR ExScript RT–PCR kit (Takara, China). The experiment was carried out in triplicate using 16S rRNA as a control gene. Dissociation analysis of the PCR products was carried out at the end of each PCR, which was used to verification that only one DNA product was successfully amplified. The comparative 2–△△CT method was occupied to compare the mRNA levels between different samples.
Molecular docking analysis
LuxO was the target of tryptanthrin in V. cholerae according to the report of Boyaci et al. (2016). Considering the conserved sequence of LuxO in Vibrio sp., the interaction between tryptanthrin and LuxOVs in V. splendidus was determined by molecular docking using the software AutoDock v4.2. Three-dimensional structure of tryptanthrin was retrieved from the data in PubChem (http://pubchem.ncbi.nlm.nih.gov). PyMOL and AutoDock were used to detect whether tryptanthrin, as well as ATP, could bind to LuxO.
Nucleotide sequence accession numbers
The nucleotide sequences have been deposited in the GenBank in NCBI. The nucleotide sequences of flhG, fliE, flgG, flhB, flgA, motY, AD, ZP1, ZP2, hemolysin I, hemolysin II, hemolysin III, luxO1, and luxO2 are under accession numbers OR743447, OR743448, OR743449, OR743450, OR743451, OR743452, OR743454, OR743455, OR743456, OR743457, OR743458, OR743459, OR743460, and OR799617, respectively.
Result
Tryptanthrin inhibited the growth and affected the cell morphology of V. splendidus AJ01
To test whether tryptanthrin could inhibit the growth of V. splendidus AJ01, an overnight culture of V. splendidus was inoculated into media with different concentrations of tryptanthrin. The results showed that V. splendidus AJ01 was highly sensitive to tryptanthrin, 80% of the growth of V. splendidus AJ01 could be reduced by 2.5 μg/mL tryptanthrin, and 10 μg/mL tryptanthrin completely inhibited the growth. Thus, 10 μg/mL was determined to be the MIC of tryptanthrin for the growth of V. splendidus AJ01 (Fig. 1A).
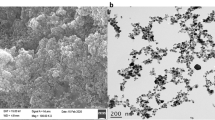
Effects of different concentrations of tryptanthrin on growth and morphology of V. splendidus AJ01. A OD600 of V. splendidus AJ01 culture grown with tryptanthrin. V. splendidus AJ01 was grown in 2216E media supplemented with 0, 0.5, 1, 2.5, 5, 10, 25, and 50 μg/mL tryptanthrin. The values are the mean ± standard errors from at least three experiments. B The representative figure of V. splendidus AJ01 cells grown with and without tryptanthrin, respectively, and morphology of V. splendidus AJ01 cells was observed under SEM. 1 represented V. splendidus cells without tryptanthrin; 2 represented V. splendidus cells with tryptanthrin
Tryptanthrin showed an obvious effect on the morphology of V. splendidus AJ01 observed under SEM. The untreated cells were intact, with vigorous growth and propagation. The long rod-shaped bacterial cells were the active cells that could divide into normal cells. After treatment with 2.5 μg/mL tryptanthrin, the cells became shorter, without the existence of extremely long rod-shaped bacterial cells. The cells also tended to cluster tightly. These results indicated that tryptanthrin could alter the morphology of bacterial cells, further interfering with their growth and survival (Fig. 1B).
Tryptanthrin affected virulence-related factors
Tryptanthrin reduced the biofilm formation
Biofilm is an important virulence factor related to bacterial virulence, so the effect of tryptanthrin on the biofilm formation of V. splendidus was determined. After the cells were incubated with tryptanthrin for 36 h statically, the inhibitory effect of tryptanthrin was not as high as that in the liquid culture. The OD600 values of the cultures with and without 2.5 μg/mL tryptanthrin were 0.62, and 0.45, respectively. Accompanied with a reduction in biomass in the presence of tryptanthrin, the biomass of biofilm was higher than that without tryptanthrin (Fig. 2A). However, the swimming ability of AJ01 reduced in the cells grown with tryptanthrin. After being incubated for 24 h, the diameters of the V. splendidus AJ01 colony without and with tryptanthrin were approximately 30 mm and 15 mm, respectively (Fig. 2B).
