Abstract
Previous studies have characterized bacterial communities in menthol versus non-menthol cigarettes. However, these studies evaluated commercial cigarettes, for which levels of chemical constituents are largely unknown, and therefore, could not assess the impact of varying nicotine and menthol concentrations on tobacco bacterial communities. To address this knowledge gap, we performed time-series experiments using SPECTRUM research cigarettes with varying nicotine and menthol levels. Cigarettes were incubated under three storage conditions for 14 days. Cigarette tobacco was then sub-sampled (n = 288), DNA extracted, and subjected to PCR amplification of the V3V4 region of the 16S rRNA gene, followed by Illumina HiSeq sequencing. Sequences were analyzed using QIIME and R. Incubation under varying conditions did not affect bacterial diversity. However, significant differences in bacterial communities were observed across varying nicotine concentrations in menthol and non-menthol products. For example, Pseudomonas spp. was negatively correlated with nicotine concentrations in menthol cigarettes. A significantly higher relative abundance of P. veronii and P. viridiflava was observed in menthols versus non-menthols, while a significantly higher relative abundance of Bacillus foraminis and B. coagulans was found in non-menthols versus menthols. Additional bacteria (e.g., Staphylococcus spp., Jeotgalicoccus psychrophilus, and B. flexus) significantly changed in relative abundance between days 0 and 14. Our findings demonstrate that nicotine and menthol levels have a significant impact on the relative abundance of potential bacterial pathogens present in cigarettes. Future work is needed to demonstrate whether these tobacco-associated bacteria could be transferred to users while smoking, ultimately contributing to adverse respiratory impacts.
Key points
• Varying nicotine levels changes bacterial composition of research cigarettes.
• Mentholation affects the tobacco bacterial microbiome.
• SPECTRUM research cigarettes are dominated by Pseudomonas and Bacillus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tobacco use and secondhand smoke are responsible for approximately 443,000 deaths per year in the USA (CDC Tobacco Free 2018). Under the Family Smoking Prevention and Tobacco Control Act (FSPTCA) of 2009, the Food and Drug Administration (FDA) established several measures to expand understanding of traditional and new tobacco products, as well as the use by and sale of these products to children, aiming to reduce the number of tobacco-related diseases and deaths (US FDA 2020). Among these measures, identifying harmful and potentially harmful constituents (HPHCs) of tobacco products, such as nicotine, has been a major focus in order to reduce the addictiveness and harm of these combustible tobacco products.
Nicotine, the main addictive agent in tobacco products, is known to contribute to acute cardiovascular events (Benowitz and Burbank 2016) and deleterious pulmonary effects, including acute inflammation, decreased lung endothelial cell proliferation, and loss of endothelial barrier function (Schweitzer et al. 2015). Nicotine is an alkaloid stimulant that acts as an agonist of nicotinic acetylcholine receptors in the peripheral and central nervous systems, driving the products’ chronic use (Benowitz 2010). Nicotine content of cigarettes varies greatly, from as low as 6 mg of nicotine per gram of tobacco to as high as 28 mg/g, with an average of 10–12 mg/g (Benowitz and Henningfield 2013). The amount of nicotine needed to make cigarettes minimally addictive is 0.4 to 0.5 mg/g of tobacco in traditional cigarettes (Donny et al. 2015). This has led to the production of “very low nicotine content” (VLNC) cigarettes (Donny et al. 2015) with a maximum nicotine concentration of 0.7 mg/g of tobacco dry weight.
VLNC and regular cigarettes are inexpensive and extremely effective nicotine delivery systems, which are made more palatable and sensory-friendly by the addition of flavors such as menthol, ultimately reinforcing the addiction potential of cigarettes. While menthol has cooling properties (Campero et al. 2009) and inhibits nicotine metabolism (Benowitz et al. 2004), it significantly increases the uptake of nicotine and N′-nitrosonornicotine (NNN) (Squier et al. 2010). To achieve a slight menthol effect, a minimum of 1–2 mg of menthol per gram of tobacco is needed (Heck 2010). However, mentholated cigarettes generally contain menthol in the range of 2.9–7.2 mg per cigarette. Non-mentholated cigarettes have also been shown to contain menthol, albeit at much lower concentrations compared to mentholated cigarettes (Ai et al. 2016).
Interestingly, menthol also has antimicrobial properties (Jirovetz et al. 2009; Singh et al. 2015) and could therefore impact a widely understudied type of constituents present in cigarette tobacco: bacteria. Several studies have shown that nicotine can also limit the growth of certain bacteria and fungi (Pavia et al. 2000; Narayanappa Athmaram 2016; El-Ezmerli and Gregory 2019). However, these studies were performed using pure nicotine tested on individual bacterial isolates and did not test nicotine concentrations normally present in commercially sold cigarettes. Previous studies by our group have characterized the diversity of bacterial communities in menthol versus non-menthol tobacco products (Chopyk et al. 2017a; Malayil et al. 2020). However, these studies evaluated commercial cigarettes, for which levels of chemical constituents are largely unknown, and therefore, could not assess the impact of varying nicotine and menthol concentrations on tobacco bacterial communities.
