Abstract
Using lignocellulosic materials as substrates, ruminal microbiota were co-inoculated with anaerobic sludge at different loading rates (LR) to study the microbial community in the semi-continuous mode. The results indicated that the highest CH4 yield reached 0.22 L/g volatile solid at LR of 4 g/L/day, which obtained 56–58% of the theoretical value. In the steady stage with LR of 2–4 g/L/day and slurry recirculation, copies of total archaea increased. Especially the Methanobacteriales increased significantly (p < 0.05) to 3.30 × 108 copies/mL. The microbial communities were examined by MiSeq 16S rRNA sequencing. Enriched hydrolytic bacteria mainly belonged to Clostridiales, including Ruminococcus, Ruminiclostridium, and Ruminofilibacter settled in the rumen. High-active cellulase and xylanase were excreted in the co-inoculated system. Acid-producing bacteria by fermentation were affiliated with Lachnospiraceae and Bacteroidales. The acidogen members were mainly Spirochaetaceae and Clostridiales. Syntrophic oxidation bacteria mainly consisted of Synergistetes, propionate oxidizers (Syntrophobacter and Pelotomaculum), and butyrate oxidizers (Syntrophus and Syntrophomonas). There had no volatile fatty acid (VFA) accumulation and the pH values varied between 6.94 and 7.35. At LR of 6 g/L/day and a recirculation ratio of 1:1, the hardly degradable components and total VFA concentrations obviously increased. The total archaea and Methanobacteriales then deceased significantly to 8.56 × 105 copies/mL and 4.14 × 103 copies/mL respectively (p < 0.05), which resulted in the inhibition of methanogenic activities. Subsequently, microbial diversity dropped, and the hydrolytic bacteria and syntrophic oxidizers obviously decreased. In contrast, the abundances of Bacteroidales increased significantly (p < 0.05). Acetate, propionate, and butyrate concentrations reached 2.02, 6.54, and 0.53 g/L, respectively, which indicated “acidification” in the anaerobic reactor. Our study illustrated that co-inoculated anaerobic sludge enriched the ruminal function consortia and hydrogenotrophic methanogens played an important role in anaerobic digestion of lignocelluloses.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Ruminal microbiota are characterized as highly efficient players for converting lignocellulosic materials to volatile fatty acids (VFAs). Then, these VFA products are absorbed by rumen epithelial cells since that microbial community only contains a small quantity of hydrogenotrophic methanogens (Dai et al. 2015; Janssen and Kirs 2008; Morrison et al. 2009). Previous studies have indicated that rumen inoculated anaerobic digestion (RUDAD) enhances the hydrolytic efficiencies of corn stover (Hu and Yu 2005), cattail (Hu et al. 2007), wheat straw (Lazuka et al. 2015), and rice straw (Deng et al. 2017b; Zhang et al. 2016). The VFAs produced by RUDAD need to be further separated from the solid residues and transferred into the methanogenic fraction (Hu et al. 2007; Hu and Yu 2005). Thus, it would reduce the degradation efficiency of lignocelluloses. Addressing the problem of whether rumen microbiota are co-inoculated with methanogens to achieve lignocellulose hydrolysis and methanogenesis in the same reactor remains to be extensively elucidated. Except for previous investigations on enrichment of ruminal microbiota in the batch experiments (Deng et al. 2017b; Ozbayram et al. 2017), studies on the co-inoculum involved in the polysaccharide-degrading consortia are rare in a semi-continuous reactor.
Hydrolytic specialists in the rumen are mainly affiliated with Ruminococcus and Fibrobacter (Dai et al. 2015; Morrison et al. 2009; Nyonyo et al. 2014). However, these species have been less frequently reported to settle in batch tests (Azman et al. 2015; Deng et al. 2017b; Sun et al. 2016). In a long-term operation reactor with the co-inoculum, whether these functional bacteria can perform the hydrolytic activities needs to further be illustrated for more details. Subsequently, intermediates have been used to produce VFAs through acidogenesis in liquid fractions. The consumption of VFAs through syntrophic acetogenesis is indispensable since they are detrimental to anaerobic digestion. Microbial populations responsible for syntrophic cooperation need to be revealed. The overloading of lignocelluloses results in VFA accumulation and subsequent CH4-producing inhibition, which has been closely related with methanogens (Azman et al. 2015; Karthikeyan and Visvanathan 2013). Thus, detailed knowledge on the functional microbes and their interactions could be essential for uncovering the lignocellulosic digestion efficiencies in an anaerobic reactor. However, only a few investigations have studied the dynamics of the methanogenic community composition provoked by the organic overload rate (Lerm et al. 2012).
Therefore, this study aimed at evaluating digestion performance seeded with ruminal microbiota and anaerobic sludge in a semi-continuous reactor. The functional consortia, enriched by the application of different lignocelluloses loading rates and slurry recirculation, were investigated with MiSeq 16S rRNA sequencing. The copies of methanogenic archaea were further examined using absolute quantitative PCR (absolute Q-PCR). Thus, the mechanisms of acidification under lignocellulose overloading conditions were analyzed based on the changes of microbial communities.
Materials and methods
Inoculum and substrate
The rumen digesta were collected from a slaughterhouse (Wuxi, China) and immediately transferred to the laboratory. The rumen fluid was then recovered by straining the whole digesta with two layers of cheesecloth, containing 2.28% total solid (TS) and 1.32% volatile solid (VS). Anaerobic sludge with 13.90% TS and 6.90% VS was obtained from an anaerobic reactor used to treat kitchen wastes at 39 °C (Jiangsu Clean Environmental Technology Co., Ltd., Suzhou, China). The rumen fluid was co-inoculated with anaerobic sludge into the reactor based on the equal VS content. Lignocellulosic feedstocks consisted of corn stalks, guinea grass, and rice straw. All the raw materials were collected and dried at 50 °C in an air-circulating oven. The dried feedstocks were ground with a pulverizer (Zhongcheng Pharmaceutical Machinery Co., Ltd., Changsha, China) to a size of 40-mesh. After mixing with an equal VS content, the substrates were then stored in a vacuum bag to prevent possible degradation. The average TS and VS were 95.30 and 88.59%, respectively.
