Abstract
The carbon and energy needed for bioconversion processes face trade-offs between cell reproduction and chemical synthesis. In most processes, microbial cells containing overexpressed pathway enzymes were accumulated in exponential phase before the productions of value-added chemicals dominate the carbon and energy fluxes in stationary phase. The pathway enzymes need to be continuously supplied to compensate their degradation, but the promoters driving their overexpressions are downregulated under stationary phase or stressed conditions. In this study, the gadA promoter enabled stress-resistant and growth phase-independent isobutanol production up to 10–28 g l−1. To investigate the activation mechanism of gadA promoter and its potential in metabolic engineering, an in vitro transcription system was established. Results showed that gadA promoter could be transcribed efficiently under environments that inhibit the transcription of ribosomal promoters, while under moderate to rapid growth conditions, the large majority of new cellular transcripts are ribosomal. This differential transcription relies on the accumulation of environmental glutamate and/or the loss of supercoiling. These results implied that the gadA promoter could be functional or even dominate the cellular transcription under an exceedingly wide range of physiological conditions. Therefore, the gadA promoter is a novel candidate for driving pathway enzyme overexpressions under not only exponential phase but also stationary phase and stressed conditions, which is important for achieving efficient biofuel production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
To minimize the cost of raw materials and production process and to survive the competition against the low petroleum price, the current process of the biofuel and bulk chemical production have to march toward its theoretical yield. This requires continuous and effective channeling of the finite carbon source to the final products. To do this, the pathway genes must be overexpressed efficiently to minimize the production of by-products and biomass.
The bioconversion processes face trade-offs between cell reproduction and chemical synthesis. The highest yield is generally achieved after the cells stop growing, usually at the beginning of the stationary phase, when most of the resources is now available for direct conversion into the target products (Huo et al. 2011). However, this period with high productivity could not persist. This is because the expressions of pathway genes are generally driven by the σ70-RNA polymerase, which is the house-keeping RNA polymerase that dominates the gene expression during the exponential phase (Battesti et al. 2011). As stresses build up during the process of cell growth (Benjaphokee et al. 2012; Goo et al. 2012), cells entering the stationary phase start to produce the stress responsive σ38. The σ38 accumulates and outcompetes the σ70 for core polymerase, leading to decreased expression of the pathway genes and an early end to the high yield stage (Sharma 2016). Therefore, to maintain continuous productions, a stress-resistant promoter that is compatible with both the σ70 and σ38 RNA polymerases is needed, and its mechanism is worth of exploring.
As one of the most commonly used host strains, Escherichia coli has an unusual ability to survive stress environments such as acid gastrointestinal tract (Foster 2004). The most important system that confers this ability depends on the decarboxylation of glutamate, which involves the glutamate decarboxylase encoded by the gadA gene (Feehily and Karatzas 2013; Zhao and Houry 2010). This gene is controlled by a special promoter that could be recognized by both the σ70 and σ38 RNA polymerases (Foster 2004; Ma et al. 2003). Therefore, the gadA promoter has the potential to express the pathway enzymes in both the exponential and stressed stationary phases. Exploring the responses of the gadA promoter toward intracellular or extracellular stressors is required to evaluate its potentials in driving gene overexpressions for the productions of biofuels and bulk chemicals.
Acid stress is one of the most common stresses that inhibit the growth and productivity of cell factories; it can originate from the preprocessing of raw materials or the accumulation of carboxylic acids released during microbial growth (Nicolaou et al. 2010; Warnecke and Gill 2005). One way to rescue the productivity is to use stress-resistant promoters to achieve prior transcription of the pathway genes over other cellular transcriptions, so the majority of the limited cellular resources can still be channeled to the productions of targeted chemicals under stressed conditions. The main flux of cellular resources is the production of ribosomes, which can account for > 90% of the total RNA synthesis under unstressed conditions (Rosenow et al. 2001). This bulk transcription is supposed to go down under osmotic shock (Gralla and Vargas 2006) or nutritional downshift (Gralla 2005). However, it remains unknown (1) whether ribosomal RNA transcription decreases under acid stress and (2) whether low pH favors transcription from the gadA promoter over the bulk transcription.
It has been reported that gadA expression was regulated by various factors. GadX was shown as the primary activator of the gadA gene (Tramonti et al. 2002), and the histone-like protein H-NS (De Biase et al. 1999) could repress gadA transcription by binding at the promoter region (Giangrossi et al. 2005). However, regulation of gadA transcription is rarely dominated by sole factor but involves complex interactions between various mechanisms. For instance, the H-NS might indirectly restrain the activity of gadA promoter by changing the DNA topology (Tupper et al. 1994), while its inhibitory effect could also be antagonized by the fusion protein MalE-GadX in a dose-dependent manner (Giangrossi et al. 2005). Therefore, investigating the synergic effects of primary regulators is the key to understand the role of gadA promoter in mediating pathway gene expressions under stressed environments. When acid or osmotic shock occurs, glutamate accumulates intracellularly, which antagonizes the osmolytes or to be converted into γ-amino butyric acid (GABA) accompanying the release of intracellular proton (Krulwich et al. 2011). Meanwhile, the chromosome becomes relaxed (less supercoiled) due to the lack of ATP for the gyrase (Dorman 2013). Since the expressions of σ70-dependent promoters generally require supercoiled template, abundant σ70-holoenzyme, and a low glutamate environment (Dorman 2013; Gralla and Vargas 2006), changes in these intracellular factors induced by stresses will inhibit the expressions of σ70-dependent genes, giving ways to the σ38-dependent promoters (Weber et al. 2005). It has been shown that the transcription by σ38 RNA polymerase is regulated by the level of glutamate and DNA supercoiling (Métris et al. 2017; Rosenthal et al. 2008). However, how these factors interactively affect σ38 transcription on gadA promoter is still not known.
Our primary aim here is to investigate whether the gadA promoter could be utilized for driving biofuel production and systematically elucidates the roles of key physiological and environmental factors on the transcription of gadA promoter upon stationary and acid stresses. To do this, the effects of gadA promoter in driving gene overexpressions throughout the whole fermentation process were first verified using the isobutanol synthetic pathway. In vitro assays were then employed to uncover the comprehensive effects of related factors on gadA transcription, including low pH, the accumulation of glutamate, the presence of σ38, the key activator GadE (Castanié-Cornet et al. 2010), and the decrease of DNA supercoiling. We hypothesized that the gadA promoter has an unusual design that allows it to be selectively transcribed at low pH under the widest range of intracellular conditions, making it a novel candidate to drive pathway gene overexpressions in not only exponential phase but also in stationary phase or stressed conditions.