To address this knowledge gap, we performed time series experiments using SPECTRUM research cigarettes with varying nicotine and menthol levels. Findings from this study will help inform future tobacco product standards that could place limits on the allowable nicotine and menthol content of commercially available cigarettes, as well as recognize and potentially regulate harmful bacterial constituents in these products.
Materials and methods
Sample collection and treatment
Twelve different SPECTRUM research cigarettes were obtained from the National Institute on Drug Abuse (NIDA, Bethesda, MD, USA) and stored in the original packaging after receipt. SPECTRUM research cigarettes were developed by NIDA in partnership with the Research Triangle Institute. These cigarettes have been well characterized in terms of their physical properties and chemical content, including menthol, nicotine, minor alkaloids, and several other major classes of HPHCs (Richter et al. 2016; Pappas et al. 2016; Ding et al. 2017). Six of the twelve were mentholated and six were non-mentholated, with varying nicotine levels (Table 1). Average nicotine concentrations in commercially available cigarettes range from 0.4 mg nicotine/g of tobacco (in VLNC cigarettes) to as high as 28 mg/g with an average of 10–12 mg/g. Our tested products were chosen to reflect the nicotine concentrations (1.0–15.0 mg/g) present in commercially available cigarettes. In addition, to keep within the range of menthol concentrations of 1–2 mg/g to obtain a slight menthol sensation, all of our tested menthol cigarettes were characterized by < 2.0 mg menthol/g of tobacco.
The cigarettes were incubated under three different temperature and relative humidity conditions for 14 days, as described in our previous studies (Chopyk et al. 2017b): (1) 25 °C and 30% relative humidity; (2) 20 °C and 50% relative humidity; and (3) 5 °C and 18% relative humidity. Tobacco from cigarettes was then sub-sampled under sterile conditions in replicates after 0, 5, 9, and 14 days of incubation (n = 288 total sub-samples).
DNA extraction
Total DNA was extracted under sterile conditions from all tobacco sub-samples (n = 288) using previously published procedures (Chopyk et al. 2017b). Briefly, 1ml of ice-cold 1X PBS buffer was added to 0.2 g of tobacco samples in lysing matrix tubes. Then the samples underwent an initial incubation step with lysozyme, mutanolysin and lysostaphin enzymes at 37 °C, and a second incubation with 10% SDS and proteinase K enzyme at 55 °C, before mechanical lysis using the MP Biomedical FastPrep 24 (Santa Ana, CA). DNA lysate was purified with the QIAmp DSP DNA mini kit 50, v2 (Qiagen, CA) using the recommended manufacturer’s protocol.
16S rRNA gene amplification and sequencing
The V3V4 region of the 16S rRNA gene was amplified and sequenced using the 319F (ACTCCTACGGGAGGCAGCAG) and 806R (GGACTACHVGGGTWTCTAAT) universal primers barcoded for each sample that also included a linker sequence required for Illumina HiSeq 300 bp paired-ends sequencing (Holm et al. 2019) and a 12-bp heterogeneity spacer index sequence. PCR amplification of sample DNA and negative controls was performed using thermocycler parameters as published previously (Chopyk et al. 2017a). Amplicon presence was confirmed with gel electrophoresis and amplicons were cleaned up with the SequelPrep Normalization Kit (Invitrogen Inc., Carlsbad, CA, USA). Samples were pooled at a concentration of 25 ng/PCR amplicon and sequenced on the Illumina HiSeq2500 (Illumina, San Diego, CA).
Sequence quality filtering
The initial sequencing dataset comprised a total of 10,806,817 reads obtained from 288 sub-samples generated with our experimental design, with an average of 37,523.67 (min. 14, max. 158,099) reads per sample. 16S rRNA reads were screened for low quality and short length and assembled using PANDAseq (Masella et al. 2012), demultiplexed, and chimera trimmed using UCHIME (Edgar et al. 2011). Quality reads were then incorporated into QIIME v1.9 (Caporaso et al. 2010) and clustered de-novo using VSEARCH, and taxonomies were assigned using the SILVA database v. 132, with a 0.97 confidence threshold. Downstream analyses were performed within RStudio (v. 0.99.473), using the Phyloseq (v. 1.19.1) and ggplot2 (v. 2.2.1) R packages. To ensure appropriate sequence coverage, the Good’s coverage cutoff was set at 0.9, and samples that received a number of sequences below that cutoff were removed for further downstream analyses. After quality filtering and removal of cyanobacterial sequences (20,425.91 sequences on average per sample [min. 71, max. 107,349]), the final sequencing dataset carried through downstream analyses comprised a total of 5,115,232 reads clustered into 1558 OTUs from 264 sub-samples, with an average of 18,138.5 (min 1085, max 107,323) reads per sample.
Statistical analysis
Alpha diversity was estimated using the Phyloseq R package (McMurdie and Holmes 2013) (v. 1.19.1) and tested for significance using ANOVA. Cumulative sum scaling (CSS) was used to normalize reads using the MetagenomeSeq (v. 1.16.0) package (Paulson et al. 2013). Beta diversity was estimated using the vegan and Phyloseq packages and tested for significance using ANOSIM. Statistical differences among bacterial OTUs relative abundances were calculated using the DESeq2 package (at alpha = 0.05) (Love et al. 2014). Unless otherwise noted, all values plotted are averages ± SE.
