Abstract
This study focuses on sirtuins, which catalyze the reaction of NAD+-dependent protein deacetylase, for riboflavin production in A. gossypii. Nicotinamide, a known inhibitor of sirtuin, made the color of A. gossypii colonies appear a deeper yellow at 5 mM. A. gossypii has 4 sirtuin genes (AgHST1, AgHST2, AgHST3, AgHST4) and these were disrupted to investigate the role of sirtuins in riboflavin production in A. gossypii. AgHST1∆, AgHST3∆, and AgHST4∆ strains were obtained, but AgHST2∆ was not. The AgHST1∆ and AgHST3∆ strains produced approximately 4.3- and 2.9-fold higher amounts of riboflavin than the WT strain. The AgHST3∆ strain showed a lower human sirtuin 6 (SIRT6)-like activity than the WT strain and only in the AgHST3∆ strain was a higher amount of acetylation of histone H3 K9 and K56 (H3K9ac and H3K56ac) observed compared to the WT strain. These results indicate that AgHst3 is SIRT6-like sirtuin in A. gossypii and the activity has an influence on the riboflavin production in A. gossypii. In the presence of 5 mM hydroxyurea and 50 µM camptothecin, which causes DNA damage, especially double-strand DNA breaks, the color of the WT strain colonies turned a deeper yellow. Additionally, hydroxyurea significantly led to the production of approximately 1.5 higher amounts of riboflavin and camptothecin also enhanced the riboflavin production even through the significant difference was not detected. Camptothecin tended to increase the amount of H3K56ac, but the amount of H3K56ac was not increased by hydroxyurea treatment. This study revealed that AgHst1 and AgHst3 are involved in the riboflavin production in A. gossypii through NAD metabolism and the acetylation of H3, respectively. This new finding is a step toward clarifying the role of sirtuins in riboflavin over-production by A. gossypii.
Key points
• Nicotinamide enhanced the riboflavin production in Ashbya gossypii.
• Disruption of AgHST1 or AgHST3 gene also enhanced the riboflavin production in Ashbya gossypii.
• Acetylation of H3K56 led to the enhancement of the riboflavin production in Ashbya gossypii.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ashbya gossypii is a natural riboflavin producer and has been utilized for the industrial production of riboflavin (Revuelta et al. 2017; Schwechheimer et al. 2016). To improve efficiency of production, some metabolic engineering has been carried out. Overexpression of RIB genes which encode enzymes in the riboflavin biosynthetic pathway enhances riboflavin production (Ledesma-Amaro et al. 2015). Moreover, deregulation of the expression of genes in its purine biosynthetic pathway also improves riboflavin production because guanosine triphosphate (GTP) is one of the precursors of riboflavin (Jimenez et al. 2005, 2008; Mateos et al. 2006).
Apart from metabolic engineering, other factors are also involved in riboflavin production by A. gossypii. Oxidative stress induced by exposure to H2O2 increases riboflavin production and light exposure also increases riboflavin production together with the accumulation of reactive oxygen species (ROS) (Silva et al. 2019; Walther and Wendland, 2012). Previously, we isolated a riboflavin-overproducing mutant by disparity mutagenesis; in this mutant, 33 homozygous and 1377 heterozygous mutations in open reading frames were found (Kato et al. 2020; Park et al. 2011). The genomic analysis of this mutant suggests that oxidative stress and the aging of cells may be involved in riboflavin over-production in this mutant because many mutations in genes involved in mitochondrial function, DNA mismatch repair, and oxidative stress response were found in addition to the increased number of ribosomal RNA gene repeat (Kato et al. 2020). These properties of cells showing compromised mitochondrial function and oxidative stress response are often shown in aged yeast and mammalian cells (Barja, 2019; Breitenbach et al. 2012) and the maintenance of the mitochondria function and oxidative stress response need several flavoproteins (Gudipati et al. 2014). These suggest that the aging may be connected with the riboflavin over-production in A. gossypii.
In Saccharomyces cerevisiae, whose genes show both homology and a particular pattern of synteny with more than 90% of A. gossypii genes (Dietrich et al. 2004), sirtuin controls aging (Wierman and Smith, 2014). Sirtuin is a member of the NAD+-dependent protein deacetylase family and is involved in longevity, energy metabolism, and stress responses (Wierman and Smith, 2014). S. cerevisiae has 5 sirtuins (Sir2, Hst1–4). Sir2 regulates transcriptional silencing in silent mating cassettes, homothallic mating left (HML) and homothallic mating right (HMR), and telomere length together with Sir3 and Sir4 (Wierman and Smith, 2014). In addition, ribosomal RNA gene repeats are silenced by Sir2 (Saka et al. 2013). This silencing is connected with replicative lifespan (Kaeberlein et al. 1999). Focusing on the relationship of sirtuin with metabolism, Sir2 deacetylates phosphoenolpyruvate carboxykinase (Pck1), leading to its inactivation and the regulation of gluconeogenesis (Casatta et al. 2013; Lin et al. 2009). Other sirtuins also deacetylate non-histone proteins and regulate metabolism and transcriptional silencing (Li et al. 2013; Madsen et al. 2015; Wierman and Smith, 2014).
In this study, the involvement of sirtuins in riboflavin production was investigated in A. gossipii. This fungus has four sirtuin genes and each was disrupted to reveal the functions of sirtuin for riboflavin production. This study describes the generation of a new type of riboflavin-overproducing mutant.