Material and methods
Proteins, strains, and plasmids
N-terminally His-tagged σ38 was obtained as described (Rosenthal et al. 2008). Core RNA polymerase and σ70-holoenzyme were purchased from Epicenter Technologies. Plasmids encoding GadE were constructed in pET14, and the His-tagged proteins were purified with standard affinity resins. Plasmid pGadA was constructed by inserting a 300 base pair fragment containing the gadA promoter (from − 150 to + 150) into pTH8 through the unique restriction sites of BamHI and HindIII. Relaxed DNA fragments were obtained by digesting the plasmids by XmnI.
The plasmids used to examine the effects of gadA promoter in driving isobutanol production were derived from the previously reported plasmids pSA69 and pSA65 (Atsumi et al. 2008b; Atsumi et al. 2010), which encode the two operons that constitute the isobutanol biosynthetic pathway. The promoter PLlacO1 located upstream of the RBS site of the alsS-ilvCD operon in pSA69 was replaced by the 183-bp gadA promoter amplified from the E. coli genomic DNA. A fragment left out the region from − 183 to − 60 upstream the transcription start site of the gadA promoter was then amplified from this plasmid with 16-bp overlap at both ends. This fragment was ligated, generating a new plasmid carrying the same alsS-ilvCD operon but controlled by a 60-bp gadA promoter (P gadA60 ). The same strategy was employed to replace the promoter PLlacO1 in pSA65 with the gadA promoter. The PLlacO1 located upstream of the RBS site of the kivd-adhA operon in pSA65 was replaced by the 183-bp gadA promoter. The fragment containing the 60-bp gadA promoter was amplified and ligated, generating a new plasmid carrying the same kivd-adhA operon but under the control of P gadA60 . The previously constructed E. coli strain JCL260 with knockouts of adhE, frdBC, fnr-ldhA, pta, and pflB (Atsumi et al. 2008a) was used as host for isobutanol fermentation. Detailed information for plasmids and strains as well as primers were listed in Tables S1 and S2.
In vivo messenger RNA analysis
Primer extension experiments were as described (Rosenthal et al. 2006). E. coli strain K12 was grown overnight in 5 ml of LB at 37 °C. The cells were diluted 1:100 with fresh medium and grown to an optical density of 0.6. Acid challenge cultures were harvested either immediately before or 5 min after the pH was decreased to 5.5, which required addition of approximately 170-mM MES. Cells were harvested by centrifugation and stored at − 70 °C. RNA was extracted from the pellets using a Qiagen RNeasy Kit. The total amount of RNA of each reaction was normalized, and the RNA was detected by extension by gadA or rrnB primer (Gralla and Vargas 2006). Ten-microliter reactions for primer extension analysis contained 10 μg of total RNA, 10 nM labeled primer, reverse transcriptase buffer (Promega), 5 U of reverse transcriptase, and 0.2 mM deoxy-NTPs. Urea stop dye was added, and samples were run at 21 W on 6% polyacrylamide sequencing gel. Radioactive bands were visualized and quantified by phosphorimager analysis. RNA samples were prepared at least three times, and the average was taken.
The alsS messenger RNA (mRNA) was quantified by qRT-PCR using a Roche LightCycler 96 System with SYBR Green I detection. The hcaT gene was used as reference as it tends to maintain steady expression upon enzyme overexpressions under various growth conditions (Zhou et al. 2011). Quantitative PCR was performed using SYBR Premix DimerEraser (TaKaRa) with an initial denaturing at 95 °C for 30 s, followed by 40 cycles of 5 s at 95 °C, 30 s at 55 °C, and 25 s at 72 °C. Melting curve analysis was performed to confirm specific amplification. Data was acquired from three biological replicates and each sample was measured in duplicate.
In vitro transcription
For single-round transcription experiments, 50 nM σ70-holoenzyme or σ38-holoenzyme (formed by pre-incubation of 200 nM σ38 and 50 nM core RNA polymerase at 37 °C for 10 min) were added into 1× Buffer B (50 mM Tris-HCL at pH 7.9, 3 mM MgCl2, 0.1 mM EDTA, 1 mM DTT, and 100 μg/ml BSA) with 100 mM KCl. The final concentration of NaCl, originally from protein storage buffer, is 37.5 mM. In some cases, 200 mM potassium glutamate was added to Buffer B + KCl when σ38 was present. Of the supercoiled or relaxed plasmid DNA, 5 nM and the designed concentration of GadE were added and incubated for 15 min at 37 °C. NTP challenge mix (final concentration 150 μM each ATP, CTP and GTP, 5 μM UTP, 0 or 200 nCi UTP, 100 μg/ml heparin) was added and incubated for the appropriate time at 37 °C. When radio-labeled UTP was present, the mRNA was autoradiographed from a 6% PAGE gel. When radio-labeled UTP was not present, the mRNA was purified by Qiagen RNeasy Purification Kit and analyzed through primer extension. The samples were run on 6% PAGE with 1× Tris-buffered EDTA at 21 W for 70 min. Radioactive bands were visualized and quantitated by phosphorimager analysis. Each experiment was conducted at least three times, and bands were standardized for each set of experiments to the condition that averaged the highest signal. For experiments at lower pH, Tris was replaced with MES.
DNA footprinting
Footprinting experiments were conducted under the same conditions as in vitro transcription with 2 μg/ml DNase I (from Sigma) added for 30 s. The reactions were stopped by the addition of 150 μl of PB buffer from Qiagen (chaotropic salts) and purified with the Qiagen PCR Purification Kit. All footprinting experiments were analyzed through primer extension. Samples were run on 8% PAGE with 1× Tris-buffered EDTA at 32 W for 100 min, and radioactive bands were visualized by phosphorimager analysis. Each experiment was conducted at least three times.
