Abstract
Clethramycin (1) and mediomycin A (2) belong to the linear polyene polyketide (LPP) family of antibiotics that exhibit potent antifungal activity. Structural similarities exist between 1 and 2, except that 2 contains an amino moiety substituted for the guanidino moiety. Herein, the draft genome sequence of Streptomyces mediocidicus ATCC23936, a strain which produces both 1 and 2, was obtained through de novo sequencing. Bioinformatic analysis of the genome revealed a clethramycin (cle) gene cluster that contained 25 open reading frames (orfs). However, amidinohydrolase for 2 formation was not found in the cle gene cluster. Further genomic analysis revealed an amidinohydrolase MedX, which can hydrolyse the guanidino form (1) into the amino form (2) via heterologous co-expression of the cle cluster in Streptomyces lividans or by in vitro catalysis. These results also suggest the feasibility of engineering novel LPPs for drug discovery by manipulating the biosynthetic machinery of S. mediocidicus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
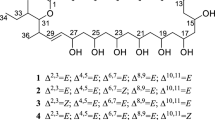
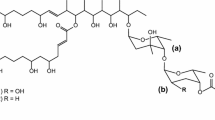
Linear polyene polyketides (LPPs), one of the important members of the polyene family, have received considerable attention because of their potent activity against various bacterial pathogens (Banskota et al. 2006; Cai et al. 2007; Mcalpine et al. 2005). Clethramycin (1) and mediomycin A (2), typical LPPs with strong antifungal activity, were first isolated from cultures of Streptomyces mediocidicus ATCC23936 or Streptomyces hygroscopicus TP-A0623 (Cai et al. 2007; Furumai et al. 2003). Apart from the guanidino moiety, 1 shares overall structural identity with 2, which represents a class of LPPs with an amino moiety (Fig. 1). Although the head of LPPs is substituted with an amino or a guanidino group, increasing evidence has shown that 4-guanidinobutanoyl-CoA derived from l-arginine provides the skeleton for all LPP biosynthesis (Mcalpine et al. 2005; Zhang et al. 2017), followed by modular type I polyketide synthase (PKS) (PKS-I) to form the polyene polyketide structural framework (Chen et al., 2015). After forming the framework, amino-containing LPPs may be derived from their guanidino-substituted counterparts at a later step in their biosynthesis through catalysis by an amidinohydrolase. The biosynthetic gene cluster of another LPP, ECO-02301 (Figs. 1 and 3), contains a potential amidinohydrolase gene (Mcalpine et al., 2005). Recently, a gene cluster of 1 was identified in the draft genome sequence of Streptomyces blastmyceticus NBRC12747, which has been validated by heterologous expression in Streptomyces lividans TK23 (Zhang et al. 2017). However, the hypothetical amidinohydrolase was not found in the clethramycin biosynthetic gene cluster (cle cluster). Therefore, a putative amidinohydrolase gene may exist outside the cle cluster.
The aim of this study is to discover the hypothetical amidinohydrolase from the genome of S. mediocidicus ATCC23936 and further investigate its function in vivo or in vitro. Through genomic sequencing and bioinformatic analysis, a hypothetical amidinohydrolase (MedX) was identified in the genome of S. mediocidicus ATCC23936. MedX converts the guanidino form 1 to the amino form 2 by heterologous co-expression of the cle cluster in S. lividans or by in vitro catalysis. Enzyme kinetics further suggests that MedX is the amidinohydrolase involved in the biosynthesis of 1.
Materials and methods
Strains, plasmids, reagents, media and culture conditions
Bacterial strains and plasmids used in the present study are listed in Table S1. Taq DNA polymerase was purchased from TaKaRa Biotechnology Co., Ltd. (Dalian, China). Restriction and other enzymes were purchased from Fermentas International INC (USA). Chemicals, biochemicals and media were purchased from Sinopharm Chemical Reagent Co., Ltd. (Shanghai, China) and Oxoid Ltd. (Basingstoke, UK) unless otherwise stated.
Genome sequencing
The complete genome sequence of S. mediocidicus ATCC23936 was determined using the Illumina HiSeq 2500 platform with paired-end sequencing and was performed at the Shanghai Biotechnology Corporation, ShanghaiBio (Shanghai, China). The genome contained 7,902,931 bp with an average G + C content of 71.50%. The gene function of the genome sequence was predicted using Prokka software, according to the Swiss-Prot database.