Semi-continuous anaerobic digestion
The co-inoculum was added into a semi-continuous reactor with a 25% volume fraction. The CH4 volume was directly measured with water column after CO2 absorption with 4 mol/L NaOH saturated solution. The reactor was set at 39 ± 1 °C with a 2.5-L working volume. Just before the reactor started, a buffer solution including K2HPO4·3H2O (1.34 g/L), KH2PO4 (0.51 g/L), and NaHCO3 (5.00 g/L) was added to maintain the initial pH value between 6.80 and 7.10.
To keep the solid content ranging within a small scale, discharging volume of the digestate increased with the improvement of loading rate (LR). The digestate was extruded and filtered through two layers of 100-mesh nylon gauze prior to feeding substrates at intervals of 2 days. The squeezed residues were used to measure the chemical compositions. The liquid fractions were returned to the system after supplementing dH2O according to the corresponding slurry recirculation ratios. As shown in Table 1, the whole digestion process was set to six operation phases. They were divided into the start-up stage, the steady stage, and the overload stage based on the discrepancy of the digestion performance.
Physicochemical analysis
The total organic carbon (TOC) and Kjeldahl nitrogen of the solid samples were measured with a TOC analyzer (SSM-5000, Shimadzu Co., Japan) and KDN-08 azotometer (Hangzhou Tuopu Instrument, Hangzhou, China) respectively and then used to calculate the C/N ratio. The cellulose, hemicellulose, and lignin content were analyzed according to a previously published principle with a slight modification of the residues being dried at 65 °C (Van Soest et al. 1991).
The slurry was centrifuged at 5000 r/min for 15 min, and then the supernatants were recovered for the triplicate assays after 10-fold dilution with dH2O. The pH values were measured with a DELTA 320 pH meter (Mettler Toledo, Shanghai, China). NH4+ concentrations were analyzed with the Nessler reagent method (APHA 2005). Dissolved TOC in the liquid fractions was detected using a TOC analyzer (TOC-VCPH, Shimadzu Co., Japan). VFA concentrations were detected by a GC-2010 Plus (Shimadzu Co., Japan) according to the previous method (Deng et al. 2017a). Xylanase and cellulase activities were measured using the dinitrosalicylic acid (DNS) method and expressed as units per milliliter following a previous study (Deng et al. 2017b). One unit of enzymatic activity (U) was defined as the amount of enzyme by which 1 μg of reducing sugar was produced per minute.
MiSeq 16S rRNA gene sequencing
The microbiome was harvested in the start-up stage (the 1st, 12th, 24th day, denoted as 1-1, 1-2, 1-3), the steady stage (the 36th, 60th, 84th, 84th day, denoted as 2, 3, 4, 5), and the overload stage (the 132nd, 144th, 156th day, denoted as 6-1, 6-2, 6-3). All 10 samples were centrifuged at 5000 r/min for 10 min and the corresponding pellets were used to extract DNA using the Power soil® DNA isolation kit (MoBio Laboratories Inc., Carlsbad, CA, USA) according to the manufacturer’s instructions. The amplified libraries were constructed with the V3-V4 regions of the 16S rRNA genes and then analyzed using the MiSeq Illumina platform (San Diego, CA, USA) by the paired-end protocol (Nelson et al. 2014). Subsequently, the raw sequences were pre-processed using the QIIME pipeline v1.8.0 (http://qiime.org/). After trimming the barcodes, the high-quality tags were classified to operational taxonomic units (OTUs) at a similarity of 97%, and subsequent statistical analysis was performed based on the OTUs. As shown in supplementary Figure S1, each sequence at the genus level was selected to a generated phylogenetic tree using MEGA 7.0 software (Kumar et al. 2016). All original sequences were deposited in the Sequence Read Archive of NCBI under the number: SRP119185.
Absolute Q-PCR on methanogens
Three samples in each stage were selected for absolute Q-PCR analysis. The primer sets of total archaea, Methanobacteriales, and Methanosarcinales are used according to Steinmetz et al. (2016). The PCR program was as follows: 95 °C initial denaturation for 3 min; followed by 40 cycles of 30 s denaturing at 95 °C, 30 s annealing at 60 °C, and 30 s extending at 72 °C; a final extension step of 72 °C for 30 s. The PCR products were analyzed electrophoretically in a 1% agarose gel and then gel-purified with an AxyPrep DNA gel-extraction kit (Axygen Corp., CA, USA). The target fragments were directly cloned into pMD19-T vector and then transformed into competent E. coli cells. The plasmid DNA were extracted and identified by sequencing results (Genewiz Inc., Suzhou, China).
Recombinant plasmid DNA concentrations were determined by an ND-1000 spectrophotometer (Thermo-Fisher, DE, USA), and copies of methanogen genes were then calculated according to Eq. 1. where MW represented genes sizes × 660 Da. DNA standards for quantification were 10-fold serially diluted with 109 to 103 copies/μL and run in triplicate alongside each Q-PCR assay.
The Q-PCR assay was set in SYBR Premix Ex Taq™ II (Takara Bio Inc., Dalian, China) with primer of a 0.20-μM final concentration and a 2.00-μL DNA template. The PCR reactions were performed in a One-Step Real-time PCR thermocycler (Applied Biosystems Inc. Carlsbad, CA, USA) following a previous study (Deng et al. 2017a). The copies of corresponding methanogens were calculated according to the Ct value and then expressed as copies per milliliter.
