Abstract
Planktonic protists, including both autotroph and heterotroph, have been recognized as a major contributor to primary production and consumers of bacteria, archaea and picophytoplankton. However, the understanding of protistan diversity is typically impeded by the large amount of metazoans when employing universal primers to environmental samples. In this study, we developed 3 blocking primers to inhibit the amplification of metazoan DNA in PCR. First, we optimized the design and concentration of blocking primers by using 2 metazoans and 3 protists in quantitative PCR. No significant difference (q > 0.05) was found in protistan community structure at phylum, family or OTU levels and genetic diversity between samples amplified with and without adding blocking primers, indicating that the blocking primer does not alter the composition of protistan community. An application to samples with a high abundance of metazoans showed that the blocking primer can reduce 42.1–72.4% of metazoan sequences, resulting the retrieval of a higher protistan richness and diversity. The use of the blocking primer helped to identify protistan community composition from a large size fraction, which is usually dominated by metazoans. Protistan community from the small (0.2–10 μm) and large (> 10 μm) size fractions exhibited a low similarity of 36.6% and shared 14.7% OTUs. About 63.8% of the OTUs were unique to the large fraction. Species from groups, such as Lepidodinium, Warnowia, Kareniaceae, Torodiniales, Phaeocystis, Chrysochromulina and Chrysophyceae, were enriched in the large fraction, indicating that they could be largely underestimated in studies that exclude the large-sized cells. Blocking primers are a promising tool to increase the efficiency for the characterization of protistan diversity in aquatic planktonic ecosystems.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Protists are an array of microbial, unicellular eukaryotes. They are ubiquitous and abundant throughout marine environments and play fundamental ecological roles in marine ecosystems. Photosynthetic protists have been recognized as a major contributor to primary production in nearly all aquatic environments due to their high biomass (Jardillier et al. 2010; Massana 2011). Together with picocyanobacteria, they are responsible for roughly half of earth’s primary production (Field et al. 1998). Heterotrophic protists are important consumers of bacteria, archaea and picophytoplankton, channelling carbon and nutrients to higher trophic levels and remineralizing organic matter to inorganic nutrients (Boenigk and Arndt 2002; Sherr and Sherr 1994). Mixotrophs contribute to ~ 50% of bacterivory in ocean systems (Zubkov and Tarran 2008). In addition, there are also some other universal and predominant inter-species relationships, such as symbiosis, parasitism, mutualism and commensalism, existing between heterotrophic protists and other marine organisms (de Vargas et al. 2015; Gast et al. 2009). Thus, protists carry out a broad range of functions in the biogeochemical cycling in the ocean.
The growing awareness of the importance of protists is paralleled by an increasing interest in studying the community composition from diverse habitats. Over the last decades, molecular approaches, particularly those sequencing the 18S rRNA gene, have provided a powerful complement to traditional microscope-based taxonomies, which are restricted to the recognition of species with conspicuous morphological features. These studies have highlighted the existence of a vast hidden diversity of novel protists at all taxonomic ranks and open new avenues for our understanding of the spatial and temporal distribution of this diversity and ecosystem functioning (de Vargas et al. 2015; Moon-van der Staay et al. 2001; Orsi et al. 2012; Pernice et al. 2016; Vaulot et al. 2008). However, these investigations mostly focus on the picoplanktonic size fraction (0.2–3 μm). The choice to target this exceptionally small entity is partially due to the dominance of metazoan sequences if larger size classes are included (Caron et al. 2012). Up to 90% of all 18S rRNA sequences could be accounted by metazoans (Sauvadet et al. 2010). This could be because of the large amount of genomic DNA contributed by body pieces, eggs or planktonic larvae (Stoeck et al. 2009). Therefore, this undesirable amplification would lead to the underestimate of diversity and cause biases, such as false-positive and false-negative results. Metazoan signal might be misinterpreted as target signal when using low-resolution methods, such as T-RFLP (terminal restriction fragment length polymorphism) and DGGE (denaturing gradient gel electrophoresis) (Dolinšek et al. 2013; Ikenaga and Sakai 2014). Amplification of protistan DNA would be masked by metazoan DNA when present in extremely high quantity (Boessenkool et al. 2012; Sauvadet et al. 2010; Vestheim and Jarman 2008). Furthermore, this prefiltration manipulation precludes us from understanding the diversity and function of protists from larger size fractions or even skews results from the true community composition. For example, it has been shown in several studies that protistan community composition from different size fractions are phylogenetically different (de Vargas et al. 2015; Díez et al. 2001b; Duret et al. 2015; Le Bescot et al. 2016; Logares et al. 2012; Parris et al. 2014). An unsuspectedly high abundance and richness of symbionts and parasites have been reported in the mesoplanktonic class (de Vargas et al. 2015; Le Bescot et al. 2016). The average cell size of MAST-1C is 4.9 μm and some cells are > 8 μm in diameter (Lin et al. 2012).