Phylogenetic analysis
The amidinohydrolase family of proteins with closely related sequences to the putative MedX was retrieved from BLASTP. Phylogenetic and molecular evolutionary analyses were conducted using MEGA 7 (Tamura et al. 2007). Briefly, for the construction of bootstrap tests of phylogenetic trees, all amino acid sequences were aligned using the Clustal W algorithm in MEGA 7. The resulting alignment (with gaps) was used to infer a neighbour-joining tree and bootstrap values (Saitou and Nei 1987). The Poisson correction model was utilised as a substitution model for construction of the tree with pairwise deletion (Nei and Kumar 2000).
Construction of genomic libraries
Construction of bacterial artificial chromosome (BAC) libraries of the S. mediocidicus ATCC23936 genome was performed using pIndigoBAC-5 vector according to the manufacturer’s protocols (Epicentre Technologies). Clones carrying the entire cle gene cluster were screened by PCR using two pairs of primers (P1/P2 and P3/P4) listed in Table S2, corresponding to the upstream and downstream regions of the gene cluster.
Accession numbers
The sequence detailed in this paper has been deposited in GenBank under the accession numbers MF139773 and MF139772.
Construction of pIndigoBAC-cle-L1/2 plasmids
The PermE-attP-oriT-aac(3)IV cassette was amplified from pSET153E by PCR using the primers P5 and P6 (Table S2). Then, pIndigoBAC-cle was introduced into Escherichia coli K-12/pDK46 (λ RED recombination plasmid) for generation of E. coli K-12/pDK46/pIndigoBAC-cle. Following induction by l-arabinose, competent cells were prepared for electroporation, and the linear gene cassette L1, up- or downstream of which are the 39-nucleotide (nt) homology arms of the chloramphenicol resistance gene chl in the pIndigoBAC-5 vector, was introduced into E. coli K-12/pDK46/ pIndigoBAC-cle by electroporation in sterile cuvettes, spread onto LB agar containing apramycin (50 μg/ml) and incubated at 37 °C overnight. The integrating vector pIndigoBAC-cle-L1 with apramycin resistance was identified by PCR to amplify the apramycin gene using the primers apr-F/apr-R. To construct the pIndigoBAC-cle-L2 vector, the medX gene was amplified by PCR using primers medX-EF/medX-ER (Table S2). Then, the recovered fragments were cloned to the NdeI/EcoRV site of pSET153E. After sequencing, the permE-medX-attP-oriT-aac(3)IV cassette was amplified from pSET153E/medX by PCR using the primers P1 and P2. The linear permE-medX-attP-oriT-aac(3)IV cassette PCR fragment was introduced into E. coli K-12/pDK46/pIndigoBAC-cle for integration into pIndigoBAC-cle DNA by homologous recombination, generating pIndigoBAC-cle-L2, which was further identified by PCR to amplify the apramycin and medX genes.
E. coli–Streptomyces conjugation
The recombinant plasmids mentioned above were introduced into S. mediocidicus ATCC23936 by E. coli–Streptomyces conjugation, as described previously (Liu et al. 2013a, b). The genotypes of these strains were further confirmed by PCR amplification using primers apr-F/R (Table S2).
Fermentation
The S. mediocidicus ATCC23936 wild-type strain and S. lividans L1/L2 were grown on ISP1 agar plates for 3 days at 30 °C. Agar pieces approximately 1 cm2 containing the wild-type or recombinant strains were then used to inoculate 250-ml flasks containing 50 ml of the fermentation medium (soluble starch, 1.5%; soybean meal, 2.0%; yeast extract, 0.5%; NaCl, 0.5%; KNO3, 0.05% and CaCO3, 0.5%) and incubated at 30 °C and 250 rpm for 96 h. Ten-millilitre samples of each culture were measured into 50-ml centrifuge tubes and extracted for 12 h with an equal volume of methanol. The extracts were then centrifuged at 12,000×g for 15 min, and the supernatant was collected.
Ultra-performance liquid chromatography–tandem quadrupole and time-of-flight high-resolution mass spectrometry analysis
Ultra-performance liquid chromatography-tandem quadrupole and
time-of-flight high-resolution mass spectrometry (UPLC-Q-TOF-HRMS) was performed on a Waters Micromass Q-TOF Premier Mass Spectrometer (Waters, Milford, Massachusetts, USA). The UV spectra were obtained using a Shimadzu Biospec-1601 spectrometer (Shimadzu, Kyoto, Japan).
