Abstract
Pseudomonas aeruginosa is known as an opportunistic pathogen whose one of the antibiotic resistance mechanisms includes biofilm formation and virulence factor production. The present study showed that the sub-minimum inhibitory concentration (sub-MIC) of streptomycin inhibited the formation of biofilm and eradicated the established mature biofilm. Streptomycin at sub-MIC was also capable of inhibiting biofilm formation on the urinary catheters. In addition, the sub-MIC of streptomycin attenuated the bacterial virulence properties as confirmed by both phenotypic and gene expression studies. The optimal conditions for streptomycin to perform anti-biofilm and anti-virulence activities were proposed as alkaline TSB media (pH 7.9) at 35 °C. However, sub-MIC of streptomycin also exhibited a comparative anti-biofilm efficacy in LB media at similar pH level and temperature. Furthermore, this condition also improved the biofilm inhibition and eradication properties of streptomycin, tobramycin and tetracycline towards the biofilm formed by a clinical isolate of P. aeruginosa. Findings from the present study provide an important insight for further studies on the mechanisms of biofilm inhibition and dispersion of pre-existing biofilm by streptomycin as well as tobramycin and tetracycline under a specific culture environment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pseudomonas aeruginosa is known as one of the common causes of nosocomial bloodstream and urinary tract infections (Miyoshi-Akiyama et al. 2017; Talwalkar and Murray 2016). The bacterial pathogenesis which results in chronic infections is governed by biofilm formation and virulence factor production (Ben Haj Khalifa et al. 2011; Jamal et al. 2018; Kannan et al. 2018). Biofilm is a community of sessile cells enclosed within extracellular polymeric substances (EPS) which is composed of exopolysaccharides, lipid, proteins, and extracellular DNA (e-DNA) (Billings et al. 2015; Flemming et al. 2016). P. aeruginosa is able to form biofilm in the living host and on several clinical devices such as ventilators and catheters (Olejnickova et al. 2014; Ramirez-Estrada et al. 2016). As compared to the planktonic counterparts, the biofilm lifestyle provides protection to the bacterial cells from the host immune system, tolerance to extreme environmental conditions, and resistance to a majority of conventional antibiotics (Flemming and Wingender 2010; Mah and O'Toole 2001). In most cases, antibiotics resistance derived from biofilm formation is proposed to attribute to (1) the biofilm architecture and thickness, which minimize drug penetration and diffusion; (2) the slow metabolic rate, which lowers susceptibility to drug; and (3) the components such as negatively charged e-DNA, exopolysaccharides, and antibiotic-modifying enzymes, which hinder the penetration of antibiotics (Hall and Mah 2017; Mulcahy et al. 2008; Penesyan et al. 2015; Shaikh et al. 2015; Singh et al. 2016). Combining with the heterogenicity and rapid gene transfer capability of the bacterial community lying within the biofilm, there is no doubt that most of the traditional antibiotic therapies were unable to combat the spread of biofilm formation as well as other antibiotic resistance mechanisms.
Due to the complications in biofilm nature, one of the recent anti-biofilm approaches is proposed to target several virulence phenotypes (Allen et al. 2014; Pattnaik et al. 2018; Qiu et al. 2019). Instead of “killing” the bacteria, this anti-virulence approach attempts to attenuate a wide range of other virulence factors, e.g., pyocyanin, rhamnolipid, iron-acquiring siderophores, proteases, and hemolysins, which are vastly produced along with biofilm formation and also regulated by quorum sensing (QS) signaling system (Khan et al. 2019a), thereby weakening the bacterial pathogenicity, reducing the infection severity and combatting the correlated capability of forming biofilm (Fleitas Martinez et al. 2019; García-Contreras et al. 2013). In fact, this approach has been reported as a highly potential replacement for traditional antibiotic treatments as to some extent, it could reduce the selective pressure caused by antibiotics, which in turn limits the possibility of developing resistance by gene mutation (Cegelski et al. 2008; Parrino et al. 2018). Therefore, attenuating virulence properties could be considered as a potential anti-biofilm strategy; hence, extensive research for a novel effective anti-virulence agent has been on demand.
Anti-virulence agents and their efficacy are extremely varied in sources and active concentrations. Among the new sources of these agents, common antibiotics at their sub-minimum inhibitory concentrations (sub-MICs) were also recognized for their inhibitory activity towards numerous Gram-negative and Gram-positive bacteria (Bala et al. 2011; El-Mowafy et al. 2017; Gupta et al. 2016; Haddadin et al. 2010; Imperi et al. 2014; Vidya et al. 2005). Aminoglycosides with polycationic nature are one of the traditional antibiotics that are currently re-examined for their virulence inhibition potentials. A majority of aminoglycoside applications are now being used in combination therapy in which the antibiotic is conjugated with other chemical compounds or immobilized onto a carrier, which was previously proposed to help minimize the side effects which resulted from high concentration and frequent use of the antibiotic (Alipour et al. 2010; Chanda et al. 2017; Furiga et al. 2015). To our updated knowledge, although gentamycin, tobramycin, and fluoroquinolone either alone or in combination have been reported as effective anti-biofilm agents against P. aeruginosa, the prevalence of resistance emergence urges the search for alternative approaches such as combination, anti-virulence, and anti-quorum sensing strategies (Antoniu 2015; Borges et al. 2017; Chanda et al. 2017; Das et al. 2016; Parai et al. 2018). Currently, despite being known as a commonly used member of aminoglycoside family, the action on P. aeruginosa biofilm at sub-MIC of streptomycin has been limitedly known. Therefore, the present study has selected streptomycin to evaluate its performance at sub-MIC under different environmental conditions (pH, culture media, and temperature) towards P. aeruginosa biofilm formation and production of several virulence properties, thus suggesting the promising role of this antibiotic as an anti-virulence agent for treating P. aeruginosa infections.
Materials and methods
Bacterial strains, chemicals, and growth conditions
Pseudomonas aeruginosa PAO1 KCTC1637 was obtained from Korean Collection for Type Cultures (KCTC, Daejeon, Korea). P. aeruginosa GNUH-NCCP 6039 (isolated from pleural fluid) was purchased from Gyeongsang National University-Hospital Branch of the National Culture Collection for Pathogens (GNUH-NCCP, Gyeongsang, Korea). Cultivation of P. aeruginosa strains was performed in tryptic soy broth (TSB; Difco Laboratory Inc., Detroit, MI, USA), Mueller-Hinton broth (MHB), Luria-Bertani (LB) broth, and TSB-agar. Streptomycin was purchased from Sigma-Aldrich Co. (St. Louis, MO, USA). The pH of each media was adjusted to 5.9, 7.2, and 7.9 using HCl and NaOH. The stock concentration of streptomycin was maintained acidic (pH 5.9) by preparing in 10−5 N HCl. The temperature used for the growth of P. aeruginosa was 25 °C, 30 °C, and 35 °C in aerobic condition. The reagents and chemicals used in the whole experiment at the present were of analytical grade.
Determination of minimum inhibitory concentration and sub-MIC levels of streptomycin
For determination of minimum inhibitory concentration (MIC) of streptomycin, the P. aeruginosa cell culture grown overnight in TSB was diluted (1:100) in fresh and sterile TSB media (pH 7.9). The diluted culture (250 μl) was transferred to a 96-well microtiter plate in triplicates. Streptomycin at the concentrations ranged from 0.0625 to 16 μg ml−1 was then added to the microtiter plate. Subsequently, the microtiter plate was incubated at 35 °C under shaking condition (567 cycle per minute (cpm)) in microplate reader for 28 h. At the end of incubation, the optical density (OD) of the grown cell culture was measured at 600 nm of wavelength using the microplate reader (BioTek, Winooski, VT, USA).
Similarly, the sub-MIC of streptomycin was also determined by incubating the overnight grown P. aeruginosa cell culture (1:100 dilution) in a 96-well microtiter plate. The concentration of streptomycin was taken in the range from 0.0625 to 8 μg ml−1. The 96-well microtiter plate was incubated under shaking condition in microplate reader at 35 °C for 28 h and the OD was measured at 600 nm wavelength at every 2-h time interval.
Biofilm inhibition assays
The formation of biofilm by P. aeruginosa was quantitatively estimated by carrying out the crystal violet staining as discussed previously (Lee et al. 2011). The P. aeruginosa biofilm formation was carried out in TSB media (pH 7.9) by placing the diluted overnight grown cell culture (1:100 dilution) in a 96-well microtiter plate. The cells were treated with sub-MIC of streptomycin (ranging from 0.0625 to 4 μg ml−1) and incubated at 35 °C for 24 h. The biofilm inhibition assay by sub-MIC of streptomycin to P. aeruginosa was also tested in MHB and LB growth media at different pH (5.9, 7.2, and 7.9) and temperature (25, 30, and 35 °C). After incubation for 24 h, the growth of both planktonic and attached bacterial cells was measured at OD at 600 nm. The procedure of biofilm inhibition assay included three-time washing of the attached cells with distilled water after planktonic cells being discarded and staining with 0.1% crystal violet dye. The residual dye was removed by three-time washing with distilled water and the attached cells were re-suspended in 95% ethanol. The total biofilm-forming cells were measured by OD measurement at 570 nm. The experiment was performed in triplicates.