Medium and cultivation
M9 medium containing 8.5% glucose, 1% yeast extract, 1 mM MgSO4, 0.1 mM CaCl2, 10−4% thiamine, 100 μg ml−1 ampicillin, and 50 μg ml−1 kanamycin was used for the production of isobutanol. The seed culture was prepared in 5 ml LB medium at 37 °C in a shaker set at 250 rpm. The overnight culture was inoculated (2% v/v) into 20 ml of fresh medium in a 250-ml screw-cap conical flask. Equal volume of oleyl alcohol was added to the flask prior to incubation for the extraction of the produced isobutanol (Connor et al. 2010). After inoculation, IPTG was added to a final concentration of 0.1 mM. Fermentation was performed under three pH conditions (pH = 7.0, 6.0, and 5.0) in triplicate at 30 °C in a shaker (250 rpm). Ten-milliliter aqueous phase and 10-ml organic phase were sampled at defined time intervals. After sampling, the fermentation broth was replenished with fresh medium and oleyl alcohol.
Detection of isobutanol
The produced isobutanol was quantified by an Agilent 6890 GC chromatograph equipped with flame ionization detector (Agilent Technologies, CA, USA). The separation of alcohol was carried out by a DB-FFAP capillary column (30 m × 0.32 mm × 0.25 μm; Agilent Technologies). For the analysis of isobutanol in aqueous phase, the GC oven temperature was initially held at 80 °C for 3 min, increased with a gradient of 115 °C min−1 until 230 °C, and held for 1 min. The isobutanol in organic phase (oleyl alcohol) was separated by first helding the oven at 90 °C for 0.5 min and heated up at a rate of 20 °C min−1 to 110 °C and held for 0.5 min. The temperature was then increased to 235 °C at 120 °C min−1 and held for 2 min. Nitrogen was used as the carrier gas. The injector and detector were maintained at 250 and 280 °C, respectively. Supernatant (0.2 μl) was sampled and injected at a split ratio of 1:50; n-pentanol was used as internal standard.
ALSS activity
Crude extracts were prepared from fresh cells using the BugBuster protein extraction reagent (Merck, Darmstadt, Germany). Total protein concentration was measured by Bradford assay. The ALSS activity was measured according to Leyval et al. (2003). Briefly, pyruvate was first converted to α-acetolactate with the addition of crude extract and then transformed into acetoin. The produced acetoin was detected by the Voges–Proskauer assay (α-naphthol and potassium hydroxide) that brings out a pink color. The absorbance of the mixture was measured at 535 nm.
Results
In vivo and in vitro gadA transcription
In vitro transcription of gadA has been established and the putative transcription start site was defined (De Biase et al. 1999). It is interesting to learn whether the intracellular environment could affect the transcription start site of the promoter by RNA polymerases under acid stress. The RNA from acid-stressed cells was isolated and the transcription start site was determined, while both of the purified σ70 and σ38 forms of RNA polymerase were used to investigate the in vitro transcription of gadA. The results showed that both forms of RNA polymerase give a common transcription start site in vitro (Fig. 1, lanes 2 and 3), which is the same as the in vivo transcription start site under acid stress (lane 1). This site was identical to the one proposed by De Biase et al. (1999), suggesting that transcription from gadA promoter was not affected by environmental conditions and the form of RNA polymerase. Only half of the − 10 region in gadA promoter shares a common sequence of CTAC as the σ38 –10 region CTACACT consensus (Fig. 1b). On the contrary, the first three nucleotides of the − 35 box has the same sequence as that of the σ70 –35 consensus (Rosenthal et al. 2006), while the rest differed from both the conserved σ70 –10 and the σ38 –10 regions. Thus, the gadA promoter has a unique design that is not similar with the classic σ70- or σ38-dependent promoters, which may confer the gadA promoter novel characteristics in the overexpressions of pathway genes.
a Determination of the in vivo and in vitro gadA transcriptional start site by primer extension. lane 1 in vivo mRNA after pH downshift. lane 2 in vitro mRNA transcribed by σ70 RNA polymerase. lane 3 in vitro mRNA transcribed by σ38 RNA polymerase. Sequencing markers were shown at the top. b The sequence features of the gadA promoter in comparison to the consensuses of the σ70- and σ38-dependent promoters
The effects of gadA promoter in driving pathway gene overexpressions
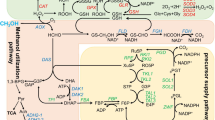
The gadA promoter was derived based on the transcription start site and the promoter region concluded from the above results. The P gadA60 truncated at the non-coding region upstream gadA at the “Gad box” in position − 60 showed the same level of strength as the one covering the entire non-coding region (183 bp) (Fig. S1), and was used in the following study. The effect of the gadA promoter in driving pathway gene overexpressions for the productions of chemicals was examined using the isobutanol biosynthetic pathway. This pathway consists of two operons. One operon includes three genes (alsS, ilvC, and ilvD) that encode the acetolactate synthase, ketol-acid reductoisomerase, and dihydroxy-acid dehydratase. These enzymes together convert pyruvate to α-ketoisovalerate, consisting the upstream of the isobutanol synthetic pathway. The last two steps that convert α-ketoisovalerate to isobutanol were catalyzed by α-ketoisovalerate decarboxylase and alcohol dehydrogenase, which were encoded by kivd and adhA, respectively, in another operon.
The fermentation conditions for isobutanol production using the gadA promoter were investigated. To alleviate the toxicity of the accumulated isobutanol on cells, oleyl alcohol was added to extract the produced isobutanol from the aqueous phase. This strategy increased the titer and yield (Figs. S2 and S3), thus was employed in the isobutanol fed-batch fermentation driven by either the strong inducible promoter PLlacO1 or P gadA60 . For both promoters, the isobutanol concentration increased dramatically during the first 48 h and then slowed down (Fig. 2a, b). The production driven by PLlacO1 produced 5.26 g l−1 isobutanol during the stationary phase from 72 to 144 h and achieved a final titer of 29.64 g l−1 at pH 7.0 (Fig. 2c). The P gadA60 retained higher productivity during the stationary phase, producing 6.60 g l−1 isobutanol from 72 to 144 h. It achieved a final titer of 28.27 g l−1 under neutral condition, which was the same level as PLlacO1. In comparison, acid stress caused dramatic decrease in titer by 43.78–81.24%. The final productions under pH 6.0 reduced to 16.66 and 14.11 g l−1 for PLlacO1 and P gadA60 , respectively. Acidification to pH 5.0 inhibited isobutanol production after 48 h of fermentation for pathway driven by PLlacO1, of which the final titer was only 5.56 g l−1. In contrast, P gadA60 resisted the total inhibition of the isobutanol synthetic pathway, achieving a titer 1.8 times higher (10.03 g l−1) than that of PLlacO1.