Results
Effect of incubation conditions on tobacco bacterial communities
Incubation under three different conditions of temperature and relative humidity for 14 days did not significantly affect alpha-diversity (observed richness and Shannon diversity; Fig. 1) (p > 0.05) and beta diversity (ANOSIM R = 0.0006, p = 0.338) (Fig. S1) measures of the tobacco-associated microbiotas. There was also no difference found across the conditions when comparisons were performed at each time point (Fig. S2). Therefore, all samples from the three incubation conditions for a given time point were combined, and we considered them as biological replicates for further downstream analyses.
Alpha diversity analysis of SPECTRUM cigarette products incubated over four time points (D0, D5, D9, and D14). Alpha diversity was measured for fridge (red), pocket (grey), and room (green) conditions using Observed richness and Shannon diversity metrics and compared using ANOVA with Tukey’s HSD post-hoc test
Bacterial composition of mentholated and non-mentholated SPECTRUM research cigarettes with varying nicotine levels
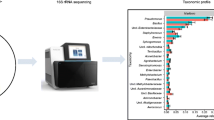
The predominant bacterial genera observed in these tobacco samples (all conditions and days combined) were Pseudomonas spp., Bacillus spp., Staphylococcus spp., Spingomonas spp., Enterobacter spp., Corynebacterium spp., and Brachybacterium spp. (Fig. 2). Comparisons of cigarettes with similar nicotine concentrations revealed that the relative abundance of Bacillus spp. was significantly (p < 0.05) different between menthol and non-mentholated products: NRC300 (3.2% ± 1.3) vs. NRC_301 (4.3% ± 2.0); NRC_400 (5.2% ± 1.1) vs. NRC_401 (8.1% ± 2.6); and NRC_404 (2.6% ± 0.4) vs. NRC_501 (4.8% ± 1.2). Relative abundance of Staphylococcus spp. was significantly (p < 0.05) different between NRC_300 (3.1% ± 0.2) vs. NRC_301 (3.4% ± 0.0.7) and NRC_600 (2.3% ± 0.3) vs. NRC_601 (3.6% ± 0.7). Relative abundance of Aerococcus spp. was found to be significantly different (p < 0.05) between NRC_300 (0.5% ± 0.2) vs. NRC_301 (3.2% ± 1.4) and NRC_404 (1.4% ± 0.5) vs. NRC_501 (3.6% ± 0.7). Relative abundance of Spingomonas spp. was also significantly (p < 0.05) different between NRC_102 (4.0% ± 0.6) vs. NRC_103 (2.8% ± 0.4) and NRC_404 (3.9% ± 0.5) vs. NRC_501 (3.2% ± 0.4) products. Relative abundance of Brachybacterium spp. was significantly different (p < 0.05) between NRC_404 (4.0% ± 0.3) vs. NRC_501 (4.0% ± 0.2). Finally, the differences in relative abundance of Pseudomonas spp. (p = 0.09) between NRC_102 vs. NRC_103 and that of Aerococcus spp. (p = 0.09) between NRC_200 vs. NRC_201 were nearly significant.
Average relative abundance (± SE) of top bacterial genera present in SPECTRUM cigarette products. Abundance was plotted for menthol (green) and non-menthol (grey) products under all incubation conditions and time points. Labels represent product names and their corresponding nicotine concentrations (mg/g). (*) represents ANOVA p value < 0.05
Effects of mentholation and duration of incubation
Alpha diversity metrics (Observed richness and Shannon diversity) were calculated on the sequencing dataset rarefied to 1085 sequences per sample. A significant effect of days of incubation was found for both Shannon diversity and Observed richness metrics (ANOVA p < 0.0001) in non-menthol products and only in Observed richness (ANOVA p = 0.004) in menthol products (Fig. S3). Comparing between days of incubation within mentholated cigarettes, the Observed species metric was statistically significantly different (Tukey HSD p < 0.05) between day 0 and day 9 samples, day 5 and day 9 samples, and day 9 and day 14 samples. Specifically, among these mentholated products, the highest diversity (Observed richness) was observed in the day 9 samples compared to the samples from the other days. There was no significant effect (Tukey HSD p > 0.05) of days of incubation on the Observed richness of species and Shannon diversity between day 0 and day 14 samples within mentholated cigarettes. Comparison within non-menthol products revealed significant (Tukey HSD p < 0.05) differences in Observed richness between day 0 and day 5 samples, as well as day 5 and day 14 samples. Specifically, among these non-mentholated products, the highest diversity was observed in the day 5 samples, when compared to that in the day 0 and day 14 samples. There was no significant effect (Tukey HSD p > 0.05) of days of incubation on Observed richness within non-menthol products between day 0 and day 14 samples. The Shannon diversity index for non-menthol products was significantly different (Tukey HSD p < 0.05) between day 0 and day 5 samples, day 0 and day 9 samples, and day 0 and day 14 samples. Specifically, day 5 samples were characterized by higher Shannon diversity when compared to the day 0 samples, and day 0 samples had higher diversity compared to day 9 and day 14 samples. Comparison of bacterial community structure using beta-diversity analysis of Bray-Curtis dissimilarity showed that mentholation (ANOSIM R: 0.060, p < 0.001) and days of incubation (ANOSIM R: 0.008, p < 0.001) had a significant effect on bacterial community composition (Fig. S4).