Materials and methods
Strains and growth conditions
Ashbya gossypii ATCC10895 was used as the WT strain. MT strain was isolated previously (Park et al. 2011). The fungus was cultivated in YD medium (1% glucose, 1% yeast extract, pH 6.8) and mycelia were kept at − 80 °C with 20% glycerol. To investigate the color and size of each strain, the glycerol stock was inoculated onto YD agar medium. Mycelia were isolated as 1 cm2 and put into medium. The additives except for camptothecin were dissolved with sterile water and added to medium at each concentration. Camptothecin was dissolved with methanol and methanol was added as a negative control at the same volume as camptothecin. For the investigation of riboflavin production in each strain, 0.3 mL of the glycerol stock was inoculated into 30 mL of liquid YD medium and cultivated for 24 h at 100 rpm. As a pre-culture, 0.3 mL of the culture medium was inoculated into 30 mL of liquid YD medium and cultivated for 24 h at 100 rpm. Then, 0.5 ml of the pre-culture medium was inoculated into 50 mL of the liquid YD medium and cultivated at 100 rpm.
Spore isolation was carried out according to our previous paper (Tajima et al. 2009). In brief, mycelia were suspended in 0.5 mL sterile water followed by the addition of 0.25 mL of 15 mg/ml Zymolyase 40-T (Seikagaku Co., Tokyo, Japan). After incubation at 37 °C for 30 min, spores were pelleted by centrifugation. The pellet was washed with 0.03% Triton X-100 two times and resuspended in 0.03% Triton X-100 containing 15% glycerol.
Transformation of A. gossypii to disrupt each sirtuin gene
To disrupt each sirtuin gene, the transformation in A. gossypii was carried out according to our previous paper (Wendland et al. 2000). In brief, 300 µL of spores were inoculated into 100 mL of complete medium (2% glucose, 1% polypeptone, 1% yeast extract) and cultivated for approximately 24 h at 100 rpm. After mycelia were collected and washed with sterile water, they were suspended in 40 mL of 50 mM potassium phosphate buffer (pH 7.5) containing 25 mM 2-mercaptoethanol. The mycelia were incubated for 30 min at 28 °C at 100 rpm and collected. Mycelia were washed with STM buffer (10 mM Tris–HCl, pH 7.5, 270 mM sucrose, and 1 mM MgCl2) and suspended in 120 µL of the same buffer. DNA (several micrograms) were put into the mycelia and electroporation performed using Gene-Pulser Xcell System (Bio-Rad, Hercules, CA, USA) with settings of 1.5 kV, 500 Ω, and 25 μF using 2 mm pre-chilled cuvettes (Bio-Rad). Mycelia were collected from the cuvette and suspended with 1 mL of full medium. Mycelia were cultivated for 20 min at 100 rpm, followed by inoculation onto TD agar medium. After the plates were incubated at 28 °C for 6 h, YD agar medium (0.6% agar) containing 300 μg/ml G418 was overlaid onto the plates. Spores were collected from mycelia grown in the presence of G418 and homozygous gene-disrupted strains were isolated. The gene disruption was confirmed by PCR using each primer set (Table 1).
To disrupt each sirtuin gene, kanamycin resistance gene expression cassette containing 50 bp of the target gene at the 5ʹ and 3ʹ ends was prepared by PCR. In this PCR, the forward primer containing 50 bp of 5ʹ region of the target gene and reverse primer containing 50 bp of 3ʹ region of the target gene were used (Table 1). As a template DNA, pYPKT vector was used (Kato and Park, 2006).
Riboflavin and protein measurement
The concentration of riboflavin produced in A. gossypii was measured according to our previous paper (Jeong et al. 2015). In brief, 0.8 mL of the culture broth was picked up and mycelia were disrupted by sonication. Then, the suspension was centrifuged and supernatant collected. The supernatant contained extracellular and intracellular riboflavin. The supernatant was thoroughly mixed with 0.2 mL of 1 N NaOH. A 0.4 mL aliquot of the solution was neutralized with 1 mL of 0.1 M potassium phosphate (pH 6.0), and its absorbance at 444 nm was measured. The riboflavin concentration was calculated using its extinction coefficient of 1.04 × 10−2 M−1 cm−1 (127 mg riboflavin/L at ABS444).
Protein concentration of the solution was measured using Pierce BCA Protein Assay Kit (Thermo Fisher Scientific K. K. Tokyo, Japan).
Sirtuin assay
SIRT6-like activity assay was carried out using CycLex® SIRT6 Deacetylase Fluorometric Assay Kit Ver.2 (MEDICAL & BIOLOGICAL LABORATORIES, Nagoya, Japan). Briefly, 200 mg of mycelia cultivated in liquid YD medium for approximately 24 h were suspended with 3 mL of 10 mM sodium phosphate buffer (pH 5.8) containing 1.2 M magnesium sulfate and 1.6 mg/mL Zymolyase-40 T to prepare its protoplasts. The protoplasts were washed twice with 10 mM Tris–HCl (pH 7.5) containing 1 M sorbitol. Then, the protoplasts were disrupted with RIPA buffer (50 mM Tris–HCl, pH 7.4, 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate). The homogenate was centrifuged and the supernatant used to measure sirtuin activity. For this assay, 8.5 µg of protein in each sample was used.