Statistical analysis
SPSS 19.0 (IBM Statistics, Chicago, IL, USA) was used to evaluate the significant difference of three stages at a 0.05 p value. Alpha diversities including (Chao1 and Shannon index) were calculated based on OTU data as described previously (Nelson et al. 2014). The general discrepancy in the microbial community structure was determined by principal component analysis (PCA) using Canoco for Windows 4.5 software (Microcomputer Power, Ithaca, NY, USA).
Results
Reactor performances
CH4 production
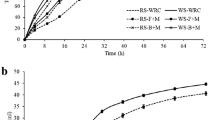
The CH4 yields of the six operation phases are shown in Fig. 1a. In the start-up stage with 1.5 g/L/day LR, the CH4 yields were on a gradual increment. However, the average yield only reached 0.14 L/g VS of lignocellulosic materials, which was attributed to the longer retention times of the methanogens in the artificial reactor (Demirel and Scherer 2008). In the steady stage, there were relatively stable CH4 yields, ranging from 0.16 L/g VS to 0.27 L/g VS. At phase 4 with 4 g/L/day LR, the average CH4 yield dropped to 0.16 L/g VS. However, the average value reached a maximum of 0.22 L/g VS when the slurry recirculation ratio was set to 1:1 from 1.5:1 at phase 5. In the overload stage, the average CH4 yield dropped sharply to 0.11 L/g VS.
The changes of TOC and VFA concentrations
As shown in Fig. 1b, the concentrations of TOC in the liquid digestate increased to 15.05 g/L from the original 2.24 g/L at the end of the experiment, especially reaching 13.67 g/L of the average concentration at the overload stage (shown in Table 2). The accumulated VFAs, mostly including acetate, propionate, and butyrate, were influenced both by the LR and recirculation ratios in this study. At phase 4, acetate began to accumulate with a concentration of 1.04 g/L. When the recirculation ratios reduced at phase 5, the acetate concentration decreased to 0.90 g/L from the initial 2.49 g/L, which was utilized through acetotrophic methanogenesis. The propionate concentration reached an average of 2.70 g/L at phase 1. Subsequently, propionate was consumed through syntrophic oxidation in the whole steady stage.
In the overload state with an LR of 6 g/L/day, the equilibrium between acidogenesis and methanogenesis was broken. Individual VFA also began to accumulate. As shown in Table 2, the average concentrations of acetate, propionate, and butyrate reached 2.02, 6.54, and 0.53 g/L, respectively. This indicated that “acidification” occurred in the anaerobic reactor.
The changes of pH values and NH4 + concentrations
As shown in Fig. 1c, the accumulation of VFAs lead to a decline in the pH values. The initial pH value was 7.21, and the values varied between 6.94 and 7.35 in the steady stage. The average pH value of the final stage was 6.99, which was still within the appropriate methanogenesis range of 6.5 to 8.5 (Azman et al. 2015). It was also shown that the pH values could not reflect the VFA accumulation timely in the anaerobic reactor, and NH4+ could also influence the pH values. The shifts of NH4+ concentrations had little effect in the first two stages, ranging from 0.21 to 0.36 g/L on average. In the overload stage, the values changed between 0.50 and 0.72 g/L with an average value of 0.63 g/L.
Characterizations of biodegradability and hydrolytic activities
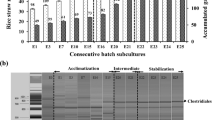
Because CH4 production originated from the lignocellulose biodegradability, it would be inevitable to analyze the composition changes of the solid residues in every phase. As shown in Fig. 2, the proportions of cellulose and hemicellulose of the six phases decreased, whereas those of lignin increased compared with the raw materials. In the steady stage, the proportions of hemicellulose and cellulose were 20.38–4.85 and 26.78–29.74%, respectively. The values of the two compositions increased to 25.18 and 31.88% in the overload stage. Correspondingly, cellulase and xylanase activities were used to assess the hydrolytic potential of the anaerobic co-inoculum (Deng et al. 2017b). The values were 36.44–50.06 and 4.13–6.45 U/mL in the steady stage, which was higher than those from the samples in the overload stage (shown in Table 2).
The diversities of bacterial communities
A total of 530,243 chimera-eliminated tags were obtained by sequence analysis with an average length of 360–480 bp. Alpha diversity analysis showed that the chao1 index in the three phases changed slightly. However, the Shannon index decreased obviously, being 5.25, 4.36, and 2.59 respectively (shown in Supplementary Table S1).
As shown in Fig. 3, PCA was evaluated based on the abundances of 505 common OTUs. In the PCo 1 coordinate, a high LR led to the failure of digestion and obvious changes of the microbial communities (6-1, 6-2, and 6-3) compared with those in the start-up stage, accounting for a 72% discrepancy. In the PCo 2 coordinate, the microbial communities of phases 3 and 4 were distinguished by 17.5% compared with those in the initial stage. However, the diversity from phase 2 was similar to those in the start-up stage, whereas the sample of phase 5 was close to those in the overload stage.
The OTUs were annotated by blasting the Silva 16S rRNA database, showing that the phyla of Bacteroidetes, Firmicutes, Synergistetes, Chloroflexi, Cloacimonetes (WWE1), Actinobacteria, Fibrobacteres, Spirochaetae, and Proteobacteria were the predominant microbial populations. The relative abundances of the phyla are shown in Fig. 4, and the total coverage was greater than 97.16% in each sample. In the steady stage, the relative abundances of Synergistetes, Chloroflexi, and Cloacimonetes increased. The Proteobacteria numbers changed slightly, while Actinobacteria, Fibrobacteres, and Spirochaetae declined in abundance. In the overload stage, the relative abundance of Firmicutes decreased from 41.44 to 18.61%, whereas the proportions of Bacteroidetes increased from 39.42 to 79.75%. Other phyla dropped obviously.