Low pH favors gadA over bulk transcription both in vitro and in vivo
The efficient production of isobutanol under the control of PgadA60 suggested that the gadA promoter could resist low pH emerged in the process of fermentation. To learn whether low pH favors the transcription of gadA over the bulk transcription, transcription of ribosomal components from the rrnB promoter was assayed.
Cells were treated with pH 5.5 MES at mid-log phase (Rosenthal et al. 2008; Weber et al. 2005), and the changes in ribosomal transcription were assayed as described (Gralla and Vargas 2006). RNA was isolated both before the acidic treatment and 5 min after. Ribosomal transcription was measured by probing for leader RNA, which has previously been shown to reflect ribosomal transcription levels (Gralla and Vargas 2006). Figure 3a (left) shows that at 5 min, post-stress ribosomal transcription has been very strongly inhibited. By contrast, gadA transcription has modestly increased at this time (Fig. 3a, right). We concluded that mild acid stress rapidly inhibits transcription from the rrnB promoter, which has the consequence of conserving cellular resources. Below, we will attempt to reproduce the differential effects on rrnB and gadA in vitro.
The effect of pH on the expression of σ70-dependent promoters. a In vivo transcription of the gadA promoters in acid-challenged cells. RNA was isolated and detected by primer extension either before or 5 min after downshift to pH 5.5, as indicated. The ribosomal rrnB (left) and gadA (right) c-DNAs are shown. b Multiple round in vitro transcription assays were on supercoiled gadA, rrnB, and lacUV5 templates as a function of pH, adjusted by MES. c Relative transcriptions of gadA (triangles), rrnB (circles), and lacUV5 (squares) with each point averaged from three individual experiments
Previous experiments on gadA used standard in vitro transcription conditions, which involve a pH of 7.9. Because acid stress leads to a lowering of intracellular pH, we explored whether transcription could continue at lower pH. Figure 3b (upper panel) shows σ70 RNA polymerase transcription of gadA as the pH was lowered below 7.9. The results showed that gadA transcription strengthened slightly as the pH was lowered to 6.0 and then crashed at lower pH. We concluded that gadA transcription is resistant as the pH lowers to 6.0 but becomes sensitive as the pH approaches 5.0.
This transcription protocol was repeated using the rrnB promoter, whose transcription was shown to be strongly inhibited upon acid stress in vivo (Fig. 3a, left). The results (Fig. 3b, middle panel) showed that in contrast to gadA transcription, rrnB transcription was strongly inhibited as the pH was lowered to 6.0. This protocol was also used on the common test promoter lacUV5, and this transcription was also very strongly inhibited at pH 6.0 (Fig. 3b, bottom panel). It appeared that gadA transcription is selectively resistant to low pH in vitro.
The results from multiple experiments were quantified in Fig. 3c, which demonstrates the extreme selectivity for gadA transcription in vitro at pH 6.0. If this effect were to apply to the intracellular environment, it would largely account for the in vivo transcription results obtained in Fig. 3a. That is, low pH alone leads to a very strong preference for gadA transcription over ribosomal rrnB transcription, largely by selectively inhibiting ribosomal transcription. Because ribosomal transcription accounts for the bulk of cellular RNA synthesis and because the lacUV5 result suggested mRNA may also be commonly inhibited, there will be a very strong preference for gadA transcription under these low pH conditions.
The potential roles of glutamate and DNA supercoiling on σ38 transcription
The concentration of glutamate can triple under some acid stress conditions (Natera et al. 2006), approaching 200 mM. To assess the effect of glutamate, we assayed σ38-dependent gadA transcription with and without added 200 mM glutamate. Figure 4a shows that gadA transcription became strong in the presence of glutamate. At pH 7.9, transcription was very weak with a low preference for the physiological start site (Fig. 4a, lane 1). But with the addition of 200 mM potassium glutamate, strong and selective transcription can occur (lane 3).
This experiment was repeated at lower pH with somewhat similar results (Fig. 4b). Without added potassium glutamate, transcription showed two weak bands that spaced closely, the bottom of which corresponds to the physiological transcript (Fig. 4b, lane 1). When potassium glutamate was added transcription became much stronger and the bottom band dominated (lane 3). These results make the following points: (1) σ38 RNA polymerase can transcribe gadA strongly and selectively in the presence of a concentration of glutamate that can occur during acid stress. (2) This transcription can occur at lower pH.
Recent information suggested that many of these regulators act to change the levels of GadE, which is thought to be the primary direct macromolecular regulator of gad transcription under most, but perhaps not all conditions (Ma et al. 2003; Sayed et al. 2007). GadE amounts are controlled at both the level of transcription and protein stability (Heuveling et al. 2008). The amount of GadE present is presumably determined by the net effects of the sum of many transcription regulators and proteases.
The σ38-dependent transcription is also affected by levels of DNA supercoiling and in the case of gadA, by the protein GadE. We did not perform systematic combinatorial experiments involving titrations of glutamate, GadE, and DNA supercoiling but did do exploratory experiments to learn the outline of how the system might respond. In initial experiments, we found that 100 mM GadE, which can inhibit σ70 transcription of gadA, had the potential to stimulate transcription when σ38 RNA polymerase was used. In Fig. 4a, lane 2 showed that a weak gadA transcript was observed when GadE was added at pH 7.9. In the absence of GadE, the transcription pattern does not show a convincing selective transcript (lane 1). However, the strength of the transcript with GadE was still much less than that observed when glutamate was present (lane 3). In fact, GadE does not obviously stimulate when glutamate was present (lane 4 vs. lane 3). We infer that at neutral pH, GadE can stimulate gadA transcription but that when glutamate is present, glutamate dominates transcriptional stimulation.
Experiments showed somewhat similar results at lower pH (Fig. 4b). Without glutamate, GadE can stimulate gadA transcription (lane 2 vs. lane 1). With glutamate, selective transcription (lane 3) was stronger and was no longer subject to stimulation by GadE (lane 4 vs. lane 3). Thus, it appeared that GadE has the potential to stimulate gadA transcription at both low and high pH but this potential is not used when glutamate is present at 200 mM.