SDS-PAGE and western blot
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) was performed using 15% polyacrylamide gel. Gels were stained with coomassie brilliant blue R250 (CBB R250). In the case of western blot, proteins were transferred onto a nitrocellulose membrane using the Mini Trans-Blot Electrophoretic Transfer Cell (Bio-Rad) after SDS-PAGE. Blocking were carried out with 5% skimmed milk in Tris-buffered saline containing 0.1% Tween 20 (TBST, pH 7.6). The PVDF membrane was incubated with TBST containing the primary antibody. As a primary antibody, 500-fold-diluted rabbit anti-Histone H3 antibody (GeneTex, Irvine, CA, USA), 2000-fold-diluted rabbit anti-Histone H3K56ac antibody (Active Motif, Carlsbad, CA, USA), and 2000-fold-diluted rabbit anti-Histone H3K9ac antibody (Active Motif) were used. After the membrane was washed with TBST three times, the membrane was incubated with 15,000 fold-diluted goat Anti-IgG (H + L chain) conjugated with horseradish peroxidase (HRP) (MEDICAL & BIOLOGICAL LABORATORIES). Detection was based on the HRP reaction and carried out using Immobilon Western Chemiluminescent HRP Substrate (Merck Millipore Japan, Tokyo, Japan). Protein bands were detected on a Fluor-S/MAX imager (Bio-Rad). Densitometry analysis was performed by ImageJ (US National Institutes of Health).
NAD measurement
The amount of intracellular total NAD was measured using a NAD/NADH Assay Kit-WST (Dojindo, Kumamoto, Japan). The homogenate was prepared by sonication of mycelia grown for 24 h, centrifuged and the supernatant used as the sample.
Statistics analysis
Statistical analysis was carried out using GraphPad Prism 8 (GraphPad Software, San Diego, CA, USA). All data were analyzed for statistical significance by unpaired Student’s t test with two-side test. Error bars in each figure indicate standard deviation.
Results
Effects of a sirtuin inhibitor on riboflavin production
Sirtuin is a NAD+-dependent histone deacetylase (class III histone deacetylase) and NAD+ is converted to nicotinamide (NAM) during the deaceylation of histones. NAM is known as an inhibitor of sirtuins (Avalos et al. 2005). To inhibit sirtuins in A. gossypii, NAM was added into the culture medium (Fig. 1A). The addition of 5 or 10 mM NAM enhanced the intensity of the yellow color of A. gossypii, but the addition of the same concentration of nicotinic acid (NA) did not change the color of A. gossypii mycelia. NAM and NA are precursors of NAD+ in its salvage pathway (Orlandi et al. 2020) and only NAM changed the mycelial color of A. gossypii. In liquid culture, an approximately fivefold higher amount of riboflavin was detected in the presence of 10 mM NAM (Fig. 1B). These results suggest that the inhibition of sirtuins may lead to the over-production of riboflavin in A. gossypii which may be controlled by epigenetic regulation.
Growth and mycelial color of A.gossypii with each additive. A Effect of NAM and NA on the growth and mycelial color of A. gossypii. A. gossypii was cultivated in YD agar medium with each concentration of NAM or NA for 3 days. B Riboflavin production of A. gossypii in YD liquid medium with each concentration of NAM after 1 day cultivation. Significant difference was indicated by asterisks (*p < 0.05, **p < 0.01, n = 3)
Disruption of sirtuin genes in A. gossypii
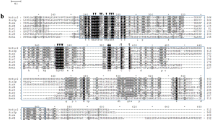
In A. gossypii, we found four sirtuin genes (AGOS_AEL013C, AGOS_AGL018C, AGOS_AEL229W, AGOS_AGL118W) using the amino acid sequences of five yeast sirtuins (SIR2, HST1, HST2, HST3, HST4) (Wierman and Smith, 2014). In Table. 1, amino acid sequences of each sirtuin gene in A. gossypii and S. cerevisiae are shown. From these sequences, AGOS_AEL013C, AGOS_AGL018C, AGOS_AEL229W, and AGOS_AGL118W genes were named as AgHST1, AgHST2, AgHST3, and AgHST4, respectively (Table 2). Then, disruption of each A. gossypii sirtuin gene was carried out by homologous recombination using a kanamycin (geneticin) resistant gene expression cassette (Fig. S1A) (Sugimoto et al.; Wendland et al. 2000). In the case of AgHST1, AgHST3, and AgHST4 genes, some homozygous geneticin-resistant colonies were grown. However, no homozygous geneticin-resistant colonies were observed when the AgHST2 gene was disrupted. To confirm each gene disruption, PCR was carried out using each primer set (Table 1) (Fig. S1A and B). The sizes of AgHST1, AgHST3, and AgHST4 genes are 1680, 1509, and 1245 bp, respectively. In each gene-disrupted strain, each sirtuin gene was replaced with a kanamycin-resistant gene expression cassette (1.5 kbp).
Riboflavin production of each gene-disrupted mutant
Colonies of AgHST1∆ and AgHST3∆ strains showed a deep yellow color compared to the WT strain, suggesting these strains produced a higher amount of riboflavin than the WT strain (Fig. 2A). The growth of AgHST1∆ and AgHST3∆ strains was slower on YD agar plates than the WT strain (Fig. 3B). AgHST1∆ and AgHST3∆ strains produced approximately 4.3- and 2.9-fold higher amounts of riboflavin at 72 h than the WT strain (Fig. 3C). Additionally, the AgHST4∆ strain also produced a slightly higher (1.3 fold) amount of riboflavin than the WT strain. The number of spores produced in each gene-disrupted mutant was almost the same as that of WT strain.