Differential abundance analysis using DeSeq2 revealed 77 OTUs with significantly different (p < 0.05) relative abundances between non-menthol and menthol products (Fig. 3, Table S1). Among these, 44 Gram-negative and 14 Gram-positive OTUs (total of 58 OTUs) were significantly higher in relative abundance in menthol products. 19 OTUs had a significantly higher relative abundance in the non-menthol products, among which 5 OTUs were Gram-negative and 14 was Gram-positive. Among menthol products, a higher relative abundance of Gram-positive Salana multivorans (OTU#91) (Fig. 3a), and Gram-negative Stenotrophomonas maltiphilia (OTU#790), Spingomonas yabuuchiae (OTU#1412), Pseudomonas viridiflava (OTU#1426), P. veronii (OTU#47), and Methylobacterium adhaesivum (OTU#32) were identified at the species level (Fig. 3b) when compared to the non-menthol products. Among non-menthol products, a higher relative abundance of Gram-positive Staphylococcus equorum (OTU#5), Oceanobacillus oncorhynchi (OTU#554), Corynebacterium stationis (OTU#14), Bacillus foraminis (OTU#708), and B. coagulans (OTU#111) (Fig. 3a) and Gram-negative Alcaligenes faecalis (OTU#174) and Acinetobacter schindleri (OTU#292) (Fig. 3b) were identified at the species level when compared to the menthol products.
Relative abundances of bacterial OTUs that were statistically significantly different (p < 0.05) between non-menthol and mentholated products. OTUs are colored by phylum and differentiated by (a) Gram positive and (b) Gram negative classification. A positive log2-fold change value denotes an OTU that is significantly higher in mentholated products, while a negative log2-fold change indicates an OTU that is significantly higher in non-mentholated products. The grey line and arrows highlight the conversion in log2-fold change from negative to positive values
Comparing within the menthol products, two OTUs were at a significantly higher relative abundance in day 14 samples, and five OTUs in the day 0 samples (Fig. 4a). Comparing within the non-menthol products, five OTUs were at a statistically significantly higher relative abundance in day 14 samples, while four OTUs were higher in the day 0 samples (Fig. 4b). Gram-positive Jeotgalicoccus psychrophilus (OTU#28) and Bacillus flexus (OTU#37) were at a statistically significantly (p < 0.05) higher relative abundance in non-menthol products in day 0 samples when compared to day 14 samples (Fig. 4b). Other OTUs at the genera level such as Oceanobacillus (OTU#99) and Staphylococcus (OTU#2350) showed higher relative abundance in day 0 samples from menthol products but lower relative abundance in day 0 samples from non-menthol products.
Relative abundances of bacterial OTUs that were statistically significantly different (p < 0.05) between day 0 and day 14 incubation. OTUs are colored by Gram classification and differentiated by (a) menthol and (b) non-menthol cigarettes. A positive log2-fold change value denotes an OTU that is significantly higher in day 0 samples, while a negative log2-fold change indicates an OTU that is significantly higher in day 14 samples. The grey line and arrows highlight the conversion in log2-fold change from negative to positive values
Among the day 0 samples, six OTUs were at a significantly (p < 0.05) higher relative abundance in the non-menthol products and 8 OTUs in the menthol products (Fig. 5a). B. coagulans (OTU#111) and B. clausii (OTU#11) were at a significantly higher relative abundance in the non-menthol products. Among the day 14 samples, 13 OTUs were at a higher relative abundance in the non-menthol products and 18 OTUs in the menthol products (Fig. 5b). S. equorum (OTU#5) and A. faecalis (OTU#174) were at a higher relative abundance in non-menthol products, while B. flexus (OTU#37) and P. viridiflava (OTU#9) were at a higher relative abundance in the menthol products.
Relative abundances of bacterial OTUs that were statistically significantly different (p < 0.05) between non-menthol and mentholated cigarettes. OTUs are colored by Gram classification and differentiated by (a) day 0 and (b) day 14 incubation. A positive log2-fold change value denotes an OTU that is significantly higher in day 0 samples, while a negative log2-fold change indicates an OTU that is significantly higher in day 14 samples. The grey line and arrows highlight the conversion in log2-fold change from negative to positive values
Effects of nicotine concentration on bacterial community composition
Alpha diversity metrics were calculated on the dataset rarefied to 1085 sequences per sample. Nicotine levels only affected the Observed richness (ANOVA p < 0.005). Comparison between menthol and non-menthol cigarettes of similar nicotine concentrations did not show significant differences (p > 0.05) in Observed richness, while comparisons between different nicotine concentrations for either menthol or non-menthol cigarettes showed significant differences (p < 0.05) in Observed richness. (Fig. 6, Table S2). Comparison of bacterial community structure using beta-diversity analysis of Bray-Curtis dissimilarity also showed that nicotine levels had a significant effect (ANOSIM R: 0.1509, p < 0.001) on bacterial community structure.