Determination of the virulence-related factors of V. splendidus AJ01 with tryptanthrin. A Quantitative measurement of the biofilm formed by V. splendidus AJ01 with tryptanthrin. V. splendidus AJ01 was grown in the presence or absence of tryptanthrin in 96-well plates for 24 h. Biofilm cells were stained with crystal violet, then dissolved using glacial acetic acid, and finally, OD590 was measured. Each growth was carried out in triplicate. *P < 0.05. B Swimming motility of V. splendidus AJ01 with tryptanthrin. Five microliters of culture with OD600 of approximately 0.4 with or without tryptanthrin was inserted into the center of the swimming agar. After being incubated for 24 h, the diameter of swimming circle was recorded. 1, the control cells without tryptanthrin; 2, the cells with tryptanthrin. C Hemolytic activity of V. splendidus AJ01 with or without tryptanthrin. Five-microliter culture of V. splendidus AJ01 (OD600 = 0.4) in the presence and absence of tryptanthrin was dropped onto the agar with sheep defatted blood. After being incubated at 28 ℃ for 24 h, the color was observed. 1, the control cells; 2, the cells with tryptanthrin. The experiment was performed in triplicate
Tryptanthrin inhibited the hemolytic activity of V. splendidus
Considering hemolysin is another important virulence factor of pathogenic bacteria, the effect of tryptanthrin on the hemolytic activity was determined. When defatted sheep blood was used, V. splendidus AJ01 exhibited a distinct α-hemolytic activity. The hemolytic activity of V. splendidus AJ01 in the presence of tryptanthrin significantly reduced (Fig. 2C).
Tryptanthrin changed the expression profiles of extracellular proteins
To explore whether tryptanthrin could affect the protein secretion of V. splendidus AJ01, supernatants collected from cultures grown with or without tryptanthrin were analyzed using SDS–PAGE. Distinctly different bands can be clearly observed on the gel. One band with a molecular mass of more than 100 kDa was significantly upregulated, whereas the other band at approximately 45 kDa was significantly downregulated (Fig. 3A). Both bands were collected, and the proteins in the bands were further determined by LC–MS. The results showed that the differentially expressed protein 1 (ZP1) was a protease, and the differentially expressed protein 2 (ZP2) was a flagellin protein involved in flagellar formation (Table 2).
Tryptanthrin affected the mRNA levels of virulence-related genes
Virulence-related genes, including genes related to flagellum, biofilm formation, and hemolysin, were chosen for real-time RT–PCR to detect the gene expression levels in cells with and without tryptanthrin. The mRNA levels of each gene in the presence of tryptanthrin are shown in Fig. 4, and the expression of each gene without tryptanthrin was used as 100%. Most of the genes involved in flagellar formation, i.e., flhG, fliE, flhB, flgG, flgA, and motY, were downregulated. Moreover, the mRNA expression of ZP2 was also downregulated. However, the expression levels of genes related to adhesion factor involved in biofilm formation in the cells with tryptanthrin increased. The mRNA expression of ZP1 protease was upregulated, exhibiting the same expression trend as the protein level in SDS–PAGE analysis.