Growth and riboflavin production in the gene-disrupted mutants. A Colonies of each gene-disrupted mutant on YD agar plates. Each strain was cultivated for 3 days. B Size of each colony on YD agar plates. Each strain was cultivated for 3 days (n = 3). C Riboflavin production in each gene-disrupted mutant. Each mutant was cultivated in the YD liquid medium and the riboflavin concentration was measured according to the Materials and methods (n = 3). In each figure, significant difference was indicated by asterisks (*p < 0.05, ***p < 0.001,) and MT indicates the riboflavin over-producing mutant isolated previously (Park et al. 2011)
Properties of each gene-disrupted mutant. A SIRT6-like activity in each gene-disrupted mutant and MT strain cultivated for 1 day. For this assay using CycLex SIRT6 Deacetylase Fluorometric Assay Kit Ver.2, 8.5 µg of proteins in each protoplast homogenate was used (n = 3). B The amount of H3K56ac and H3K9ac in each gene-disrupted mutant cultivated for 1 day. For SDS-PAGE, 80 µg of proteins in the protoplast homogenate was used, followed by western blot. H3, H3K56ac, and H3K9ac were detected using anti-H3 antibody, anti-H3K56ac and anti-H3K9ac antibodies, respectively, according to the “Materials and methods” section. C Total NAD amount in each gene-disrupted mutant. Total NAD amount was measured using NAD/NADH Assay Kit-WST (n = 3). In each figure, significant difference was indicated by asterisks (*p < 0.05) and MT indicates the riboflavin over-producing mutant isolated previously (Park et al. 2011)
Properties of each gene-disrupted mutant
Sirtuin assay was also performed to confirm the gene disruption. We used the SIRT6 assay kit in this experiment because Hst3 and Hst4 have human SIRT6-like activity which catalyzes the acetylation of histone H3 lysine 56 (H3K56) (Bosch-Presegué and Vaquero, 2015; Wierman and Smith, 2014). SIRT6-like activities in AgHST3∆ and AgHST4∆ strains were reduced to 65% and 75%, respectively, compared to that in the WT strain, but the AgHST1∆ strain showed almost the same specific SIRT6 activity as the WT strain (Fig. 3A). This result indicates that AgHst3 and AgHst4 have SIRT6-like deacetylase activity and AgHst1 has other sirtuin deacetylase activity. The specific SIRT6 activity in the MT strain, which was isolated as a riboflavin-overproducing mutant previously (Park et al. 2011), was also lower than that in the WT strain. MT has a heterozygous missense mutation (I70T) in the AgHST3 gene (Kato et al. 2020). This mutation may cause the lower SIRT6-like deacetylase activity of the MT strain. In S. cerevisiae, Hst3 and Hst4 catalyzed the deacetylation of H3K56ac to maintain genome integrity (Wierman and Smith, 2014). Additionally, the deacetylation of H3K9ac is catalyzed by SIRT6 in mammalian cells (Bosch-Presegué and Vaquero, 2015). We investigated the amount of H3K56ac and H3K9ac using specific antibodies in each gene-disrupted strain (Fig. 3B). In AgHST3∆ strain, increased amount of both H3K9ac and H3K56ac was detected in AgHST3∆ strain compared to the WT strain, as well as in MT strain. This result indicates that AgHst3 catalyzes the deacetylation of both H3K56ac and H3K9ac and further suggests its disruption may lead to improved riboflavin production by increasing the amount of both H3K56ac and H3K9ac.
In S. cerevisiae, Hst1 controls the amount of intracellular NAD+ regulating expression of the BNA2 gene encoding indoleamine 2,3-dioxygenase, which catalyzes the first reaction in de novo NAD+ biosynthesis from tryptophan (Bedalov et al. 2003). We measured the total intracellular amount of NAD (NAD+ and NADH) in each strain. The AgHST1∆ strain had approximately a 1.3-fold higher amount of total NAD than the WT strain, but AgHST3∆ strain had almost the same amount as WT strain (Fig. 3C). On the other hand, the total amount of NAD in AgHST4∆ was reduced to 47% compared to that in the WT strain. This result indicates that the disruption of the AgHST1 gene led to the increase of total intracellular NAD.
Riboflavin production in the presence of hydroxyurea and camptothecin
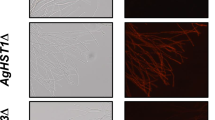
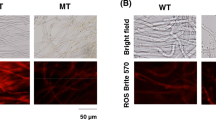
AgHST3∆ strain had higher amount of both H3K9ac and H3K56ac than WT strain, but other gene-disrupted strain did not (Fig. 3B). In S. cerevisiae, the disruption of HST3 and HST4 genes causes the hyperacetylation of H3K56 and the accumulation of spontaneous DNA damage (Celic et al. 2008). In mammalian cells, hydroxyurea, which inhibits ribonucleotide reductase, leading to stalled replication forks, the collapse of the forks and double-strand DNA breaks, also increases the amount of H3K56ac (Petermann et al. 2010; Singh and Xu, 2016; Yuan et al. 2009). In yeasts, sirtuin gene-disrupted mutants showed sensitivity to hydroxyurea (Konada et al. 2018; Simoneau et al. 2015). To investigate the involvement of DNA damage in riboflavin production in A. gossypii, it was cultivated in the presence of hydroxyurea. In solid medium, colonies of A. gossypii showed a more yellowish color following cultivation for a week in the presence of 5 mM hydroxyurea, compared to the control (0 mM), even though growth in the presence of 5 mM hydroxyurea was slower (Fig. 4A). In liquid medium, addition of 5 mM hydroxyurea allowed A. gossypii to produce approximately a 1.5-fold higher amount of riboflavin than the control (0 mM) (Fig. 4B). Along with hydroxyurea, camptothecin known as a topoisomerase I inhibitor that causes double-strand DNA breaks increases the amount of H3K56ac in yeast and mammalian cells (Masumoto et al. 2005; Yuan et al. 2009). In the presence of 50 µM of camptothecin, the color of the mycelia also showed a deeper yellow, indicating that camptothecin induces the production of riboflavin in A. gossypii (Fig. 4C). Additionally, in YD liquid medium, 50 µM camptothecin enhanced riboflavin production by 1.4-fold even through the difference was not statistically significant (p = 0.054) (Fig. 4D). Acetylation of H3K56 tended to be increased by 50 µM camptothecin, suggesting that DNA double-strand breaks may enhance riboflavin production in A. gossypii (Fig. 4E). However, 5 mM hydroxyurea did not increase the acetylation of H3K56, suggesting that hydroxyurea enhances riboflavin production in A. gossypii by unknown mechanism. The mycelia grown in the presence of hydroxyurea showed pale yellow color when N-acetyl-l-cysteine, which is a precursor of intracellular cysteine and glutathione and known as an anti-oxidant, was added (Fig. 4F) (Sun, 2010). These results suggest that the generation of reactive oxygen species (ROS) may enhance the riboflavin production in A. gossypii by hydroxyurea instead of DNA double-strand breaks. However, the quantification of ROS in the mycelia using some specific dyes was not successful (Data not shown).
