Abstract
The goal of bacterial engineering is to rewire metabolic pathways to generate high-value molecules for various applications. However, the production of recombinant proteins is constrained by the complexity of the connections between cellular physiology and recombinant protein synthesis. Here, we used a rational and highly efficient approach to improve bacterial engineering. Based on the complete genome and annotation information of the Escherichia coli ER2566 strain, we compared the transcriptomic profiles of the strain under leaky expression and low temperature-induced stress. Combining the gene ontology (GO) enrichment terms and differentially expressed genes (DEGs) with higher expression, we selected and knocked out 36 genes to determine the potential impact of these genes on protein production. Deletion of bluF, cydA, mngR, and udp led to a significant decrease in soluble recombinant protein production. Moreover, at low-temperature induction, 4 DEGs (gntK, flgH, flgK, flgL) were associated with enhanced expression of the recombinant protein. Knocking out several motility-related DEGs (ER2666-ΔflgH-ΔflgL-ΔflgK) simultaneously improved the protein yield by 1.5-fold at 24 °C induction, and the recombinant strain had the potential to be applied in the expression studies of different exogenous proteins, aiming to improve the yields of soluble form to varying degrees in comparison to the ER2566 strain. Totally, this study focused on the anabolic and stress-responsive hub genes of the adaptation of E. coli to recombinant protein overexpression on the transcriptome level and constructs a series of engineering strains increasing the soluble protein yield of recombinant proteins which lays a solid foundation for the engineering of bacterial strains for recombinant technological advances.
Key points
• Comparative transcriptome analysis shows host responses with altered induction stress.
• Deletion of bluF, cydA, mngR, and udp genes was identified to significantly decrease the soluble recombinant protein productions.
• Synchronal knockout of flagellar genes in E. coli can enhance recombinant protein yield up to ~ 1.5-fold at 24 °C induction.
• Non-model bacterial strains can be re-engineered for recombinant protein expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbial cell engineering for the improved performance of microbial platforms has emerged as a powerful tool for the production of metabolites. Major successes in this area include the improved production/synthesis of 2,3-butanediol, fumarate, L-proline, hyaluronic acid, and human O-linked glycoprotein, among others (Chen et al. 2020; Long et al. 2020; Meng et al. 2020; Natarajan et al. 2020; Wang et al. 2020). In this regard, a range of host organisms has been tested, among which the Escherichia coli bacterial expression system continues to be the preferred system for laboratory investigations and for the early-stage development of commercial applications. Indeed, more than 60% of recombinant proteins and nearly 30% of approved recombinant therapeutic proteins are produced by the E. coli expression system (Correa and Oppezzo 2015). The Escherichia coli ER2566 strain (NC_CP014268.2) was developed as a BL21 (DE3) derivative strain and had been widely used in recombinant protein expression (Fomenkov et al. 2017). Although E. coli is a well-established host that offers short culturing time, easy genetic manipulation, and low cost, some bottlenecks in the production process, such as heterologous protein aggregation, prevent its more wide-scale use (Chen 2012). For instance, many eukaryotic proteins fail to fold properly when expressed using the E. coli system and form insoluble aggregates (Sahdev et al. 2008). Thus, from a host perspective, microbial cell engineering is becoming an important consideration to facilitate the production of native-state recombinant proteins with high efficiency. Unlike with typical metabolic pathways, recombinant protein expression is intricately linked to the cellular machinery, with multiple factors determining the flux through the pathway (Mahalik et al. 2014). Furthermore, the complex linkages between cellular physiology and heterogeneous expression make it difficult to break those bottlenecks in the successful production of proteins. Fortunately, knowledge from the increasing number of transcriptomic, genomic, and metabolomic studies has helped to provide a deeper understanding of the cellular factors responsible for exogeneous protein expression, and this will benefit rational host cell engineering (Brunk et al. 2016; Tan et al. 2020).
Protein aggregation (inclusion bodies) is the most frequently encountered issue during the overexpression of exogeneous proteins using the E. coli expression system (Ami et al. 2009). How to effectively avoid the formation of inclusion bodies and improve soluble expression yields remains a topic of discussion. Ranging from inheritance to environment, soluble expression can be affected by numerous factors: nucleotide sequence, protein size, the presence of post-translational modifications or cytoplasmic enzyme interactions, the vectors and promoters used in the reactions, and outside factors, such as the pH and temperature (Fink 1998; Idicula-Thomas and Balaji 2005; Peterson 2012). Indeed, induction at a lower temperature is known to increase protein yield in most cases and improve the biological activity of the product (Qing et al. 2004; Le and Schumann 2007; Vera et al. 2007; Caspeta et al. 2009). However, a systematic understanding of how a lower temperature contributes to high yield is lacking, and this lack of knowledge hinders the application of this temperature modification to bacterial engineering. Meanwhile, leaky expression—i.e., expression that occurs in the absence of proper induction—is another major concern in the use of the E. coli expression system. Whereas leaky expression can be optimized and is beneficial in some cases (e.g., production of toxic proteins, technical enzymes, and biopharmaceutical products) (Ding et al. 2019), it can also reduce growth rates, cause cell death, or result in plasmid instability, which is an issue for large-scale industrial production (Saida et al. 2006).
In this study, we adopted a rational and highly efficient strategy to re-engineer E. coli for better exogeneous protein expression. To this end, we compared the transcriptomic profiles of blank and plasmid-transferred bacteria without induction (leaky expression) and at two induction temperatures: 37 °C (inclusion body) and 24 °C (soluble expression). We used bioinformatics and genome editing technologies to mine differentially expressed genes and relevant metabolic pathways associated with the different cellular phenotypes grown under these variable conditions. This study seeks to provide a comprehensive understanding of the biological processes underlying low temperature–induced high yield and offers a flexible way to reengineer other bacteria that are used for recombinant expression.