mRNA levels of the specific gene. V. splendidus AJ01 were cultured with or without tryptanthrin to an OD600 of approximately 0.4. Real-time RT–PCR was performed following RNA extraction and reverse transcription. 16S rRNA was used as a reference gene. The expression of the genes in the absence of tryptanthrin was defined as “1.” The average value from at least three experiments was presented as the average ± SE. *P < 0.05
Sigma-54-dependent transcriptional regulators are the potential target of tryptanthrin in V. splendidus
Cloning and characterization of two putative LuxOVs
The genomic DNA of V. splendidus AJ01 has two annotated luxO genes (unpublished data), named luxO1 and luxO2. The open-reading frame of the luxO1 gene was 1347 bp, and it codes LuxO1 with a pI of 5.8 and a molecular mass of 51.5 kDa. The open-reading frame of the luxO2 gene was 1512 bp, and it codes LuxO2 with a pI of 5.89 and a molecular mass of 56.5 kDa. Phylogenetic analysis showed that the LuxO1 and LuxO2 in V. splendidus AJ01 had high homology to the LuxOs of other Vibrio spp. (Fig. 5A). Based on the result of BLAST in NCBI, both LuxOs are the regulators of NtrC homolog, which are sigma-54-dependent transcriptional regulators based on the result of BLAST in NCBI, with the same modular architectures predicted using the SMART tools (Supplementary Data Fig. S1). However, the BLAST result of LuxO1 indicated that it was more likely a quorum sensing sigma-54-dependent transcriptional regulator. The protein structure of LuxO1 and LuxO2 in V. splendidus predicted by SWISS is shown in Fig. 5B. Both showed binding capacity to the substrate of ATP (Supplementary Data Fig. S2 and Table S1). Two pairs of specific primers that corresponded to luxO1 and luxO2 were synthesized in accordance with the inconsistent DNA sequences of both genes, and further RT–PCR showed that the expression levels of both genes responded to cell densities. Their expression levels were relatively higher in V. splendidus AJ01 cells with lower OD600 than those in V. splendidus AJ01 cells with higher OD600. However, a slight difference was observed in the detailed expression profile, in which the mRNA level of luxO1 was consistent at lower cell density, whereas that of luxO2 was initially upregulated from the start to the maximum level and then decreased (Fig. 6).
A Molecular evolutionary tree constructed by the NJ method using MEGA software, based on the amino acid sequences of LuxOs from different Vibrio species. All protein sequences are obtained from the NCBI. The sequence numbers from top to bottom are as follows: CDT55873.1, LuxO2, CAH7011493.1, ACY51693.1, WP_268853584.1, CAH0536155, CCO53396, ETT11540, WP_152429975.1, GLT18841, WP_334511995, WP_156845775.1, RBM47670, BAF43696.1, EDL52722, LuxO1, WP_332234102, PIB16600, AEX22661, EGF45372, AEL22994, EEZ87274, ABF50762.1, BDR17646, EEX31766.1, ANS84831, CAH8199495.1, WP_026028983.1, ACP05294.1, WP_059120319, and EEX42264. B The structure of LuxO1 modeled using SWISS. C The structure of LuxO2 modeled using SWISS
mRNA level of luxO1 (A) and luxO2 (B) in the cells at different cell densities. V. splendidus AJ01 was cultured to OD600 of 0.2, 0.7, and 1.2, respectively. Real-time RT–PCR was performed following RNA extraction and reverse transcription. 16S rRNA was used as a reference gene. The expression of both luxOs in the cells at the first of inoculation was defined as “1.” The average value from at least three experiments was presented as the average ± SE. *P < 0.05
Molecular docking of two putative LuxOVs and tryptanthrin
The amino acid residues of LuxO and tryptanthrin bound through hydrophobic interactions or the benzene of amino acid, and the heterocycles of tryptanthrin formed P-P stacking interactions. In the present study, tryptanthrin could bind to LuxO1 and LuxO2, but the predicted binding sites in the two cases were different. Tryptanthrin bound to the pocket structure of LuxO1 by hydrogen bonding, through forming single hydrogen bonds with Arg293, Gln218, Ser121, Val69, Thr146, Asn120, or Lys195, respectively (Fig. 7A and B, Supplementary Data Table S2), while it bound to LuxO2 through forming single hydrogen bonds with the amino acids at amino acids of Lys253, Asp191, Arg360, Lys338, Phe194, Lys227, Arg119, or Thr348, respectively (Fig. 7C, D; Supplementary Data Table S2), significantly different from the binding to LuxO1. The model prediction showed that ATP and tryptanthrin possibly bind to the same amino acids of LuxO1, i.e., Arg293 or Gln218, but they could bind to different amino acids of LuxO2 (Fig. 7E, F). This result revealed that tryptanthrin could clearly serve as a competitive inhibitor of ATPase in LuxO1.