Several approaches are possible to avoid detecting metazoans. In addition to prefiltration, other methods include enzymatic digestion of non-target DNA (Blankenship and Yayanos 2005), designing group- or species-specific primer (Sauvadet et al. 2010), and PCR clamping (Karkare and Bhatnagar 2006). However, restriction enzymatic digestion requires that there is a unique restriction site. Using group- or species-specific primers can be problematic when it inhibits amplification of species of interest and lead to bias (Sauvadet et al. 2010). PCR clamping, such as locked nucleic acids (LNAs), peptide nucleic acids (PNAs) and morpholinos, suppresses the amplification of dominant DNA by specifically and stably binding target DNA but not priming DNA polymerisation (O'Rorke et al. 2012). Moreover, it takes several weeks and is quite expensive to synthesize these probes. Recently new methods have been developed, such as catalytically active oligonucleotides containing LNAs (LNAzymes) (Dolinšek et al. 2013) and depletion of abundant sequences by hybridization (DASH) (Gu et al. 2016). Although they have been shown to be capable of deleting unwanted DNA, they typically require more experimental handling, for example, in vitro transcription required for LNAzymes (Dolinšek et al. 2013) and recombinant Cas9 proteins for DASH (Gu et al. 2016).
A simple method is to use a metazoan-specific blocking primer, which binds to metazoan DNA but has the 3′-end labelled with dideoxynucleotides that inhibits enzymatic elongation of the primer (Seyama et al. 1992). This primer can be designed to overlap with or lie between the amplification primers and, thus, compete annealing (called annealing inhibiting blocking primers) or prevent elongation (called elongation arrest blocking primers), respectively (Vestheim et al. 2011). Blocking primers have been applied in environmental microbiology studies (Gofton et al. 2015), ancient DNA (Boessenkool et al. 2012), diet characterization (Vestheim and Jarman 2008) and clinic diagnosis and prognosis (Dominguez and Kolodney 2005). In this study, we first developed 3 blocking primers specific to metazoan 18S rRNA genes and then tested the inhibition specificity and efficiency by using pure cultures and environmental DNA. Secondly, we applied this method to environmental samples with abundant metazoan DNA to compare protistan community structures generated with and without a blocking primer. Thirdly, protistan community from the largely understudied fraction, > 10 μm, was characterized and compared with that from the small fraction, 0.2–10 μm.
Materials and methods
Design of blocking primers
In a pilot profiling study on protistan community in the western North Pacific Ocean, up to 91% of 454 quality sequences in our datasets were metazoan. To inhibit amplification of these overwhelming sequences during PCR, three blocking primers were designed for the fragment targeted by the 18S universal primers Euk82F (López-García et al. 2003) and Euk516R (Díez et al. 2001a), which generate a product with a predicted length of 435 bp. Because of a lack of regions conservative in metazoans but variable in protistan 18S rRNA gene, only the representative sequences of the 3 most abundant OTUs (operational taxonomic units) were used for alignment (Fig. 1). The OTUs were clustered using the sample 4_10 (Table 1) following the method stated in the part “Sequence processing”. The top 3 abundant OTUs accounted for 81.3% of the reads in the sample.
Alignment of the 3 blocking primers used in this study, the amplification primers and the representative sequences of the top 3 abundant metazoan OTUs and the top 10 abundant non-metazoan protistan OTUs. These representative sequences were retrieved from a samples in a pilot profiling study on protistan community in the western North Pacific Ocean. The positions of the bases are shown above the alignment
The 18S_blkMetaz1 and 18S_blkMetaz3 blocking primers annealed to the template in the region lying between the 2 universal primers to prevent the read-through of the Taq polymerase and this is determined as an elongation arrest. The 18S_blkMetaz2 is an annealing inhibiting primer, which overlapped with the 3′ end of the forward universal primer Euk82F but extended into metazoan-specific sequences. All 3 blocking primers were modified with a C3 spacer (3 hydrocarbons, 1-dimethoxytrityloxy-propanediol-3-succinoyl-long chain alkylamino) at the 3′ end, which is stable in terms of degradation and enzymatic removing. It is very difficult to design annealing inhibiting primers next to the binding site of the reverse universal primer Euk516R because of self-complementarity and complementarity with the universal primers.
Test of blocking primers
To evaluate the specificity and efficiency of the 3 different blocking primers, pure cultures of a variety of protists, including 2 copepods (Apocyclops royi and Tigriopus japonicus), 1 diatom (Thalassiosira weissflogii), 1 nanoflagellate (Caecitellus sp.) and 1 dinoflagellate (Lepidodinium sp.), were subjected to quantitative PCR (qPCR). 1–5 ml cultures or 10 copepods were used to isolate DNA following the method by Puchooa (2004). Briefly, cells were suspended with an extraction buffer (100 mM Tris pH 8.0, 20 mM EDTA pH 8.0, 2 M NaCl, 2% CTAB, 2% PVP, 10 μg/ml RNase A, 5% β-mercaptoethanol and 0.1 mg/ml proteinase K) and incubated at 65 °C overnight. DNA was extracted using equal volume of chloroform:isoamylalcohol (24:1), precipitated using isopropanol and washed using 70% ethanol. DNA was finally dissolved in 100 μl TE buffer and stored at − 20 °C.
All DNA samples were diluted to 0.1 ng/μl, quantified by using a Quant-iT PicoGreen dsDNA Assay Kit (Life Technology, Eugene, Oregon, USA), and measured in duplicates. All samples for qPCR were measured in triplicates. Each qPCR reaction (10 μl) contained 0.5 μl DNA, 5 μl 2 × SYBR Green Master Mix (Takara, Otsu, Japan), 0.2 μl ROX II, 0.5 μM each universal primer, and 2 μM blocking primer (suggested as the optimal concentration by Boessenkool et al. (2012)). Reactions were carried out on a LightCycler 480 Instrument II (Roche, Basel, Switzerland) with the following reaction profile: 94 °C for 5 min, 55 cycles of 94 °C for 30 s, 55 °C for 30 s and 72 °C for 60 s.