Materials and methods
Strain and bacterial cultures
E. coli strain ER2566 (strain ATCC#PTA-3420), BL21(DE3) (OneShot® BL21 Star™ (DE3) chemically competent cells, #C601003, Thermo Fisher Scientific, Waltham, USA) and other derivative strains, obtained by λ-red homology recombination genome editing, were used as hosts for the expression of exogenous proteins (Fomenkov et al. 2017). The strain was stored in Luria–Bertani (LB) broth (5 g/L NaCl, 10 g/L tryptone, 5 g/L yeast extract, pH 7.0) containing 10% glycerol at − 80 °C. The gene encoding human papillomavirus 16 type L1 (HPV16-L1) was cloned into the pTO-T7 expression vector with kanamycin resistance gene by Gibson assembly and then transferred into the recombinant strain (Luo et al. 2000). The strain was grown at 37 °C in 100 mL of LB medium with kanamycin overnight. Then, 1 mL of culture was transferred into a flask containing 500 mL LB medium until an optical density at 600 nm (OD600) 0.8–1.0 was reached. HPV16-L1 protein expression was then induced with a final concentration of 10 mM isopropyl β-D-thiogalactoside (IPTG) for 6 h at 24 °C or 37 °C.
RNA extraction
Cells were harvested by centrifugation at room temperature for 10 min at 7000 rpm and the total RNA was extracted using the MasterPure RNA Purification Kit, according to the manufacturer’s protocol (Lucigen, Wisconsin, USA). DNase was added to avoid genomic DNA contamination. Total RNA was extracted in 50 μl of RNase-free diethyl pyrocarbonate-treated water (DEPC). RNA concentration was determined using an RNA Assay Kit in a Qubit2.0 Fluorometer (Thermo Fisher Scientific, Waltham, USA).
cDNA library construction and sequencing
For each sample, 3 µg RNA was used for library construction. Libraries of RNA-seq templates were constructed with NEBNext Ultra RNA Library Prep Kit for Illumina following the manufacturer’s recommendations (Illumina, San Diego, CA, USA). Sequencing was performed on an Illumina Hiseq 2500 platform and 125-bp paired-end reads were generated (Novogene, Beijing, China).
Bioinformatics analysis of transcriptome data
Raw reads were handled by running Trimmomatic to remove low-quality reads, poly-N, and adapters with the following parameters: SLIDINGWINDOW:4:20 and MINLEN:50 (Bolger et al. 2014). The index of the ER2566 genome was built from NZ_CP014268.2 using Burrows-Wheeler Aligner (bio-bwa.sourceforge.net), and the clean data were aligned to the reference genome using the MEM algorithm with the default setting (Jo and Koh 2015). FeatureCounts software was used to calculate read mapping to each gene (Liao et al. 2014). Based on the gene annotation of the transcripts, the aligned-read values were calculated to determine the gene expression levels. The expression values were normalized by DESeq2 methods previously described based on the negative binomial distribution (Love et al. 2014). Genes with fold change ≥ 2 and p-adjusted ≤ 0.05 were designated as differentially expressed. Gene ontology enrichment analysis of differentially expressed genes (DEGs) was implemented by the ClusterProfiler package, and GO terms with corrected p-values less than 0.01 were considered significantly enriched (Yu et al. 2012). The biological replicates were checked for any batch effects before the raw counts were generated using the Bioconductor Rsubread package (Liao et al. 2019).
Real-time PCR (qPCR)
To further validate the RNA-seq analysis, qPCR was conducted to determine gene expression patterns. The total RNA isolation was performed as described above and purified using the RNAprep Pure cell/Bacteria kit. One Step qPCR was performed using the primers and probes listed in Supplemental Table S1 in the following reaction volumes: 1 μl template, 0.5 μl F/R primers, 64 μl of 10 × buffer, 2 μl of 2.5 mM dNTP, 0.4 μl HS Taq, 0.2 μl TransScript II Reverse Transcriptase (TransGen Biotech, Beijing, China), 10.4 μl DEPC H2O, and 1 μl probe. The PCR program was run at 50 °C for 10 min, 95 °C for 10 min, followed by 45 cycles of 95 °C for 15 s and 55 °C for 50 s. The relative mRNA levels were evaluated using the comparative Cycle Threshold (2 − △△Ct) method, with 16S rRNA used as the internal reference (Brosius et al. 1978).
Strain construction
All ER2566 derivative strains used in this study are listed in Supplemental Table S2. E. coli DH5α (Takara, Dalian, China) was used for plasmid construction and ER2566 cells harboring pKD46 were used as the host strain for gene knock-out homologous recombination with the λ Red recombinase system (Datsenko and Wanner 2000). In brief, donor DNA with the selectable marker kan flanked by the Flp recognition target (FRT) site were integrated into the E. coli chromosome, and the kanamycin cassette was removed from the pCP20 plasmid using Flp recombinase (Doublet et al. 2008). After recombination, the correct recombinant colony was confirmed by sequencing.
ELISA quantitative detection
The HPV16-L1 monoclonal antibody 22E4 was coated into the wells of 96-well microplates (200 ng/well), as described previously (Gu et al. 2017). The wells were blocked and then incubated with 100 μL of two-fold serially diluted HPV16-L1 antigen in 1 × ED11 (Innovax, Beijing, China). HRP-conjugated HPV16L1 antibody 21A5 (diluted 1:3000 in 1 × ED11) was used as a secondary antibody. The known concentrations of HPV16-L1 antigens were used to make standard curves to detect the content of HPV16-L1 in the supernatant. The absorbance (450 nm; reference, 620 nm) was recorded using an automated ELISA reader (TECAN, Männedorf, Switzerland).
Relative quantification of recombinant protein
Soluble recombinant protein fractions (HPV69-51–26, HPV58, GFP, and HEV P239) were purified and determined by densitometry analysis of the SDS-PAGE gel using ImageJ (Rueden et al. 2017).
Statistical analysis
Significant differences between samples were analyzed using SPSS 20 (IBM, Chicago, USA). p < 0.05 was considered statically significant. Significant differences between the two means are presented as p < 0.05 (*).