The characterizations of functional bacteria
The common polysaccharide-degrading bacteria are shown in Table 3. The main hydrolytic bacteria belonged to the Clostridiales order within Firmicutes, including Ruminococcus, Ruminiclostridium, and Ruminofilibacter. Except for Ruminococcus flavefaciens, the other hydrolytic bacteria had a similar change tendency. There was high abundances in the first two stages. The proportions deceased obviously in the overload stage, especially Fibrobacter disappearing from the reactor, although there was no significant difference in the steady and overload stage (p < 0.05). VFA-producing bacteria by fermentation mainly included Lachnospiraceae and Bacteroidales. In the overload stage, the relative abundance of Lachnospiraceae gradually diminished, whereas Bacteroidales increased significantly (p < 0.05), accounting for 79.75%. The members of Clostridiales (Caldicoprobacter, Sedimentibacter, Anaerovorax) and Spirochaetaceae were regarded as acidogens (Azman et al. 2015; FitzGerald et al. 2015; Wang et al. 2010). The relative abundance was higher in the first two stages than in the overload stage, although there was a slight discrepancy. In the steady stage, the abundances of the Synergistetes phylum increased to 9.45%. Propionate oxidizers belonged to Syntrophobacterales in the Proteobacteria phylum, including Syntrophobacter and Pelotomaculum. Syntrophus and Syntrophomonas, attaching to Syntrophomonadaceae of Firmicutes phylum, were the butyrate oxidizers.
The changes of methanogenic community composition in the three stages
The proportion of methanogens was relatively low in the functional microbiota because sequencing primer sets were not designed specifically for the methanogenic community. Therefore, Q-PCR analysis was performed for the total archaea, Methanobacteriales, and Methanosarcinales, according to Steinmetz et al. (2016). The actual fragment sizes were identified by sequencing and had 100% similarity with the known methanogenic populations in the GenBank database (shown in Supplementary Figure S2). The standard regression equation was constructed with 10-fold serially diluted plasmid DNA as follows. Total archaea: Y = − 3.436 lg X + 41.072 (R2 = 0.9986); Methanobacteriales: Y = − 3.472 lg X + 40.885 (R2 = 0.9958); Methanosarcinales: Y = − 3.135lgX + 39.172 (R2 = 0.9873), where X represents the plasmid DNA concentrations with 109–106 copies/μL, and Y is the Ct value. Consequently, the copies of the methanogenic archaea were calculated and shown in Table 4. In the steady stage, the total archaea and Methanosarcinales increased to 6.20 × 108 and 1.75 × 107 copies/mL, respectively. The Methanobacteriales increased significantly (p < 0.05), being 3.30 × 108 copies/mL. In the overload stage, the total archaea and Methanobacteriales decreased significantly (p < 0.05). The copies of Methanosarcinales dropped to 2.08 × 104 copies/mL.
Discussion
The changes of digestion performance
The theoretical CH4 yield of cellulose is 0.415 L/g VS based on the structural formula of (C6H10O5)n, and the theoretical value of hemicellulose is 0.424 L/g VS with the structural formula of (C5H8O4)n (Karthikeyan and Visvanathan 2013). Thus, the feedstock achieves 0.35–0.36 L/g VS of theoretical values based on the proportions of cellulose and hemicellulose in this study. The maximum yield at phase 5 reached 56–58% of the theoretical value, which had obvious advantages compared with the lignocellulose digestion after physicochemical pretreatments (Mussoline et al. 2013). The higher CH4 yields could be explained from three respects. First, mixed substrates, including easily digestible corn stalks and guinea grass, were beneficial to lignocellulose hydrolysis. In a previous study, Fuma et al. (2012) confirmed that bean husks facilitated the ability of the hydrolytic bacteria to colonize within the residues, to stimulate the fibrolytic enzymes of Fibrobacter succinogenes and then accelerate the biodegradation of recalcitrant rice straw. Second, the ruminal microbiota contained high amounts of lignocellulose-degrading consortia, which benefitted the hydrolytic efficiency (Dai et al. 2015). In addition, the methanogenic sludge enhanced the beneficial cooperation since the microbial interactions were involved in the hydrolysis of lignocelluloses (Azman et al. 2015; Miller et al. 2000; Rychlik and May 2000). Our previous study on the rumen co-inoculated anaerobic digestion of rice straw indicated that co-inoculation enhanced the hydrolysis and CH4 yields, making digestion more efficient (Deng et al. 2017a).
In a previous study on single corn stalk degradation with recirculation, Hu et al. (2014) found that refractory degradable TOC components accumulating in the digestate resulted in the malfunction of the anaerobic digester. In addition, excessively high concentrations of VFAs will inhibit methanogenic activities (Hu et al. 2014; Nordberg et al. 2007; Shakeri Yekta et al. 2012; Xiao et al. 2013; Zhang et al. 2016). The concentrations over 8 g/L were noted as the threshold value of inhibition in mesophilic conditions (Karthikeyan and Visvanathan 2013). In the present study, the total VFA concentration of 9.09 g/L was accompanied by the inhibition of methanogenesis in the overload stage, making the CH4 yield drop sharply (shown in Table 2).
Acetate composition performed a “multi-faceted” role, not only as the substrate of the acetotrophic methanogenesis but also as the inhibitor of methanogens when it is higher than the inhibition threshold. The investigation by Xiao et al. (2013) indicated that the threshold value was 3 g/L, which completely inhibited CH4-producing activity. In the present study, the acetate concentration was lower than the inhibition value in the whole digestion process. This also indicated that the acetotrophic methanogenesis was enhanced by co-inoculation. In addition, Lerm et al. (2012) believed that syntrophic oxidation accelerated with increment of LRs. Once exceeding the metabolic capacities of the methanogens, the reducing H2 could not be consumed and then metabolized via propionate and butyrate accumulation. In the present study, the average concentrations of propionate and butyrate reached 6.54 and 0.53 g/L respectively in the overload stage, which indicated that anaerobic digestion was inhibited (shown in Table 2).