Expression and purification of MedX
Plasmid pET28a was selected as an expression vector for MedX, and a His6-tagged sequence was fused to the N terminus of MedX. The medX DNA fragment was amplified from the genome of S. mediocidicus ATCC23936 using the primers medX-EF2/medX-ER (Table S2), and the recovered fragments were digested and cloned into the NdeI/HindIII site of pET28a, generating pET28a-medX. The pET28a-medX plasmid was transformed into E. coli BL21 (DE3) for protein expression at 18 °C, 220 rpm and 0.5 mM IPTG for 12 h. Cells were harvested, washed with 50 mM potassium phosphate buffer (pH 7.5) and disrupted using high-pressure cell-disruption system. The lysate was centrifuged for 30 min at 10,000×g at 4 °C and loaded onto a Ni-NTA column (2.0 cm × 20 cm, Novagen, USA) for affinity chromatography. The column was washed with a 20-ml linear gradient of imidazole (0–0.5 M) in 50 mM potassium phosphate buffer (pH 7.5). The eluate was subjected to 10% SDS-PAGE, and the eluate containing the MedX protein was concentrated and desalted using a centrifugal filtration device (Millipore Corp., USA).
Kinetic studies for MedX
The reaction mixtures (1 ml) contained 400 μg MedX, 1 mM CoCl2 and various concentrations of 1 (90–360 μM) in 50 mM Tris-HCl buffer (pH 9.0). After incubation at 220 rpm and 30 °C for 30 min, all reaction mixtures were terminated by addition of an equal volume of methanol. After removing the precipitate by centrifugation, the supernatant was subjected to HPLC (Hong et al. 2016).
Results
Genomic-driven discovery of the cle biosynthetic gene cluster
To probe the novel amidinohydrolase involved in 2 biosynthesis, S. mediocidicus ATCC23936, a strain known to produce 1 and 2, was selected for genome sequencing. The genome of S. mediocidicus ATCC23936 comprises a 7,902,931-bp circular chromosome (Fig. 2); the cle gene cluster was identified by analysis of this genome and was found to contain 25 orfs (Fig. 3 and Table S3). Bioinformatic analysis showed that the cle gene cluster in S. mediocidicus ATCC23936 is highly similar to previously reported gene cluster in S. blastmyceticus NBRC12747 (Zhang et al. 2017). Isotope feeding experiments have demonstrated that the guanidino-containing skeleton of LPPs originates from 4-aminobutyric acid (Igarashi et al., 2003). During biosynthesis of 1, 4-aminobutyric acid may be converted from l-arginine by CleD, a putative l-arginine mono-oxygenase. Thus, CleO and CleE, a putative 4-guanidinobutyramide hydrolase and 4-guanidinylbutanoate acyl-CoA ligase, respectively, may be responsible for γ-guanidinebutyryl CoA formation. CleG, an acyl carrier protein (ACP) acyltransferase, was proposed to be playing a role in loading the activated γ-aminobutyryl group onto the ACP domain before the initiation of polyene polyketide structural framework formation by PKS-I modules (Cle 1-9). After polyene polyketide structural framework formation, cleB, which encodes a protein similar to sulphotransferase, was proposed to be playing a role in sulphate group formation (Fig. 3). However, a putative amidinohydrolase outside the cle cluster may catalyse the hydrolysis of guanidino form 1 into the amino form 2 during post-PKS modifications.
Circular genome map of S. mediocidicus ATCC23936. The scales at the outermost layer indicate the location in million base pairs; each scale is 0.1 Mb. Circle (2) and circle (3): CDSs of the forward strand and backward strand, respectively; differently coloured CDSs indicate the different COG functional categories; circle (4): rRNAs and tRNAs; circle (5): GC content; circle (6): GC skew (G − C/G + C)
Organisation of CLE biosynthetic genes and biosynthetic hypothesis for CLE (deduced functions of which are summarised in Table S1)
Genomic mining of a putative amidinohydrolase outside the cle cluster
To confirm the existence of a putative amidinohydrolase in the genome sequence of S. mediocidicus ATCC23936, genome scanning was used in the present study. A predicted orf (medX) that is not co-located with the cle cluster region was selected for further analysis. Sequence alignments indicated that medX encodes a protein similar to DstH (31% identity), a known amidinohydrolase in the biosynthesis of desertomycin A (Hong et al. 2016). Sequence alignments with four known amidinohydrolases of various compounds revealed that MedX contains an agmatinase proclavaminic acid amidinohydrolase (PAH) domain that was associated with the putative active site and Mn2+-ion binding site of proclavaminic acid amidinohydrolase (PAH, Fig. 4a). MedX contains the conserved sequence motifs xGGDH, DAHxD and SxDIDVxDPxxAPGTGTP (where x = any amino acid) (Fig. 4b), which are implicated in the binding and catalysis of this enzyme superfamily (Dowling et al. 2008; Elkins et al. 2002; Lee et al. 2011). The preferred method for ascertaining the role of MedX in situ requires inactivation of the medX gene; however, in spite of extensive efforts, S. mediocidicus ATCC23936 was resistant to the available genetic methods for in situ inactivation (data not shown).