Results
Host omics engineering strategy and stress effects on cell growth and yields of recombinant protein products
Here, we proposed a workflow that integrates transcriptomic data to study the effects of heterologous protein production within the engineering host (Fig. 1). Based on improved genome annotation (Zhou et al. 2020), host dynamic flux variations could be obtained from transcriptomic comparative analyses, which provided detailed molecular insight into the biological process and aided in the identification of key genes and pathways. These key pathways and genes could then be addressed by a single-gene editing strategy (λ RED-ET system) or via high-throughput genome engineering methods (clustered regularly interspaced short palindromic repeats cas9 system) to generate new strains (Jiang et al. 2015). In turn, these strains were tested in a microbioreactor format to select the best clones and optimize the subsequent scale-up. This strategy made it possible to explore specific phenotype differences and developed microbial cell factories with superior performance.
Overview for host omics-engineering strategies. a The complete genome assembly and accurate annotation pipelines for the engineering host. b The dynamic flux variations are obtained from the analysis of omics studies (transcriptomics) and the systems level bottlenecks are identified. c Bottlenecks can be addressed by generating derived/recombinant strains through single gene modification or high-throughput gene editing methods. These recombinant strains are tested in a microbioreactor format to select the best clones for further scale-up
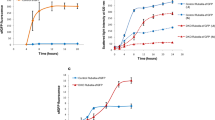
In this study, we designated the ER2566 strain cultured at 37 °C (blank; B37, three replicates) as the negative control and used human papillomavirus 16 type L1 capsid protein as a candidate that one important ingredient of HPV vaccines protects against infection with human papillomaviruses. Three experimental groups were set up: (i) IPTG-induced expression of HPV16-L1 (pTO-T7-HPV16L1) at 24 °C, producing a soluble product (Y24, three replicates); (ii) IPTG-induced expression of HPV16-L1 at 37 °C, producing inclusion bodies (Y37, three replicates); (iii) non-induced expression of HPV16-L1 at 37 °C (N37, three replicates) (Fig. 2a). Shake flask experiments were prepared to investigate the physiological impact of the different environmental conditions on recombinant protein expression. From the growth curves (OD600), we noted a significant growth inhibition after IPTG induction in both the Y24 and Y37 groups (Fig. 2b). There was no significant difference in cell density profiles between the blank (B37) and non-induced (N37) groups. In the Y37 group, cell growth quickly returned to baseline levels 2 h after IPTG addition, whereas cells in the low-temperature group (Y24) exhibited slower growth rates and remained at a lower cell density over the next 3 to 10 h. Significant improvements in target protein yields were observed in the Y24 group compared with the Y37 group, as measured by enzyme-linked immunosorbent assay (ELISA) (Fig. 2b, Supplemental Fig. S1). The Y24 group had a maximum protein yield of 10.8 μg per gram of wet cell weight (WCW), which was nearly 100-fold higher than that measured for the Y37 group. Of note, the N37 group without IPTG induction also showed a twofold increase in heterologous protein yield.
RNA sequencing analysis. a Strategies for exploring the exogeneous expression of HPV16-L1 in the E. coli ER2566 strain under different growth and induction conditions. Recombinant proteins are produced as soluble proteins or inclusion bodies under different stress. b Growth curves (left) and production bars (right) of E. coli cells treated with different induction conditions. The OD values on the growth curve represent the average calculated from three biological repetitions. HPV16-L1 proteins from the lysate of same wet cell weight (WCW) for each inoculum were quantified with double antibody sandwich ELISA. Values represent average + SD (n = 3). c Principal Components Analysis (PCA) of each group were generated by R package plotPCA. d Number of the differentially expressed genes (DEGs) in different comparisons
RNA-seq and data analysis
RNA sequencing was used to determine the expression profiles of the varied responses of ER2566 cells under different growth conditions. A total of 12 sequencing libraries mapping to the 4 groups (with 3 replicates) were constructed, each yielding about 22.52 to 31.68 million raw reads of 125-bp paired-end sequences (Illumina sequencing). All FASTQ files had good quality scores over 30. A total of 303,543,228 raw reads were harvested, and 96.46% (up to 292,801,439) of these were obtained as clean reads after trimming (Supplemental Table S3). For the N37, Y37, and Y24 overexpression samples, only 30.86 ~ 44.51% reads were mapped to the reference genome, which was significantly lower than that of the B37 samples: this reduction was due to the large amount of HPV16-L1 mRNA transcription, which was not related to the bacterial genome sequence.
A correlation analysis was performed to investigate reproducibility among the biological replicates and to determine the similarities between the different samples. Principal component analysis (PCA) based on gene expression abundance indicated high reliability of the biological triplicates (Fig. 2c). Next, we identified the genes that were differentially expressed in response to the different environments (Fig. 2d). In terms of leaky expression (N37 vs B37), 540 genes were significantly differentially expressed (log2|FoldChange|≥ 2.0 and p-adj ≤ 0.05), with 268 upregulated genes and 272 downregulated genes. Growth in a lower temperature (Y24 vs Y37) resulted in 620 DEGs (log2|FoldChange|≥ 1.0 and p-adj ≤ 0.05), with 261 upregulated genes and 359 downregulated genes. The DEGs were confirmed by real-time PCR (qPCR) analysis using the 2−ΔΔCt method and 16S ribosomal RNA for normalization. Five selected DEGs were submitted to the confirmation analysis, and the fold change in each gene expression was calculated. We found a high correlation between the assayed genes and those in the RNA-Seq data, confirming the reliability of the RNA-Seq analysis (Supplemental Fig. S2).
Molecular responses to leaky expression
A leaky expression can be problematic for protein production using the E. coli expression system. To obtain a broad overview of the molecular responses activated under conditions of leaky expression (N37 vs B37), gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were undertaken using the ClusterProfiler R package (Kanehisa and Goto 2000; Harris et al. 2004). All 540 DEGs identified for leaky conditions were associated with the following three GO categories: BP, biological process; CC, cellular component; and MF, molecular function, and clustered by whether the gene was upregulated or downregulated (Fig. 2d). DEGs that showed downregulation were significantly enriched in 103 terms, including “Ribosome assembly,” “Proton transmembrane transport,” “Membrane protein complex,” and “Disaccharide transport” (Supplemental Table S4a, Fig. 3a). Comparatively, the upregulated DEGs were mainly enriched across 68 terms related to biological processes (“Transcription,” “RNA biosynthetic process,” “Regulation of transcription,” “Fatty acid oxidation,” etc.) and 11 terms associated with a molecular function (“Oxidoreductase activity,” “Transcription regulator activity,” “DNA binding,” etc.); 3 terms were associated with cellular components (“Pilus,” “Integral component of cell outer membrane,” “Cell projection,”) (Supplemental Table S4b, Fig. 3b).