Nitrogen is indispensable to microbial synthesis and metabolism. The NH4+ concentration with 0.20 g/L was of vital importance for anaerobic bacteria (Karthikeyan and Visvanathan 2013). However, the slurry recirculation could also cause an accumulation of NH4+, as well as the TOC components, which eventually affected digestion performance. In a previous study on the anaerobic digestion of alfalfa silage, NH4+ inhibition eventually appeared at 100% liquid recirculation (Nordberg et al. 2007). It is generally believed that the NH4+ concentration at more than 1.50 g/L has obvious toxicity effects on the anaerobic reactor (Karthikeyan and Visvanathan 2013). In the present study, the initial nitrogen source mainly came from the co-inocula. The raw materials hardly brought nitrogen into the reactor because the lignocelluloses had a high C/N ratio of 46.82. The accumulated NH4+ was not the inhibition factor because of the low concentration. The increment of concentrations just reflected the organic nitrogen releasing from dying microbes in the overload stage.
Biodegradability of lignocellulosic compositions
It is generally believed that CH4 is produced from the degradation of cellulose and hemicellulose rather than lignin composition in the anaerobic digestion process (Lazuka et al. 2015). This is similar to the shifts of lignocellulosic compositions witnessed in the present study. As shown in Table 2, the corresponding hydrolytic activities in the steady phase were higher than those of the other two stages. The xylanase and cellulase activities at phase 5 had the highest values of 6.45 and 50.06 U/mL respectively, which were consistent with the CH4 production rate. It also suggested that the hydrolysis process was still the rate-limiting step regulating lignocelluloses degradation, and the high hydrolytic activities were beneficial to the CH4-producing efficiency.
The composition degradation rate dropped significantly in the overload stage, which illustrated that the hydrolytic activities were restrained. This phenomenon was explained from two respects. On the one hand, hydrolytic bacteria were inhibited by high VFA concentrations. In a previous study, Romsaiyud et al. (2009) found that high acetate concentration of over 1.80 g/L would affect the enzymatic cellulose hydrolysis, which led to the inhibition of the hydrolytic bacteria (Romsaiyud et al. 2009). In the present study, the acetate concentration of this stage was 2.02 g/L, higher than the inhibitory threshold (shown in Table 2). On the other hand, the methanogens were more easily inhibited by high VFA concentrations than cellulolytic microbes, resulting that H2 produced during the hydrolysis process could be consumed by hydrogenotrophic methanogenesis. This would decrease hydrolytic efficiencies of lignocelluloses (Azman et al. 2015; Miller et al. 2000; Rychlik and May 2000).
The characterizations of microbial community composition
The phyla Bacteroidetes and Firmicutes mostly dominated in the samples, which agreed with previous studies on the anaerobic digestion of lignocelluloses (Lazuka et al. 2015; Sun et al. 2016). Within the Firmicutes, Clostridiales members served predominant roles in polysaccharide digestion (Azman et al. 2015). In the steady stage, higher abundances of Synergistetes indicated that efficient syntrophic oxidations were achieved (Liu et al. 2009). Chloroflexi populations were known to metabolize carbohydrate and microbial detritus (Miura and Okabe 2008; Sun et al. 2016). The Cloacimonetes bacteria were discovered to hydrolyze the celluloses or utilize fermentation products (Limam et al. 2014). The phylum Proteobacteria contained the genes coding for cellulolytic enzymes (Azman et al. 2015). Thus, the analysis of the dominant community composition accompanied with high CH4 yields and hydrolytic activities suggested that these bacteria played an important role in the process of lignocellulosic hydrolysis and fermentation metabolism (shown in Table 2).
Within the main hydrolytic bacteria of start-up and steady stages, Ruminococcus, Butyrivibrio fibrisolvens, and Fibrobacter were highly efficient hydrolytic bacteria that settled in the rumen niche (Dai et al. 2015; Morrison et al. 2009; Nyonyo et al. 2014). Ruminiclostridium displayed anaerobic hydrolytic activities while some species of Ruminofilibacter (e.g., Ruminofilibacter xylanolyticum) processed hemi-cellulolytic activities (Azman et al. 2015; Kröber et al. 2009). Higher activities of cellulase and xylanase were then achieved in the reactor. Within the VFA-producing bacteria, many unclassified species of Lachnospiraceae belonged to soluble polysaccharide-degrading bacteria (Nyonyo et al. 2014). Bacteroidales, usually present in the anaerobic reactor, degraded various polysaccharides with the production of VFAs (Hatamoto et al. 2014). Prevotellaceae numbers, which were affiliated with the Bacteroidales order, can utilize pectin to produce succinate or butyrate. It was suggested that the degradation was beneficial to the exposure of the cellulose portions (Fondevila and Dehority 1996; Lazuka et al. 2015). A significant increment of Bacteroidales in the abundance indicated that this order was involved in the acidification in the overload stage. In contrast, the higher abundances of acetogens (Clostridiales and Spirochaetaceae) illustrated efficient fermentation activities in the first two stages (FitzGerald et al. 2015).
It was regarded that syntrophic oxidation accelerated with the increment of LR within a certain range (Lerm et al. 2012). In the steady stage, the appearance of Synergistetes indicated that efficiently acetotrophic methanogenesis was attained (FitzGerald et al. 2015; Liu et al. 2009). Syntrophic oxidizers relieved the accumulation of propionate and butyrate. This also explained the performances of individual VFA in the co-inoculated system. In the overload stage, the proportions of propionate and butyrate oxidizers significantly decreased, especially Syntrophobacter disappeared from the reactor. This was attributed to the inhibition of oxidization and associated methanogenesis because of the overloading of lignocelluloses (Lerm et al. 2012; Müller et al. 2010).