Alpha diversity analysis of menthol (green) and non-menthol (grey) SPECTRUM cigarette products containing different nicotine concentrations (mg/g). Diversity was measured after combining all incubation conditions (fridge, room and pocket) across all-time points together. Statistical significance between diversity indices was measured with ANOVA and Tukey’s HSD post-hoc test
Combining all conditions together, Pseudomonas spp. was the most abundant genus across all the samples at day 0 (Fig. 7). While the relative abundance of Pseudomonas spp. remained mostly constant with increasing nicotine levels in non-menthol cigarettes, Pseudomonas spp. decreased in relative abundance with increasing nicotine levels in the menthol products. There was no clear trend in the relative abundance of the rest of the seven bacterial genera with increasing nicotine concentrations on day 0. Bacillus spp. was present at a higher relative abundance in the day 14 mentholated products when compared to their day 0 counterparts. While Bacillus spp. increased in relative abundance at lower nicotine concentrations (< 2.38 mg/ml), it showed a decrease in relative abundance at higher nicotine concentrations (5.0–15.0 mg/ml). A slight increase in relative abundance of Corynebacterium spp. was also observed in day 14 mentholated products when compared to day 0 mentholated samples at nicotine concentrations of 2.38 mg/ml. The bacterial species that were found to be significantly (p < 0.05) differentially abundant between nicotine concentrations when comparing menthol and non-menthol cigarettes are listed in Tables S3 and S4, respectively.
Effect of tobacco characteristics on bacterial species
Pseudomonas spp. and bacteria from the Aurantimonadaceae family were found to be negatively correlated with nicotine concentration, and the Enterobacteriaceae family was negatively correlated with menthol (p < 0.05) (Fig. S5). Aurantimonadaceae was also negatively correlated (p < 0.05) with nornicotine and high-nicotine blends of tobacco (p < 0.05). While Pseudomonas spp. showed a significant positive correlation (p < 0.05) with low-nicotine tobacco blends, Enterobacter spp. showed a significant positive correlation (p < 0.05) with tar and total particulate matter (TPM) from tobacco smoke. Pseudomonas spp. showed a negative correlation with TPM and tar, but a positive correlation with mainstream smoke CO. Enterobacter spp. was also found to be significantly positively correlated with mainstream smoke, TPM and tar. Staphylococcus spp., Corynebacterium spp., Brachybacterium spp., and Bacillus spp. was found to be negatively correlated with mainstream smoke tar, TPM and CO, and positively correlated with menthol content on tobacco, although these results were not statistically significant.
Discussion
Multiple studies have characterized the complex array of bacterial communities in commercially available cigarette brands, and these studies have highlighted a potential role of chronic exposure to tobacco-associated bacteria in adverse health effects linked to tobacco use. However, because the exact composition of commercially available cigarettes is unknown and may even vary across products of the same brand, assessing the impact of specific constituents, such as nicotine and menthol, on shifts in tobacco-associated bacteria is challenging. Research SPECTRUM cigarettes have a known and standardized chemical composition, making them the ideal research tool to study the effect of individual constituents on tobacco bacterial communities. Our study provides a comprehensive characterization of the bacterial communities in mentholated and non-mentholated SPECTRUM research cigarettes with varying amounts of nicotine.
Confirming our previous studies with commercially available cigarettes (Chopyk et al. 2017a; Malayil et al. 2020), Pseudomonas bacteria were identified in all the tested SPECTRUM cigarettes. While Pseudomonas relative abundance was not affected by increasing nicotine levels in non-menthol cigarettes, a slight decrease in relative abundance was found in the menthol cigarettes as nicotine levels increased. The genus Pseudomonas comprises bacteria normally present in the environment, as well as those that are known opportunistic human pathogens, causing highly invasive diseases that can sometimes prove to be lethal. P. aeruginosa is known to persist for decades in individuals with cystic fibrosis, causing chronic lung infections (Faure et al. 2018), while multi-drug resistant P. putida is prevalent in nosocomial infections and exhibits antibiotic resistance genes that can be transferred to other bacteria in hospital environments (Li et al. 2010). Multiple strains of P. putida have also been identified as nicotine degrading, including those found on the surface of tobacco leaves (Sarris et al. 2012). P. viridiflava and P. veronii are two Pseudomonas species that were identified in our tested menthol cigarettes. P. viridiflava has been traditionally known as a multi-host phytopathogenic type of bacteria. Given that tobacco is a plant product, the presence of P. viridiflava was not unexpected. Additionally, we found that Pseudomonas spp. are negatively correlated (p < 0.05) with high nicotine blends of tobacco and nicotine concentrations. Nicotine-degrading bacteria (like Pseudomonas spp. Nic22) play an important role in reducing the harmful effects of nicotine and simultaneously maintain the desirable taste and flavor of the manufactured tobacco (Savini 2016). P. stutzeri, a rare opportunistic pathogen commonly found in soil and water, also was found in our tested products.
Our tested menthol cigarettes also harbored B. cereus across all of the tested nicotine concentrations. Bacillus cereus is a common non-pathogenic human gastrointestinal colonizer (Bottone 2010). Recent studies have also shown that it can cause pneumonia and resulting lung destruction, particularly in immunocompromised patients, including a case of necrotizing pneumonia (Savini 2016; Krupka et al. 2019). Other identified Bacillus species were B. clausii, B. foraminis, B. flexus, B. coagulans, and B. hortii. Other species were also identified in the tested products, such as Lactococcus garvieae, an emerging zoonotic pathogen that has been associated with endocarditis (Malek et al. 2019), as well as a case of meningitis (Tandel et al. 2017).