During stress, the level of DNA supercoiling often changes, typically transiently, and this may also be the case for acid stress (Huo et al. 2008). Figure 5a shows that gadA transcription by σ70 relied completely on DNA supercoils as it was lost when the DNA was linearized. In contrast, σ38 transcription of gadA was not very sensitive to changes in DNA supercoiling (Fig. 5b). That is, there was a slight reduction in signal when supercoiling was completely eliminated by linearizing the DNA. This suggested that σ38 transcription of gadA can occur whatever the state of DNA supercoiling (but the highest level of transcription still depends on glutamate as in Fig. 4). In both cases, GadE stimulation was not detectable on linear DNA (data not shown and see footprinting studies in Fig. S4). We infer that reductions in DNA supercoiling will strongly favor a switch from σ70- to σ38-dependent transcription of gadA when both are present. This switch to σ38 RNA polymerase transcription will also be favored by the accumulation of glutamate, which stimulates σ38-dependent gadA transcription (Fig. 4) but not σ70-dependent gadA transcription (data not shown). The same shift would be expected to occur if GadE accumulates to high levels where it inhibits σ70-dependent gadA transcription (Fig. S5) but stimulates σ38-dependent transcription (Fig. 4).
Effects of loss of supercoils on gadA transcription in vitro using σ70 RNA polymerase (a) and σ38 RNA polymerase (b). The DNA was linearized in the right panels, as indicated. The gradients indicate increasing RNA polymerase concentrations of 50, 100, and 200 nM. Single round transcription was as pH 6.5 with 200 mM potassium glutamate added for the σ38 transcription. Eσ38 σ38-RNA polymerase, Eσ70 σ70-RNA polymerase
Discussion
In order to achieve robust productions for the biosynthesis of fuels and bulk chemicals, this study uncovered the mechanisms that enable stress-selective and growth phase-independent transcription of the gadA promoter. Results showed that low pH favors gadA transcription over that from the rrnB promoter, which dominates cellular transcription under exponential phase. The activation of gadA promoter relies on the accumulation of glutamate and/or the loss of DNA supercoiling. Our study suggested that the gadA promoter is a novel candidate to drive pathway gene overexpressions throughout the exponential to the stressed stationary phase.
Increase the stress tolerance and bypass the growth control poses a big challenge for biorefinery. Strategies focusing on the σ38 factor have been proposed to overcome acid stress. For instance, overexpressing a set of non-coding sRNAs (DsrA, RprA, or ArcA) could disrupt the inhibitory loop of the rpoS mRNA and enhance the acid tolerance for cells in the stationary phase (Gaida et al. 2013). However, this kind of global regulation will work only if the desired genes were regulated by σ38-dependent promoters, mostly those of the stress-resistant genes. For functional modules constructed using σ70-dependent promoters, their expression may still be suppressed even under enhanced acid tolerance. In contrast, the replacement of a highly adaptable promoter with selective σ factor dependence appears to be a more flexible approach to maintain the overexpressions of desired genes under stressed conditions. Thus, gadA promoter serves as a promising alternative to σ38 regulations in dealing with acid and other stresses.
As expected, the gadA promoter was able to drive the overexpressions of multiple pathway genes for the productions of biofuels. It maintained isobutanol production from the exponential phase to the stationary phase (Figs. 2 and S1) and remained active under acidified environment, ensuring a biofuel titer > 10 g l−1 (Fig. 2b, c) without the aid of inducers. This promoter is comparable to the strong inducible promoter PLlacO1 (Lutz and Bujard 1997) and even wins out under acid stress. The efficient production of isobutanol under low pH could reduce the risk of contamination and the cost of pH control. Compared with neutral condition, the decrease in isobutanol production under acid stress could be attributed to lowered enzyme activities (Atsumi et al. 2009; Cahn et al. 2016). For instance, the activity of acetolactate synthase decreased by half from 119.48 nmol α-acetolactate formed per mg protein per min for cells grown at pH 7.0 to 52.14 nmol α-acetolactate formed per mg protein per min for cells grown at pH 5.0.
The direct contributions of low pH and GadE to gadA expression upon initial acid shock
The pH titrations in vitro showed that gadA transcription rose slightly to its maximum as the pH was lowered to 6.0. By contrast, the response of the ribosomal rrnB promoter was opposite as it was virtually silenced at pH 6.0 (Fig. 3c). Thus, switching to acidic pH can induce very strong selectivity for gadA transcription over ribosomal RNA transcription. This switch also occurred rapidly in vivo. Five minutes after lowering the external pH to 5.5, rrnB transcription was shut down and gadA transcription was induced (Fig. 3a). Because external acid conditions lower the internal pH (although not as low as the external pH), the acid-resistant properties of the gadA promoter are likely to be relevant to its regulation. This response can be fast because it does not require productions of new macromolecules. Thus, it can contribute to the immediate induction of protective Gad enzymes when acid stress results in a lowering of intracellular pH. The differential sensitivity to low pH also contributes to the conservation of resources under stress as ribosomal RNA dominates transcription in vivo. The activation of gadA under low pH could facilitate the transcription of pathway genes under acid stress. When cells were collected after 48 h of isobutanol production for the quantification of mRNAs driven by the gadA promoter, the amount of alsS mRNA at pH 5.0 was 8.6-fold higher than that at pH 7.0 (Fig. S6).
The low pH behavior of gadA transcription is unprecedented. Because the experiments were done in vitro, it is clear that this can occur without the involvement of other factors. RNA polymerase and DNA were the only macromolecules presented in the transcription reaction. RNA polymerase appears to undergo transitions that interfere with its ability to form functional complexes in a reaction with a pKa near 6.0 (Lew and Gralla 2004). Both rrnB transcription and that of the common test promoter lacUV5 were shown here to be very strongly inhibited at this pH. Thus, the gadA promoter appears to act as a chaperone in protecting RNA polymerase against low pH inactivation. This feature could be attributed to the unique sequence in its key elements (Fig. 1).
This study also showed that the protein GadE can stimulate σ70-dependent gadA transcription at either low or normal pH (Fig. S5). Upon acid shock, a rapid contribution from GadE might also be expected as acid conditions block GadE turnover (Heuveling et al. 2008) and thus increase its concentration. How GadE works is still unclear. Our transcription data showed that as GadE concentration rises, it switches from activator to inhibitor. This is a classic behavior for dual regulatory proteins such as lambda repressor that occupies a series of sites (Manzo et al. 2012). The footprinting data do show a series of GadE interaction sites along much of the gadA upstream region, the strongest of which is near position − 111 (Fig. S4). It would be interesting to learn if relative site occupancy changes as GadE level change, with the level of glutamate and the inclusion of the σ70 and σ38 forms of RNA polymerase as additional required variables. It is possible that some combinations of these conditions might change the ability of GadE to occupy the various sites and hence change its function, as occurs with many other dual regulators (Collado-Vides et al. 1991; Feng and Cronan 2012).