Domain structure and amino acid alignments of MedX. a Predicted domain structure of MedX. Agmatinase PAH domain contains putative active site and Mn2+ binding site. b Alignments of the PAH domain of MedX with related proteins. Numbers indicate amino acid residues from the N terminus of the protein. ECO-02301-ORF32 putative amidinohydrolase from S. aizunensis NRRL B-11277, NmdS putative amidinohydrolase from Streptomyces sp. RK95-74, PAH proclavaminate amidinohydrolase from S. clavuligerus, DstH amidinohydrolase from S. olivaceus Tü4018. Identical amino acid residues are highlighted in black
Heterologous expression of the cle cluster in S. lividans TK24
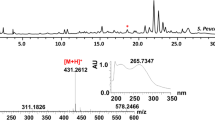
Heterologous co-expression of the cle cluster and medX in S. lividans TK24 is an alternative option for studying the function of MedX. To obtain the whole gene cluster of 1, the BAC DNA library was constructed using the pIndigoBAC-5 vector, according to the manufacturer’s instructions, and the vector pIndigoBAC-cle containing the 180-kb DNA fragment encompassing the entire cle gene cluster was selected for further investigation. pIndigoBAC-cle plasmid does not contain PhiC31 integrase system for integration of exogenous gene to S. mediocidicus ATCC23936 chromosome through recombination between the attachment sites of phage and bacteria genomes, known as attB and attP sites, respectively. Moreover, the chloramphenicol resistance of pIndigoBAC plasmid is not sensitive for Streptomyces. Therefore, it is essential to modify pIndigoBAC-cle plasmid for stable and integrated expression of cle gene cluster in S. lividans TK24. Then, the ermP-attP-aac(3)IV-oriT cassette L1, up- or downstream of which are the 39-nt homology arms of the chloramphenicol resistance gene in the pIndigoBAC-5 vector, was amplified by PCR using plasmid pSET153E as a template. Introduction of the linear gene cassette L1 into E. coli K-12/pDK46/pIndigoBAC-cle was performed through electroporation in sterile cuvettes (Gust et al. 2003). Following induction by l-arabinose, the gene cassette L1 was inserted into the vector pIndigoBAC-cle by replacing the chloramphenicol resistance gene chl and generating an integrating vector pIndigoBAC-cle-L1 with apramycin resistance (Fig. S1). To identify the product of the cle cluster, the vector pIndigoBAC-cle-L1 was introduced to S. lividans TK24 to produce S. lividans L1 (Fig. 5a). The resulting S. lividans L1, upon UPLC-Q-TOF-HRMS analysis of the fermentation extracts (Fig. 5b), was unable to produce 2, and 1 was generated instead, similar to that observed in the wild-type strain S. mediocidicus ATCC23936. Compound 1 from S. lividans L1 was subsequently identified by high-resolution electrospray ionisation mass spectrometry (HRESI-MS) at m/z 1216.6266 [M-H]− (Fig. S2) and was found to have a molecular formula of C63H99N3O18S (m/z calculated 1216.6566). HRESI-MS analysis of the culture extract of S. mediocidicus ATCC23936 revealed a product peak corresponding to 1 (Fig. 5b, Fig. S3) with a m/z 1216.6465 [M-H]−. These results demonstrated that heterologous expression of the cle cluster in S. lividans TK24 can generate 1 rather than 2, suggesting that the putative amidinohydrolase was outside the cle cluster.