Biofilm assays on the surfaces of urinary catheter
Urinary catheter which is a medical device was also used to examine the effect of sub-MIC of streptomycin on P. aeruginosa biofilm formation as described earlier with a slight modification (Al-Mathkhury et al. 2011; Kart et al. 2017). Polyvinyl chloride urinary catheter was purchased from Hyupsung Medical Co., Ltd., Korea, and was aseptically cut into coupons (15 mm × 15 mm). The catheter coupons were placed in a 6-well plate containing overnight grown cell culture (with an initial OD600 of 0.05) in TSB (pH 7.9) and sub-MICs of streptomycin. The titer plate was incubated at 35 °C for 24 h. Two methods were adopted to quantify the biofilm cells: 1-crystal violet staining and 2-bacterial viable cells count. In crystal violet staining, the planktonic cells were removed and the cells attached on the coupons were washed three times with distilled water and stained with 0.1% aqueous crystal violet. After 20 min of staining, the residual crystal violet was discarded and the catheter coupons were washed thrice with distilled water. The stained cells present on the catheter coupons were dissolved into 95% ethyl alcohol in test tubes and the tubes were sonicated (10 min) and vortexed at high speed in order to detach the residual attached cells into solution. The OD of cell suspension was measured at 570 nm. In viable cell count method, the colony-forming unit (CFU) of biofilm cells was determined. Briefly, catheter coupons along with overnight grown P. aeruginosa cell culture and sub-MIC of streptomycin in 6-well plate were incubated at 35 °C for 24 h. After 24 h of incubation, the free-floating planktonic cells were discarded and the catheter coupons were washed three times with fresh TSB. Each catheter coupon was then placed in a sterile test tube containing 1 ml fresh TSB and was vortexed at high speed for 10 min. A serial dilution of cell suspension was carried out up to 10−7 dilutions in the fresh and sterile TSB. The diluted cell suspension (100 μl) was spread onto TSA agar plate, followed by incubation at 35 °C for 24. The colonies formed on the TSA agar plate were counted. The experiment was performed in triplicates.
Eradication of established mature biofilm
The dispersion of old established biofilm formed by P. aeruginosa was performed by inoculating the diluted (100-fold dilution) overnight grown cell culture in a 96-well microtiter plate. The experiment was performed in TSB media (pH 7.9) in three separated 96-well microtiter plates for the evaluation of dispersion at different time periods. The inoculated cells were allowed to establish biofilm without streptomycin during incubation at 35 °C up to 24 h under static condition (Li et al. 2017). After 6, 12, and 24 h of incubation, the planktonic cells were removed while the adhered cells were washed twice with TSB media. Each well was loaded with fresh TSB along with various concentrations of streptomycin ranging from 0.5 to 128 μg ml−1. After further incubation at 35 °C for 24 h, dispersion of the adhered cells was determined by staining with 0.1% aqueous crystal violet dye followed by OD measurement at 570 nm. The experiment was carried in triplicates.
Microscopic observation of biofilm cells
Scanning electron microscope
The biofilm inhibition property of streptomycin in P. aeruginosa was visualized by scanning electron microscope (SEM) following the previous protocol (Lee et al. 2011). Briefly, the overnight grown culture of P. aeruginosa cells at 100-fold dilution was allowed to grow over the nylon membrane surface (0.5 × 0.5 cm) which were positioned in a 24-well plate containing sub-MIC (2 μg ml−1) streptomycin. After incubation at 35 °C for 24 h, the cells were subjected to fixation with formaldehyde and glutaraldehyde for overnight at 4 °C. The planktonic cells were gently discarded and the attached cells were washed with phosphate buffer saline (PBS) (pH 7.4) for three times. These cells were further dehydrated with different concentrations of ethanol (50, 70, 80, 90, 95, and 100%), each for 20 min. The membranes with adhered cells were freeze-dried using a freeze dryer (model no. FD8518, ilShinBiobase Co. Ltd., Korea). The dried membranes were affixed to SEM stubs, followed by coating with white gold for 120 s using an ion sputter (E-1010, Hitachi, Tokyo, Japan). The biofilms formed on the nylon membrane surface were examined using the JSM-6490LV microscope (JEOL, Tokyo, Japan) at magnification of × 5000 and a voltage of 15 kV.
Fluorescence microscope
Similarly, the biofilm inhibition effect of streptomycin was also examined using fluorescence microscope. Visualization of cells was performed by using the acridine orange dye (Sigma-Aldrich St. Louis, MO, USA). The cells (1:100 dilution) were allowed to form biofilm over the surface of glass pieces (1 × 1 cm) kept in a 12-well plate along with sub-MIC (2 μg ml−1) streptomycin at 35 °C. The cells attached on the glass pieces were washed thrice with PBS and stained using acridine orange (10 μg ml−1). The residual dye was discarded by washing three times with PBS. The cells adhered to the glass pieces were examined by the Leica DMI300B fluorescence microscope (Leica Microsystems, Wetzlar, Germany) with × 40 magnification.
Assays of P. aeruginosa virulence factors
Hemolysis assay
The effect of streptomycin on P. aeruginosa hemolytic property was determined by following the methodology discussed previously (Lee et al. 2012). Firstly, the P. aeruginosa cell culture (1:100 dilutions with TSB) grown overnight was added in the 96-well microtiter plate along with sub-MICs of streptomycin. The microtiter plate was incubated overnight at 35 °C under shaking at 567 cpm. To perform the hemolysis assay, the grown cell culture (50 μl) was added into 950 μl of diluted sheep red blood cells (RBCs) (obtained from MBcell Ltd., Seoul, Korea) in centrifuge tubes (1.5 ml). The tubes were incubated at 35 °C with shaking (250 rpm). After incubation for 1 h, the hemolysis of the RBCs was quantitatively determined by measuring the OD of supernatant at 543 nm.
Assays for the virulence factor productions
The impacts of streptomycin on the production of numerous virulence factors such as rhamnolipid, pyocyanin, and siderophore, e.g., pyoverdine and LasA protease, synthesized by P. aeruginosa were also studied. To determine the production of rhamnolipid and pyocyanin, the overnight grown cell culture at the dilution of 1:100 in TSB was treated with sub-MIC streptomycin and incubated for 12 h at 35 °C under shaking (250 rpm). The orcinol colorimetric assay was performed for the quantification of rhamnolipid production at 421-nm wavelength following the previous protocol (Wilhelm et al. 2007). Determination of pyocyanin was carried out using the supernatant after extraction with chloroform according to the previous methodology (Essar et al. 1990). The extracted green-blue color turned into pink color as a result of acidification with 0.2 N HCl and the OD of the pink color was measured at 520 nm. For the quantification of pyoverdine production, iron-limited minimal salt media and 2% sodium succinate were used for the cultivation of P. aeruginosa in the presence and absence of streptomycin at sub-MIC. The OD of the culture supernatant was then measured at 405 nm (Stintzi et al. 1998). Production of each virulence factor from P. aeruginosa was estimated in triplicates.
Protease activity assay
The secretion and activity of P. aeruginosa protease were performed spectrophotometrically as well as on the casein agar plates following the previous protocol (Lee et al. 2012; Luo et al. 2017). The content of LasA protease released from supernatant of the samples treated with sub-MICs of streptomycin was measured by the azocasein assay according to previous studies (Luo et al. 2017). Briefly, the streptomycin-treated samples after 12 h of incubation at 35 °C were centrifuged and filtered. The filtered supernatant (150 μl) was added to the tube containing 250 μl azocasein (2%) (prepared in 50 mM Tris-HCl, pH 7.8) and the mixture was incubated at 4 °C for 4 h. After incubation, the reaction was terminated with 10% trichloroacetic acid, followed by incubating at 4 °C for 15 min. The reaction mixture was centrifuged (10,000 rpm for 10 min) and the obtained supernatant was neutralized with NaOH (1 M). The activity of LasA protease was determined by OD measurement at 440-nm wavelength using microtiter plate reader and the relative value was compared to control. For testing the protease activity on the agar plate, the agar plate was prepared from 2% Bacto agar and 10 g casein powder. The overnight grown P. aeruginosa cell culture was diluted at 1:100 dilutions in TSB media (pH 7.9) and grown along with sub-MIC of streptomycin. After 12 h of incubation at 35 °C under shaking condition, the cell culture was centrifuged and filtered using sterile 0.2-μm filter. The cell-free filtered culture supernatant was added in the holes which were made in casein-agar plate and incubated at 35 °C overnight. After incubation, the zone of clearance (which indicated the digestive activity of casein protein) around the hole was assumed as the positive protease activity.
Motility assays
The motility properties such as swimming, swarming, and twitching of P. aeruginosa were evaluated in the presence of streptomycin at sub-MIC levels. The methods used for the study of motility were adopted from the previous methods (Lee et al. 2012; Luo et al. 2017). The swarming assay was performed on agar plates prepared from LB broth containing 0.5% casamino acids, 0.5% glucose, and 0.4% Bacto agar according to the procedure developed by Luo et al. (2017). The swimming motility was also performed on agar plate which are prepared from 1% tryptone, 0.25% NaCl, and 0.3% Bacto agar according to the protocol described previously (Lee et al. 2012). To perform the motility assays, the P. aeruginosa cell culture grown overnight (5 μl) was positioned in the center of each agar plate. The twitching experiment was performed by placing the overnight grown cell culture in the center of the Petri plate using toothpick following the previous protocol by Luo et al. (2017). The placed cell culture was embedded by pouring LB medium containing 0.2% casamino acids, 30 mM glucose, and 1.5% Bacto agar. The agar plates used for swimming, swarming, and twitching assays were supplemented with sub-MICs (from 0.25 to 2.0 μg ml−1) of streptomycin. After incubation at 35 °C for 24 h, the diameters (cm) of cells traveling on the swarming, swimming, and twitching agar plates were measured.