To test the efficiency of the blocking primer in blocking the amplification of metazoan DNA, qPCR was performed on a serial concentrations of T. japonicus DNA: 0.001, 0.01, 0.1 and 1 ng/μl. All dilutions were carried out from a serial dilution of a stock DNA solution of ~ 3.4 ng/μl.
To determine the minimal concentration required to sufficiently supress the amplification of metazoan DNA, different concentrations of the best blocking primer were added to qPCR reactions with 0.1 ng/μl T. japonicus DNA: 0.5, 1.0, 1.5, 2.0., 2.5 and 3 μM, corresponding to 1, 2, 3, 4, 5 and 6 times of the universal primer concentration.
Application of the blocking primer to environmental samples in pyrosequencing
Sample collection
Surface water samples (~ 4 L) were collected from the western North Pacific Ocean during a cruise in July 2013 (Table 1). Prefiltered (200-μm mesh) water samples were filtered sequentially through a 10- and 2-μm polycarbonate membrane to get the 2 size fractions, > 10 μm and 0.2–10 μm, respectively. Membranes were soaked in the RNAlater (Ambion, Austin, Texas, USA) and stored at − 80 °C before DNA extraction. First we applied the best blocking primer to the small fraction (0.2–10 μm) that had relatively few metazoan DNA to further validate the specificity. Then we investigated the performance of the best blocking primer on the > 10 μm fraction samples which had knowingly abundant metazoan DNA (Table 1). The western North Pacific Ocean was selected because it is one of the least studied areas on earth and harbours multiple environmental gradients.
DNA extraction, PCR and 454 pyrosequencing
DNA was extracted using the TRIzol reagent (Invitrogen, Austin, Texas, USA) in combination with the Genomic DNA Mini Kit (Invitrogen, Grand Island, Nebraska, USA), and stored at − 20 °C. Then DNA was amplified with and without the blocking primer, 18S_blkMetaz2, using barcoded universal primers. All samples were carried out in duplicate PCRs with a 20-μl reaction containing 0.75 U Platinum Taq DNA polymerase (Invitrogen, Austin, Texas, USA), 1 × PCR buffer, 1.75 mM MgCl2, 0.25 mM dNTPs, 0.5 μM each barcoded universal primer, 2 μM blocking primer and 1–50 ng DNA. The thermal profile consisted of 95 °C for 5 min, 30 cycles of 95 °C for 30 s, 55 °C for 60 s and 72 °C for 60 s, followed with a final extension at 72 °C for 7 min. Duplicate PCR reactions were combined and purified from gel using a Universal DNA Purification Kit (TIANGEN, Beijing, China) and followed with a further purification using a Agencourt AMPure XP kit (Beckman Coulter, Brea, California, USA). An equimolar mix of each sample was created and subjected to 454 GS Junior pyrosequencing (Roche, Bradford, Pennsylvania, USA).
Sequence processing
Sequences were quality-controlled following the protocol from the Mothur package (Schloss et al. 2011). Primer and tag sequences were removed. Ambiguous base calls were not allowed. Sequences remained only if the average quality score was > 20 and read length was > 250 bp. Reads were then de-replicated and aligned against the SILVA eukaryote reference database. Sequences that are outside the desired range of the alignment were removed. The alignment was filtered out to remove any columns that had only gaps or had gaps at the terminals. Chimeras were checked using the UCHIME method (Edgar et al. 2011) and removed. To account for PCR and sequencing errors, sequences represented only once in the total dataset were discarded. Sequences were assigned with a taxonomic identity using the PR2 database (Guillou et al. 2013) at a cutoff of 60% and clustered into OTUs using the default setting. A subsample to the number of sequences in the smallest sample was taken from all samples. Sobs richness and Shannon evenness and diversity of protists were calculated at 97% similarity.
Phylogenetic analyses
In order to compare the distribution of phylogenetic groups in the 2 fractions, phylogenetic trees were built using representative sequences of the top 50 abundant OTUs of all samples. Highly divergent sequences were removed after preliminary phylogenetic analyses by neighbour-joining to avoid long-branch attraction effects. These sequences were aligned with reference sequences from the GenBank using the MAFFT v.7 software (Katoh and Standley 2013) under the G-INS-i method and then manually curated. For maximum likelihood (ML) analyses, the best fit model was obtained from jModelTest v2.1.7 (Darriba et al. 2012) among 88 models. A total of 100 bootstrapped trees were constructed using the PhyML v3.0 software (Guindon et al. 2010).
All sequences used in this study are deposited in the NCBI short reads archive database under the BioProject PRJNA384701.
Statistical analyses
Comparisons across samples were conducted using analysis of variance (ANOVA) if normality and equal variance are met. If not, they were based on the generalized linear model conducted using the R packages, robust and multcomp (Hothorn et al. 2008). The Shapiro-Wilk test was used to test normality. The Bartlett’s test was used to test if samples are with equal variance. All analyses were conducted using the R software (Team RC 2014). P values were corrected using the Storey’s false discovery rate method. Weighted UniFrac and analysis of molecular variance (AMOVA) embedded in the Mothur were used to assess community structure and genetic diversity, respectively, between the samples amplified with and without using the blocking primer. Thetayc distance was used to calculate the similarity in community structure.