Gene ontology (GO) enrichment analysis of differentially expressed genes (DEGs) between non-induced ER2566 strain expressing HPV16-L1 protein cultured at 37 °C (N37) and blank ER2566 strain at the same temperature (B37). a Downregulated genes are totally enriched into 103 GO terms covering all three categories: BP, biological process; CC, cellular component; and MF, molecular function, and the 15 most significantly enriched BP terms, 10 CC terms and 15 MF terms are presented. The y-axis shows the GO enrichment terms across the three categories, whereas the x-axis shows the gene numbers. Enrichment criteria: pvalueCutoff < 0.01 and qvalueCutoff < 0.01. b Upregulated genes are enriched into 82 GO terms covering 3 categories and the 26 most significantly enriched BP terms, 3 CC terms and 11 MF terms are presented
Next, DEGs derived from a comparison of the non-induced and blank cultures (N37 vs B37) were submitted to KEGG enrichment analysis (p < 0.05; Supplemental Table S5). The results identified a downregulation in pathways associated with “Ribosome assembly” and “Oxidative phosphorylation,” and an upregulation in pathways that focused predominantly on “Pentose and glucuronate interconversions,” “Fatty acid metabolism,” “Tryptophan metabolism,” and “Propanoate metabolism.” These findings suggest that leaky expression can accelerate host energy metabolism and alter multiple host regulatory factors, with decreased ribosome synthesis assembly and transport pathways. This knowledge may help to establish a mechanism that describes the adaptative changes that occur under leaky expression stress.
Overall analysis of DEGs response to low-temperature induction
GO analysis was used to explore how low-temperature induction improves the soluble expression of the recombinant protein (Y24 vs Y37). The 359 downregulated DEGs identified were enriched into 6 clusters by GO terms: 5 terms belonged to biological processes (oxidation–reduction process, energy derivation by oxidation of organic compounds, Cellular respiration, anaerobic respiration, and carbohydrate transmembrane transport), and 1 term to cellular components (Ni–Fe hydrogenase complex) (Fig. 4a). The 261 upregulated DEGs were enriched in 11 terms (Fig. 4b), with 7 terms related to biological processes (polysaccharide transport, polysaccharide localization, maltodextrin transport, movement of cell or subcellular component, cell motility, localization of cell, and maltose transport), and 4 terms related to cellular components (bacterial-type flagellum, bacterial-type flagellum hock, bacterial-type flagellum, and cell projection). There was no enrichment of terms related to molecular function for either up- or downregulated genes associated with temperature-regulated induction. Comprehensive consideration of the BaseMean, fold changes, p-adj values, and genes which crossed multiple GO terms—for example, hybC, hybA, napA, cydA, aceK, malK, malF, flgH, among others—could potentially play important roles in promoting HPV16-L1 capsid protein soluble expression (Supplemental Table S6). In total, we concluded that 18 up- and 19 downregulated DEGs could be involved in host adaption for improved protein production under low-temperature induction (Fig. 5).
Volcano plot showing gene expression differences under induction temperatures of 24 °C and 37 °C. For each gene, the –log10(p-adj) was plotted against its log2(FoldChange). Genes with p-adj ≤ 0.05 and log2(FoldChange) ≤ -1 were designated as downregulated, with the others designated as upregulated. The key regulated genes are either in red (upregulated) or blue (downregulated) and are also represented in Supplemental Table S6a and S6b, respectively
Genetic engineering of ER2566 strain
To investigate the involvement of these 18 up- and 19 downregulated DEGs, we successfully constructed 36 singly gene knocked out strains using the λ-Red homologous recombination system except one of downregulated lethal gene, cydB, and compared recombinant HPV16-L1 expression levels between the wild-type and DEG-knock out strains at 24 °C and 37 °C (Fig. 6a, b). Among the knock-out recombinant strains, three downregulated genes (cydA, mngR, udp) and one upregulated gene (bluF) altered HPV16-L1 protein expression and resulted in a 30 ~ 70% decrease in protein production. From the growth curves (OD600), we noted that the knockout of cydA gene affects the bacterial growth and the production of the target protein. However, there was no significant difference in cell density profiles between wild-type and recombinant strains (ER2566-ΔmngR, Δudp, ΔbluF) (Fig. 6c), and we speculate that genes could act as key roles, affecting and regulating the solubility of the recombinant protein.