The shifts of methanogenic compositions were used to illustrate how the operation parameters influenced the methanogenic community structure and bioconversion pathways (Steinmetz et al. 2016). Hydrogenotrophic Methanobacteriales, including Methanobacterium, Methanobrevibacter, and Methanosphaera, were the most abundant methanogens in the rumen (Dai et al. 2015; Janssen and Kirs 2008). The members within this order could form functional consortia with hydrolytic bacteria and syntrophic oxidizers to accelerate the interspecies H2 transfer (Miller et al. 2000; Rychlik and May 2000). The methylotrophic and aceticlastic Methanosarcinales (Methanosarcina and Methanosaeta) were indispensable for anaerobic digestion due to the rapid consumption of acetate (Demirel and Scherer 2008; FitzGerald et al. 2015). The increments of methanogen copies were achieved for the enhancement of methanogenic activities in the steady stage.
Enrichments of functional consortia and mechanisms of acidification
As shown in Fig. 5, the ruminal microbiota co-inoculated with anaerobic sludge were enriched under the LR of 1.5–4 g/L/day and slurry recirculation. In view of the strengthening mechanism, co-inoculum by increasing the proportion of methanogens was beneficial to the formation of the functional consortia. First, hydrogenotrophic methanogens consumed the H2 produced during the lignocellulosic digestion process, which made the NAD+ of ruminal hydrolytic bacteria (Ruminococcus, Fibrobacter) regenerate, and then more ATP was produced by the cell. As a result, the digestion efficiency of the hydrolytic consortia was enhanced (Miller et al. 2000; Rychlik and May 2000). In a study on the switchgrass digestion in the bovine rumen, Piao et al. (2014) found that Methanobrevibacter was inevitable for cellulolytic activities in situ. On the investigation of digestion performance enhanced by sheep rumen microbiota, microbial profiles suggested that enhancement cultures increased the abundances of Ruminococcaceae and hydrogenotrophic Methanobacteriales, which could enhance the CH4 yields from wheat straw in batch reactors (Ozbayram et al. 2017). In our previous study of rumen-derived digestion of rice straw, it was indicated that microbial cooperation involving Methanobrevibacter and Ruminococcus was pivotal for digestion activities (Deng et al. 2017a). Thus, the high hydrolytic activities and CH4 yields would contribute to the effect of the methanogenic sludge co-inoculated into the artificial reactor in the present study. Consequently, the predominant ruminal bacteria involving Ruminococcus and Fibrobacter settled down in the start-up and steady stages, which was different from other studies in biogas plants (Azman et al. 2015; Sun et al. 2016).
In the liquid phase, VFAs were consumed only when syntrophic oxidizing consortia formed because the methanogens could overcome the unfeasible thermodynamics by lowering the H2 concentrations. Hydrogenotrophic Methanospirillum and Methanobacterium were usually regarded as syntrophic partners (Müller et al. 2010). In the present study, the obvious increments of Methanobacteriales could accelerate H2 transfer and then made the syntrophic oxidations more efficient. This also explained the reactor performance of no propionate and butyrate accumulation in the steady stage.
In the overload stage with 6 g/L/day LR, the TOC concentrations in the slurry increased rapidly to 13.67 from 9.28 g/L, and the total VFA concentration reached 9.09 g/L. Subsequently, the copies of total archaea, especially Methanobacteriales, decreased significantly (p < 0.05), and the CH4 yield dropped to 0.11 L/g VS. Lerm et al. (2012) also found that hydrogenotrophic populations were especially important for the reactor, and the VFAs accumulated along with disappearance of Methanobacterium. As a result, the unconsumed H2 then suppressed the hydrolytic activities and syntrophic oxidations, making the corresponding functional bacteria decline in abundance. At the same time, Bacteroidales increased significantly (p < 0.05), and the Prevotellaceae and Rikenellaceae members within the order could produce excessive VFAs. These acid-producing bacteria were found to be influenced slightly by the inhibiting components in the liquid fractions (Hu et al. 2007; Hu and Yu 2005; Lerm et al. 2012; Zhang et al. 2016). Thus, the inhibition of hydrogenotrophic methanogens provoked by overloading lignocelluloses eventually decreased the digestion efficiency in the semi-continuous reactor.