Production of 2 by heterologous expression of the CLE gene cluster and a putative amidinohydrolase MedX. a S. mediocidicus ATCC23936 is the wild-type strain of 1 and 2 production; S.lividans L1 contains the whole gene cluster of 1 and a gene cassette containing ermP promoter, attP, aac(3)IV and oriT, which is integrated into the attB site of the S. lividans TK24 chromosome by PhiC31-directed site-specific recombination; S. lividans L2 contains the whole gene cluster of 1 and a gene cassette containing ermP promoter, medX gene, attP, aac(3)IV and oriT, which is integrated into the attB site of the S. lividans TK24 chromosome by PhiC31-directed site-specific recombination. b HPLC analyses of fermentation products from S. mediocidicus ATCC23936, S.lividans L1 and S. lividans L2
Function determination of MedX by heterologous expression
To study the possible role of MedX in the biosynthesis of 2, the medX gene from S. mediocidicus ATCC23936 was cloned and inserted into pSET153E under the control of the ermP promoter. The gene cassette containing the ermP promoter, medX gene, attP, aac(3)IV and oriT was also inserted into the vector pIndigoBAC-cle by replacing the chloramphenicol resistance gene chl to generate an integrating vector pIndigoBAC-cle-L2 with apramycin resistance (Fig. 5a, Fig. S4). To confirm the role of MedX, vector pIndigoBAC-cle-L2 was introduced into S. lividans TK24 through E. coli–Streptomyces conjugation. The resulting recombinant strain S. lividans L2 was grown in liquid culture and analysed for the production of 2 by UPLC-Q-TOF-HRMS. S. lividans L2, upon UPLC-Q-TOF-HRMS analysis of the fermentation extracts (Fig. 5b), produced a compound with a m/z 1174.6991 [M-H]− (Fig. S5), corresponding to a molecular formula of C62H90NO18S (m/z calculated 1174.6348), which is similar to 2 (Fig. 5b). A similar compound was subsequently identified in the culture extract of S. mediocidicus ATCC23936 with a m/z 1174.7239 [M-H]− on HRESI-MS analysis (Fig. 5b and Fig. S3).
Confirmation of the catalytic activity and characteristics in vitro
To confirm the catalytic activity of MedX in vitro, MedX was expressed in E. coli BL21 (DE3), and the His6-tagged MedX was purified to homogeneity by Ni-Sepharose, concentrated and desalted using a centrifugal filter device. Compounds 1 and 2 were purified from S. mediocidicus ATCC23936, according to previously reported methods (Cai et al. 2007). The enzymatic reaction was then performed by incubating 1 and 2 with MedX and Co2+. It was reported that Co2+ can completely convert guanidino primycins into the corresponding amino forms compared with the other metal ions used in DstH-catalysed reactions (Hong et al. 2016). HPLC analysis revealed that the recombinant MedX can convert the guanidino form 1 into the corresponding amino form 2 (Fig. 6). Subsequently, the relative kinetic parameters of MedX in the hydrolysis of 1 were measured with varied substrate concentrations from 90 to 360 μM. The reciprocal Lineweaver–Burk method was performed to analyse experimental data, and the apparent kinetic constants were obtained (Table 1). This finding strongly suggests that MedX is the amidinohydrolase of S. mediocidicus ATCC23936 which is responsible for the production of 2 by hydrolysing the guanidino form 1 into the amino form 2 during post-PKS modifications (Fig. 7).
Discussion
Actinomycetes are known to produce many secondary metabolites that are encoded by biosynthetic gene clusters (Hornung et al., 2007). In the past few decades, a large number of antibiotics have been identified from natural sources, particularly Streptomyces. However, the biosynthetic gene clusters of only a small proportion of natural compounds have been discovered thus far. Because of their low cost, genome-sequencing projects have been demonstrated as an efficient method of identifying gene clusters involved in the biosynthesis of such products in microbial genomes (Kusserow and Tam 2017; Park et al. 2017; Sbaraini et al. 2017). In this study, the cle gene cluster was obtained through analysis of the draft genome sequence of S. mediocidicus ATCC23936. However, the amidinohydrolase for 2 formation was not found in the cle gene cluster. Heterogeneous expression of the speculative cle gene cluster could produce 1 rather than 2. Then, an amidinohydrolase (MedX) was found which could hydrolyse the guanidino form 1 into the amino form 2 via heterologous co-expression of the cle cluster in S. lividans or by in vitro catalysis.