Fold change in the expression of HPV16-L1 recombinant protein in knock-out ER2566 engineered strains in comparison to wild type (WT) strain ER2566. a Downregulated and b upregulated DEGs were singly knocked-out to compare protein expression with the WT cells using the double antibody sandwich ELISA. The dashed lines represent a 20% increase or decrease in expression. Data are the mean and standard deviation calculated from two biological repetitions. c Growth curves of different E. coli recombinant strains. The OD values on the growth curve represent the average calculated from three biological repetitions. d The protein yield of different reconstruct strains in flasks (left) and fermentation (right). Quantification of recombinant protein was measured by double antibody sandwich ELISA. These results show the average values from the experiments independently repeated three times. Error bars represent the standard error. The asterisk corresponds to p < 0.05, related to ER2566 by t-test. e The particle morphology of HPV 16 L1 VLP from ER2566 strain and recombinant strain (ER2566-ΔflgL-ΔflgH-ΔflgK) were characterized by TEM and HPSEC, respectively. f Soluble recombinant protein fractions of different recombinant proteins in E. coli ER2566 strain and ER2566-ΔflgL-ΔflgH-ΔflgK were determined by densitometry analysis of SDS-PAGE gel using ImageJ
Some of the knock-out strains for upregulated genes, ΔgntK, ΔflgH, ΔflgL, ΔflgK, led to a 20% increase in the production of the target protein at 24 °C but not at 37 °C. Further engineering of the host (ER2566-ΔflgH-ΔflgL-ΔflgK) by combining multiple flagellar motor DEGs improved the recombinant protein yield to 1.5-fold than that of the wild-type strain with statistical differences (p < 0.05) (Fig. 6d). To further investigate the recombinant protein expression in high-density fermentation process between different strains, we cultured the cells in fermentation of 5-L scale; the strains ER2566 and BL21(DE3), harboring an HPV type 16 L1 protein expression plasmid (pTO-T7-H16), respectively served as a control, the recombinant strains (ER2566-ΔflgH-ΔflgL-ΔflgK) as an experimental group. Next, we evaluated the HPV16 L1 protein expression capability and found that the ER2566(pTO-T7-H16) strain and BL21(DE3)(pTO-T7-H16) strain had similar protein yield (~ 16 mg/g). Like the result in the flask culture, the recombinant strains (ER2566-ΔflgH-ΔflgL-ΔflgK) had a 1.3-fold higher protein yield (~ 20.3 mg/g) than the control groups (Fig. 6d). Additionally, as visualized by negative-stain, transmission electron microscopy (TEM) confirmed that the HPV L1 proteins expressed from the ER2566 E. coli, and recombinant strains were able to self-assemble into the form of virus-like particles (VLPs) with a similar diameter of 50 nm (Fig. 6e). Meanwhile, high-performance size exclusion chromatography (HPSEC) also confirmed a similar molecular weight for VLPs achieved by the two expression hosts in terms of relative retention time (Fig. 6e). Meanwhile, we also selected a series of recombinant proteins such as HPV 69–51-26, HPV58, green fluorescent protein (GFP) and HEV P239 protein, to assess the soluble expression of the target protein. Soluble expression levels of HPV 69–51-26, HPV58, and GFP protein in recombinant strain ER2566-ΔflgL-ΔflgH-ΔflgK were detected by ImageJ. In terms of the soluble recombinant protein fractions of different recombinant proteins by pixel-counting quantification approach, we found that the protein yields of reconstruct strains increased up to ~ 1.3 fold in comparison to the WT strain, except for HEV P239, which has been found to be expressed in nearly exclusively inclusion body form in our previous study (Li et al. 2015), and we also show the difficulty in soluble expression in this study as well, and the engineering in strain ER2566-ΔflgL-ΔflgH-ΔflgK has no observable effect for HEV P239 soluble expression (Fig. 6f).
Discussion
Omics-guided bacterial engineering for recombinant protein expression had been developed to improve strain performance (Choi et al. 2019). Despite massive research efforts and much recent progress (Choi and Lee 2013; d’Espaux et al. 2017; Pontrelli et al. 2018), only a few model strains have been applied in industrial production, mainly due to the lack of comprehensive knowledge on related genome information and corresponding biosynthetic pathways. In this study, our workflow integrates a common method based on our previous publication which demonstrated a universal pipeline for genome assembly and reannotation for various bacteria, including the non-model engineering hosts without the complete genome and annotation information, which greatly increases the plasticity of industrial strains. In addition, our reannotation pipeline with high speed and accuracy could be extrapolated for the reannotation of other bacterial genomes to provide a better understanding of gene function under the external burden and provide more clues to engineer bacteria for biotechnological applications. In the future, we will develop a web-based genomic annotation analysis website to facilitate the annotation and analysis of industrial strains for more users.
Inclusion body formation is often a troubling issue during the overexpression of exogenous proteins using the E. coli expression system (Ami et al. 2009). Lower-temperature induction is known to increase protein yield in most cases and improve the biological activity of the product (Qing et al. 2004). In this study, we showcase an over-expression of HPV16-L1 by comparative transcriptomics of the strains under low-temperature induction results in the upregulation of polysaccharide transport, maltodextrin transport, and cell motility, along with a downregulation in the processes surrounding oxidation–reduction, cellular respiration, and anaerobic respiration, among others. The analysis suggests that a critical pathway could play the key role in regulating the balance between heterologous protein production and host metabolism.
In addition, we found that the knock-out of three motility-related DEGs (ER2666-ΔflgH-ΔflgL-ΔflgK) could improve the expression yield for various interest proteins up to 1.5-folds in comparison to the prototype strain either in shake-flask culture or high-density fermentation. Consistent with our results, some studies had shown that the flagellar production is energy-intensive and that the flg gene knock-out or mutation strains could provide more energy and substrate for the synthesis of recombinant protein production by preventing cell motility and the assembly of the flagellum. Taken together, the knock-out of flg-related genes could also be applied to other E. coli strains for improving the expression level of recombinant protein. Notably, a large proportion of the knocked-out strains showed different responses at the two temperatures, suggestive of more complicated interactions under low-temperature stress. The challenges to improve the performance of the expression systems will thus require a comprehensive strategy that includes “balancing” the yield and activity of the protein, as well as the product quality and associated metabolite toxicity. Meanwhile, some hub genes were first identified to significantly influence recombinant protein productions, such as the deletion of bluF, cydA, mngR, and udp, which led to a significant decrease in soluble recombinant protein production. Other studies indicated that the bluF gene mainly binds to and releases the bluR repressor from its bound DNA target and also may serve as a thermometer (Hasegawa et al. 2006; Nakasone et al. 2010). According to this knowledge, we assumed that bluF knock-out might decrease the soluble expression of HPV16 L1 by losing a thermometer for E. coli to sense the low temperature. cydA is the component of the aerobic respiratory chain of E. coli that predominates when cells are grown at low aeration, and there was a significant difference in cell density profiles of bacterial cultures between wild-type and knock-out strains (ER2566-ΔcydA) which might affect cellular energy metabolism to decrease the expression of soluble heterologous proteins (Borisov et al. 2011). mngR might involve in the regulation of the acid citric cycle in response to fatty acids, a suggestion based on in vitro experiments as a transcription regulation factor to influence the expression of recombinant protein (Sampaio et al. 2004). However, how these genes clearly regulate the soluble expression needs to be further investigated by a variety of approaches such as additional regulatory gene over-expression and multi-omics in the future.