Results
Selection of blocking primers
Of the 3 blocking primers tested, 18S_blkMetaz2 was the most effective in blocking the amplification of metazoans (A. royi and T. japonicus) (Fig. 2a). The addition of 18S_blkMetaz2 increased the Ct values of A. royi and T. japonicus from 23.1 to 29.6 and from 19.7 to 33.8, respectively. 18S_blkMetaz3 increased the Ct values slightly, but not as much as 18S_blkMetaz2. 18S_blkMetaz1 did not show any inhibition on both copepods. In addition, 18S_blkMetaz2 showed a negligible inhibition on the amplification of all non-metazoan species (Fig. 2a).
Test of the specificity and efficiency of the blocking primers based on Ct values generated from qPCR. Performance of the 3 blocking primers on 2 metazoans (Apocyclops royi and Tigriopus japonicus), 1 diatom (Thalassiosira weissflogii), 1 nanoflagellate (Caecitellus sp.) and 1 dinoflagellate (Lepidodinium sp.) (a). Inhibition of different concentrations of the blocking primer 18S_blkMetaz2 on the amplification of metazoan (Tigriopus japonicus) DNA (b). Ability of the blocking primer 18S_blkMetaz2 (2 μM) of inhibiting different concentrations of metazoan (Tigriopus japonicus) DNA (c). BP represents that blocking primers were added. BP represents no blocking primer added. ** indicates a statistical significance (P < 0.01) compared to the non-BP
Performance of blocking primer
We then tested the minimal concentration of the blocking primer 18S_blkMetaz2 that can perform an effective inhibition of metazoan DNA amplification in PCR. The inhibition exhibited a dose-dependent response (Fig. 2b). The Ct value increased with the increasing concentration of 18S_blkMetaz2. There was no significant difference (P > 0.05) in Ct values when the blocking primer concentration increased from 2 to 2.5 μM. The inhibiting effect even decreased when the concentration went to 3 μM though not significantly (P > 0.05) compared to when the concentration of 2.5 μM was used. Therefore 2 μM was the minimal concentration of 18S_blkMetaz2 for effective inhibition and used in sections thereafter.
In order to test the ability of the blocking primer 18S_blkMetaz2 of inhibiting the amplification of metazoan DNA, we made 10-fold serial dilutions of T. japonicus DNA, ranging from 0.001 to 1 ng/μl. Significant blocking was possible until the concentration reached 1 ng/μl (Fig. 2c).
Test of specificity on environmental samples
To further examine the specificity, three seawater samples from the western North Pacific Ocean from the small size fraction (0.2–10 μm), where the fraction of metazoans is knowingly low, comprising 4.9–20.5% of total 18S rRNA sequences (Table 1), were selected and subjected to pyrosequencing with the use of the blocking primer 18S_blkMetaz2. This fraction is chosen because amplification may be biased by a high abundance of metazoan DNA, which would lead to a false evaluation of the specificity. Specificity was tested by comparing the relative abundance of different protistan groups between samples amplified with the blocking primer and the samples amplified without it. Overall, there was not a significant difference (q > 0.05 on the means with false discovery rate correction) in the relative abundance of individual phylum (or super group) (Fig. 3a), family (Fig. 3b) and OTU (Fig. 3c). At the OTU level, when top 100 abundant OTUs were considered, community composition of the 3 samples amplified with and without the blocking primer shared a mean similarity of 88.6%, which, empirically, is a typical similarity among experimental replicates. Weighted UniFrac and AMOVA further indicated respectively that the community structure and genetic diversity between the samples amplified with and without the blocking primer showed no significant differences (P > 0.05).
Application to environmental samples
To test the utility of this method, we applied the blocking primer 18S_blkMetaz2 to 3 samples collected in the western North Pacific Ocean from the large size fraction (> 10 μm). Metazoan reads accounted for 68.6 to 84.1% of all reads in the 18S rRNA gene library (Table 1). As a result, the sequencing effort should be increased markedly in order to get a full spectrum of the protistan diversity. Therefore removal of unwanted metazoan sequences in PCR would largely save both finance and time on analysis. In order to rate the performance of the blocking primer, we define “blocking efficiency” as (X-Y)/Y, where X is the ratio of the number of metazoan sequences to the number of protistan sequences in the 18S rRNA gene library when the sample is amplified without the blocking primer; Y is the ratio the number of metazoan sequences to the number of protistan sequences when the same sample is amplified with the blocking primer. This calculation is based on the assumption that the blocking primer is specific to metazoan DNA and amplification of non-metazoan DNA will not be inhibited. The result showed that the efficiency of the blocking primer 18S_blkMetaz2 varied across the samples, ranging from 42.1 to 72.4% (Table 1).
A total of 119,229 18S rRNA reads were generated, resulting in an average of 6624 sequences for each sample. Community indexes, such as Sobs richness, Shannon evenness and Shannon diversity, were applied to evaluate the results of the blocking primer on protistan community structure. As expected, more phylotypes (a higher richness) and a higher diversity were recovered when samples were amplified with the blocking primer than amplified without it under the same sequencing effort. On average, the use of the blocking primer helped to get an increase by 65.6% in richness and 2.9% in diversity (Fig. 4). A slight decrease was found with evenness by an average of 5.9%. This result indicates that more information about the community, particularly about those less abundant species, can be obtained with the using of the blocking primer.
Comparison of protistan community structure between the small and large size fractions
Protistan community has been found with size partitioning (de Vargas et al. 2015; Díez et al. 2001b; Duret et al. 2015; Le Bescot et al. 2016; Logares et al. 2012; Parris et al. 2014). With using the blocking primer, we are able to identify the shared and unique phylogenetic groups between the small and large fractions although there was a high abundance of metazoans from the large fraction (Table 1). The result showed that the 2 fractions from the 3 stations harboured a strikingly different protistan community, showing a mean similarity of 36.6% when top 100 OTUs were considered.