Effects of hydroxyurea and camptothecin on the growth and the riboflavin production in A. gossypii. A Growth on the YD agar medium in the presence of hydroxyurea. A. gossypii was cultivated for a week. B Riboflavin production in the YD liquid medium with 5 mM hydroxyurea. WT strain was cultivated in the YD liquid medium with 5 mM hydroxyurea for 2 days and the riboflavin concentration and the dry mycelial weight were measured (n = 3). C Growth on the YD agar medium in the presence of camptothecin. As a control, methanol was added to the YD agar medium at the same volume (5%) as 50 µM of camptothecin. A. gossypii was cultivated for a week. D Riboflavin production in the YD liquid medium with 50 µM camptothecin. WT strain was cultivated in the YD liquid medium with 50 µM camptothecin for 2 days and the riboflavin concentration and the dry mycelial weight were measured (n = 7). As a control, methanol was added to the YD agar medium at the same volume (5%) as 50 µM of camptothecin. The significant difference is not detected (p = 0.054). E Acetylation of H3K56 in the presence of 5 mM hydroxyurea or 50 µM camptothecin. For SDS-PAGE, 80 µg of proteins in the protoplast homogenate was used and western blot was carried out by the same method as Fig. 3B. In the case of camptothecin, methanol was added to the YD agar medium at the same volume (5%) as 50 µM of camptothecin as a control (0 mM). Densitometry analysis was carried out using ImageJ (n = 3). F Growth of A. gossypii in the presence of hydroxyurea and N-acetyl-l-cysteine. Each concentration of N-acetyl-l-cysteine was added to the YD agar medium with 5 mM hydroxyurea and A. gossypii was grown for 4 days
Discussion
Sirtuins are known as NAD+-dependent histone deacetylases involved in metabolism, DNA repair, and aging (Fiorino et al. 2014). In particular, yeast Sir2 has been identified as a longevity factor (Wierman and Smith, 2014). We previously reported that a riboflavin-overproducing mutant isolated by disparity mutagenesis has approximately 1400 homo- and heterozygous mutations in protein-encoding regions and exhibits the features of aging (Kato et al. 2020). In this study, to reveal the relationship of aging with riboflavin production in A. gossypii, we focused on sirtuins. Four sirtuin genes (AgHST1, AgHST2, AgHST3, AgHST4) were found in A. gossypii and these genes were individually disrupted. AgHST1∆, AgHST3∆, AgHST4∆ strains were isolated, but AgHST2∆ strain was not. AgHST1∆ and AgHST3∆ strains produced a higher amount of riboflavin in liquid YD medium than the WT strain, but the AgHST4∆ strain produced almost the same amount of riboflavin as the WT strain (Fig. 2C). In the case of the AgHST1∆ strain, the total intracellular amount of NAD was increased compared to the WT strain and other gene-disrupted strains (Fig. 3C). On the other hand, AgHST3∆ strain had lower SIRT6-like deacetylase activity than the WT strain and the other gene-disrupted strains (Fig. 3A). This indicates that the disruption of AgHST1 and AgHST3 genes increased riboflavin production via different routes. In the AgHST3∆ strain, the amount of H3K9ac and H3K56ac was increased by the reduction of SIRT6-like deacetylase activity as well as in the riboflavin-overproducing MT strain (Fig. 3B). However, in AgHST4∆ strain, lower SIRT6-like deacetylase activity was detected compared to in WT strain, but no increase of the amount of H3K9ac and H3K56ac was observed. Additionally, little enhancement of the riboflavin production was observed on the contrary to AgHST3∆ strain (Fig. 2C). These results indicate that the acetylation of H3K9 and H3K56 is involved in the riboflavin over-production in AgHST3∆ strain showing the deeper yellow color compared to the WT strain (Fig. 2A).
H3K56ac is a post-translational modification of histone H3 responsive to DNA damage. Increase of H3K56ac is observed when DNA double-strand breaks (and ultraviolet radiation) induce DNA damage that occurs in yeast and mammalian cells (Celic et al. 2008; Masumoto et al. 2005; Miller et al. 2010; Petermann et al. 2010; Singh and Xu, 2016; Yuan et al. 2009; Zhu et al. 2015). In addition to acetylation of H3K56, the acetylation of H3K9 is also involved in double-strand DNA breaks in fission yeast and mammalian cells (Bosch-Presegué and Vaquero, 2015; Yamada et al. 2013). Camptothecin, which causes DNA double-strand breaks, also induced riboflavin production in A. gossypii (Fig. 4C). A previous report described that riboflavin-overproducing mutants are sensitive to photo-induced DNA damage (Silva et al. 2019). These results suggest that DNA double-strand breaks and the acetylation of H3K56 may be important factors inducing riboflavin production in A. gossypii. Regarding camptothecin, A. gossypii is less sensitive than S. cerevisiae which cannot grow normally in the presence of 50 µM camptothecin (Puddu et al. 2017). This result suggests that riboflavin production in A. gosyypii may be involved in its resistance to camptothecin.