Based on the above findings including transcriptomic data and knock-out assay, we proposed a possible model to illustrate the response of the ER2566 strain under low-temperature induction (Fig. 7). GO and KEGG analyses show that those genes which are downregulated upon the induction at lower temperature might associate with the slowdown of carbohydrate transmembrane transport and cellular respiration. In contrast, the maltose transporter-related genes are significantly upregulated. Indeed, the maltose transporter knock-out assays indicate a significant decrease in the expression of the soluble recombinant protein at low-temperature induction as compared with the wild type ER2566 cells. We speculate that maltose transportation and metabolism might have a role in maintaining energy metabolism in the host cell at lower temperatures and are benefitting from the soluble expression of recombinant proteins. In another primary mechanistic branch, genes related to cell motility, localization, and flagellum terms were significantly enriched under low-temperature induction; these findings are consistent with previous research that the loss of E. coli motility at 37 °C is recovered when the temperature is reduced to 24 °C (Noor et al. 2013). Some authors have proposed that flagellar biosynthesis and assembly are an energy‐intensive process (Yoon et al. 2009). This mechanism was further validated in our knock-out assays, where deleting the DEG association with the bacterial-type flagellum (flgH, flgK, flgL) resulted in an increase in the production of the target protein. This increase in production was further exacerbated to ~ 1.5-fold by the combined knockout of flgH, flgK, and flgL genes under 24 °C induction in shake-flask or high-density fermentation.
Suggested model of the major overexpression response mechanism for recombinant protein expression under low-temperature induction. Arrows indicate primary processes or direct interactions; dotted arrows indicate secondary or indirect interactions. Green and red arrows, respectively, represent down- and upregulated pathways under low-temperature induction. In this model, the overexpression of the heterologous protein under low-temperature induction results in the upregulation of polysaccharide transport, maltodextrin transport and cell motility, along with a downregulation in the processes surrounding carbohydrate transport (e.g., glucose influx), cellular respiration, and anaerobic respiration, among others. Cells consume lots of energy for cell motility and the assembly of flagellum under low-temperature induction. Motility-related knock-out strains may have better energy optimization and substrate distribution, which, in turn, would promote the soluble expression of the recombinant protein. Meanwhile, some hub genes, such as bluF and cydA among others, were identified to significantly influence recombinant protein production
In summary, we proposed a model to interpret the response of the ER2566 strain under low-temperature induction. The low temperature–induced high yield may derive from comprehensive interactions, among cell motility and maltose transportation, and a reduction in pathways associated with carbohydrate transmembrane transportation and cellular respiration. Meanwhile, independent DEGs may also influence host adaptations and metabolic networks. Totally, this study focused on the anabolic and stress-responsive hub genes of the adaptation of E. coli to recombinant protein overexpression on the transcriptome level and constructed a series of engineering strains increasing the soluble protein yield of recombinant proteins which lays a solid foundation for the engineering of bacterial strains for recombinant technological advances.
Data availability
All data are provided in the manuscript, and the author promises to support the availability of the data. The RNA-Seq raw data used in this study have been deposited in the NIH Sequence Read Archive under accession number: PRJNA679153.
References
Ami D, Natalello A, Schultz T, Gatti-Lafranconi P, Lotti M, Doglia SM, de Marco A (2009) Effects of recombinant protein misfolding and aggregation on bacterial membranes. Biochim Biophys Acta - Proteins Proteomics 1794:263–269. https://doi.org/10.1016/j.bbapap.2008.10.015
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Borisov VB, Murali R, Verkhovskaya ML, Bloch DA, Han H, Gennis RB, Verkhovsky MI (2011) Aerobic respiratory chain of Escherichia coli is not allowed to work in fully uncoupled mode. Proc Natl Acad Sci U S A 108:17320–17324. https://doi.org/10.1073/pnas.1108217108
Brosius J, Palmer ML, Kennedy PJ, Noller HF (1978) Complete nucleotide sequence of a 16S ribosomal RNA gene from Escherichia coli. Proc Natl Acad Sci U S A 75:4801–4805. https://doi.org/10.1073/pnas.75.10.4801
Brunk E, George KW, Alonso-Gutierrez J, Thompson M, Baidoo E, Wang G, Petzold CJ, McCloskey D, Monk J, Yang L, O’Brien EJ, Batth TS, Martin HG, Feist A, Adams PD, Keasling JD, Palsson BO, Lee TS (2016) Characterizing strain variation in engineered E. coli using a multi-omics-based workflow. Cell Syst 2:335–346. https://doi.org/10.1016/j.cels.2016.04.004
Caspeta L, Flores N, Pérez NO, Bolívar F, Ramírez OT (2009) The effect of heating rate on Escherichia coli metabolism, physiological stress, transcriptional response, and production of temperature-induced recombinant protein: a scale-down study. Biotechnol Bioeng 102:468–482. https://doi.org/10.1002/bit.22084