Then we compared the community indexes between the 2 fractions. Although lower, richness from the large fraction was as much as half of that from the small fraction. Evenness and diversity were approaching to those from the small fraction, reaching 89.2 and 75.9%, respectively (Fig. 5). This result indicates that there is a high diversity of protists from the large fraction, which can be potentially underestimated or even missed in studies focused on picoplanktonic protists.
On average, about 14.7% of the OTUs were shared between the small and large fractions with a respective proportion of 50.6% and 74.2% of reads in each fraction, indicating that these common OTUs are the abundant ones. A mean of 49.4 and 25.8% of the reads were unique to the small and large fractions, respectively. This result suggests that these unique OTUs were numerically abundant in the small fraction but less abundant in the large fraction (Table 2).
Certain OTUs were differentially detected in the 2 fractions. Therefore phylogenetic trees using the representative sequences of the top 50 OTUs were constructed to assign these major OTUs to the phylogenetic affiliations (Figs. 6 and 7). Alveolata was the most abundant group in both fractions (40/47 OTUs). No alveolate groups contained OTUs that were only abundant in the small or large fraction. Sequences from groups, such as Lepidodinium, Warnowia, Kareniaceae, and Torodiniales, mostly had a higher relative abundance in the large fraction. Syndiniales groups retained species that were mostly more abundant in the small fraction with exceptions such as OTUs 38 and 45 (Fig. 6). Stramenopile groups, including diatoms (Bacillariophyta), were mostly retained in the small fraction except species from Chrysophyceae (Fig. 7b). Haptophytes, namely species close to Phaeocystis and Chrysochromulina, were more represented in the large fraction. Only one ciliate OTU, closest to Stenosemella, was retrieved and was only observed in the small fraction (Fig. 7a).
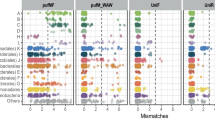
Maximum-likelihood phylogenetic tree of dinoflagellates. A heatmap shows the relative abundance of the OTUs between the small (0.2–10 μm) and large (> 10 μm) size fractions among 3 stations. The tree was constructed under the JC method. Bootstrap values based on 100 iterations are shown at the nodes. The colour gradient denotes the relative abundance of the OTUs in each sample
Maximum-likelihood phylogenetic trees of ciliates (a) and haptophytes and stramenopiles (b). A heatmap shows the relative abundance of the OTUs between the small (0.2–10 μm) and large (> 10 μm) size fractions among 3 stations. The tree of ciliates was constructed under the TrNef method. The tree of haptophytes and stramenopiles was constructed under the TPM1 + G method. Bootstrap values based on 100 iterations are shown at the nodes. The colour gradient denotes the relative abundance of the OTUs in each sample
Discussion
The major advantages of using blocking primers to inhibit unwanted DNA are the simplicity and efficiency. This method is as simple as a standard PCR and allows for simultaneous amplification of target DNA and inhibition of non-target DNA. It is efficient enough to detect minor DNA with the presence of 103~104-fold excess of non-target template (Craig et al. 2014; Seyama et al. 1992; Vestheim and Jarman 2008).
Evaluation of blocking primers
Because the annealing inhibiting primers targeting the reverse universal primer Euk516R form intra- and inter-primer dimers we designed one annealing inhibiting primer targeting the forward universal primer and 2 elongation arrest primers. Compared to the annealing inhibiting primer, the elongation arrest primer was more effective with a high specificity since it resulted in more increase in the Ct values of amplifying metazoan DNA but not much changes in the Ct values of non-metazoan DNA (Fig. 2a). Similar results have been found in other studies (Chow et al. 2011; Vestheim and Jarman 2008; von Wintzingerode et al. 2000). Elongation arrest once was not taken as a reliable way of inhibiting the amplification of non-target DNA (O'Rorke et al. 2012). This was suggested because of the complex interactions among the primer, DNA and Taq polymerase (von Wintzingerode et al. 2000). However, elongation arrest primers worked in some cases (Peano et al. 2005; von Wintzingerode et al. 2000; Yu et al. 1997). It was found that PCR parameters could be optimized to make elongation arrest work, such as designing a blocker with a higher Tm than the primer Tm (O'Rorke et al. 2012). However, the Tm of the 2 elongation arrest primers in this study were slightly lower than the Tm of the amplification primers, which may be the reason for their failure.
Although primers have been developed to amplify 18S rRNA genes of metazoan species (Machida and Knowlton 2012), it is very difficult, if not impossible, to design a good universal blocking primer that can block the amplification of all metazoans. The high diversity of metazoan and the short length of genes amplified by the universal primers impeded the selection of a single blocking primer. Thus, in this study, the blocking primers were designed using the dominant metazoans as the template. A plausible approach of achieving a higher efficiency is to design multiple blocking primers targeting the dominant metazoans in order to achieve a higher blocking efficiency. Generally there are more sites available for the design of elongation arrest primers than for annealing inhibiting primers. The efficiency of elongation arrest primers can be improved by adding an extra modification on the 5′-end (Vestheim et al. 2011).