The addition of N-acetyl-l-cysteine led to the loss of the yellow color of mycelia in the presence of hydroxyurea, which induced the riboflavin production (Fig. 4F). Hydroxyurea also generates ROS and activate Yap and Arf regulons, which regulate redox and iron homeostasis, in S. cerevisiae (Dubacq et al. 2006; Singh and Xu, 2016). Hydroxyurea also induces DNA double-strand breaks, but H3K56ac was not increased in A. gossypii by hydroxyurea in this study even though the riboflavin production was enhanced by hydroxyurea (Fig. 4A, B, and E). The enhancement of the riboflavin production may not be caused by DNA double-strand breaks, but by ROS produced by hydroxyurea. It was reported previously that ROS is involved in the riboflavin production in A. gossypii (Silva et al. 2019). ROS is also one of important factors for the riboflavin production in A. gossypii.
Based on the identity of the amino acid sequences, AgHst1 is a homolog of yeast Sir2 and Hst1 (Table 2), but A. gossypii has only a single type of sirtuin, AgHst1. As well as yeast Hst1, AgHst1 controls the amount of intracellular NAD (Fig. 3C). NAD+ biosynthesis is regulated by purine metabolism and ATP synthesis in yeast and Bas1, a Myb-related transcription factor, upregulates de novo NAD+ and purine biosynthesis (Pinson et al. 2019; Zhang et al. 1998). In A. gossypii, the disruption of the AgBAS1 gene leads to enhanced riboflavin production even though the gene disruption promotes adenine-auxotrophy (Mateos et al. 2006). The purine biosynthesis pathway is important for riboflavin production in A. gossypii because guanosine triphosphate (GTP) is a precursor of riboflavin (Jimenez et al. 2005, 2008). In addition, NAD metabolism is regulated in human cells by epigenetic control (Anderson et al. 2017; Etchegaray and Mostoslavsky, 2016). Therefore, NAD and purine biosynthesis may be connected with riboflavin production in A. gossypii.
This study revealed first that two sirtuins, AgHst1 and AgHst3, control riboflavin production in A. gossypii via two routes, the acetylation of H3 and the enhancement of NAD biosynthesis. This finding leads to the elucidation of the mechanism of the riboflavin production in A. gossypii and the generation of new riboflavin-overproducing mutants of A. gossypii.
Data availability
All data analyzed in this study are shown in this published article including its supplementary information files.
References
Anderson KA, Madsen AS, Olsen CA, Hirschey MD (2017) Metabolic control by sirtuins and other enzymes that sense NAD+, NADH, or their ratio. Biochim Biophys Acta Bioenerg 1858:991–998. https://doi.org/10.1016/j.bbabio.2017.09.005
Avalos JL, Bever KM, Wolberger C (2005) Mechanism of sirtuin inhibition by nicotinamide: altering the NAD+ cosubstrate specificity of a Sir2 enzyme. Mol Cell 17:855–868. https://doi.org/10.1016/j.molcel.2005.02.022
Barja G (2019) Towards a unified mechanistic theory of aging. Exp Gerontol 124:110627. https://doi.org/10.1016/j.exger.2019.05.016
Bedalov A, Hirao M, Posakony J, Nelson M, Simon JA (2003) NAD+-dependent deacetylase Hst1p controls biosynthesis and cellular NAD+ levels in Saccharomyces cerevisiae. Mol Cell Biol 23:7044–7054. https://doi.org/10.1128/mcb.23.19.7044-7054.2003
Bosch-Presegué L, Vaquero A (2015) Sirtuin-dependent epigenetic regulation in the maintenance of genome integrity. FEBS J 282:1745–1767. https://doi.org/10.1111/febs.13053
Breitenbach M, Laun P, Dickinson JR, Klocker A, Rinnerthaler M, Dawes IW, Aung-Htut MT, Breitenbach-Koller L, Caballero A, Nyström T, Büttner S, Eisenberg T, Madeo F, Ralser M (2012) The role of mitochondria in the aging processes of yeast. Subcell Biochem 57:55–78. https://doi.org/10.1007/978-94-007-2561-43
Casatta N, Porro A, Orlandi I, Brambilla L, Vai M (2013) Lack of Sir2 increases acetate consumption and decreases extracellular pro-aging factors. Biochim Biophys Acta 1833:593–601. https://doi.org/10.1016/j.bbamcr.2012.11.008
Celic I, Verreault A, Boeke JD (2008) Histone H3 K56 hyperacetylation perturbs replisomes and causes DNA damage. Genetics 179:1769–1784. https://doi.org/10.1534/genetics.108.088914
Dietrich FS, Voegeli S, Brachat S, Lerch A, Gates K, Steiner S, Mohr C, Pöhlmann R, Luedi P, Choi S, Wing RA, Flavier A, Gaffney TD, Philippsen P (2004) The Ashbya gossypii genome as a tool for mapping the ancient Saccharomyces cerevisiae genome. Science 304:304–307. https://doi.org/10.1126/science.1095781
Dubacq C, Chevalier A, Courbeyrette R, Petat C, Gidrol X, Mann C (2006) Role of the iron mobilization and oxidative stress regulons in the genomic response of yeast to hydroxyurea. Mol Genet Genomics 275:114–124. https://doi.org/10.1007/s00438-005-0077-5