Moreover, initial experiments (data not shown) showed that removal of DNA upstream from position − 85 led to an increase in transcription, even in the absence of GadE. This raises the possibility that GadE activation might have a component of anti-inhibition; that is, it may overcome inhibition imposed by upstream DNA. The extremely AT-rich nature of upstream DNA could provide non-productive binding sites for RNA polymerase and inhibit proper interactions with the promoter. It also contains binding sites for potential inhibitors. For instance, the sequence between − 89 and − 133 provides a binding site for H-NS (Giangrossi et al. 2005), which could directly silence the gadA promoter (De Biase et al. 1999). Exclusion of this binding site would dramatically reduce the inhibition of H-NS on gadA expression both in vivo and in vitro (Giangrossi et al. 2005). Therefore, GadE binding over the upstream region could suppress these non-productive interactions, thus favoring proper binding and transcription. This explained why the PgadA60 without GadE binding sites showed the same level of activity as that of the 183-bp gadA promoter (Fig. S1).
gadA expression via σ38
After an initial acid stress or if other stresses have been applied (for example stationary phase), σ38 begins to accumulate. Many factors can contribute to favoring σ38 transcription over σ70 transcription (Battesti et al. 2011). The current experiments showed that the conditions that apply after the initial adaptive response should strongly favor gadA transcription by the σ38 form of RNA polymerase. That is, high concentrations of GadE and glutamate, both of which can accumulate during acid shock (Natera et al. 2006; Sayed et al. 2007) and selectively activate gadA transcription by σ38 RNA polymerase. These effects would ensure that gadA transcription would continue as the cellular transcription activity switched to reliance on σ38.
Stress in general is accompanied by increase in the amount of σ38 and changes in DNA supercoiling. DNA supercoiling regulated by proteins such as H-NS can respond to various environmental signals (Hulton et al. 1990). It decreases during stationary phase and nutritional downshift (Dorman 2013; Gutiérrez-Estrada et al. 2014), transiently increases upon osmotic shock, and transiently decreases upon acid shock (Cameron et al. 2011; Reyes-Domínguez et al. 2003; Rui and Tse-Dinh 2003). Although DNA topology affects prokaryotic transcription (Hulton et al. 1990), our data showed that σ38 RNA polymerase transcription of gadA is not very sensitive to changes in DNA supercoiling, in contrast to the behavior of σ70 RNA polymerase. This would seem to ensure that gadA promoter could remain active under the widest possible range of physiological conditions.
In vivo studies have attempted to distinguish which RNA polymerase is used under which physiological condition, with some success when specific conditions were tested (Ma et al. 2003). It is known that the mix of σ70 and σ38 depends on the degree of stress, the type of stress (Typas et al. 2007), and the type of media (Tao et al. 1999). Thus, one need not expect a clean partitioning of transcription between the two RNA polymerases. Instead, one expects that both can transcribe gadA in vivo, with the relative contribution of each set by their relative amounts and are influenced by the accumulation of glutamate, changes in DNA supercoiling, and the amount of GadE and other effectors such as GadX. The net effect is that the design of gadA promoter allows the cell to mount an effective response to acid stress under an exceptionally wide range of conditions. This explains the phenomena of continuous isobutanol production throughout the whole growth stage when driven by gadA promoter.
Growth phase-independent and environmental shock-resistant gene overexpressions
This study approached the mechanism behind the growth phase-independent and environmental shock-resistant gadA overexpression by reconstituting gadA transcription in vitro and studying how it may be induced. Two general conclusions are most prominent. First, and most surprising, is the observation that the gadA promoter is selectively resistant to inactivation at low pH in vitro. That is, when the pH is lowered from near 8.0 to near 6.0, gadA transcription continues unabated whereas ribosomal RNA production (which accounts for the bulk of cellular RNA) is shut down. Thus, low pH can directly contribute to selective gadA expression without the participation of other factors. Second, the σ38 form of RNA polymerase that becomes prominent during most stresses is shown to direct gadA transcription in vitro and to be favored by a variety of effectors. That is, high concentrations of the activator GadE or the physiological effector glutamate or the loss of DNA supercoils all favor σ38 over σ70 transcription of gadA. Taken together, these results implied that the gadA promoter has an unusual design that allows it to be selectively transcribed at low pH under the widest range of intracellular conditions (Table 1), including stresses imposed by various external agents.
In all, the application of gadA promoter could be a powerful tool to bridge the gap between practical and ideal productivities in the productions of value-added chemicals (Fig. 6). The adaptability of gadA promoter makes it a novel candidate to drive pathway gene overexpressions for biofuel productions in not only exponential phase but also in stationary phase or stressed conditions.