To minimize unspecific inhibition of the amplification of target DNA, minimal blocking primer concentration should be used. We showed that the optimal concentration for 18S_blkMetaz2 was 2 μM (Fig. 2b), a ratio of 4:1 compared to the concentration of the amplification primers, which is similar to the ratio (5:1) used in the study by Boessenkool et al. (2012), where they used singular human DNA at the same concentration (0.1 ng/μl) as we did. However, this ratio varies from 1:1 to 25:1 in different studies (Deagle et al. 2009, 2010; Gigli et al. 2009; Gofton et al. 2015; Vestheim and Jarman 2008; Yi et al. 2014; Yu et al. 1997). The blocking efficiency depends on not only the concentration of the non-target DNA, as shown in this study that the blocking primer can suppress the amplification of metazoan DNA up to the concentration of 1 ng/μl (Fig. 2c), but also the ratio of target/non-target DNA (Craig et al. 2014; Vestheim and Jarman 2008). Therefore it is not appropriate, if not impossible, to compare the ratios from different studies. In this case, it is necessary to determine the optimal blocker concentration (Vestheim et al. 2011).
It is important to examine the specificity of the blocking primers since biases could be caused by unintended inhibition on the amplification of target DNA, particularly for studies about species diversity. This is insufficiently investigated in previous studies (Vestheim and Jarman 2008). Thus, we validated the specificity of the blocking primer 18S_blkMetaz2 using environmental samples with knowingly few metazoan DNA. An overall non-significant inhibition was observed on the amplification of protistan DNA with the using of the blocking primer (Fig. 3), indicating a high specificity to metazoans. This is further confirmed by using analyses of community structure and genetic diversity. Only one group of protists, alveolates at station 6, the relative abundance of which significantly (q < 0.05) decreased by 3.2% after using the blocking primer, resulting in a minor underestimate. Nevertheless, when looking at finer levels, groups belonging to alveolates did not show significant changes in the relative abundance.
Blocking primer allows insights into the protistan diversity from the large size fraction
Large size fractions, such as nanoplankton and microplankton, have been reported with a high relative richness of dinoflagellates (Le Bescot et al. 2016) and a high abundance of protists (de Vargas et al. 2015). Therefore a focus on the picoplanktonic protists would lead to an underestimate of protistan diversity. In our study, the application of the blocking primer helped the characterization of protists from the long-neglected fraction. We showed that there was a high diversity of protists from the large fraction (> 10 μm), close to that from the small fraction (0.2–10 μm) (Fig. 5), confirming the missing protistan diversity. This result is further validated by the small number of shared OTUs (14.7%) between the 2 fractions (Table 2). These results confirm the finding that there is size partitioning in protistan community (de Vargas et al. 2015; Díez et al. 2001b; Duret et al. 2015; Le Bescot et al. 2016; Logares et al. 2012; Parris et al. 2014). Thus, the use of blocking primers in surveys to other ecosystems would help to bridge this knowledge gap.
Dinoflagellates tend to be large and conspicuous organisms. The high representation of dinoflagellates in the small fraction (59.0%) as compared to that in the large fraction (71.4%) (Fig. 6) may be explained by filtration artefacts and amplification of dissolved DNA. Another possible reason is the existence of this-size dinoflagellates as their known minimal size is 5–10 μm. Species might be underrated if they are from groups such as Lepidodinium, Warnowia, Kareniaceae, and Torodiniales when prefiltration is processed because OTUs from these groups showed a higher relative abundance in the large fraction. This result is generally coherent with morphological information. For example, Lepidodinium include species with sizes ranging from 22 to 53 μm (Watanabe et al. 1990). Members from Warnowia have cell sizes between 26 and 67 μm (Reñé et al. 2015). Genera identified in our study belonging to family Kareniaceae include Karenia, Takayama and Karlodinium all contain species of > 10 μm in size (Reñé et al. 2015). The commonly reported dimension of Torodinium is 100–130 μm in length (Gómez 2009). Syndiniales is an order composed of a variety of parasites. Most of the Syndiniales OTUs were found in the small fraction, indicating that a majority of these parasites are present in the free-living form. The 2 Syndiniales OTUs detected with a higher abundance in the large fraction is an indication that they are likely in their infectious stage within large-size hosts, such as mesoplanktonic protists and metazoans. Thus, the application of blocking primer to protists in large fractions makes it possible to dissect parasitic interactions.
Other OTUs enriched in the large fraction belong to Chrysophyceae (Chromulinales) and Haptophyta (Phaeocystis and Chrysochromulina) (Fig. 7). Spumella and Paraphysomonas typically have a size in length smaller than 10 μm (Vaulot et al. 2008). Therefore this chrysophyte sequence is likely to represent a new species belonging to Chromulinales, particularly when considering that it has a similarity of 95% with its closest relative Spumella (AY651091). Phaeocystis exhibit phase alteration between free-living cells of a picoplanktonic size and colonies reaching several millimetre (Vaulot et al. 2008). The higher relative abundance of Phaeocystis in the larger fraction, particularly at station 6 (11.0% in the large fraction compared to 0.2% in the small fraction), suggests that Phaeocystis may bloom and form colonies. Chrysochromulina is composed of species with a size mostly < 10 μm. However, taxa such as C. strobilus (Leadbeater and Manton 1969b) and C. camella (Leadbeater and Manton 1969a) can be larger than this size.
Protists from > 10 μm are relatively well known because 10 μm in size represents the lower limit of morphological identification by light microscopy. However, this morphology-based approach has difficulties in taxonomizing, e.g., unarmoured dinoflagellates, which tend to be easily damaged by sampling and distorted by fixation. Some studies using DNA sequencing have demonstrated the presence of cryptic species within morphospecies (Caron 2013). Additionally, molecular approaches have the advantage of analysing a large number of samples at the community level, allowing to examine the response of the whole protistan community to environmental forces (Caron 2013). Moreover, nanoplanktonic dinoflagellates have been shown with the highest relative richness of the dinoflagellates from the whole size spectrum (Le Bescot et al. 2016), highlighting the potential of applying molecular approaches to protists of a size larger than the picosize.