Etchegaray JP, Mostoslavsky R (2016) Interplay between metabolism and epigenetics: a nuclear adaptation to environmental changes. Mol Cell 62:695–711. https://doi.org/10.1016/j.molcel.2016.05.029
Fiorino E, Giudici M, Ferrari A, Mitro N, Caruso D, De Fabiani E, Crestani M (2014) The sirtuin class of histone deacetylases: regulation and roles in lipid metabolism. IUBMB Life 66:89–99. https://doi.org/10.1002/iub.1246
Gudipati V, Koch K, Lienhart WD, Macheroux P (2014) The flavoproteome of the yeast Saccharomyces cerevisiae. Biochim Biophys Acta 1844:535–544. https://doi.org/10.1016/j.bbapap.2013.12.015
Jeong BY, Wittmann C, Kato T, Park EY (2015) Comparative metabolic flux analysis of an Ashbya gossypii wild type strain and a high riboflavin-producing mutant strain. J Biosci Bioeng 119:101–106. https://doi.org/10.1016/j.jbiosc.2014.06.014
Jimenez A, Santos MA, Pompejus M, Revuelta JL (2005) Metabolic engineering of the purine pathway for riboflavin production in Ashbya gossypii. Appl Environ Microbiol 71:5743–5751. https://doi.org/10.1128/AEM.71.10.5743-5751.2005
Jimenez A, Santos MA, Revuelta JL (2008) Phosphoribosyl pyrophosphate synthetase activity affects growth and riboflavin production in Ashbya gossypii. BMC Biotechnol 8:67. https://doi.org/10.1186/1472-6750-8-67
Kaeberlein M, McVey M, Guarente L (1999) The SIR2/3/4 complex and SIR2 alone promote longevity in Saccharomyces cerevisiae by two different mechanisms. Genes Dev 13:2570–2580. https://doi.org/10.1101/gad.13.19.2570
Kato T, Park EY (2006) Expression of alanine:glyoxylate aminotransferase gene from Saccharomyces cerevisiae in Ashbya gossypii. Appl Microbiol Biotechnol 71:46–52. https://doi.org/10.1007/s00253-005-0124-5
Kato T, Azegami J, Yokomori A, Dohra H, El Enshasy HA, Park EY (2020) Genomic analysis of a riboflavin-overproducing Ashbya gossypii mutant isolated by disparity mutagenesis. BMC Genomics 21:319. https://doi.org/10.1186/s12864-020-6709-7
Konada L, Aricthota S, Vadla R, Haldar D (2018) Fission yeast sirtuin Hst4 functions in preserving genomic integrity by regulating replisome component Mcl1. Sci Rep 8:8496. https://doi.org/10.1038/s41598-018-26476-4
Ledesma-Amaro R, Serrano-Amatriain C, Jiménez A, Revuelta JL (2015) Metabolic engineering of riboflavin production in Ashbya gossypii through pathway optimization. Microb Cell Fact 14:163. https://doi.org/10.1186/s12934-015-0354-x
Li M, Valsakumar V, Poorey K, Bekiranov S, Smith JS (2013) Genome-wide analysis of functional sirtuin chromatin targets in yeast. Genome Biol 14:R48. https://doi.org/10.1186/gb-2013-14-5-r48
Lin Y, Lu J, Zhang J, Walter W, Dang W, Wan J, Tao SC, Qian J, Zhao Y, Boeke JD, Berger SL, Zhu H (2009) Protein acetylation microarray reveals that NuA4 controls key metabolic target regulating gluconeogenesis. Cell 136:1073–1084. https://doi.org/10.1016/j.cell.2009.01.033
Madsen CT, Sylvestersen KB, Young C, Larsen SC, Poulsen JW, Andersen MA, Palmqvist EA, Hey-Mogensen M, Jensen PB, Treebak JT, Lisby M, Nielsen ML (2015) Biotin starvation causes mitochondrial protein hyperacetylation and partial rescue by the SIRT3-like deacetylase Hst4p. Nat Commun 6:7726. https://doi.org/10.1038/ncomms8726
Masumoto H, Hawke D, Kobayashi R, Verreault A (2005) A role for cell-cycle-regulated histone H3 lysine 56 acetylation in the DNA damage response. Nature 436:294–298. https://doi.org/10.1038/nature03714
Mateos L, Jimenez A, Revuelta JL, Santos MA (2006) Purine biosynthesis, riboflavin production, and trophic-phase span are controlled by a Myb-related transcription factor in the fungus Ashbya gossypii. Appl Environ Microbiol 72:5052–5060. https://doi.org/10.1128/AEM.00424-06
Miller KM, Tjeertes JV, Coates J, Legube G, Polo SE, Britton S, Jackson SP (2010) Human HDAC1 and HDAC2 function in the DNA-damage response to promote DNA nonhomologous end-joining. Nat Struct Mol Biol 17:1144–1151. https://doi.org/10.1038/nsmb.1899
Orlandi I, Alberghina L, Vai M (2020) Nicotinamide, nicotinamide riboside and nicotinic acid-emerging roles in replicative and chronological aging in yeast. Biomolecules 10:604. https://doi.org/10.3390/biom10040604
Park EY, Ito Y, Nariyama M, Sugimoto T, Lies D, Kato T (2011) The improvement of riboflavin production in Ashbya gossypii via disparity mutagenesis and DNA microarray analysis. Appl Microbiol Biotechnol 91:1315–1326. https://doi.org/10.1007/s00253-011-3325-0
Petermann E, Orta ML, Issaeva N, Schultz N, Helleday T (2010) Hydroxyurea-stalled replication forks become progressively inactivated and require two different RAD51-mediated pathways for restart and repair. Mol Cell 37:492–502. https://doi.org/10.1016/j.molcel.2010.01.021