The head of most LPPs is represented by guanidino or amino groups. Compound 2 probably possesses an amino group which originates from the guanidino form 1. MedX, an amidinohydrolase sequence exhibiting similarity with PAH, has been identified as a candidate for modifying the head of 1. The catalysis mechanism of MedX may be similar to that of PAH, which hydrolyses the guanidino intermediate to proclavaminic acid and urea (Elkins et al. 2002). Among known LPPs, ECO-0501 demonstrates strong antibacterial activity against many gram-positive pathogens. ECO-0501 has an N-methyl-guanidino group, and its gene cluster does not contain amidinohydrolase (Banskota et al. 2006). Except for LPPs, many marginolactone antibiotics also possess guanidino groups, such as primycin A1 (Frank et al. 1987), azalomycin F (Chandra and Nair 1995) and kanchanamycin C (Stephan et al. 1996). Whether MedX can catalyse other guanidino-containing natural compounds to generate the amino-containing products remains unclear. If so, it may be exploited through combinatorial biosynthesis for structural diversity in LPP natural products.
In summary, the amidinohydrolase MedX has been identified, for the first time, by genome scanning of the genome of S. mediocidicus ATCC23936. MedX can hydrolyse the guanidino form LPP 1 into the amino form 2 during post-PKS modifications. The present findings not only represent a significant step towards the complete elucidation of the biosynthetic pathway of 2 but also provide a possibility of using the newly identified amidinohydrolases to generate structural diversity antibiotics by combinatorial biosynthesis in future studies.
References
Banskota AH, Mcalpine JD, Ibrahim A, Aouidate M, Piraee M, Alarco AM, Farnet CM, Zazopoulos E (2006) Genomic analyses lead to novel secondary metabolites. Part 3. ECO-0501, a novel antibacterial of a new class. J Antibiot 59(9):533–542. https://doi.org/10.1038/ja.2006.74
Cai P, Kong F, Fink P, Ruppen ME, Williamson RT, Keiko T (2007) Polyene antibiotics from Streptomyces mediocidicus. J Nat Prod 70(2):215–219. https://doi.org/10.1021/np060542f
Chandra A, Nair MG (1995) Azalomycin F complex from Streptomyces hygroscopicus, MSU/MN-4-75B. J Antibiot 48(8):896–898. https://doi.org/10.7164/antibiotics.48.896
Chen S, Wu Q, Shen Q, Wang H (2015) Progress in understanding the genetic information and biosynthetic pathways behind Amycolatopsis antibiotics, with implications for the continued discovery of novel drugs. Chembiochem 17:119–128
Dowling DP, Di CL, Gennadios HA, Christianson DW (2008) Evolution of the arginase fold and functional diversity. Cell Mol Life Sci 65(13):2039–2055. https://doi.org/10.1007/s00018-008-7554-z
Elkins JM, Clifton IJ, Hernández H, Doan LX, Robinson CV, Schofield CJ, Hewitson KS (2002) Oligomeric structure of proclavaminic acid amidino hydrolase: evolution of a hydrolytic enzyme in clavulanic acid biosynthesis. Biochem J 366(2):423–434. https://doi.org/10.1042/bj20020125
Frank J, Dékány G, Pelczer I, ApSimon JW (1987) The composition of primycin. Tetrahedron Lett 28(24):2759–2762. https://doi.org/10.1016/S0040-4039(00)96202-6
Furumai T, Yamakawa T, Yoshida R, Igarashi Y (2003) Clethramycin, a new inhibitor of pollen tube growth with antifungal activity from Streptomyces hygroscopicus TP-A0623. I. Screening, taxonomy, fermentation, isolation and biological properties. J Antibiot 56(8):700–704. https://doi.org/10.7164/antibiotics.56.700
Gust B, Challis GL, Fowler K, Kieser T, Chater KF (2003) PCR-targeted Streptomyces gene replacement identifies a protein domain needed for biosynthesis of the sesquiterpene soil odor geosmin. Proc Natl Acad Sci U S A 100(4):1541–1546. https://doi.org/10.1073/pnas.0337542100
Hong H, Samborskyy M, Lindner F, Leadlay PF (2016) An amidinohydrolase provides the missing link in the biosynthesis of amino marginolactone antibiotics. Angew Chem Int Ed Engl 55(3):1118–1123. https://doi.org/10.1002/anie.201509300