Predicted interaction of tryptanthrin with the LuxO1 and LuxO2 of V. splendidus AJ01 using the AutoDock software. A One representative prediction of LuxO1 interacting with tryptanthrin through a hydrogen bond. B One representative prediction of LuxO1 interacting with tryptanthrin through two hydrogen bonds. C One representative prediction of LuxO2 interacting with tryptanthrin through a hydrogen bond. D One representative prediction of LuxO2 interacting with tryptanthrin through two hydrogen bonds. E Prediction of the simultaneous binding of ATP and tryptanthrin to LuxO1. F Prediction of the simultaneous binding of ATP and tryptanthrin to LuxO2
Discussion
The environmental problems caused by residual antibiotics in the environment have led to the exploration of antibiotic alternatives for treating bacterial infection (Polianciuc et al. 2020). Antibiotic alternatives have been reported to reduce mortality caused by several marine pathogens, including Vibrio spp. (Yilmaz et al. 2022). Tryptanthrin is a natural phytochemical that can be used as a bacterial inhibitor (Kawakami et al. 2011), with the merits of easy availability and high safety, which facilitate its application (Kaur et al. 2017; Narendrakumar et al. 2019). Here, the antibacterial activity of tryptanthrin was explored. Tryptanthrin has been reported to possess antibacterial activity against Mycobacterium tuberculosis (Mitscher and Baker 1998), Bacillus subtilis (Fickenscher and Zähner 1971), and E. coli (Bandekar et al. 2010). Therefore, the present study is the first attempt to explore the inhibitory effect of tryptanthrin on the growth of marine isolates of Vibrio sp. as a new antibacterial agent with an MIC of 10 μg/mL, thereby widening the antibacterial spectrum of tryptanthrin. Similar to its inhibition on the growth of E. coli (Bandekar et al. 2010), tryptanthrin could also inhibit the biomass of V. splendidus, which facilitates its application in aquaculture.
V. cholerae forms biofilm at low cell density to survive the acidic condition under gastric environment (Rothenbacher and Zhu 2014). However, in V. splendidus, the biofilm is increased when there were more cells in the culture (Yang et al. 2023). In the presence of tryptanthrin, the growth was inhibited, but the biofilm formation increased. Considering the nature of the biofilm (Mah and O’Toole 2001), the increased biofilm cells suggested that the remaining cells survived the killing effect of tryptanthrin might through the biofilm formation. This was supported by the significant upregulation of one gene coding an inner membrane complex protein involved in cell attachment. Studies have shown that flagellar movement plays an important role in the formation of biofilm in the life cycle of Vibrio spp. (Echazarreta and Klose 2019; Teschler et al. 2015), and the initial step of biofilm development is generally accepted to be quick adherence of non-swimming cells (Belas 2013). In the present study, effects of tryptanthrin on the biofilm formation and cell motility were the opposite, which was similar to the effect of c-di-GMP on this two bacterial behaviors (Wolfe and Visick 2008). The swimming motility and the mRNA levels of genes that participated in flagellar formation of the remaining cells decreased, which contribute to the biofilm formation as described in V. cholerae and Salmonella enterica serovar Typhi, in which a negative relationship between motility and biofilm forming behavior was present (Guttenplan and Kearns 2013; Kalai Chelvam et al. 2014; Liu et al. 2010; Wolfe and Visick 2008). Although no significant effect was found on the extracellular protease activity in the presence of tryptanthrin, one extracellular protease determined as immune inhibitor A was upregulated. Immune inhibitor A is well studied in Bacillus sp., and it is involved in the breakage of host proteins as its pathogenesis (Pflughoeft et al. 2014). The upregulated immune inhibitor A in the presence of tryptanthrin indicated that tryptanthrin could mediate the virulence-related factors of Vibrio sp.