References
Atsumi S, Cann AF, Connor MR, Shen CR, Smith KM, Brynildsen MP, Chou KJY, Hanai T, Liao JC (2008a) Metabolic engineering of Escherichia coli for 1-butanol production. Metab Eng 10(6):305–311. https://doi.org/10.1016/j.ymben.2007.08.003
Atsumi S, Hanai T, Liao JC (2008b) Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 451(7174):86–89. https://doi.org/10.1038/nature06450
Atsumi S, Li Z, Liao JC (2009) Acetolactate synthase from Bacillus subtilis serves as a 2-ketoisovalerate decarboxylase for isobutanol biosynthesis in Escherichia coli. Appl Environ Microbiol 75(19):6306–6311. https://doi.org/10.1128/AEM.01160-09
Atsumi S, Wu T-Y, Eckl E-M, Hawkins SD, Buelter T, Liao JC (2010) Engineering the isobutanol biosynthetic pathway in Escherichia coli by comparison of three aldehyde reductase/alcohol dehydrogenase genes. Appl Microbiol Biotechnol 85(3):651–657. https://doi.org/10.1007/s00253-009-2085-6
Battesti A, Majdalani N, Gottesman S (2011) The RpoS-mediated general stress response in Escherichia coli. Annu Rev Microbiol 65(1):189–213. https://doi.org/10.1146/annurev-micro-090110-102946
Benjaphokee S, Hasegawa D, Yokota D, Asvarak T, Auesukaree C, Sugiyama M, Kaneko Y, Boonchird C, Harashima S (2012) Highly efficient bioethanol production by a Saccharomyces cerevisiae strain with multiple stress tolerance to high temperature, acid and ethanol. New Biotechnol 29(3):379–386. https://doi.org/10.1016/j.nbt.2011.07.002
Cahn JKB, Baumschlager A, Brinkmann-Chen S, Arnold FH (2016) Mutations in adenine-binding pockets enhance catalytic properties of NAD(P)H-dependent enzymes. Protein Eng Des Sel 29:31–38
Cameron ADS, Stoebel DM, Dorman CJ (2011) DNA supercoiling is differentially regulated by environmental factors and FIS in Escherichia coli and Salmonella enterica. Mol Microbiol 80(1):85–101. https://doi.org/10.1111/j.1365-2958.2011.07560.x
Castanié-Cornet M-P, Cam K, Bastiat B, Cros A, Bordes P, Gutierrez C (2010) Acid stress response in Escherichia coli: mechanism of regulation of gadA transcription by RcsB and GadE. Nucleic Acids Res 38(11):3546–3554. https://doi.org/10.1093/nar/gkq097
Collado-Vides J, Magasanik B, Gralla JD (1991) Control site location and transcriptional regulation in Escherichia coli. Microbiol Rev 55(3):371–394
Connor MR, Cann AF, Liao JC (2010) 3-Methyl-1-butanol production in Escherichia coli: random mutagenesis and two-phase fermentation. Appl Microbiol Biotechnol 86(4):1155–1164. https://doi.org/10.1007/s00253-009-2401-1
De Biase D, Tramonti A, Bossa F, Visca P (1999) The response to stationary-phase stress conditions in Escherichia coli: role and regulation of the glutamic acid decarboxylase system. Mol Microbiol 32(6):1198–1211. https://doi.org/10.1046/j.1365-2958.1999.01430.x
Dorman CJ (2013) Genome architecture and global gene regulation in bacteria: making progress towards a unified model? Nat Rev Microbiol 11(5):349–355. https://doi.org/10.1038/nrmicro3007
Feehily C, Karatzas K (2013) Role of glutamate metabolism in bacterial responses towards acid and other stresses. J Appl Microbiol 114(1):11–24. https://doi.org/10.1111/j.1365-2672.2012.05434.x
Feng Y, Cronan JE (2012) Crosstalk of Escherichia coli FadR with global regulators in expression of fatty acid transport genes. PLoS One 7(9):e46275. https://doi.org/10.1371/journal.pone.0046275
Foster JW (2004) Escherichia coli acid resistance: tales of an amateur acidophile. Nat Rev Microbiol 2(11):898–907. https://doi.org/10.1038/nrmicro1021
Gaida SM, Al-Hinai MA, Indurthi DC, Nicolaou SA, Papoutsakis ET (2013) Synthetic tolerance: three noncoding small RNAs, DsrA, ArcZ and RprA, acting supra-additively against acid stress. Nucleic Acids Res 41(18):8726–8737. https://doi.org/10.1093/nar/gkt651
Giangrossi M, Zattoni S, Tramonti A, De Biase D, Falconi M (2005) Antagonistic role of H-NS and GadX in the regulation of the glutamate decarboxylase-dependent acid resistance system in Escherichia coli. J Biol Chem 280(22):21498–21505. https://doi.org/10.1074/jbc.M413255200
Goo E, Majerczyk CD, An JH, Chandler JR, Seo Y-S, Ham H, Lim JY, Kim H, Lee B, Jang MS, Greenberg EP, Hwang I (2012) Bacterial quorum sensing, cooperativity, and anticipation of stationary-phase stress. Proc Natl Acad Sci USA 109(48):19775–19780. https://doi.org/10.1073/pnas.1218092109
Gralla JD (2005) Escherichia coli ribosomal RNA transcription: regulatory roles for ppGpp, NTPs, architectural proteins and a polymerase-binding protein. Mol Microbiol 55(4):973–977. https://doi.org/10.1111/j.1365-2958.2004.04455.x
Gralla JD, Vargas DR (2006) Potassium glutamate as a transcriptional inhibitor during bacterial osmoregulation. EMBO J 25(7):1515–1521. https://doi.org/10.1038/sj.emboj.7601041
Gutiérrez-Estrada A, Ramírez-Santos J, Gómez-Eichelmann MC (2014) Role of chaperones and ATP synthase in DNA gyrase reactivation in Escherichia coli stationary-phase cells after nutrient addition. SpringerPlus 3(1):656. https://doi.org/10.1186/2193-1801-3-656
Heuveling J, Possling A, Hengge R (2008) A role for Lon protease in the control of the acid resistance genes of Escherichia coli. Mol Microbiol 69(2):534–547. https://doi.org/10.1111/j.1365-2958.2008.06306.x
Hulton CSJ, Seirafi A, Hinton JCD, Sidebotham JM, Waddell L, Pavitt GD, Owen-Hughes T, Spassky A, Buc H, Higgins CF (1990) Histone-like protein H1 (H-NS), DNA supercoiling, and gene expression in bacteria. Cell 63(3):631–642. https://doi.org/10.1016/0092-8674(90)90458-Q
Huo Y-X, Cho KM, Rivera JGL, Monte E, Shen CR, Yan Y, Liao JC (2011) Conversion of proteins into biofuels by engineering nitrogen flux. Nat Biotech 29(4):346–351. https://doi.org/10.1038/nbt.1789
Huo YX, Rosenthal AZ, Gralla JD (2008) General stress response signalling: unwrapping transcription complexes by DNA relaxation via the sigma38 C-terminal domain. Mol Microbiol 70(2):369–378. https://doi.org/10.1111/j.1365-2958.2008.06412.x
Krulwich TA, Sachs G, Padan E (2011) Molecular aspects of bacterial pH sensing and homeostasis. Nat Rev Microbiol 9(5):330–343. https://doi.org/10.1038/nrmicro2549
Lew CM, Gralla JD (2004) Nucleotide-dependent isomerization of Escherichia coli RNA polymerase. Biochemistry 43(39):12660–12666. https://doi.org/10.1021/bi0492814