Real-time polymerase chain reaction
The expression of genes associated with biofilm formation, QS signaling, virulence factor production, and motilities was accessed using real-time polymerase chain reaction (qRT-PCR). The primers used in this study were taken from the earlier studies reported elsewhere such as QS signaling genes (rhlR-rhlI and lasR-lasI systems) (Furiga et al. 2015), biofilm matrix-forming exopolysaccharide genes (algA, algU, and pslM) (Fu et al. 2017), virulence genes (phzC, phzE, pvdA, pvcC, and lasB) (Furiga et al. 2015; Lee et al. 2011, 2014), and motility gene (flgG) (Lee et al. 2011; Oura et al. 2015). In this study, the pyrroline-5-carboxylate reductase (proC) housekeeping gene (Lee et al. 2014) was selected as the control for normalization of expression data of all the genes mentioned above. RNA extraction was carried out by growing the P. aeruginosa cell culture overnight (1:100 dilution) in 25 ml TSB (pH 7.9) in the 250-ml Erlenmeyer flask in the presence and absence of sub-MIC (2 μg ml−1) streptomycin under shaking (250 rpm) at 35 °C for 12 h following the previous protocol with slight modifications (Lee et al. 2011). The grown cell culture was firstly chilled in dry ice for 30 s, followed by centrifugation (13,000 rpm; 5 min). From the cell pellets, total RNA was extracted using AccuZol™ reagent kit (Bioneer, South Korea). The concentration at 260 nm and purity of RNA at 260/280 nm were determined by NanoDrop spectrophotometer. From the extracted RNA, the first complementary DNA strand (cDNA) was synthesized by using AccuPower® GreenStar™ qRT-PCR premix kit (Bioneer, South Korea). The cDNA acted as a template and the primers mentioned above were used for the qRT-PCR by using DNA engine Chromo4 real-time Detector (Bio-Rad, Hercules, CA, USA). The PCR program and procedure were adopted from the method described earlier (Luo et al. 2017). After normalizing the gene expression level with the proC housekeeping gene, the relative expression level of each gene was measured following the methodology described earlier (Schmittgen and Livak 2008).
Statistical analysis
All the presented graphs were plotted by using GraphPad Prism 7.0 (GraphPad Software Inc., San Diego, CA) and the results were shown as means ± SD. Similarly, analysis of the data in this study was also carried out by performing one-way ANOVA.
Results
Determination of minimum and sub-minimum inhibitory concentration of streptomycin
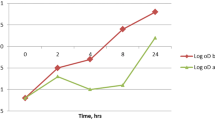
Prior to the study of biofilm inhibition and suppression of virulence factors by streptomycin, firstly we studied the growth of P. aeruginosa for establishing the MIC values of streptomycin. The MIC of streptomycin against P. aeruginosa was found as 8 μg ml−1 (Fig. 1a). Furthermore, to determine the sub-MIC of streptomycin, P. aeruginosa was incubated with streptomycin at different concentrations below the MIC. The bacterial growth at 4 μg ml−1 after 28 h of incubation was found to be slightly reduced as compared to the control or below 4 μg ml−1 (Fig. 1b). However, the lag phase at 4 μg ml−1 concentration was found to prolong up to 16 h. Although at 2 μg ml−1 concentration there was also 6 h of lag phase, after 28 h of incubation, the growth was found to be almost equal to 4 μg ml−1 of concentration. Above 4 μg ml−1 concentration of streptomycin, which is 8 μg ml−1, there was a complete growth inhibition. Based on the above results, the sub-MIC ranging from 0.0625 to 4 μg ml−1 was selected for the subsequent studies of biofilm inhibition and virulence factor production of P. aeruginosa.
MIC determination of streptomycin against P. aeruginosa PAO1. a Bactericidal effect of streptomycin against P. aeruginosa. The absorbance of negative control (only TSB) was found to 0.093 ± 0.003; hence, based on this value, the positive growth of the culture was considered when absorbance was found to > 0.2. b Growth curve of P. aeruginosa in the presence of sub-MIC of streptomycin. **P < 0.01 was accepted as statistically significant and ns indicated non-significance
Streptomycin inhibited biofilm formation and eradicated the mature biofilm of P. aeruginosa
With the attempts of investigating the P. aeruginosa biofilm formation in response to streptomycin activity, three sub-MIC levels of streptomycin (0.5, 1, and 2 μg ml−1) were selected to culture with the bacterial cells along with several different culture conditions, using three different culture media (TSB, MHB, and LB), three pH levels (5.9, 7.2, and 7.9), and three temperature levels (25, 30, and 35 °C). The biofilm inhibition results obtained from the aforementioned conditions are summarized in the Supplementary Fig. S1. In terms of media types, the culture conditions of MHB media along with all tested temperature and pH levels were unable to suppress the bacterial biofilm formation. Furthermore, it was found that the acidic pH (pH 5.9) at all temperatures from 25 to 35 °C has promoted biofilm formation. In contrast, when environmental pH became alkaline (pH 7.9) and at 35 °C of temperature, a reduction in bacterial biofilm formation was observed in both LB and TSB media. As compared to LB media, the anti-biofilm effect was slightly improved in TSB culture media. Particularly, in TSB media with pH 7.9 at 35 °C, treatment of streptomycin at sub-MIC (2 μg ml−1) and sub-MIC (4 μg ml−1) inhibited 78% and 76% of bacterial biofilm formation, respectively (Fig. 2). Considering all results from pH, temperature, and media types, TSB culture media, pH 7.9, and 35 °C of temperature were selected for further detailed study of biofilm inhibition and several other phenotypic properties.
The analysis of the biofilm architecture in the presence of streptomycin at sub-MIC (2 μg ml−1) was evaluated by SEM and fluorescence microscope (Fig. 3a, b). The SEM analysis was carried out to check the impact of streptomycin on the biofilm architecture. As shown in the SEM image, in the presence of sub-MIC of streptomycin, P. aeruginosa cells became wrinkled (Fig. 3a) as compared to the control, whereas the non-treated cells clearly showed a dense colonization and highly compact biofilm architecture. The fluorescence microscopy results also showed a reduction in fluorescence intensity, which reflected the inhibitory activity of streptomycin at the sub-MIC against the bacterial biofilm. The streptomycin-treated cells showed a scanty architecture while the non-treated control showed a dense colonization and highly compact architecture of the cells on the glass slide surface (Fig. 3b). The relative fluorescence intensity determination of the cells also showed a clear difference between the treated samples and non-treated sample (control) (Fig. 3c).
Visualization of biofilm structure by SEM and fluorescence microscopes in the presence of sub-MIC (2 μg ml−1) of streptomycin. a SEM image of the biofilm cells cultured on the nylon membrane. b Fluorescence image of the biofilm cells cultured on the glass pieces. c Relative fluorescence intensity of biofilm cells
In most cases, eradication of pre-existing bacterial biofilm is considered highly challenging as the structure has (1) attached to a surface, (2) produced a wide array of virulence factors and adhesins, and (3) formed a more complicated morphological structure that enhances the level of resistance (Das et al. 2014; Jamal et al. 2018; Sheraton et al. 2018). Previous studies have found several anti-biofilm agents which were highly active in inhibiting biofilm yet were unable to disperse the pre-existing mature one (Otto 2013; Zhao et al. 2017). However, results from the present study showed a significant disruption of 24-h mature biofilm by all tested concentrations of streptomycin (Fig. 4). The effectiveness of streptomycin application reached the highest level during the first 6-h period and gradually decreased at the end of 24 h of treatment.