Hornung A, Bertazzo M, Dziarnowski A, Schneider K, Welzel K, Wohlert SE, Holzenkämpfer M, Nicholson GJ, Bechthold A, Süssmuth RD (2007) A genomic screening approach to the structure-guided identification of drug candidates from natural sources. Chembiochem 8(7):757–766. https://doi.org/10.1002/cbic.200600375
Igarashi Y, Iwashita T, Fujita T, Naoki H, Yamakawa T, Yoshida R, Furuma T (2003) Clethramycin, a new inhibitor of pollen tube growth with antifungal activity from Streptomyces hygroscopicus TP-A0623. II. Physico-chemical properties and structure determination. J Antibiot 56(8):705–708. https://doi.org/10.7164/antibiotics.56.705
Kusserow K, Tam G (2017) Complete genome sequence of Actinomadura parvosata subsp. kistnae, a rich source of novel natural product (bio-)chemistry. J Genomics 5:75–76. https://doi.org/10.7150/jgen.19673
Lee SJ, Kim DJ, Kim HS, Lee BI, Yoon HJ, Yoon JY, Kim KH, Jang JY, Im HN, An DR (2011) Crystal structures of Pseudomonas aeruginosa guanidinobutyrase and guanidinopropionase, members of the ureohydrolase superfamily. J Struct Biol 175(3):329–338. https://doi.org/10.1016/j.jsb.2011.05.002
Liu W, Min M, Xue Y, Nan L, Wang S, Chen Y (2013a) The C-terminal extended serine residue is absolutely required in nosiheptide maturation. Chembiochem 14(5):573–576. https://doi.org/10.1002/cbic.201200681
Liu W, Xue Y, Ma M, Wang S, Liu N, Chen Y (2013b) Multiple oxidative routes towards the maturation of nosiheptide. Chembiochem 14(13):1544–1547. https://doi.org/10.1002/cbic.201300427
Mcalpine JB, Bachmann BO, Piraee M, Tremblay S, Alarco AM, Zazopoulos E, Farnet CM (2005) Microbial genomics as a guide to drug discovery and structural elucidation: ECO-02301, a novel antifungal agent, as an example. J Nat Prod 68(4):493–496. https://doi.org/10.1021/np0401664
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, USA
Park HB, Perez CE, Barber KW, Rinehart J, Crawford JM (2017) Genome mining unearths a hybrid nonribosomal peptide synthetase-like-pteridine synthase biosynthetic gene cluster. elife 6:e25229
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425
Sbaraini N, Andreis FC, Thompson CE, Guedes RLM, Junges Â, Campos T, Staats CC, Vainstein MH, Vasconcelos ATRD, Schrank A (2017) Genome-wide analysis of secondary metabolite gene clusters in ophiostoma ulmi and ophiostoma novo-ulmi reveals a fujikurin-like gene cluster with a putative role in infection. Front Microbiol 8:1063. https://doi.org/10.3389/fmicb.2017.01063
Stephan H, Kempter C, Metzger JW, Jung G, Potterat O, Pfefferle C, Fiedler H (1996) Kanchanamycins, new polyol macrolide antibiotics produced by Streptomyces olivaceus Tü 4018. J Antibiot 49(8):765–769. https://doi.org/10.7164/antibiotics.49.765
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24(8):1596–1599. https://doi.org/10.1093/molbev/msm092
Zhang L, Hashimoto T, Qin B, Hashimoto J, Kozone I, Kawahara T, Okada M, Awakawa T, Ito T, Asakawa Y, Ueki M, Takahashi S, Osada H, Wakimoto T, Ikeda H, Shin-Ya K, Abe I (2017) Characterization of giant modular PKSs provides insight into genetic mechanism for structural diversification of aminopolyol polyketides. Angew Chem Int Ed Engl 56(7):1740–1745. https://doi.org/10.1002/anie.201611371
Acknowledgements
We thank Dr. Mei Ge, Shanghai Laiyi Center for Biopharmaceuticals R & D, for providing S. mediocidicus ATCC23936.
Funding
This work was supported by grants from Tianjin science and technology plan projects (no. 16YFZCSY01000).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 489 kb)
Rights and permissions
About this article
Cite this article
Sun, F., Xu, S., Jiang, F. et al. Genomic-driven discovery of an amidinohydrolase involved in the biosynthesis of mediomycin A. Appl Microbiol Biotechnol 102, 2225–2234 (2018). https://doi.org/10.1007/s00253-017-8729-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8729-z