References
APHA (2005) Standard methods for the examination of water and wastewater, 21th edn. American Public Health Association (APHA), New York
Azman S, Khadem AF, van Lier JB, Zeeman G, Plugge CM (2015) Presence and role of anaerobic hydrolytic microbes in conversion of lignocellulosic biomass for biogas production. Crit Rev Env Sci Tech 45:2523–2564. https://doi.org/10.1080/10643389.2015.1053727
Dai X, Tian Y, Li J, Su X, Wang X, Zhao S, Liu L, Luo Y, Liu D, Zheng H, Wang J, Dong Z, Hu S, Huang L (2015) Metatranscriptomic analyses of plant cell wall polysaccharide degradation by microorganisms in the cow rumen. Appl Environ Microbiol 81:1375–1386. https://doi.org/10.1128/AEM.03682-14
De Bok FAM, Harmsen HJM, Plugge CM, De Vries MC, Akkermans ADL, De Vos WM, Stams AJM (2005) The first true obligately syntrophic propionate-oxidizing bacterium, Pelotomaculum schinkii sp. nov., co-cultured with Methanospirillum hungatei, and emended description of the genus Pelotomaculum. Int J Syst Evol Micr 55:1697–1703. https://doi.org/10.1099/ijs.0.02880-0
Demirel B, Scherer P (2008) The roles of acetotrophic and hydrogenotrophic methanogens during anaerobic conversion of biomass to methane: a review. Rev Environ Sci Biotechnol 7:173–190. https://doi.org/10.1007/s11157-008-9131-1
Deng Y, Huang Z, Ruan W, Zhao M, Miao H, Ren H (2017a) Co-inoculation of cellulolytic rumen bacteria with methanogenic sludge to enhance methanogenesis of rice straw. Int Biodeterior Biodegrad 117:224–235. https://doi.org/10.1016/j.ibiod.2017.01.017
Deng Y, Huang Z, Zhao M, Ruan W, Miao H, Ren H (2017b) Effects of co-inoculating rice straw with ruminal microbiota and anaerobic sludge: digestion performance and spatial distribution of microbial communities. Appl Microbiol Biot 101:5937–5948. https://doi.org/10.1007/s00253-017-8332-3
FitzGerald JA, Allen E, Wall DM, Jackson SA, Murphy JD, Dobson AD (2015) Methanosarcina play an important role in anaerobic co-digestion of the seaweed Ulva lactuca: taxonomy and predicted metabolism of functional microbial communities. PLoS One 10:e0142603. https://doi.org/10.1371/journal.pone.0142603
Fondevila M, Dehority BA (1996) Interactions between Fibrobacter succinogenes, Prevotella ruminicola, and Ruminococcus flavefaciens in the digestion of cellulose from forages. J Anim Sci 74(3):678–684. https://doi.org/10.2527/1996.743678x
Fuma R, Oyaizu S, Nukui Y, Ngwe T, Shinkai T, Koike S, Kobayashi Y (2012) Use of bean husk as an easily digestible fiber source for activating the fibrolytic rumen bacterium Fibrobacter succinogenes and rice straw digestion. Anim Sci J 83(10):696–703. https://doi.org/10.1111/j.1740-0929.2012.01017.x
Hatamoto M, Kaneshige M, Nakamura A, Yamaguchi T (2014) Bacteroides luti sp. nov., an anaerobic, cellulolytic and xylanolytic bacterium isolated from methanogenic sludge. Int J Syst Evol Micr 64:1770–1774. https://doi.org/10.1099/ijs.0.056630-0
Hu Y, Shen F, Yuan H, Zou D, Pang Y, Liu Y, Zhu B, Chufo WA, Jaffar M, Li X (2014) Influence of recirculation of liquid fraction of the digestate (LFD) on maize stover anaerobic digestion. Biosyst Eng 127:189–196. https://doi.org/10.1016/j.biosystemseng.2014.09.006
Hu ZH, Yu HQ, Yue ZB, Harada H, Li YY (2007) Kinetic analysis of anaerobic digestion of cattail by rumen microbes in a modified UASB reactor. Biochem Eng J 37:219–225. https://doi.org/10.1016/j.bej.2007.04.013
Hu ZH, Yu HQ (2005) Application of rumen microorganisms for enhanced anaerobic fermentation of corn stover. Process Biochem 40(7):2371–2377. https://doi.org/10.1016/j.procbio.2004.09.021
Janssen PH, Kirs M (2008) Structure of the archaeal community of the rumen. Appl Environ Microbiol 74:3619–3625. https://doi.org/10.1128/AEM.02812-07
Karthikeyan OP, Visvanathan C (2013) Bio-energy recovery from high-solid organic substrates by dry anaerobic bio-conversion processes: a review. Rev Environ Sci Biotechnol 12:257–284. https://doi.org/10.1007/s11157-012-9304-9
Kröber M, Bekel T, Diaz NN, Goesmann A, Jaenicke S, Krause L, Miller D, Runte KJ, Viehöver P, Pühler A, Schlüter A (2009) Phylogenetic characterization of a biogas plant microbial community integrating clone library 16S-rDNA sequences and metagenome sequence data obtained by 454-pyrosequencing. J Biotechnol 142:38–49. https://doi.org/10.1016/j.jbiotec.2009.02.010
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lazuka A, Auer L, Bozonnet S, Morgavi DP, O'Donohue M, Hernandez-Raquet G (2015) Efficient anaerobic transformation of raw wheat straw by a robust cow rumen-derived microbial consortium. Bioresour Technol 196:241–249. https://doi.org/10.1016/j.biortech.2015.07.084
Lerm S, Kleyböcker A, Miethling-Graff R, Alawi M, Kasina M, Liebrich M, Würdemann H (2012) Archaeal community composition affects the function of anaerobic co-digesters in response to organic overload. Waste Manag 32:389–399. https://doi.org/10.1016/j.wasman.2011.11.013
Limam RD, Chouari R, Mazéas L, Wu TD, Li T, Grossin-debattista J, Guerquin-Kern JL, Saidi M, Landoulsi A, Sghir A, Bouchez T (2014) Members of the uncultured bacterial candidate division WWE1 are implicated in anaerobic digestion of cellulose. MicrobiologyOpen 3:157–167. https://doi.org/10.1002/mbo3.144
Liu FH, Wang SB, Zhang JS, Zhang J, Yan X, Zhou HK, Zhao GP, Zhou ZH (2009) The structure of the bacterial and archaeal community in a biogas digester as revealed by denaturing gradient gel electrophoresis and 16S rDNA sequencing analysis. J Appl Microbiol 106:952–966. https://doi.org/10.1111/j.1365-2672.2008.04064.x
Müller N, Worm P, Schink B, Stams AJ, Plugge CM (2010) Syntrophic butyrate and propionate oxidation processes: from genomes to reaction mechanisms. Env Microbiol Rep 2:489–499. https://doi.org/10.1111/j.1758-2229.2010.00147.x