For further confirmation the anti-biofilm activity of streptomycin, we have determined the MIC levels, followed by biofilm inhibition and mature biofilm eradication activities of tetracycline and tobramycin, using the same protocols as streptomycin. The culture conditions were also maintained similar to streptomycin, which were proposed as pH 7.9, 35 °C of temperature, and TSB media. As shown in Fig. 5a, the MIC levels of tetracycline and tobramycin were determined as 8 μg ml−1 and 0.5 μg ml−1, respectively. Although at sub-MIC levels tetracycline and tobramycin did not show the concentration-depended biofilm inhibition, both exhibited statistically significant inhibition of biofilm under the similar conditions as specified for the streptomycin (Fig. 5b, c). However, both of them showed a comparative significant concentration-dependent dispersal of preformed mature biofilm of P. aeruginosa (Fig. 5d).
a MIC determination of tobramycin and tetracycline against P. aeruginosa PAO1 in TSB and pH 7.9. b Inhibitory effects of sub-MICs of tetracycline on P. aeruginosa PAO1 biofilm formation and cell growth without shaking condition. c Inhibitory effects of tobramycin at sub-MICs on P. aeruginosa PAO1 biofilm formation and cell growth without shaking condition. d Eradication of 24-h mature P. aeruginosa PAO1 biofilm by 6-h treatments of tobramycin and tetracycline using crystal violet staining assay **P < 0.01 was accepted as statistically significant and ns indicated non-significance
Due to the variations of P. aeruginosa biofilm and virulence phenotypes, in addition to the standard laboratory strain PAO1, a clinical strain isolated from pleural fluid of a patient in hospital (i.e., GNUH-NCCP 6039) was also employed to evaluate the anti-biofilm effects of streptomycin, tobramycin, and tetracycline. A similar sequence of experiments was performed under the proposed culture conditions (pH 7.9, 35 °C, and TSB) for all three antibiotics, including MIC determination, biofilm inhibition assay, and eradication of preformed mature biofilm. The results are shown in Fig. S2 and S3. Firstly, the MIC values of streptomycin, tetracycline, and tobramycin were found as 16 μg ml−1, 32 μg ml−1, and 0.5 μg ml−1, respectively. Secondly, the anti-biofilm and eradication activities of the three antibiotics were evaluated using crystal violet staining. The biofilm inhibition by streptomycin, tetracycline, and tobramycin of the clinical strain GNUH-NCCP 6039 was significant but not in a concentration-dependence manner as compared to PAO1 (Fig. S2b, c, d). However, streptomycin, tetracycline, and tobramycin exhibited a concentration-dependent eradication towards 24-h mature biofilm of GNUH-NCCP 6039 when treated for 12 h (Fig. S3).
Streptomycin inhibits biofilm formation on the surfaces of urinary catheter
Urinary catheter is one of the commonly used medical devices, which may readily acquire the biofilm formation by pathogenic bacteria on either outer or inner surfaces when it is inserted into the patient (Al-Mathkhury et al. 2011). Reports have showed that P. aeruginosa commonly colonizes and establishes biofilms on the surface of urinary catheter, which have become one of its resistance properties against the anti-biofilm agents (Shigemura et al. 2006; Stickler 1996). For this reason, we have also studied the impact of sub-MICs of streptomycin on the biofilm of P. aeruginosa formed on the urinary catheter. The impact of streptomycin on biofilm was determined by viable cell counting and crystal violet staining of bacterial cells attached to the surface of urinary catheter. The results from crystal violet staining showed that the sub-MICs of streptomycin inhibited P. aeruginosa biofilm formation in a dose-dependent manner as shown in Fig. 6. Approximately 66% of biofilm biomass was inhibited at the sub-MIC concentration (2 μg ml−1) of streptomycin (Fig. 6a). This result has also been confirmed by determining the bacterial cell count of attached biofilm cells (Fig. 6b). The result of the bacterial cell count was also relatively concentration-dependent in response to streptomycin.
Streptomycin attenuates the virulence properties of P. aeruginosa
P. aeruginosa virulence properties which contribute to the bacterial pathogenicity were found by the hemolysis of RBCs and also production of several virulence factors (Allen et al. 2014). The impact of streptomycin on the hemolytic properties was studied at sub-MIC levels. Streptomycin at sub-MIC level (2 μg ml−1) has reduced the P. aeruginosa hemolytic properties by about 54%, as compared to the control (Fig. 7a).
Inhibitory effects of sub-MICs of streptomycin on hemolytic activity and production of virulence factors of P. aeruginosa PAO1. a Hemolysis assay, b pyocyanin production, c rhamnolipid production, and d pyoverdine production. The hemolysis and production of each virulence factor from streptomycin-treated sample are represented a relative value with respect to the control. All the experiments were performed in triplicates. **P < 0.01 was considered statistically significant
The biosynthesis of virulence factors such as pyocyanin, rhamnolipid, and siderophores such as pyoverdine was evaluated in the presence of streptomycin at sub-MICs. Under sub-MICs of streptomycin, approximately 77.8% of pyocyanin production was reduced in comparison with the control (Fig. 7b). Similarly, the production of rhamnolipid and pyoverdine was also reduced by about 65% and 88%, respectively, by streptomycin at sub-MIC (2 μg ml−1) (Fig. 7c, d).
Synthesis of hydrolytic protease enzyme such as LasA protease is also crucial for P. aeruginosa pathogenicity towards the host cells (Andrejko et al. 2013; Chanda et al. 2017). The present study examined the effect of sub-MICs streptomycin (0.125–4 μg ml−1) in attenuating the protease activity of the bacteria. Results from proteolytic assays using both azocasein protein digestion and casein agar plates showed that in comparison to the control, streptomycin treatment was able to reduce the protease activity (Fig. 8a, b). Thus, similar to the hemolytic and several virulence factors, P. aeruginosa protease can also be suppressed by applying treatment of sub-MICs streptomycin.
Inhibition of protease activity in P. aeruginosa PAO1 by sub-MICs of streptomycin. a Effect of sub-MICs of streptomycin on azocasein degrading protease enzymes secreted by P. aeruginosa. b Effect of sub-MICs of streptomycin on protease activity on the casein agar plate. **P < 0.01 was considered statistically significant
It is well studied that the motility towards a biotic or abiotic surface is linked to the bacterial virulence and is governed by different mechanisms (Harshey 2003; Josenhans and Suerbaum 2002; Shi and Sun 2002). In the present study, we have checked P. aeruginosa swarming, swimming, and twitching movements in the presence of the sub-MIC of streptomycin. The results indicated that streptomycin at sub-MIC (2 μg ml−1) inhibited these motions significantly, with 99% and 67% of bacterial swimming and swarming motions was reduced as compared to the control (Fig. 9a, b, c, d). Likewise, the bacterial twitching motility was also significantly inhibited as shown in Fig. 9e, f. Combining with the growth studies of P. aeruginosa at different concentrations (Fig. 1a, b), inhibition of motility at sub-MICs of streptomycin was not due to the defect in the bacterial cell growth.
Inhibition of motility properties of P. aeruginosa PAO1 by sub-MICs of streptomycin. a Swimming motility on agar plate. b Diameter value (cm) of swimming motility. c Swarming motility on agar plate. d Diameter value (cm) of swarming motility. e Twitching motility. f Diameter value (cm) of twitching motility. A representative image of each assay is presented and each experiment was carried out three times. **P < 0.01 was considered statistically significant
Effect of sub-MIC of streptomycin on biofilm-associated and QS-related virulence genes expression
In P. aeruginosa, the establishment of biofilm and production of other virulence factors which contribute to the bacterial pathogenesis are majorly regulated by LasI/LasR and RhlI/RhlR QS systems associated with PQS (Pseudomonas quinolone sensing signal) and IQS (integrating quorum sensing signal) (Martinez et al. 2019). The LasI/LasR system which uses 3-oxo-C12-hemoserine lactone (3O-C12-HSL) autoinducer and the RhlI/RhlR system which uses butanoyl homoserine lactone (C4-HSL) autoinducer determine the production and expression of pyocyanin, siderophores, rhamnolipid, and protease enzymes as well as swarming motility (Holm and Vikstrom 2014; Rutherford and Bassler 2012). With promising final effects from virulence phenotypic assays performed earlier, the expression of each specific virulence gene of P. aeruginosa under sub-MIC of streptomycin (2 μg ml−1) was then further studied using qRT-PCR. Results from qRT-PCR revealed that streptomycin at sub-MIC significantly suppressed the expression of genes encoding for virulence factors (pyocyanin—phzE and phzC; pyoverdine—pvdA and pvcC; elastase—lasB), exopolysaccharide (algA) and flagella (flgG), in comparison with the non-treated control (Fig. 10). A similar effect was observed in the group of QS-regulated genes (lasI, lasR, rhlI, and rhlR) in the presence of sub-MIC of streptomycin. In contrast, streptomycin at sub-MIC enhanced the expression of the remaining exopolysaccharide genes, which are algU and pslM. Combining the results from phenotypic and gene expression analysis of virulence properties, streptomycin at sub-MIC can be regarded as a promising anti-virulence agent against P. aeruginosa.
qRT-PCR analysis for the determination of relative expression levels of biofilm-forming, virulence, and motility genes in the presence of sub-MIC of streptomycin. The relative expression of genes represents transcriptional levels after treatment with sub-MIC streptomycin versus non-treated control. The experiment was carried out in triplicate
Discussion
Aminoglycoside is a common antibiotic family owing to the polycationic nature, which was previously hypothesized to limit them from penetrating through the bacterial cells without concentration loss (Li et al. 2015). In fact, these antibiotics have been reported to be unable to control the growth of several bacteria as their efficacy is encountered by multiple barriers, including bacterial cell wall permeability (Bansal-Mutalik and Nikaido 2014; Needham and Trent 2013; Sarathy et al. 2012), degrading enzymes (Ramirez and Tolmasky 2010), modified-binding target (Wilson 2014), and efflux pumps (Fernandez and Hancock 2012). However, for the complications of a bacterial community formed within a biofilm structure, along with varied results obtained in different bacteria, a general understanding about the resistant responses of P. aeruginosa towards aminoglycosides currently remained lacking (Garneau-Tsodikova and Labby 2016; Poole 2012). Although at present, fluoroquinolone is known as the most frequently used antibiotic against P. aeruginosa, the prevalence of resistance emergence had also been recorded in several studies (Guss et al. 2009; Wu et al. 2018; Yang et al. 2015), urging further exploitation for alternative approaches such as combination strategy and diversifying the available antibiotic options (Khan et al. 2019b, c, d; Kotra et al. 2000). For these reasons, streptomycin, which is a common member of the aminoglycoside antibiotic family, has been selected in the current study to evaluate the effects on P. aeruginosa growth, biofilm formation, and virulence properties. In addition, owing to the unique polycationic nature of streptomycin, the pH value, the culture media, and temperature were also taken into consideration.