Previous studies have demonstrated that Spingomonas spp. can act as another human opportunistic pathogen (Angelakis et al. 2009). While S. melonis can utilize nicotine as its sole source of carbon, nitrogen, and energy, S. paucimobilis has been associated with nosocomial infections (Ryan and Adley 2010), particularly in immunocompromised patients. In our study, Spingomonas spp. was found to be negatively correlated with nicotine concentrations. Methylobacterium spp., another Gram-negative bacteria detected in our menthol products, has also been associated with healthcare-associated infections including systemic infections and pneumonia (Lai et al. 2011). Methylobacterium spp. has been isolated from variable environments including soil, dust, and aquatic systems and has been shown to be chlorine resistant (Gray et al. 1983). Alcaligenes faecalis, another bacterial pathogen identified in the non-menthol products, has been traditionally associated with respiratory diseases in chickens. But in a 2016 case report, this bacterial species was isolated from bronchoalveolar lavage of a dengue patient (Agarwal et al. 2016). A. faecalis, usually a member of gastrointestinal commensals, can sometimes cause rare infections that can prove to be fatal. It has been shown to be the cause of pneumonia, bacteremia, and meningitis (Mordi et al. 2013). Acinetobacter schindleri, also identified in the tested cigarette products, is an emerging opportunistic human pathogen which is mostly associated with hospital acquired infections, particularly nosocomial pneumonia. Several Acinetobacter species (A. baumannii, A. lwoffii, A. johnsonii, A. junii) have been implicated in a wide range of infectious diseases including meningitis, endocarditis, and urinary tract infections (Joly-Guillou 2005), and some are emerging fish pathogens. A. baumannii, another bacterial species identified in the tested cigarettes, is frequently multi-drug resistant (Dijkshoorn et al. 2007).
In summary, this study provides a detailed characterization of the bacterial communities residing within menthol and non-menthol SPECTRUM cigarettes with varying levels of nicotine incubated over 14 days, reflecting a normal user storage period. Our findings demonstrate that nicotine concentration and mentholation have a significant impact on the relative abundance of several potential bacterial pathogens present in cigarettes. Many of these microorganisms have been shown to cause respiratory illnesses; therefore, future work is needed to demonstrate whether these tobacco-associated bacteria could be transferred to users while smoking, ultimately contributing to adverse respiratory impacts.
Data availability
Data concerning the samples included in this study are deposited in the NCBI BioProject database under BioProject accession numbers PRJNA635703.
References
Agarwal A, Sharma S, Vivek B, Vibha B, Agarwal M, Ajrun M (2016) First reported case of Alcaligenes faecalis isolated from bronchoalveolar lavage in a patient with dengue hemorrhagic fever. J Assoc Chest Physicians 5:51–55
Ai J, Taylor KM, Lisko JG, Tran H, Watson CH, Holman MR (2016) Menthol content in US marketed cigarettes. Nicotine Tob Res Off J Soc Res Nicotine Tob 18:1575–1580. https://doi.org/10.1093/ntr/ntv162
Angelakis E, Roux V, Raoult D (2009) Sphingomonas mucosissima bacteremia in patient with sickle cell disease. Emerging Infectious Diseases Journal 15(1). https://doi.org/10.3201/eid1501.080465
Benowitz NL (2010) Nicotine Addiction. N Engl J Med 9
Benowitz NL, Burbank AD (2016) Cardiovascular toxicity of nicotine: implications for electronic cigarette use. Trends Cardiovasc Med 26:515–523. https://doi.org/10.1016/j.tcm.2016.03.001
Benowitz NL, Henningfield JE (2013) Reducing the nicotine content to make cigarettes less addictive. Tob Control 22:i14–i17. https://doi.org/10.1136/tobaccocontrol-2012-050860
Benowitz NL, Herrera B, Jacob P (2004) Mentholated cigarette smoking inhibits nicotine metabolism. J Pharmacol Exp Ther 310:1208–1215. https://doi.org/10.1124/jpet.104.066902
Bottone EJ (2010) Bacillus cereus, a volatile human pathogen. Clin Microbiol Rev 23:382–398. https://doi.org/10.1128/CMR.00073-09
Campero M, Baumann TK, Bostock H, Ochoa JL (2009) Human cutaneous C fibres activated by cooling, heating and menthol. J Physiol 587:5633–5652. https://doi.org/10.1113/jphysiol.2009.176040
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
CDC Tobacco Free (2018) 2014 SGR: The health consequences of smoking—50 Years of Progress. In: Cent. Dis. Control Prev. https://www.cdc.gov/tobacco/data_statistics/sgr/50th-anniversary/index.htm