Miller T, Currenti E, Wolin M (2000) Anaerobic bioconversion of cellulose by Ruminococcus albus, Methanobrevibacter smithii, and Methanosarcina barkeri. Appl Microbiol Biot 54:494–498. https://doi.org/10.1007/s002530000430
Miura Y, Okabe S (2008) Quantification of cell specific uptake activity of microbial products by uncultured Chloroflexi by microautoradiography combined with fluorescence in situ hybridization. Environ Sci Technol 42:7380–7386. https://doi.org/10.1021/es800566e
Morrison M, Pope PB, Denman SE, Mcsweeney CS (2009) Plant biomass degradation by gut microbiomes: more of the same or something new? Curr Opin Biotech 20:358–363. https://doi.org/10.1016/j.copbio.2009.05.004
Mussoline W, Esposito G, Giordano A, Lens P (2013) The anaerobic digestion of rice straw: a review. Crit Rev Env Sci Tec 43:895–915. https://doi.org/10.1080/10643389.2011.627018
Nelson MC, Morrison HG, Benjamino J, Grim SL, Graf J (2014) Analysis, optimization and verification of Illumina–generated 16S rRNA gene amplicon surveys. PLoS One 9:e94249–e94249. https://doi.org/10.1371/journal.pone.0094249
Nordberg Å, Jarvis Å, Stenberg B, Mathisen B, Svensson BH (2007) Anaerobic digestion of alfalfa silage with recirculation of process liquid. Bioresour Technol 98:104–111. https://doi.org/10.1016/j.biortech.2005.11.027
Nyonyo T, Shinkai T, Mitsumori M (2014) Improved culturability of cellulolytic rumen bacteria and phylogenetic diversity of culturable cellulolytic and xylanolytic bacteria newly isolated from the bovine rumen. FEMS Microbiol Ecol 88:528–537. https://doi.org/10.1111/1574-6941.12318
Ozbayram EG, Kleinsteuber S, Nikolausz M, Ince B, Ince O (2017) Effect of bioaugmentation by cellulolytic bacteria enriched from sheep rumen on methane production from wheat straw. Anaerobe 46:122–130. https://doi.org/10.1016/j.anaerobe.2017.03.013
Piao H, Lachman M, Malfatti S, Sczyrba A, Knierim B, Auer M, Tringe SG, Mackie RI, Yeoman CJ, Hess M (2014) Temporal dynamics of fibrolytic and methanogenic rumen microorganisms during in situ incubation of switchgrass determined by 16S rRNA gene profiling. Front Microbiol 5:307. https://doi.org/10.3389/fmicb.2014.00307
Romsaiyud A, Songkasiri W, Nopharatana A, Chaiprasert P (2009) Combination effect of pH and acetate on enzymatic cellulose hydrolysis. J Environ Sci 21:965–970. https://doi.org/10.1016/S1001-0742(08)62369-4
Rychlik JL, May T (2000) The effect of a methanogen, Methanobrevibacter smithii, on the growth rate, organic acid production, and specific ATP activity of three predominant ruminal cellulolytic bacteria. Curr Microbiol 40:176–180. https://doi.org/10.1007/s002849910035
Shakeri Yekta S, Gonsior M, Schmitt-Kopplin P, Svensson BH (2012) Characterization of dissolved organic matter in full scale continuous stirred tank biogas reactors using ultrahigh resolution mass spectrometry: a qualitative overview. Environ Sci Technol 46:12711–12719. https://doi.org/10.1021/es3024447
Steinmetz RLR, Mezzari MP, da Silva MLB, Kunz A, do Amaral AC, Tápparo DC, Soares HM (2016) Enrichment and acclimation of an anaerobic mesophilic microorganism’s inoculum for standardization of BMP assays. Bioresour Technol 219:21–28. https://doi.org/10.1016/j.biortech.2016.07.031
Sun L, Liu T, Müller B, Schnürer A (2016) The microbial community structure in industrial biogas plants influences the degradation rate of straw and cellulose in batch tests. Biotechnol Biofuels 9:128. https://doi.org/10.1186/s13068-016-0543-9
Van Soest PJ, Robertson JB, Lewis BA (1991) Methods for dietary fiber, neutral detergent fiber, and nonstarch polysaccharides in relation to animal nutrition. J Dairy Sci 74:3583–3597. https://doi.org/10.3168/jds.S0022-0302(91)78551-2
Wang H, Vuorela M, Keränen AL, Lehtinen TM, Lensu A, Lehtomäki A, Rintala J (2010) Development of microbial populations in the anaerobic hydrolysis of grass silage for methane production. FEMS Microbiol Ecol 72:496–506. https://doi.org/10.1111/j.1574-6941.2010.00850.x
Xiao KK, Guo CH, Zhou Y, Maspolim Y, Wang JY, Ng WJ (2013) Acetic acid inhibition on methanogens in a two-phase anaerobic process. Biochem Eng J 75:1–7. https://doi.org/10.1016/j.bej.2013.03.011
Zhang H, Zhang P, Ye J, Wu Y, Fang W, Gou X, Zeng G (2016) Improvement of methane production from rice straw with rumen fluid pretreatment: a feasibility study. Int Biodeterior Biodegrad 113:9–16. https://doi.org/10.1016/j.ibiod.2016.03.022
Acknowledgments
The authors were grateful to the Enzyme Engineering Technology Center of Jiangsu for the help in the experiments of this work
Funding
This work was funded by the National Natural Science Foundation of China [grant numbers 21506076, 51678279, and 51508230], National Science and Technological Support of China [grant number 2014BAC25B01], Fundamental Research Funds for the Central Universities [grant number JUSRP1703XNC], the fund of Jiangsu Key Laboratory of Anaerobic Biotechnology [grant number JKLAB201607], and the funds for green catalysis and applied enzyme team of Changzhou Vocational Institute of Engineering [111308002216006, 201713102015Y].
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 827 kb)
Rights and permissions
About this article
Cite this article
Deng, Y., Huang, Z., Ruan, W. et al. Enriching ruminal polysaccharide-degrading consortia via co-inoculation with methanogenic sludge and microbial mechanisms of acidification across lignocellulose loading gradients. Appl Microbiol Biotechnol 102, 3819–3830 (2018). https://doi.org/10.1007/s00253-018-8877-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-8877-9