The sub-MIC values were selected as the working concentrations for subsequent tests, as no inhibition in P. aeruginosa growth was recorded at these concentrations. Previous studies have reported that the antimicrobial activity of some antibiotics such as aminoglycosides was hindered by their electrostatic interaction with the negatively charged components present in the biofilm extracellular matrix (Rabin et al. 2015, Stewart 2002). Therefore, the present study proposed one possible approach to improve aminoglycoside activity is by changing the bacterial culture environment, which included temperature, pH, and media type. Three levels of pH, temperature, and three culture media types were used to generate various culture conditions for P. aeruginosa biofilm formation under streptomycin treatment at sub-MICs. Results have shown that the biofilm inhibition was correlated to the environmental pH, temperature, and media type. Overall, streptomycin anti-biofilm activity against P. aeruginosa was maximized under the condition of 35 °C of temperature, alkaline pH, and TSB media. One of the purposes of forming biofilm in bacteria, particularly P. aeruginosa, is to resist and survive through extreme conditions both inside the host system and outside the environment system. Firstly, pH was known to affect biofilm formation and EPS synthesis (Henry-Stanley et al. 2014). Furthermore, previous studies have shown that the acidic pH even promoted the biofilm establishment of P. aeruginosa and reduced the efficacy of anti-biofilm agents (Schlessinger 1988; Wilton et al. 2016). A similar observation was also obtained from the present study, in which acidic pH when combined with all tested temperature and media types did not support streptomycin anti-biofilm activity. In contrast, under alkaline pH, gentamycin and its combination with l-arginine have reduced P. aeruginosa and S. aureus biofilm formation (Lebeuax et al. 2014). Secondly, temperature was known to affect biofilm structure, thickness, virulence, and genetic transfer in biofilm (LaBauve and Wargo 2012; MacFadden et al. 2018). In fact, high temperature (37 °C) could accelerate the maturation of P. aeruginosa biofilm (Kannan and Gautam 2015). Finally, media type could act as a nutrient source as well as the environment for movements and biofilm thickness as well as multiple physiochemical properties (Vrany et al. 1997). For instance, the biofilm inhibition activity of several aminoglycoside, including streptomycin against Staphylococcus aureus, was dependent on the culture media (Henry-Stanley et al. 2014). Of all three media tested, which are LB, MHB, and TSB, streptomycin performed the highest inhibitory activity against P. aeruginosa biofilm in TSB media. Along with alkaline pH and temperature of 35 °C, this culture condition can be used to improve the aminoglycoside penetration into P. aeruginosa biofilm. Interestingly, the anti-biofilm activity of streptomycin was also comparatively noteworthy in LB media with alkaline pH and 35 °C of temperature. Results have revealed that sub-MIC levels of streptomycin exhibited inhibitory activity on these biofilms under alkaline pH and in TSB media. Eradication activity of streptomycin was also shown at sub-MICs, as well as above MIC levels. Several reports showed that P. aeruginosa developed different types of resistance mechanisms against the action of antibiotics such as (1) modification and mutation of the antibiotic binding site, (2) modification of antibiotic chemical structure, (3) modification of the bacterial membrane, and (4) efflux pump (Al-Wrafy et al. 2017; Garneau-Tsodikova and Labby 2016). The effectiveness of sub-MIC streptomycin on P. aeruginosa biofilm was also supported by SEM and fluorescence microscopic visualization. The results from microscopic visualization showed that the biofilm structure genuinely appeared wrinkled and less dense in the presence of streptomycin at sub-MIC. Compared to the previous studies in which streptomycin was recognized as a weak biofilm inhibitor to P. aeruginosa, the sub-MIC levels of the antibiotic turned out to be limitedly studied (Mu et al. 2016; Zhang et al. 2013). Overall, the findings from the present study suggested a specific concentration range, environmental pH, and temperature, which is sub-MIC levels, pH 7.9, 35 °C, and TSB as bacterial culture media to improve the anti-biofilm function of streptomycin.
In order to compare the anti-biofilm effects of streptomycin with different antibiotics, tobramycin and tetracycline which also have similar inhibitory effect on protein synthesis as streptomycin were employed. Under similar culture conditions as proposed for streptomycin, both antibiotics were more active in eradicating the pre-formed mature biofilm formed by P. aeruginosa PAO1 strain. The modifications in terms of pH, temperature, and media type of the culture environment have possibly caused a noticeable change in the anti-biofilm activities of all tested antibiotics. According to previous reports by Kumar and Ting (2016) and Hoffman et al. (2005), the sub-MICs of streptomycin and tobramycin promoted P. aeruginosa biofilm formation. However, in these studies, different culture conditions were used such as [TSB, pH 7.0, and 37 °C] as for streptomycin and [MHB, 37 °C] as for tobramycin. Furthermore, the condition used for PAO1 strain was also tested on a clinical isolate (GNUH-NCCP 6039). Results indicated that streptomycin, tetracycline, and tobramycin showed a significant biofilm inhibition and a concentration-dependent dispersion towards the preformed mature biofilm.
Furthermore, the application of streptomycin at sub-MIC for the biofilm inhibition on medical devices such as urinary catheter has also been examined using similar media and conditions. An effective biofilm inhibition by streptomycin was found in a concentration-dependent manner as evaluated by crystal violet staining. The viable cell count of the cells which formed biofilm on the surface of urinary catheter also showed significant reduction in the presence of streptomycin at sub-MIC.
Along with biofilm formation, secretion of virulence factors also primarily characterizes for the pathogenicity, infection, and survival of P. aeruginosa in adverse environmental conditions (Luo et al. 2017; Meirelles and Newman 2018; Orgad et al. 2011; Poppe et al. 2018). Phenotypic studies revealed that as compared to the control and other sub-MIC levels, the concentration of 2 μg ml−1 of streptomycin has reduced the production of all QS-related virulence factors and inhibited the hemolytic activity of P. aeruginosa. Therefore, this sub-MIC of streptomycin was selected for genetic analysis of the virulence phenotypes, along with (1) their QS regulation genes, (2) exopolysaccharide genes, (3) protease-encoded gene, and (4) flagella-encoded gene. Except algU and pslM, the expression of all tested genes was downregulated by sub-MIC of streptomycin, which reasonably explained for the reduction in these phenotypes as observed earlier. Similarly, in previous studies, the sub-MICs of ciprofloxacin and azithromycin had also reduced biofilm formation by attenuating these factors (Bala et al. 2011; Gupta et al. 2016; Imperi et al. 2014). In fact, recently, the sub-MIC level of common antibiotics such as quinolone and beta-lactam has gained more attention in the modern anti-biofilm therapies against a variety of biofilm-forming bacteria (El-Mowafy et al. 2017; Haddadin et al. 2010; Otto et al. 2013; Vidya et al. 2005; Viedma et al. 2018). At sub-MIC levels, these antibiotics most likely targeted the virulence factor production as well as the virulence-regulated system, which is termed “quorum sensing.” As aforementioned, the virulence factors play an equally important role as biofilm formation in characterizing the P. aeruginosa pathogenesis; thus, using antibiotics as anti-virulence agents can be considered a promising alternative strategy for combating the bacterial biofilm formation (Gupta et al. 2016).
The present study also investigated the effects of sub-MIC of streptomycin on inhibiting P. aeruginosa motility, including swimming, swarming, and twitching (Davies et al. 1998; Glessner et al. 1999; O'Toole and Kolter 1998). Results have shown that sub-MIC of streptomycin could impair the flagella-mediated swimming, swarming and type IV pili-regulated twitching motility of P. aeruginosa (Fig. 9). This result was supported by the suppression of flagella-encoded gene (flgG) expression under streptomycin at sub-MIC treatment using qRT-PCR (Fig. 10). Similar observations were reported by Saroj and Rather (2013), in which sub-MIC levels of streptomycin effectively inhibited the genes encoded for motility in Acinetobacter baumannii, which was also regulated by QS system. However, further research is required to identify the mechanism behind the anti-motility effect of streptomycin at sub-MIC levels.
In conclusion, this study aimed to revitalize the potentials of a common antibiotic at sub-MIC levels, where streptomycin showed the inhibitory effects against P. aeruginosa biofilm formation, eradicated the mature biofilm and attenuated various bacterial virulence properties. The alkaline pH and 35 °C of temperature as well as TSB culture media were proposed to play a crucial role in improving the anti-biofilm activity of streptomycin. Moreover, the ability of streptomycin at sub-MIC level to reduce the bacterial biofilm formation on the surface of urinary catheter proposed its potential application in preventing the infections associated with this medical device. Overall, from the obtained results, streptomycin can be considered as a promising candidate for the up-to-date anti-biofilm approaches which employ the sub-MIC levels of common antibiotics to target the bacterial virulence properties. Nevertheless, future research is in demand in order to understand the specific molecular interactions between streptomycin antibiotic and the virulence compounds, as well as the drug pharmacokinetics and optimal environmental conditions for the antibiotics to perform its inhibitory activity against P. aeruginosa biofilm.