Leyval D, Uy D, Delaunay S, Goergen J, Engasser J (2003) Characterisation of the enzyme activities involved in the valine biosynthetic pathway in a valine-producing strain of Corynebacterium glutamicum. J Biotechnol 104(1-3):241–252. https://doi.org/10.1016/S0168-1656(03)00162-7
Lutz R, Bujard H (1997) Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements. Nucleic Acids Res 25(6):1203–1210. https://doi.org/10.1093/nar/25.6.1203
Ma Z, Gong S, Richard H, Tucker DL, Conway T, Foster JW (2003) GadE (YhiE) activates glutamate decarboxylase-dependent acid resistance in Escherichia coli K-12. Mol Microbiol 49(5):1309–1320. https://doi.org/10.1046/j.1365-2958.2003.03633.x
Manzo C, Zurla C, Dunlap David D, Finzi L (2012) The effect of nonspecific binding of lambda repressor on DNA looping dynamics. Biophys J 103(8):1753–1761. https://doi.org/10.1016/j.bpj.2012.09.006
Métris A, George SM, Ropers D (2017) Piecewise linear approximations to model the dynamics of adaptation to osmotic stress by food-borne pathogens. Int J Food Microbiol 240:63–74. https://doi.org/10.1016/j.ijfoodmicro.2016.06.022
Natera V, Sobrevals L, Fabra A, Castro S (2006) Glutamate is involved in acid stress response in Bradyrhizobium sp. SEMIA 6144 (Arachis hypogaea L.) microsymbiont. Curr Microbiol 53(6):479–482. https://doi.org/10.1007/s00284-006-0146-y
Nicolaou SA, Gaida SM, Papoutsakis ET (2010) A comparative view of metabolite and substrate stress and tolerance in microbial bioprocessing: from biofuels and chemicals, to biocatalysis and bioremediation. Metab Eng 12(4):307–331. https://doi.org/10.1016/j.ymben.2010.03.004
Reyes-Domínguez Y, Contreras-Ferrat G, Ramírez-Santos J, Membrillo-Hernández J, Gómez-Eichelmann MC (2003) Plasmid DNA supercoiling and gyrase activity in Escherichia coli wild-type and rpoS stationary-phase cells. J Bacteriol 185(3):1097–1100. https://doi.org/10.1128/JB.185.3.1097-1100.2003
Rosenow C, Saxena RM, Durst M, Gingeras TR (2001) Prokaryotic RNA preparation methods useful for high density array analysis: comparison of two approaches. Nucleic Acids Res 29(22):e112–e112. https://doi.org/10.1093/nar/29.22.e112
Rosenthal AZ, Hu M, Gralla JD (2006) Osmolyte-induced transcription: − 35 region elements and recognition by sigma38 (rpoS). Mol Microbiol 59(3):1052–1061. https://doi.org/10.1111/j.1365-2958.2005.04999.x
Rosenthal AZ, Kim Y, Gralla JD (2008) Poising of Escherichia coli RNA polymerase and its release from the σ38 C-terminal tail for osmY transcription. J Mol Biol 376(4):938–949. https://doi.org/10.1016/j.jmb.2007.12.037
Rui S, Tse-Dinh YC (2003) Topoisomerase function during bacterial responses to environmental challenge. Front Biosci 8:d256–d263
Sayed AK, Odom C, Foster JW (2007) The Escherichia coli AraC-family regulators GadX and GadW activate gadE, the central activator of glutamate-dependent acid resistance. Microbiology 153(8):2584–2592. https://doi.org/10.1099/mic.0.2007/007005-0
Sharma UK (2016) Role of changes in σ70-driven transcription in adaptation of E. coli to conditions of stress or starvation. In: Bruijn FJ (ed) Stress and environmental regulation of gene expression and adaptation in bacteria. John Wiley & Sons, Inc., New Jersey, pp 37–47. https://doi.org/10.1002/9781119004813.ch5
Tao H, Bausch C, Richmond C, Blattner FR, Conway T (1999) Functional genomics: expression analysis of Escherichia coli growing on minimal and rich media. J Bacteriol 181(20):6425–6440
Tramonti A, Visca P, De Canio M, Falconi M, De Biase D (2002) Functional characterization and regulation of gadX, a gene encoding an AraC/XylS-like transcriptional activator of the Escherichia coli glutamic acid decarboxylase system. J Bacteriol 184(10):2603–2613. https://doi.org/10.1128/JB.184.10.2603-2613.2002
Tupper AE, Owen-Hughes TA, Ussery D, Santos D, Ferguson D, Sidebotham J, Hinton J, Higgins C (1994) The chromatin-associated protein H-NS alters DNA topology in vitro. EMBO J 13:258
Typas A, Becker G, Hengge R (2007) The molecular basis of selective promoter activation by the σS subunit of RNA polymerase. Mol Microbiol 63(5):1296–1306. https://doi.org/10.1111/j.1365-2958.2007.05601.x
Warnecke T, Gill RT (2005) Organic acid toxicity, tolerance, and production in Escherichia coli biorefining applications. Microb Cell Factories 4(1):25. https://doi.org/10.1186/1475-2859-4-25
Weber H, Polen T, Heuveling J, Wendisch VF, Hengge R (2005) Genome-wide analysis of the general stress response network in Escherichia coli: σS-dependent genes, promoters, and sigma factor selectivity. J Bacteriol 187(5):1591–1603. https://doi.org/10.1128/JB.187.5.1591-1603.2005
Zhao B, Houry WA (2010) Acid stress response in enteropathogenic gammaproteobacteria: an aptitude for survival. Biochem Cell Biol 88(2):301–314. https://doi.org/10.1139/O09-182
Zhou K, Zhou L, Lim QE, Zou R, Stephanopoulos G, Too H-P (2011) Novel reference genes for quantifying transcriptional responses of Escherichia coli to protein overexpression by quantitative PCR. BMC Mol Biol 12(1):18. https://doi.org/10.1186/1471-2199-12-18
Acknowledgements
We would like to show our gratitude to Prof. Jay Gralla and Dr. Adam Z. Rosenthal for the initial supports and experiments regarding this project. We also thank Ms. Jiao Zhang and Mr. Kenao Lv for the initial fermentation experiments. The work finished in Beijing Institute of Technology was supported by the National Natural Science Foundation of China (grant No. 21676026) and the National Key R↦D Program of China (grant No. 2017YFD0201400). The work finished in University of California, Los Angeles (UCLA) was supported by NIH grant GM35754.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical statement
This article does not contain any studies with human or animal subjects.
Electronic supplementary material
ESM 1
(PDF 2658 kb).
Rights and permissions
About this article
Cite this article
Huo, YX., Guo, L. & Ma, X. Biofuel production with a stress-resistant and growth phase-independent promoter: mechanism revealed by in vitro transcription assays. Appl Microbiol Biotechnol 102, 2929–2940 (2018). https://doi.org/10.1007/s00253-018-8809-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-8809-8