Little is known about the mechanism on the antimicrobial activity of tryptanthrin. Several proposed theories in different bacteria have been proposed. The inhibition mechanism of tryptanthrin was proposed to act as DNA intercalators, as confirmed in E. coli (Bandekar et al. 2010). In Vibrio sp., quorum sensing is the central regulatory system that controls collective behavior, including the biofilm formation, virulence, and biogeochemical cycling (Israel et al. 2023; Lami 2019; Milton 2006). Among the pathways, the highly conserved LuxO as the central component of the quorum sensing pathway was confirmed in Vibrio sp. (Boyaci et al. 2016). In V. cholerae, the binding of tryptanthrin to LuxO showed antibiofilm activity (Narendrakumar et al. 2019). In the present study, two annotated LuxOs were proposed to be the potential target of tryptanthrin on the basis of bioinformatic analysis. The same binding sites of ATP and tryptanthrin to LuxO1 indicated direct competitive binding between the two chemicals, and the inconsistent binding of ATP and tryptanthrin to LuxO2 may work through the conformation affection as indicated previously (Boyaci et al. 2016). This consistence in Vibrio sp. could facilitate the exploration of the inhibitory mechanism of tryptanthrin. A previous study showed that the biofilm formation of V. splendidus was positively regulated by quorum sensing (Yang et al. 2023), which indicated that the phosphorylated LuxO negatively regulated biofilm formation at lower cell density. Tryptanthrin not only could maintain V. splendidus at relative lower cell density but also potentially inhibit the phosphorylation of LuxO, thus upregulating the biofilm formation.
Data availability
The datasets generated and/or analyzed during the current study are available from one of the corresponding author, Weiwei Zhang, upon reasonable request.
References
Bandekar PP, Roopnarine KA, Parekh VJ, Mitchell TR, Novak MJ, Sinden RR (2010) Antimicrobial activity of tryptanthrins in Escherichia coli. J Med Chem 53(9):3558–3565. https://doi.org/10.1021/jm901847f
Belas R (2013) When the swimming gets tough, the tough form a biofilm. Mol Microbiol 90(1):1–5. https://doi.org/10.1111/mmi.12354
Bigoniya P, Rana AC (2009) Antidiarrheal and antispasmodic activity of Wrightia tinctoria bark and its steroidal alkaloid fraction. Pharmacologyonline 3:298–310
Boyaci H, Shah T, Hurley A, Kokona B, Li Z, Ventocilla C, Jeffrey PD, Semmelhack MF, Fairman R, Bassler BL, Hughson FM (2016) Structure, regulation, and inhibition of the quorum-sensing signal integrator LuxO. PloS Biol 14(5):e1002464. https://doi.org/10.1371/journal.pbio.1002464
De SN, Bhattacharyya K, Roychandhury PK (1954) The haemolytic activities of Vibrio cholerae and related Vibrios. J Pathol Bacteriol 67(1):117–127. https://doi.org/10.1002/path.1700670116
Echazarreta MA, Klose KE (2019) Vibrio flagellar synthesis. Front Cell Infect Microbiol 9:131. https://doi.org/10.3389/fcimb.2019.00131
Fickenscher U, Zähner H (1971) Metabolic products of microorganisms. 8.8 Effect of L-2,5-dihydrophenylalanine–a phenylalanine antagonist. Arch Mikrobiol 76(1):28–46
Guttenplan SB, Kearns DB (2013) Regulation of flagellar motility during biofilm formation. FEMS Microbiol Rev 37(6):849–871. https://doi.org/10.1111/1574-6976.12018
Han Q, Keesing JK, Liu D (2016) A review of sea cucumber aquaculture, ranching, and stock enhancement in China. Rev Fish Sci Aquac 24(4):326–341. https://doi.org/10.1080/23308249.2016.1193472