References
Al-Mathkhury HJ, Ali AS, Ghafil JA (2011) Antagonistic effect of bacteriocin against urinary catheter associated Pseudomonas aeruginosa biofilm. N Am J Med Sci 3:367–370. https://doi.org/10.4297/najms.2011.3367
Al-Wrafy F, Brzozowska E, Gorska S, Gamian A (2017) Pathogenic factors of Pseudomonas aeruginosa - the role of biofilm in pathogenicity and as a target for phage therapy. Postepy Hig Med Dosw (Online) 71:78–91
Alipour M, Suntres ZE, Lafrenie RM, Omri A (2010) Attenuation of Pseudomonas aeruginosa virulence factors and biofilms by co-encapsulation of bismuth-ethanedithiol with tobramycin in liposomes. J Antimicrob Chemother 65:684–693. https://doi.org/10.1093/jac/dkq036
Allen RC, Popat R, Diggle SP, Brown SP (2014) Targeting virulence: can we make evolution-proof drugs? Nat Rev Microbiol 12:300–308. https://doi.org/10.1038/nrmicro3232
Andrejko M, Zdybicka-Barabas A, Janczarek M, Cytrynska M (2013) Three Pseudomonas aeruginosa strains with different protease profiles. Acta Biochim Pol 60:83–90
Antoniu S (2015) Novel inhaled combined antibiotic formulations in the treatment of Pseudomonas aeruginosa airways infections in cystic fibrosis. Expert Rev Anti-Infect Ther 13:897–905. https://doi.org/10.1586/14787210.2015.1041925
Bala A, Kumar R, Harjai K (2011) Inhibition of quorum sensing in Pseudomonas aeruginosa by azithromycin and its effectiveness in urinary tract infections. J Med Microbiol 60:300–306. https://doi.org/10.1099/jmm.0.025387-0
Bansal-Mutalik R, Nikaido H (2014) Mycobacterial outer membrane is a lipid bilayer and the inner membrane is unusually rich in diacyl phosphatidylinositol dimannosides. Proc Natl Acad Sci U S A 111:4958–4963. https://doi.org/10.1073/pnas.1403078111
Ben Haj Khalifa A, Moissenet D, Vu Thien H, Khedher M (2011) Virulence factors in Pseudomonas aeruginosa: mechanisms and modes of regulation. Ann Biol Clin (Paris) 69:393–403. https://doi.org/10.1684/abc.2011.0589
Billings N, Birjiniuk A, Samad TS, Doyle PS, Ribbeck K (2015) Material properties of biofilms-a review of methods for understanding permeability and mechanics. Rep Prog Phys 78:036601. https://doi.org/10.1088/0034-4885/78/3/036601
Borges A, Sousa P, Gaspar A, Vilar S, Borges F, Simoes M (2017) Furvina inhibits the 3-oxo-C12-HSL-based quorum sensing system of Pseudomonas aeruginosa and QS-dependent phenotypes. Biofouling 33:156–168. https://doi.org/10.1080/08927014.2017.1280732
Cegelski L, Marshall GR, Eldridge GR, Hultgren SJ (2008) The biology and future prospects of antivirulence therapies. Nat Rev Microbiol 6:17–27. https://doi.org/10.1038/nrmicro1818
Chanda W, Joseph TP, Padhiar AA, Guo X, Min L, Wang W, Lolokote S, Ning A, Cao J, Huang M, Zhong M (2017) Combined effect of linolenic acid and tobramycin on Pseudomonas aeruginosa biofilm formation and quorum sensing. Exp Ther Med 14:4328–4338. https://doi.org/10.3892/etm.2017.5110
Das T, Sehar S, Koop L, Wong YK, Ahmed S, Siddiqui KS, Manefield M (2016) Attenuation of Pseudomonas aeruginosa biofilm formation by Vitexin: a combinatorial study with azithromycin and gentamicin. Sci Rep 6:23347. https://doi.org/10.1038/srep23347
Das T, Sehar S, Koop L, Wong YK, Ahmed S, Siddiqui KS, Manefield M (2014) Influence of calcium in extracellular DNA mediated bacterial aggregation and biofilm formation. PLoS One 9:e91935. https://doi.org/10.1371/journal.pone.0091935
Davies DG, Parsek MR, Pearson JP, Iglewski BH, Costerton JW, Greenberg EP (1998) The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science 280:295–298
El-Mowafy SA, Abd El Galil KH, Habib EE, Shaaban MI (2017) Quorum sensing inhibitory activity of sub-inhibitory concentrations of beta-lactams. Afr Health Sci 17:199–207. https://doi.org/10.4314/ahs.v17i1.25
Essar DW, Eberly L, Hadero A, Crawford IP (1990) Identification and characterization of genes for a second anthranilate synthase in Pseudomonas aeruginosa: interchangeability of the two anthranilate synthases and evolutionary implications. J Bacteriol 172:884–900
Fernandez L, Hancock RE (2012) Adaptive and mutational resistance: role of porins and efflux pumps in drug resistance. Clin Microbiol Rev 25:661–681. https://doi.org/10.1128/CMR.00043-12
Fleitas Martinez O, Cardoso MH, Ribeiro SM, Franco OL (2019) Recent advances in anti-virulence therapeutic strategies with a focus on dismantling bacterial membrane microdomains, toxin neutralization, quorum-sensing interference and biofilm inhibition. Front Cell Infect Microbiol 9:74. https://doi.org/10.3389/fcimb.2019.00074
Flemming HC, Wingender J (2010) The biofilm matrix. Nat Rev Microbiol 8:623–633. https://doi.org/10.1038/nrmicro2415
Flemming HC, Wingender J, Szewzyk U, Steinberg P, Rice SA, Kjelleberg S (2016) Biofilms: an emergent form of bacterial life. Nat Rev Microbiol 14:563–575. https://doi.org/10.1038/nrmicro.2016.94
Fu B, Wu Q, Dang M, Bai D, Guo Q, Shen L, Duan K (2017) Inhibition of Pseudomonas aeruginosa biofilm formation by traditional chinese medicinal herb Herba patriniae. Biomed Res Int 2017:9584703. https://doi.org/10.1155/2017/9584703
Furiga A, Lajoie B, El Hage S, Baziard G, Roques C (2015) Impairment of Pseudomonas aeruginosa biofilm resistance to antibiotics by combining the drugs with a new quorum-sensing inhibitor. Antimicrob Agents Chemother 60:1676–1686. https://doi.org/10.1128/AAC.02533-15
García-Contreras R, Maeda T, Wood TK (2013) Resistance to quorum-quenching compounds. Appl Environ Microbiol 79:6840–6846. https://doi.org/10.1128/AEM.02378-13
Garneau-Tsodikova S, Labby KJ (2016) Mechanisms of resistance to aminoglycoside antibiotics: overview and perspectives. Medchemcomm 7:11–27. https://doi.org/10.1039/C5MD00344J
Glessner A, Smith RS, Iglewski BH, Robinson JB (1999) Roles of Pseudomonas aeruginosa las and rhl quorum-sensing systems in control of twitching motility. J Bacteriol 181:1623–1629
Gupta P, Chhibber S, Harjai K (2016) Subinhibitory concentration of ciprofloxacin targets quorum sensing system of Pseudomonas aeruginosa causing inhibition of biofilm formation & reduction of virulence. Indian J Med Res 143:643–651. https://doi.org/10.4103/0971-5916.187114
Guss J, Abuzeid WM, Doghramji L, Edelstein PH, Chiu AG (2009) Fluoroquinolone-resistant Pseudomonas aeruginosa in chronic rhinosinusitis. ORL J Otorhinolaryngol Relat Spec 71:263–267. https://doi.org/10.1159/000242428
Haddadin RN, Saleh S, Al-Adham IS, Buultjens TE, Collier PJ (2010) The effect of subminimal inhibitory concentrations of antibiotics on virulence factors expressed by Staphylococcus aureus biofilms. J Appl Microbiol 108:1281–1291. https://doi.org/10.1111/j.1365-2672.2009.04529.x
Hall CW, Mah TF (2017) Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiol Rev 41:276–301. https://doi.org/10.1093/femsre/fux010
Harshey RM (2003) Bacterial motility on a surface: many ways to a common goal. Annu Rev Microbiol 57:249–273. https://doi.org/10.1146/annurev.micro.57.030502.091014
Henry-Stanley MJ, Hess DJ, Wells CL (2014) Aminoglycoside inhibition of Staphylococcus aureus biofilm formation is nutrient dependent. J Med Microbiol 63:861–869. https://doi.org/10.1099/jmm.0.068130-0
Hoffman LR, D’Argenio DA, MacCoss MJ, Zhang Z, Jones RA, Miller SI (2005) Aminoglycoside antibiotics induce bacterial biofilm formation. Nature 436:1171–1175. https://doi.org/10.1038/nature03912
Holm A, Vikstrom E (2014) Quorum sensing communication between bacteria and human cells: signals, targets, and functions. Front Plant Sci 5:309. https://doi.org/10.3389/fpls.2014.00309
Imperi F, Leoni L, Visca P (2014) Antivirulence activity of azithromycin in Pseudomonas aeruginosa. Front Microbiol 5:178. https://doi.org/10.3389/fmicb.2014.00178