Pinson B, Ceschin J, Saint-Marc C, Daignan-Fornier B (2019) Dual control of NAD+ synthesis by purine metabolites in yeast. Elife 8:e43808. https://doi.org/10.7554/eLife.43808
Puddu F, Salguero I, Herzog M, Geisler NJ, Costanzo V, Jackson SP (2017) Chromatin determinants impart camptothecin sensitivity. EMBO Rep 18:1000–1012. https://doi.org/10.15252/embr.201643560
Revuelta JL, Ledesma-Amaro R, Lozano-Martinez P, Díaz-Fernández D, Buey RM, Jiménez A (2017) Bioproduction of riboflavin: a bright yellow history. J Ind Microbiol Biotechnol 44:659–665. https://doi.org/10.1007/s10295-016-1842-7
Saka K, Ide S, Ganley ARD, Kobayashi T (2013) Cellular senescence in yeast is regulated by rDNA noncoding transcription. Curr Biol 23:1794–1798. https://doi.org/10.1016/j.cub.2013.07.048
Schwechheimer SK, Park EY, Revuelta JL, Becker J, Wittmann C (2016) Biotechnology of riboflavin. Appl Microbiol Biotechnol 100:2107–2119. https://doi.org/10.1007/s00253-015-7256-z
Silva R, Aguiar TQ, Oliveira R, Domingues L (2019) Light exposure during growth increases riboflavin production, reactive oxygen species accumulation and DNA damage in Ashbya gossypii riboflavin-overproducing strains. FEMS Yeast Res 19: foy114. https://doi.org/10.1093/femsyr/foy114
Simoneau A, Delgoshaie N, Celic I, Dai J, Abshiru N, Costantino S, Thibault P, Boeke JD, Verreault A, Wurtele H (2015) Interplay between histone H3 lysine 56 deacetylation and chromatin modifiers in response to DNA damage. Genetics 200:185–205. https://doi.org/10.1534/genetics.115.175919
Singh A, Xu JY (2016) The cell killing mechanisms of hydroxyurea. Genes (basel) 7:99. https://doi.org/10.3390/genes7110099
Sugimoto T, Kanamasa S, Kato T, Park EY (2009) Importance of malate synthase in the glyoxylate cycle of Ashbya gossypii for the efficient production of riboflavin. Appl Microbiol Biotechnol 83:529–539. https://doi.org/10.1007/s00253-009-1972-1
Sun SY (2010) N-acetylcysteine, reactive oxygen species and beyond. Cancer Biol Ther 9:109–110. https://doi.org/10.4161/cbt.9.2.10583
Tajima S, Itoh Y, Sugimoto T, Kato T, Park EY (2009) Increased riboflavin production from activated bleaching earth by a mutant strain of Ashbya gossypii. J Biosci Bioeng 108:325–329. https://doi.org/10.1016/j.jbiosc.2009.04.021
Walther A, Wendland J (2012) Yap1-dependent oxidative stress response provides a link to riboflavin production in Ashbya gossypii. Fungal Genet Biol 49:697–707. https://doi.org/10.1016/j.fgb.2012.06.006
Wendland J, Ayad-Durieux Y, Knechtle P, Rebischung C, Philippsen P (2000) PCR-based gene targeting in the filamentous fungus Ashbya gossypii. Gene 242:381–391. https://doi.org/10.1016/s0378-1119(99)00509-0
Wierman MB, Smith JS (2014) Yeast sirtuins and the regulation of aging. FEMS Yeast Res 14:73–88. https://doi.org/10.1111/1567-1364.12115
Yamada S, Ohta K, Yamada T (2013) Acetylated Histone H3K9 is associated with meiotic recombination hotspots, and plays a role in recombination redundantly with other factors including the H3K4 methylase Set1 in fission yeast. Nucleic Acids Res 41:3504–3517. https://doi.org/10.1093/nar/gkt049
Yuan J, Pu M, Zhang Z, Lou Z (2009) Histone H3–K56 acetylation is important for genomic stability in mammals. Cell Cycle 8:1747–1753. https://doi.org/10.4161/cc.8.11.8620
Zhang F, Kirouac M, Zhu N, Hinnebusch AG, Rolfes RJ (1998) Evidence that complex formation by Bas1p and Bas2p (Pho2p) unmasks the activation function of Bas1p in an adenine-repressible step of ADE gene transcription. Mol Cell Biol 17:3272–3283. https://doi.org/10.1128/mcb.17.6.3272
Zhu Q, Battu A, Ray A, Wani G, Qian J, He J, Wang Q, Wani AA (2015) Damaged DNA-binding protein down-regulates epigenetic mark H3K56Ac through histone deacetylase 1 and 2. Mutat Res 776:16–23. https://doi.org/10.1016/j.mrfmmm.2015.01.005
Funding
This work was supported by JSPS KAKENHI (Grant-in-Aid for Scientific Research (C) Grant Number JP21K05390).
Author information
Authors and Affiliations
Contributions
TK, HAE, and EYP conceived and designed this research and the experiments. JA and MK performed all experiments. TK, HAE, and EYP wrote this manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
The authors declare that no human participants or animals were used for the purpose of this study.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kato, T., Azegami, J., Kano, M. et al. Effects of sirtuins on the riboflavin production in Ashbya gossypii. Appl Microbiol Biotechnol 105, 7813–7823 (2021). https://doi.org/10.1007/s00253-021-11595-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11595-2