Honda G, Tabata M, Tsuda M (1979) The antimicrobial specificity of tryptanthrin. Planta Med 37(2):172–174. https://doi.org/10.1055/s-0028-1097320
Israel E, Ramganesh S, Abia ALK, Chikere CB (2023) Quorum sensing: unravelling the intricacies of microbial communication for biofilm formation, biogeochemical cycling, and biotechnological applications. J Mar Sci Eng 11(8):1586. https://doi.org/10.3390/jmse11081586
Jensen S, Samuelsen OB, Andersen K, Torkildsen L, Lambert C, Choquet G, Paillard C, Bergh O (2003) Characterization of strains of Vibrio splendidus and V. tapetis isolated from corkwing wrasse Symphodus melops suffering vibriosis. Dis Aquat Organ 53(1):25–31. https://doi.org/10.3354/dao053025
KalaiChelvam K, Chai LC, Thong KL (2014) Variations in motility and biofilm formation of Salmonella enterica serovar Typhi. Gut Pathog 6(1):2. https://doi.org/10.1186/1757-4749-6-2
Kaur R, Manjal SK, Rawal RK, Kumar K (2017) Recent synthetic and medicinal perspectives of tryptanthrin. Bioorg Med Chem 25(17):4533–4552. https://doi.org/10.1016/j.bmc.2017.07.003
Kawakami J, Matsushima N, Ogawa Y, Kakinami H, Nakane A, Kitahara H, Nagaki M, Ito S (2011) Antibacterial and antifungal activities of tryptanthrin derivatives. Trans Mater Res Soc J 36(4):603–606. https://doi.org/10.14723/tmrsj.36.603
Kirpotina LN, Schepetkin IA, Hammaker D, Kuhs A, Khlebnikov AI, Quinn MT (2020) Therapeutic effects of tryptanthrin and tryptanthrin-6-oxime in models of rheumatoid arthritis. Front Pharmacol 11:1145. https://doi.org/10.3389/fphar.2020.01145
Lami R (2019) Quorum sensing in marine biofilms and environments. Quorum Sensing 3:55–96. https://doi.org/10.1016/B978-0-12-814905-8.00003-4
Le Roux F, Gay M, Lambert C, Nicolas JL, Gouy M, Berthe F (2004) Phylogenetic study and identification of Vibrio splendidus -related strains based on gyrB gene sequences. Dis Aquat Organ 58(2–3):143–150. https://doi.org/10.3354/dao058143
Li Y, Yang H, Zhang J, Shi W, Li W, Zhang W (2023) VspC from Vibrio splendidus is responsible for collagen degradation in Apostichopus japonicus. Aquaculture 571:739489. https://doi.org/10.1016/j.aquaculture.2023.739489
Liu X, Beyhan S, Lim B, Linington RG, Yildiz FH (2010) Identification and characterization of a phosphodiesterase that inversely regulates motility and biofilm formation in Vibrio cholerae. J Bacteriol 192(18):4541–4552. https://doi.org/10.1128/jb.00209-10
Mah TFC, O’Toole GA (2001) Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol 9(1):34–39. https://doi.org/10.1016/s0966-842x(00)01913-2
Milton DL (2006) Quorum sensing in vibrios: complexity for diversification. Int J Med Microbiol 296(2–3):61–71. https://doi.org/10.1016/j.ijmm.2006.01.044
Mitscher LA, Baker W (1998) Tuberculosis: a search for novel therapy starting with natural products. Med Res Rev 18(6):363–374. https://doi.org/10.1002/(sici)1098-1128(199811)18:6%3c363::Aid-med1%3e3.0.Co;2-i
Narendrakumar L, Theresa M, Chandrika SK, Thomas S (2019) Tryptanthrin, a potential biofilm inhibitor against toxigenic Vibrio cholerae, modulating the global quorum sensing regulator. LuxO Biofouling 35(10):1093–1103. https://doi.org/10.1080/08927014.2019.1696315
O’Toole GA (2011) Microtiter dish biofilm formation assay. J vis Exp 47:2437. https://doi.org/10.3791/2437
Pflughoeft KJ, Swick MC, Engler DA, Yeo H-J, Koehler TM (2014) Modulation of the Bacillus anthracis secretome by the immune inhibitor A1 protease. J Bacteriol 196(2):424–435. https://doi.org/10.1128/jb.00690-13
Polianciuc SI, Gurzau AE, Kiss B, Stefan MG, Loghin F (2020) Antibiotics in the environment: causes and consequences. Med Pharm Rep 93(3):231–240. https://doi.org/10.15386/mpr-1742