Jamal M, Ahmad W, Andleeb S, Jalil F, Imran M, Nawaz MA, Hussain T, Ali M, Rafiq M, Kamil MA (2018) Bacterial biofilm and associated infections. J Chin Med Assoc 81:7–11. https://doi.org/10.1016/j.jcma.2017.07.012
Josenhans C, Suerbaum S (2002) The role of motility as a virulence factor in bacteria. Int J Med Microbiol 291:605–614. https://doi.org/10.1078/1438-4221-00173
Kannan A, Gautam P (2015) A quantitative study on the formation of Pseudomonas aeruginosa biofilm. Springerplus 4:379. https://doi.org/10.1186/s40064-015-1029-0
Kannan S, Sathasivam G, Marudhamuthu M (2018) Decrease of growth, biofilm and secreted virulence in opportunistic nosocomial Pseudomonas aeruginosa ATCC 25619 by glycyrrhetinic acid. Microb Pathog 126:332–342. https://doi.org/10.1016/j.micpath.2018.11.026
Kart D, Kustimur AS, Sagiroglu M, Kalkanci A (2017) Evaluation of antimicrobial durability and anti-biofilm effects in urinary catheters against Enterococcus faecalis clinical isolates and reference strains. Balkan Med J 34:546–552. https://doi.org/10.4274/balkanmedj.2016.1853
Khan F, Javaid A, Kim YM (2019a) Functional diversity of quorum sensing receptors in pathogenic bacteria: interspecies, intraspecies and interkingdom level. Curr Drug Targets 20:655–667. https://doi.org/10.2174/1389450120666181123123333
Khan F, Manivasagan P, Lee JW, Pham DTN, Oh J, Kim YM (2019b) Fucoidan-stabilized gold nanoparticle-mediated biofilm inhibition, attenuation of virulence and motility properties in Pseudomonas aeruginosa PAO1. Mar Drugs 17. https://doi.org/10.3390/md17040208
Khan F, Manivasagan P, Pham DTN, Oh J, Kim SK, Kim YM (2019c) Antibiofilm and antivirulence properties of chitosan-polypyrrole nanocomposites to Pseudomonas aeruginosa. Microb Pathog 128:363–373. https://doi.org/10.1016/j.micpath.2019.01.033
Khan F, Oloketuyi SF, Kim YM (2019d) Diversity of bacteria and bacterial products as antibiofilm and antiquorum sensing drugs against pathogenic bacteria. Curr Drug Targets. https://doi.org/10.2174/1389450120666190423161249
Kotra LP, Haddad J, Mobashery S (2000) Aminoglycosides: perspectives on mechanisms of action and resistance and strategies to counter resistance. Antimicrob Agents Chemother 44:3249–3256. https://doi.org/10.1128/aac.44.12.3249-3256.2000
Kumar A, Ting Y-P (2016) Streptomycin favors biofilm formation by altering cell surface properties. Appl Microbiol Biotechnol 100:8843–8853. https://doi.org/10.1007/s00253-016-7793-0
LaBauve AE, Wargo MJ (2012) Growth and laboratory maintenance of Pseudomonas aeruginosa. Curr Protoc Microbiol Chapter 6:Unit 6E 1. https://doi.org/10.1002/9780471729259.mc06e01s25
Lebeaux D, Chauhan A, Létoffé S, Fischer F, de Reuse H, Beloin C, Ghigo JM (2014) pH-Mediated potentiation of aminoglycosides kills bacterial persisters and eradicates in vivo biofilms. J Infect Dis 210:1357–1366. https://doi.org/10.1093/infdis/jiu286
Lee JH, Cho MH, Lee J (2011) 3-Indolylacetonitrile decreases Escherichia coli O157:H7 biofilm formation and Pseudomonas aeruginosa virulence. Environ Microbiol 13:62–73. https://doi.org/10.1111/j.1462-2920.2010.02308.x
Lee JH, Kim YG, Cho MH, Kim JA, Lee J (2012) 7-Fluoroindole as an antivirulence compound against Pseudomonas aeruginosa. FEMS Microbiol Lett 329:36–44. https://doi.org/10.1111/j.1574-6968.2012.02500.x
Lee JH, Kim YG, Cho MH, Lee J (2014) ZnO nanoparticles inhibit Pseudomonas aeruginosa biofilm formation and virulence factor production. Microbiol Res 169:888–896. https://doi.org/10.1016/j.micres.2014.05.005
Li XH, Kim SK, Lee JH (2017) Anti-biofilm effects of anthranilate on a broad range of bacteria. Sci Rep 7:8604. https://doi.org/10.1038/s41598-017-06540-1
Li XZ, Plesiat P, Nikaido H (2015) The challenge of efflux-mediated antibiotic resistance in Gram-negative bacteria. Clin Microbiol Rev 28:337–418. https://doi.org/10.1128/CMR.00117-14
Luo J, Dong B, Wang K, Cai S, Liu T, Cheng X, Lei D, Chen Y, Li Y, Kong J, Chen Y (2017) Baicalin inhibits biofilm formation, attenuates the quorum sensing-controlled virulence and enhances Pseudomonas aeruginosa clearance in a mouse peritoneal implant infection model. PLoS One 12:e0176883. https://doi.org/10.1371/journal.pone.0176883
MacFadden DR, McGough SF, Fisman D, Santillana M, Brownstein JS (2018) Antibiotic resistance increases with local temperature. Nat Clim Chang 8:510–514. https://doi.org/10.1038/s41558-018-0161-6
Mah TF, O'Toole GA (2001) Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol 9:34–39
Martinez E, Cosnahan RK, Wu M, Gadila SK, Quick EB, Mobley JA, Campos-Gomez J (2019) Oxylipins mediate cell-to-cell communication in Pseudomonas aeruginosa. Commun Biol 2:66. https://doi.org/10.1038/s42003-019-0310-0
Meirelles LA, Newman DK (2018) Both toxic and beneficial effects of pyocyanin contribute to the lifecycle of Pseudomonas aeruginosa. Mol Microbiol 110:995–1010. https://doi.org/10.1111/mmi.14132
Miyoshi-Akiyama T, Tada T, Ohmagari N, Viet Hung N, Tharavichitkul P, Pokhrel BM, Gniadkowski M, Shimojima M, Kirikae T (2017) Emergence and spread of epidemic multidrug-resistant Pseudomonas aeruginosa. Genome Biol Evol 9:3238–3245. https://doi.org/10.1093/gbe/evx243
Mu H, Liu Q, Niu H, Sun Y, Duan J (2016) Gold nanoparticles make chitosan–streptomycin conjugates effective towards Gram-negative bacterial biofilm. RSC Adv 6:8714–8721. https://doi.org/10.1039/C5RA22803D
Mulcahy H, Charron-Mazenod L, Lewenza S (2008) Extracellular DNA chelates cations and induces antibiotic resistance in Pseudomonas aeruginosa biofilms. PLoS Pathog 4:e1000213. https://doi.org/10.1371/journal.ppat.1000213
Needham BD, Trent MS (2013) Fortifying the barrier: the impact of lipid A remodelling on bacterial pathogenesis. Nat Rev Microbiol 11:467–481. https://doi.org/10.1038/nrmicro3047
O'Toole GA, Kolter R (1998) Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol Microbiol 30:295–304
Olejnickova K, Hola V, Ruzicka F (2014) Catheter-related infections caused by Pseudomonas aeruginosa: virulence factors involved and their relationships. Pathog Dis 72:87–94. https://doi.org/10.1111/2049-632X.12188
Orgad O, Oren Y, Walker SL, Herzberg M (2011) The role of alginate in Pseudomonas aeruginosa EPS adherence, viscoelastic properties and cell attachment. Biofouling 27:787–798. https://doi.org/10.1080/08927014.2011.603145
Otto M (2013) Staphylococcal infections: mechanisms of biofilm maturation and detachment as critical determinants of pathogenicity. Annu Rev Med 64:175–188. https://doi.org/10.1146/annurev-med-042711-140023
Otto MP, Martin E, Badiou C, Lebrun S, Bes M, Vandenesch F, Etienne J, Lina G, Dumitrescu O (2013) Effects of subinhibitory concentrations of antibiotics on virulence factor expression by community-acquired methicillin-resistant Staphylococcus aureus. J Antimicrob Chemother 68:1524–1532. https://doi.org/10.1093/jac/dkt073
Oura H, Tashiro Y, Toyofuku M, Ueda K, Kiyokawa T, Ito S, Takahashi Y, Lee S, Nojiri H, Nakajima-Kambe T, Uchiyama H, Futamata H, Nomura N (2015) Inhibition of Pseudomonas aeruginosa swarming motility by 1-naphthol and other bicyclic compounds bearing hydroxyl groups. Appl Environ Microbiol 81:2808–2818. https://doi.org/10.1128/AEM.04220-14
Parai D, Banerjee M, Dey P, Chakraborty A, Islam E, Mukherjee SK (2018) Effect of reserpine on Pseudomonas aeruginosa quorum sensing mediated virulence factors and biofilm formation. Biofouling 34:320–334. https://doi.org/10.1080/08927014.2018.1437910
Parrino B, Diana P, Cirrincione G, Cascioferro S (2018) Bacterial biofilm inhibition in the development of effective anti-virulence strategy. Open Med Chem J 12:84–87. https://doi.org/10.2174/1874104501812010084