This work constitutes a proof-of-concept study that blocking primers can be used to inhibit the amplification of predominant metazoan DNA, allowing the detection of less abundant protistan DNA. The advantage of this technique lies with the capability of detecting novel protists with universal primers and saving efforts through constructing the library within one PCR for one sample. Although this study is based on 454 pyrosequencing, this method can be readily applied to other high-throughput sequencing platforms involving library preparation based on PCR, such as Illumina and PacBio. Adding blocking primers can reduce the representation of 42.1–72.4% of metazoan sequences in the library. The application of blocking primers helps the study of protistan diversity in the large size fraction (> 10 μm), which has not been well studied using molecular approaches. In this study we found 63.8% of the OTUs were unique to the large fraction. Species from groups, such as Lepidodinium, Warnowia, Kareniaceae, Torodiniales, Phaeocystis, Chrysochromulina and Chrysophyceae, were enriched in the large fraction, lending them to be largely underestimated in studies filtering out large-size cells.
References
Blankenship LE, Yayanos AA (2005) Universal primers and PCR of gut contents to study marine invertebrate diets. Mol Ecol 14(3):891–899. https://doi.org/10.1111/j.1365-294X.2005.02448.x
Boenigk J, Arndt H (2002) Bacterivory by heterotrophic flagellates: community structure and feeding strategies. Antonie Van Leeuwenhoek 81(1):465–480. https://doi.org/10.1023/a:1020509305868
Boessenkool S, Epp LS, Haile J, Bellemain E, Edwards M, Coissac E, Willerslev E, Brochmann C (2012) Blocking human contaminant DNA during PCR allows amplification of rare mammal species from sedimentary ancient DNA. Mol Ecol 21(8):1806–1815. https://doi.org/10.1111/j.1365-294X.2011.05306.x
Caron DA (2013) Towards a molecular taxonomy for protists: benefits, risks, and applications in plankton ecology. J Eukaryot Microbiol 60(4):407–413. https://doi.org/10.1111/jeu.12044
Caron DA, Countway PD, Jones AC, Kim DY, Schnetzer A (2012) Marine protistan diversity. Annu Rev Mar Sci 4(1):467–493. https://doi.org/10.1146/annurev-marine-120709-142802
Chow S, Suzuki S, Matsunaga T, Lavery S, Jeffs A, Takeyama H (2011) Investigation on natural diets of larval marine animals using peptide nucleic acid-directed polymerase chain reaction clamping. Mar Biotechnol 13(2):305–313. https://doi.org/10.1007/s10126-010-9301-3
Craig C, Kimmerer WJ, Cohen CS (2014) A DNA-based method for investigating feeding by copepod nauplii. J Plankton Res 36(1):271–275. https://doi.org/10.1093/plankt/fbt104
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9(8):772–772. https://doi.org/10.1038/nmeth.2109
de Vargas C, Audic S, Henry N, Decelle J, Mahé F, Logares R, Lara E, Berney C, Le Bescot N, Probert I, Carmichael M, Poulain J, Romac S, Colin S, Aury J-M, Bittner L, Chaffron S, Dunthorn M, Engelen S, Flegontova O, Guidi L, Horák A, Jaillon O, Lima-Mendez G, Lukeš J, Malviya S, Morard R, Mulot M, Scalco E, Siano R, Vincent F, Zingone A, Dimier C, Picheral M, Searson S, Kandels-Lewis S, Acinas SG, Bork P, Bowler C, Gorsky G, Grimsley N, Hingamp P, Iudicone D, Not F, Ogata H, Pesant S, Raes J, Sieracki ME, Speich S, Stemmann L, Sunagawa S, Weissenbach J, Wincker P, Karsenti E, Boss E, Follows M, Karp-Boss L, Krzic U, Reynaud EG, Sardet C, Sullivan MB, Velayoudon D (2015) Eukaryotic plankton diversity in the sunlit ocean. Science 348(6237):1261605. https://doi.org/10.1126/science.1261605
Deagle BE, Chiaradia A, McInnes J, Jarman SN (2010) Pyrosequencing faecal DNA to determine diet of little penguins: is what goes in what comes out? Conserv Genet 11(5):2039–2048. https://doi.org/10.1007/s10592-010-0096-6
Deagle BE, Kirkwood R, Jarman SN (2009) Analysis of Australian fur seal diet by pyrosequencing prey DNA in faeces. Mol Ecol 18(9):2022–2038. https://doi.org/10.1111/j.1365-294X.2009.04158.x
Díez B, Pedrós-Alió C, Marsh TL, Massana R (2001a) Application of denaturing gradient gel electrophoresis (DGGE) to study the diversity of marine picoeukaryotic assemblages and comparison of DGGE with other molecular techniques. Appl Environ Microbiol 67(7):2942–2951. https://doi.org/10.1128/aem.67.7.2942-2951.2001
Díez B, Pedrós-Alió C, Massana R (2001b) Study of genetic diversity of eukaryotic picoplankton in different oceanic regions by small-subunit rRNA gene cloning and sequencing. Appl Environ Microbiol 67(7):2932–2941. https://doi.org/10.1128/aem.67.7.2932-2941.2001
Dolinšek J, Dorninger C, Lagkouvardos I, Wagner M, Daims H (2013) Depletion of unwanted nucleic acid templates by selective cleavage: LNAzymes, catalytically active oligonucleotides containing locked nucleic acids, open a new window for detecting rare microbial community members. Appl Environ Microbiol 79(5):1534–1544. https://doi.org/10.1128/aem.03392-12