Chen R (2012) Bacterial expression systems for recombinant protein production: E. coli and beyond. Biotechnol Adv 30:1102–1107. https://doi.org/10.1016/j.biotechadv.2011.09.013
Chen X, Ma D, Liu J, Luo Q, Liu L (2020) Engineering the transmission efficiency of the noncyclic glyoxylate pathway for fumarate production in Escherichia coli. Biotechnol Biofuels 13:1–10. https://doi.org/10.1186/s13068-020-01771-3
Choi KR, Jang WD, Yang D, Cho JS, Park D, Lee SY (2019) Systems metabolic engineering strategies: integrating systems and synthetic biology with metabolic engineering. Trends Biotechnol 37:817–837. https://doi.org/10.1016/j.tibtech.2019.01.003
Choi YJ, Lee SY (2013) Microbial production of short-chain alkanes. Nature 502:571–574. https://doi.org/10.1038/nature12536
Correa A, Oppezzo P (2015) Overcoming the solubility problem in E. coli: available approaches for recombinant protein production. Methods Mol Biol 1258:27–44. https://doi.org/10.1007/978-1-4939-2205-5_2
d’Espaux L, Ghosh A, Runguphan W, Wehrs M, Xu F, Konzock O, Dev I, Nhan M, Gin J, Reider Apel A, Petzold CJ, Singh S, Simmons BA, Mukhopadhyay A, García Martín H, Keasling JD (2017) Engineering high-level production of fatty alcohols by Saccharomyces cerevisiae from lignocellulosic feedstocks. Metab Eng 42:115–125. https://doi.org/10.1016/j.ymben.2017.06.004
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A 97:6640–6645. https://doi.org/10.1073/pnas.120163297
Ding N, Ruan Y, Fu X, Lin Y, Yu H, Han L, Fu C, Zhang J, Hu X (2019) Improving production of N-glycosylated recombinant proteins by leaky Escherichia coli. 3 Biotech 9:1–9. https://doi.org/10.1007/s13205-019-1830-5
Doublet B, Douard G, Targant H, Meunier D, Madec JY, Cloeckaert A (2008) Antibiotic marker modifications of λ Red and FLP helper plasmids, pKD46 and pCP20, for inactivation of chromosomal genes using PCR products in multidrug-resistant strains. J Microbiol Methods 75:359–361. https://doi.org/10.1016/j.mimet.2008.06.010
Fink AL (1998) Protein aggregation: folding aggregates, inclusion bodies and amyloid. Fold Des 3:9–23. https://doi.org/10.1016/S1359-0278(98)00002-9
Fomenkov A, Sun Z, Dila DK, Anton BP, Roberts RJ, Raleigh EA (2017) EcoBLMcrX, a classical modification-dependent restriction enzyme in Escherichia coli B: characterization in vivo and in vitro with a new approach to cleavage site determination. PLoS ONE 12:1–21. https://doi.org/10.1371/journal.pone.0179853
Gu Y, Wei M, Wang D, Li Z, Xie M, Pan H, Wu T, Zhang J, Li S, Xia N (2017) Characterization of an Escherichia coli-derived human papillomavirus type 16 and 18 bivalent vaccine. Vaccine 35:4637–4645. https://doi.org/10.1016/j.vaccine.2017.06.084
Harris MA, Clark J, Ireland A, Lomax J, Ashburner M, Foulger R, Eilbeck K, Lewis S, Marshall B, Mungall C, Richter J, Rubin GM, Blake JA, Bult C, Dolan M, Drabkin H, Eppig JT, Hill DP, Ni L, Ringwald M, Balakrishnan R, Cherry JM, Christie KR, Costanzo MC, Dwight SS, Engel S, Fisk DG, Hirschman JE, Hong EL, Nash RS, Sethuraman A, Theesfeld CL, Botstein D, Dolinski K, Feierbach B, Berardini T, Mundodi S, Rhee SY, Apweiler R, Barrell D, Camon E, Dimmer E, Lee V, Chisholm R, Gaudet P, Kibbe W, Kishore R, Schwarz EM, Sternberg P, Gwinn M, Hannick L, Wortman J, Berriman M, Wood V, de la Cruz N, Tonellato P, Jaiswal P, Seigfried T, White R (2004) The gene oncology (GO) database and informatics resource. Nucleic Acids Res 32:258–261. https://doi.org/10.1093/nar/gkh036
Hasegawa K, Masuda S, Ono TA (2006) Light induced structural changes of a full-length protein and its bluF domain in ycgF(blrP), a blue-light sensing protein that uses FAD (bluF). Biochem 45:3785–3793. https://doi.org/10.1021/bi051820x
Idicula-Thomas S, Balaji PV (2005) Understanding the relationship between the primary structure of proteins and its propensity to be soluble on overexpression in Escherichia coli. Protein Sci 14:582–592. https://doi.org/10.1110/ps.041009005
Jiang Y, Chen B, Duan C, Sun B, Yang J, Yang S (2015) Multigene editing in the Escherichia coli genome via the CRISPR-Cas9 system. Appl Environ Microbiol 81:2506–2514. https://doi.org/10.1128/AEM.04023-14
Jo H, Koh G (2015) Faster single-end alignment generation utilizing multi-thread for BWA. Biomed Mater Eng 26:S1791–S1796. https://doi.org/10.3233/BME-151480
Kanehisa M, Goto S (2000) KEGG: Kyoto Encyclopedia Of Genes And Genomes. Nucleic Acids Res 28:27–30. https://doi.org/10.1093/nar/28.1.27
Le ATT, Schumann W (2007) A novel cold-inducible expression system for Bacillus subtilis. Protein Expr Purif 53:264–269. https://doi.org/10.1016/j.pep.2006.12.023
Li SW, Zhao Q, Wu T, Chen S, Zhang J, Xia NS (2015) The development of a recombinant hepatitis E vaccine HEV 239. Hum Vaccines Immunother 11:908–914. https://doi.org/10.1080/21645515.2015.1008870
Liao Y, Smyth GK, Shi W (2014) FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30:923–930. https://doi.org/10.1093/bioinformatics/btt656
Liao Y, Smyth GK, Shi W (2019) The R package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nucleic Acids Res 47:1–9. https://doi.org/10.1093/nar/gkz114
Long M, Xu M, Qiao Z, Ma Z, Osire T, Yang T, Zhang X, Shao M, Rao Z (2020) Directed evolution of ornithine cyclodeaminase using an evolvR-based growth-coupling strategy for efficient biosynthesis of l -Proline. ACS Synth Biol 9:1855–1863. https://doi.org/10.1021/acssynbio.0c00198
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:1–21. https://doi.org/10.1186/s13059-014-0550-8