Pattnaik S, Ahmed T, Ranganathan SK, Ampasala DR, Sarma VV, Busi S (2018) Aspergillus ochraceopetaliformis SSP13 modulates quorum sensing regulated virulence and biofilm formation in Pseudomonas aeruginosa PAO1. Biofouling 34:410–425. https://doi.org/10.1080/08927014.2018.1460748
Penesyan A, Gillings M, Paulsen IT (2015) Antibiotic discovery: combatting bacterial resistance in cells and in biofilm communities. Molecules 20:5286–5298. https://doi.org/10.3390/molecules20045286
Poole K (2012) Bacterial stress responses as determinants of antimicrobial resistance. J Antimicrob Chemother 67:2069–2089. https://doi.org/10.1093/jac/dks196
Poppe J, Reichelt J, Blankenfeldt W (2018) Pseudomonas aeruginosa pyoverdine maturation enzyme PvdP has a noncanonical domain architecture and affords insight into a new subclass of tyrosinases. J Biol Chem 293:14926–14936. https://doi.org/10.1074/jbc.RA118.002560
Qiu MN, Wang F, Chen SY, Wang PC, Fu YH, Liu YY, Wang X, Wang FB, Wang C, Yang HW, Wu Y, Zhu SY, Zhou HB, Chen WM, Lin J, Zheng JX, Sun PH (2019) Novel 2, 8-bit derivatives of quinolines attenuate Pseudomonas aeruginosa virulence and biofilm formation. Bioorg Med Chem Lett 29:749–754. https://doi.org/10.1016/j.bmcl.2018.12.068
Rabin N, Zheng Y, Opoku-Temeng C, Du Y, Bonsu E, Sintim HO (2015) Biofilm formation mechanisms and targets for developing antibiofilm agents. Future Med Chem 7:493–512. https://doi.org/10.4155/fmc.15.6
Ramirez-Estrada S, Borgatta B, Rello J (2016) Pseudomonas aeruginosa ventilator-associated pneumonia management. Infect Drug Resist 9:7–18. https://doi.org/10.2147/IDR.S50669
Ramirez MS, Tolmasky ME (2010) Aminoglycoside modifying enzymes. Drug Resist Updat 13:151–171. https://doi.org/10.1016/j.drup.2010.08.003
Rutherford ST, Bassler BL (2012) Bacterial quorum sensing: its role in virulence and possibilities for its control. Cold Spring Harb Perspect Med 2. https://doi.org/10.1101/cshperspect.a012427
Sarathy JP, Dartois V, Lee EJ (2012) The role of transport mechanisms in Mycobacterium tuberculosis drug resistance and tolerance. Pharmaceuticals (Basel) 5:1210–1235. https://doi.org/10.3390/ph5111210
Saroj SD, Rather PN (2013) Streptomycin inhibits quorum sensing in Acinetobacter baumannii. Antimicrob Agents Chemother 57:1926–1929. https://doi.org/10.1128/AAC.02161-12
Schlessinger D (1988) Failure of aminoglycoside antibiotics to kill anaerobic, low-pH, and resistant cultures. Clin Microbiol Rev 1:54–59
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3:1101–1108
Shaikh S, Fatima J, Shakil S, Rizvi SM, Kamal MA (2015) Antibiotic resistance and extended spectrum beta-lactamases: types, epidemiology and treatment. Saudi J Biol Sci 22:90–101. https://doi.org/10.1016/j.sjbs.2014.08.002
Sheraton MV, Yam JKH, Tan CH, Oh HS, Mancini E, Yang L, Rice SA, Sloot PMA (2018) Mesoscopic energy minimization drives Pseudomonas aeruginosa biofilm morphologies and consequent stratification of antibiotic activity based on cell metabolism. Antimicrob Agents Chemother 62. https://doi.org/10.1128/AAC.02544-17
Shi W, Sun H (2002) Type IV pilus-dependent motility and its possible role in bacterial pathogenesis. Infect Immun 70:1–4
Shigemura K, Arakawa S, Sakai Y, Kinoshita S, Tanaka K, Fujisawa M (2006) Complicated urinary tract infection caused by Pseudomonas aeruginosa in a single institution (1999-2003). Int J Urol 13:538–542. https://doi.org/10.1111/j.1442-2042.2006.01359.x
Singh R, Sahore S, Kaur P, Rani A, Ray P (2016) Penetration barrier contributes to bacterial biofilm-associated resistance against only select antibiotics, and exhibits genus-, strain- and antibiotic-specific differences. Pathog Dis 74. https://doi.org/10.1093/femspd/ftw056
Stewart PS (2002) Mechanisms of antibiotic resistance in bacterial biofilms. Int J Med Microbiol 292:107–113. https://doi.org/10.1078/1438-4221-00196
Stickler DJ (1996) Bacterial biofilms and the encrustation of urethral catheters. Biofouling 9:293–305. https://doi.org/10.1080/08927019609378311
Stintzi A, Evans K, Meyer JM, Poole K (1998) Quorum-sensing and siderophore biosynthesis in Pseudomonas aeruginosa: lasR/lasI mutants exhibit reduced pyoverdine biosynthesis. FEMS Microbiol Lett 166:341–345. https://doi.org/10.1111/j.1574-6968.1998.tb13910.x
Talwalkar JS, Murray TS (2016) The approach to Pseudomonas aeruginosa in cystic fibrosis. Clin Chest Med 37:69–81. https://doi.org/10.1016/j.ccm.2015.10.004
Vidya KC, Mallya PS, Rao PS (2005) Inhibition of bacterial adhesion by subinhibitory concentrations of antibiotics. Indian J Med Microbiol 23:102–105
Viedma E, Pérez-Montarelo D, Villa J, Muñoz-Gallego I, Larrosa N, Fernández-Hidalgo N, Gavaldà J, Almirante B, Chaves F (2018) Sub-inhibitory concentrations of oxacillin modify the expression of agr locus in Staphylococcus aureus clinical strains belonging to different clonal complexes. BMC Infect Dis 18:177. https://doi.org/10.1186/s12879-018-3088-7
Vrany JD, Stewart PS, Suci PA (1997) Comparison of recalcitrance to ciprofloxacin and levofloxacin exhibited by Pseudomonas aeruginosa bofilms displaying rapid-transport characteristics. Antimicrob Agents Chemother 41:1352–1358
Wilhelm S, Gdynia A, Tielen P, Rosenau F, Jaeger KE (2007) The autotransporter esterase EstA of Pseudomonas aeruginosa is required for rhamnolipid production, cell motility, and biofilm formation. J Bacteriol 189:6695–6703. https://doi.org/10.1128/JB.00023-07
Wilson DN (2014) Ribosome-targeting antibiotics and mechanisms of bacterial resistance. Nat Rev Microbiol 12:35–48. https://doi.org/10.1038/nrmicro3155
Wilton M, Charron-Mazenod L, Moore R, Lewenza S (2016) Extracellular DNA acidifies biofilms and induces aminoglycoside resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother 60:544–553. https://doi.org/10.1128/AAC.01650-15
Wu PF, Lin YT, Wang FD, Yang TC, Fung CP (2018) Is fluoroquinolone monotherapy a useful alternative treatment for Pseudomonas aeruginosa bacteraemia? Infection 46:365–373. https://doi.org/10.1007/s15010-018-1131-7
Yang X, Xing B, Liang C, Ye Z, Zhang Y (2015) Prevalence and fluoroquinolone resistance of Pseudomonas aeruginosa in a hospital of South China. Int J Clin Exp Med 8:1386–1390
Zhang A, Mu H, Zhang W, Cui G, Zhu J, Duan J (2013) Chitosan coupling makes microbial biofilms susceptible to antibiotics. Sci Rep 3:3364. https://doi.org/10.1038/srep03364
Zhao X, Zhao F, Wang J, Zhong N (2017) Biofilm formation and control strategies of foodborne pathogens: food safety perspectives. RSC Adv 7:36670–36683. https://doi.org/10.1039/C7RA02497E
Funding
This work was supported by the Marine Biotechnology Program (Grant number 20150220) funded by Ministry of Oceans and Fisheries, Republic of Korea. Also, this work was financially supported by the National Institute of Fisheries Science (Grant number R2019053), Republic of Korea.
Author information
Authors and Affiliations
Contributions
FK, JWL, JHL, and DTNP performed the experiment. FK, HWK, YKK, and YK designed the experiment and analyzed the data. All authors were involved in the writing and correction of the manuscript.
Corresponding author
Ethics declarations
This article does not contain any studies with human participants or animals.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regards to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 585 kb)
Rights and permissions
About this article
Cite this article
Khan, F., Lee, JW., Pham, D.T.N. et al. Streptomycin mediated biofilm inhibition and suppression of virulence properties in Pseudomonas aeruginosa PAO1. Appl Microbiol Biotechnol 104, 799–816 (2020). https://doi.org/10.1007/s00253-019-10190-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-10190-w