Pu Z, Tang H, Long N, Qiu M, Gao M, Sun F, Dai M (2020) Assessment of the anti-virulence potential of extracts from four plants used in traditional Chinese medicine against multidrug-resistant pathogens. BMC Complement Med Ther 20(1):318. https://doi.org/10.1186/s12906-020-03114-z
Rothenbacher FP, Zhu J (2014) Efficient responses to host and bacterial signals during Vibrio cholerae colonization. Gut Microbes 5(1):120–128. https://doi.org/10.4161/gmic.26944
Teschler JK, Zamorano-Sánchez D, Utada AS, Warner CJ, Wong GC, Linington RG, Yildiz FH (2015) Living in the matrix: assembly and control of Vibrio cholerae biofilms. Nat Rev Microbiol 13(5):255–268. https://doi.org/10.1038/nrmicro3433
Wang W, Sun J, Liu C, Xue Z (2017) Application of immunostimulants in aquaculture: current knowledge and future perspectives. Aquac Res 48(1):1–23. https://doi.org/10.1111/are.13161
Wang JJ, Li Q, Huang LX (2023) Effect of the zinc transporter ZupT on virulence mechanism of mesophilic Aeromonas salmonicida SRW-OG1. Anim Res One Health 1:30–42. https://doi.org/10.1002/aro2.17
Wolfe AJ, Visick KL (2008) Get the message out: cyclic-Di-GMP regulates multiple levels of flagellum-based motility. J Bacteriol 190(2):463–475. https://doi.org/10.1128/JB.01418-07
Yang Y, Li W, Li Y, Shi W, Zhang J, Dang W, Zhang W (2023) Exogenous c-di-GMP inhibited the biofilm formation of Vibrio splendidus. Microb Pathog 175:105981. https://doi.org/10.1016/j.micpath.2023.105981
Yilmaz S, Yilmaz E, Dawood MAO, Ringo E, Ahmadifar E, Abdel-Latif HMR (2022) Probiotics, prebiotics, and synbiotics used to control vibriosis in fish: a review. Aquaculture 547:737514. https://doi.org/10.1016/j.aquaculture.2021.737514
Zhang W, Li C (2021) Virulence mechanisms of vibrios belonging to the splendidus clade as aquaculture pathogens, from case studies and genome data. Rev Aquacult 13(4):2004–2026. https://doi.org/10.1111/raq.12555
Zhang WW, Liang WK, Li CH (2016) Inhibition of marine Vibrio sp. by pyoverdine from Pseudomonas aeruginosa PA1. J Hazard Mater 302:217–224. https://doi.org/10.1016/j.jhazmat.2015.10.003
Zhang S, Liu N, Liang W, Han Q, Zhang W, Li C (2017) Quorum sensing-disrupting coumarin suppressing virulence phenotypes in Vibrio splendidus. Appl Microbiol Biotechnol 101(8):3371–3378. https://doi.org/10.1007/s00253-016-8009-3
Funding
This study was funded by the Zhejiang Provincial Natural Science Foundation for Distinguished Young Scholars (LR20C190001), the National Natural Science Foundation of China (31972833), the Natural Science Foundation of Ningbo City (2021J062), and the K.C. Wong Magna Fund in Ningbo University.
Author information
Authors and Affiliations
Contributions
HY designed and conducted the most part of the experiments, analyzed the data, wrote the original manuscript, and revised the manuscript. YL conducted parts of the experiments and analyzed the data. WS conducted some of the experiments and analyzed parts of the data. WZ conceived and planned the research, supervised the research, revised the manuscript, and acquired funding.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics approval
This article does not contain any studies with human participates or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, H., Li, Y., Shi, W. et al. Characterization of tryptanthrin as an antibacterial reagent inhibiting Vibrio splendidus. Appl Microbiol Biotechnol 108, 343 (2024). https://doi.org/10.1007/s00253-024-13158-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13158-7