Luo WX, Zhang J, Yang HJ, Li SW, Xie XY, Pang SQ, Li SJ, Xia NS (2000) Construction and application of an Escherichia coli high effective expression vector with an enhancer. Sheng Wu Gong Cheng Xue Bao Chinese J Biotechnol 16:578–581. https://doi.org/10.3321/j.issn:1000-3061.2000.05.009
Mahalik S, Sharma AK, Mukherjee KJ (2014) Genome engineering for improved recombinant protein expression in Escherichia coli. Microb Cell Fact 13:1–13. https://doi.org/10.1186/s12934-014-0177-1
Meng W, Zhang Y, Cao M, Zhang W, Lü C, Yang C, Gao C, Xu P, Ma C (2020) Efficient 2,3-butanediol production from whey powder using metabolically engineered Klebsiella oxytoca. Microb Cell Fact 19:1–10. https://doi.org/10.1186/s12934-020-01420-2
Nakasone Y, Ono TA, Ishii A, Masuda S, Terazima M (2010) Temperature-sensitive reaction of a photosensor protein YcgF: possibility of a role of temperature sensor. Biochem 49:2288–2296. https://doi.org/10.1021/bi902121z
Natarajan A, Jaroentomeechai T, Cabrera-Sánchez M, Mohammed JC, Cox EC, Young O, Shajahan A, Vilkhovoy M, Vadhin S, Varner JD, Azadi P, DeLisa MP (2020) Engineering orthogonal human O-linked glycoprotein biosynthesis in bacteria. Nat Chem Biol 16:1062–1070. https://doi.org/10.1038/s41589-020-0595-9
Noor R, Islam Z, Munshi SK, Rahman F (2013) Influence of temperature on Escherichia coli growth in different culture media. J Pure Appl Microbiol 7:899–904. https://doi.org/10.1099/vir.0.049619-0
Peterson SN (2012) The HaloTag: improving soluble expression and applications in protein functional analysis. Curr Chem Genomics 6:8–17. https://doi.org/10.2174/1875397301206010008
Pontrelli S, Chiu TY, Lan EI, Chen FYH, Chang P, Liao JC (2018) Escherichia coli as a host for metabolic engineering. Metab Eng 50:16–46. https://doi.org/10.1016/j.ymben.2018.04.008
Qing G, Ma LC, Khorchid A, Swapna GVT, Mal TK, Takayama MM, Xia B, Phadtare S, Ke H, Acton T, Montelione GT, Ikura M, Inouye M (2004) Cold-shock induced high-yield protein production in Escherichia coli. Nat Biotechnol 22:877–882. https://doi.org/10.1038/nbt984
Rueden CT, Schindelin J, Hiner MC, DeZonia BE, Walter AE, Arena ET, Eliceiri KW (2017) Image J2: ImageJ for the next generation of scientific image data. BMC Bioinformatics 18:1–26. https://doi.org/10.1186/s12859-017-1934-z
Sahdev S, Khattar SK, Saini KS (2008) Production of active eukaryotic proteins through bacterial expression systems: a review of the existing biotechnology strategies. Mol Cell Biochem 307:249–264. https://doi.org/10.1007/s11010-007-9603-6
Saida F, Uzan M, Odaert B, Bontems F (2006) Expression of highly toxic genes in E. coli: special strategies and genetic tools. Curr Protein Pept Sci 7:47–56. https://doi.org/10.2174/138920306775474095
Sampaio MM, Chevance F, Dippel R, Eppler T, Schlegel A, Boos W, Lu YJ, Rock CO (2004) Phosphotransferase-mediated transport of the osmolyte 2-O-α -mannosyl-D-glycerate in Escherichia coli occurs by the product of the mngA (hrsA) gene and is regulated by the mngR (farR) gene product acting as repressor. J Biol Chem 279:5537–5548. https://doi.org/10.1074/jbc.M310980200
Tan J, Sastry AV, Fremming KS, Bjørn SP, Hoffmeyer A, Seo S, Voldborg BG, Palsson BO (2020) Independent component analysis of E. coli’s transcriptome reveals the cellular processes that respond to heterologous gene expression. Metab Eng 61:360–368. https://doi.org/10.1016/j.ymben.2020.07.002
Vera A, González-Montalbán N, Arís A, Villaverde A (2007) The conformational quality of insoluble recombinant proteins is enhanced at low growth temperatures. Biotechnol Bioeng 96:1101–1106. https://doi.org/10.1002/bit.21218
Wang Y, Hu L, Huang H, Wang H, Zhang T, Chen J, Du G, Kang Z (2020) Eliminating the capsule-like layer to promote glucose uptake for hyaluronan production by engineered Corynebacterium glutamicum. Nat Commun 11:1–10. https://doi.org/10.1038/s41467-020-16962-7
Yoon SH, Jeong H, Kwon SK, Kim JF (2009) Genomics, biological features, and biotechnological applications of Escherichia coli B: Is B for better? Systems Biology and Biotechnology of Escherichia coli. Springer, Dordrecht, pp 1–17
Yu G, Wang LG, Han Y, He QY (2012) ClusterProfiler: an R package for comparing biological themes among gene clusters. Omi A J Integr Biol 16:284–287. https://doi.org/10.1089/omi.2011.0118
Zhou L, Yu H, Wang K, Chen T, Ma Y, Huang Y, Li J, Liu L, Li Y, Kong Z, Zheng Q, Wang Y, Gu Y, Xia N, Li S (2020) Genome re-sequencing and reannotation of the Escherichia coli ER2566 strain and transcriptome sequencing under overexpression conditions. BMC Genomics 21:1–11. https://doi.org/10.1186/s12864-020-06818-1
Acknowledgements
We thank Mr. Rongwei Zhang for his support in sanger sequencing.
Funding
This work was supported by the National Natural Science Foundation of China (grant no. U1705283), the Natural Science Foundation of Fujian Province (grant no. 2017J07005) and the Major Research Plan of Xiamen City (grant no. 3502Z20203023).
Author information
Authors and Affiliations
Contributions
LZ, HY, and SL: conceived, designed research, and wrote the manuscript. LZ, YM, KW, and TC conducted the experiments and analyzed data. YH, LL, YL, JS, YH, and QZ analyzed the data. TL, ZK, and YW contributed new reagents or analytical tools. YG, JZ, and NX conceived or designed the study. All authors had read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, L., Ma, Y., Wang, K. et al. Omics-guided bacterial engineering of Escherichia coli ER2566 for recombinant protein expression. Appl Microbiol Biotechnol 107, 853–865 (2023). https://doi.org/10.1007/s00253-022-12339-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12339-6