Chopyk J, Chattopadhyay S, Kulkarni P, Claye E, Babik KR, Reid MC, Smyth EM, Hittle LE, Paulson JN, Cruz-Cano R, Pop M, Buehler SS, Clark PI, Sapkota AR, Mongodin EF (2017a) Mentholation affects the cigarette microbiota by selecting for bacteria resistant to harsh environmental conditions and selecting against potential bacterial pathogens. Microbiome 5:22. https://doi.org/10.1186/s40168-017-0235-0
Chopyk J, Chattopadhyay S, Kulkarni P, Smyth EM, Hittle LE, Paulson JN, Pop M, Buehler SS, Clark PI, Mongodin EF, Sapkota AR (2017b) Temporal variations in cigarette tobacco bacterial community composition and tobacco-specific nitrosamine content are influenced by brand and storage conditions. Front Microbiol 8 https://doi.org/10.3389/fmicb.2017.00358
Dijkshoorn L, Nemec A, Seifert H (2007) An increasing threat in hospitals: multidrug-resistant Acinetobacter baumannii. Nat Rev Microbiol 5:939–951. https://doi.org/10.1038/nrmicro1789
Ding YS, Richter P, Hearn B (2017) Chemical characterization of mainstream smoke from SPECTRUM variable nicotine research cigarettes. https://www.ingentaconnect.com/content/trsg/trs/2017/00000003/00000001/art00008
Donny EC, Denlinger RL, Tidey JW, Koopmeiners JS, Benowitz NL, Vandrey RG, Al’Absi M, Carmella SG, Cinciripini PM, Dermody SS, Drobes DJ, Hecht SS, Jensen J, Lane T, Le CT MCFJ, Montoya ID, Murphy SE, Robinson JD, Stitzer ML, Strasser AA, Tindle H, Hatsukami DK (2015) Randomized trial of reduced-nicotine standards for cigarettes. N Engl J Med 373:1340–1349. https://doi.org/10.1056/NEJMsa1502403
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinforma Oxf Engl 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
El-Ezmerli NF, Gregory RL (2019) Effect of nicotine on biofilm formation of Streptococcus mutans isolates from smoking and non-smoking subjects. J Oral Microbiol 11:1662275. https://doi.org/10.1080/20002297.2019.1662275
Faure E, Kwong K, Nguyen D (2018) Pseudomonas aeruginosa in chronic lung infections: How to adapt within the host? Front Immunol 9. https://doi.org/10.3389/fimmu.2018.02416
Gray JG, Roberts JF, Dillman RC, Simmons DG (1983) Pathogenesis of change in the upper respiratory tracts of turkeys experimentally infected with an Alcaligenes faecalis isolate. Infect Immun 42:350–355. https://doi.org/10.1128/IAI.42.1.350-355.1983
Heck JD (2010) A review and assessment of menthol employed as a cigarette flavoring ingredient. Food Chem Toxicol 48:S1–S38. https://doi.org/10.1016/j.fct.2009.11.002
Holm JB, Humphrys MS, Robinson CK, Settles ML, Ott S, Fu L, Yang H, Gajer P, He X, McComb E, Gravitt PE, Ghanem KG, Brotman RM, Ravel J (2019) Ultrahigh-throughput multiplexing and sequencing of >500-base-pair amplicon regions on the Illumina HiSeq 2500 platform mSystems 4 https://doi.org/10.1128/mSystems.00029-19
Jirovetz L, Buchbauer G, Bail S, Denkova Z, Slavchev A, Stoyanova A, Schmidt E, Geissler M (2009) Antimicrobial activities of essential oils of mint and peppermint as well as some of their main compounds. J Essent Oil Res 21:363–366. https://doi.org/10.1080/10412905.2009.9700193
Joly-Guillou M-L (2005) Clinical impact and pathogenicity of Acinetobacter. Clin Microbiol Infect 11:868–873. https://doi.org/10.1111/j.1469-0691.2005.01227.x
Krupka PN, Henry L, Shah N, Manek G, Perkins ME (2019) Bacillus Cereus pneumonia with lung destruction in an immunocompetent patient. In: D48. CRITICAL CARE CASE REPORTS: INFECTION AND SEPSIS II. American Thoracic Society A6591–A6591
Lai C-C, Cheng A, Liu W-L, Tan C-K, Huang Y-T, Chung K-P, Lee M-R, Hsueh P-R (2011) Infections caused by unusual Methylobacterium species. J Clin Microbiol 49:3329–3331. https://doi.org/10.1128/JCM.01241-11
Li H, Li X, Duan Y, Zhang K-Q, Yang J (2010) Biotransformation of nicotine by microorganism: the case of Pseudomonas spp. Appl Microbiol Biotechnol 86:11–17. https://doi.org/10.1007/s00253-009-2427-4
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Malayil L, Chattopadhyay S, Kulkarni P, Hittle L, Clark PI, Mongodin EF, Sapkota AR (2020) Mentholation triggers brand-specific shifts in the bacterial microbiota of commercial cigarette products. Appl Microbiol Biotechnol 104:6287–6297. https://doi.org/10.1007/s00253-020-10681-1
Malek A, De la Hoz A, Gomez-Villegas SI, Nowbakht C, Arias CA (2019) Lactococcus garvieae, an unusual pathogen in infective endocarditis: case report and review of the literature. BMC Infect Dis 19:301. https://doi.org/10.1186/s12879-019-3912-8
Masella AP, Bartram AK, Truszkowski JM, Brown DG, Neufeld JD (2012) PANDAseq: paired-end assembler for illumina sequences. BMC Bioinformatics 13:31. https://doi.org/10.1186/1471-2105-13-31
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217. https://doi.org/10.1371/journal.pone.0061217
Mordi RM, Yusuf EO, Onemu SO, Igeleke CL, Odjadjare EE (2013) The prevalence of Alcaligenes faecalis, in bacteremia, meningitis and wound sepsis in a tertiary health care institution in western part of Nigeria. Int J Biotechnol 7
Narayanappa Athmaram T (2016) Evaluation of novel nicotine analogues for their anti-bacterial and anti-fungal activity. J Microbiol Exp 3 https://doi.org/10.15406/jmen.2016.03.00079
Pappas RS, Gray N, Gonzalez-Jimenez N, Fresquez M, Watson CH (2016) Triple quad-ICP-MS measurement of toxic metals in mainstream cigarette smoke from spectrum research cigarettes. J Anal Toxicol 40:43–48. https://doi.org/10.1093/jat/bkv109
Paulson JN, Stine OC, Bravo HC, Pop M (2013) Differential abundance analysis for microbial marker-gene surveys. Nat Methods 10:1200–1202. https://doi.org/10.1038/nmeth.2658
Pavia CS, Pierre A, Nowakowski J (2000) Antimicrobial activity of nicotine against a spectrum of bacterial and fungal pathogens. J Med Microbiol 49:675–676. https://doi.org/10.1099/0022-1317-49-7-675
Richter P, Steven PR, Bravo R, Lisko JG, Damian M, Gonzalez-Jimenez N, Gray N, Keong LM, Kimbrell JB, Kuklenyik P, Lawler TS, Lee GE, Mendez M, Perez J, Smith S, Tran H, Tyx R, Watson CH (2016) Characterization of SPECTRUM variable nicotine research cigarettes. Tob Regul Sci 2:94–105. https://doi.org/10.18001/TRS.2.2.1
Ryan MP, Adley CC (2010) Sphingomonas paucimobilis: a persistent gram-negative nosocomial infectious organism. J Hosp Infect 75:153–157. https://doi.org/10.1016/j.jhin.2010.03.007
Sarris PF, Trantas EA, Mpalantinaki E, Ververidis F, Goumas DE (2012) Pseudomonas viridiflava, a multi host plant pathogen with significant genetic variation at the molecular level. PLoS One 7:e36090. https://doi.org/10.1371/journal.pone.0036090
Savini V (2016) Chapter 6 - Bacillus cereus pneumonia. In: Savini V (ed) The Diverse Faces of Bacillus cereus. Academic Press 73–84
Schweitzer KS, Chen SX, Law S, Van Demark M, Poirier C, Justice MJ, Hubbard WC, Kim ES, Lai X, Wang M, Kranz WD, Carroll CJ, Ray BD, Bittman R, Goodpaster J, Petrache I (2015) Endothelial disruptive proinflammatory effects of nicotine and e-cigarette vapor exposures. Am J Phys Lung Cell Mol Phys 309:L175–L187. https://doi.org/10.1152/ajplung.00411.2014
Singh R, Shushni MAM, Belkheir A (2015) Antibacterial and antioxidant activities of Mentha piperita L. Arab J Chem 8:322–328. https://doi.org/10.1016/j.arabjc.2011.01.019
Squier CA, Mantz MJ, Wertz PW (2010) Effect of menthol on the penetration of tobacco carcinogens and nicotine across porcine oral mucosa ex vivo. Nicotine Tob Res 12:763–767. https://doi.org/10.1093/ntr/ntq084
Tandel K, Bhatt P, Ranjan P, Rathi KR (2017) Meningitis caused by Lactococcus garvieae. Med J Armed Forces India 73:94–96. https://doi.org/10.1016/j.mjafi.2015.08.004
U.S. FDA (2020) FDA announces comprehensive regulatory plan to shift trajectory of tobacco-related disease, death. In: FDA. http://www.fda.gov/news-events/press-announcements/fda-announces-comprehensive-regulatory-plan-shift-trajectory-tobacco-related-disease-death
Funding
This work was performed under Project 3 “Exploring Tobacco Microbial Constituents and the Oral Microbiome of Tobacco Users” (PI: AS and EM) as part of the University of Maryland Tobacco Center of Regulatory Science (UMD TCORS) “Rapid Response Characterization of New and Manipulated Tobacco Products” funded by the National Institute of Health (NIH) and the Food and Drug Administration (FDA) – Award # P50-CA-180523-01 (P. Clark, Center PI).
Author information
Authors and Affiliations
Contributions
SC performed bioinformatics analysis, wrote, and edited the manuscript. LM and SC performed laboratory analyses. ARS and EFM contributed to the study design, protocol development, data analysis, and manuscript preparation. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Disclaimer
Dr. Mongodin contributed to this article as an employee of the University of Maryland School of Medicine. The views expressed are his own and do not necessarily represent the views of the National Institutes of Health or the US Government
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Emmanuel F. Mongodin and Amy R. Sapkota shared senior co-authorship
Rights and permissions
About this article
Cite this article
Chattopadhyay, S., Malayil, L., Mongodin, E.F. et al. Nicotine concentration and mentholation affect bacterial community diversity in SPECTRUM research cigarettes. Appl Microbiol Biotechnol 105, 4241–4253 (2021). https://doi.org/10.1007/s00253-021-11327-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11327-6