Dominguez PL, Kolodney MS (2005) Wild-type blocking polymerase chain reaction for detection of single nucleotide minority mutations from clinical specimens. Oncogene 24(45):6830–6834. https://doi.org/10.1038/sj.onc.1208832
Duret MT, Pachiadaki MG, Stewart FJ, Sarode N, Christaki U, Monchy S, Srivastava A, Edgcomb VP (2015) Size-fractionated diversity of eukaryotic microbial communities in the Eastern Tropical North Pacific oxygen minimum zone. FEMS Microbiol Ecol 91(5):fiv037. https://doi.org/10.1093/femsec/fiv037
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27(16):2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Field CB, Behrenfeld MJ, Randerson JT, Falkowski P (1998) Primary production of the biosphere: integrating terrestrial and oceanic components. Science 281(5374):237–240. https://doi.org/10.1126/science.281.5374.237
Gast RJ, Sanders RW, Caron DA (2009) Ecological strategies of protists and their symbiotic relationships with prokaryotic microbes. Trends Microbiol 17(12):563–569. https://doi.org/10.1016/j.tim.2009.09.001
Gigli E, Rasmussen M, Civit S, Rosas A, de la Rasilla M, Fortea J, Gilbert MTP, Willerslev E, Lalueza-Fox C (2009) An improved PCR method for endogenous DNA retrieval in contaminated Neandertal samples based on the use of blocking primers. J Archaeol Sci 36(12):2676–2679. https://doi.org/10.1016/j.jas.2009.08.004
Gofton AW, Oskam CL, Lo N, Beninati T, Wei H, McCarl V, Murray DC, Paparini A, Greay TL, Holmes AJ, Bunce M, Ryan U, Irwin P (2015) Inhibition of the endosymbiont “Candidatus Midichloria mitochondrii” during 16S rRNA gene profiling reveals potential pathogens in Ixodes ticks from Australia. Parasit Vectors 8(1):1–11. https://doi.org/10.1186/s13071-015-0958-3
Gómez F (2009) Torodinium and Pavillardia (Gymnodiniales, Dinophyceae): two unarmoured dinoflagellates with a body extension, collected from the open Pacific Ocean. Protistology 6(2):131–135
Gu W, Crawford ED, O'Donovan BD, Wilson MR, Chow ED, Retallack H, DeRisi JL (2016) Depletion of abundant sequences by hybridization (DASH): using Cas9 to remove unwanted high-abundance species in sequencing libraries and molecular counting applications. Genome Biol. https://doi.org/10.1101/031708
Guillou L, Bachar D, Audic S, Bass D, Berney C, Bittner L, Boutte C, Burgaud G, de Vargas C, Decelle J, del Campo J, Dolan JR, Dunthorn M, Edvardsen B, Holzmann M, Kooistra WHCF, Lara E, Le Bescot N, Logares R, Mahé F, Massana R, Montresor M, Morard R, Not F, Pawlowski J, Probert I, Sauvadet A-L, Siano R, Stoeck T, Vaulot D, Zimmermann P, Christen R (2013) The protist ribosomal reference database (PR2): a catalog of unicellular eukaryote small sub-unit rRNA sequences with curated taxonomy. Nucleic Acids Res 41(D1):D597–D604. https://doi.org/10.1093/nar/gks1160
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59(3):307–321. https://doi.org/10.1093/sysbio/syq010
Hothorn T, Bretz F, Westfall P (2008) Simultaneous inference in general parametric models. Biom J 50(3):346–363. https://doi.org/10.1002/bimj.200810425
Ikenaga M, Sakai M (2014) Application of locked nucleic acid (LNA) oligonucleotide-PCR clamping technique to selectively PCR amplify the SSU rRNA genes of bacteria in investigating the plant-associated community structures. Microbes Environ 29(3):286–295. https://doi.org/10.1264/jsme2.ME14061
Jardillier L, Zubkov MV, Pearman J, Scanlan DJ (2010) Significant CO2 fixation by small prymnesiophytes in the subtropical and tropical northeast Atlantic Ocean. ISME J 4(9):1180–1192. https://doi.org/10.1038/ismej.2010.36
Karkare S, Bhatnagar D (2006) Promising nucleic acid analogs and mimics: characteristic features and applications of PNA, LNA, and morpholino. Appl Microbiol Biotechnol 71(5):575–586. https://doi.org/10.1007/s00253-006-0434-2
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(4):772–780. https://doi.org/10.1093/molbev/mst010
Le Bescot N, Mahé F, Audic S, Dimier C, Garet M-J, Poulain J, Wincker P, de Vargas C, Siano R (2016) Global patterns of pelagic dinoflagellate diversity across protist size classes unveiled by metabarcoding. Environ Microbiol 18(2):609–626. https://doi.org/10.1111/1462-2920.13039
Leadbeater BSC, Manton I (1969a) Chrysochromulina camella sp. nov. and C. cymbium sp. nov., two new relatives of C. strobilus Parke and Manton. Arch Mikrobiol 68(2):116–132. https://doi.org/10.1007/bf00413872
Leadbeater BSC, Manton I (1969b) New observations on the fine structure of Chrysochromulina strobilus Parke and Manton with special reference to some unusual features of the haptonema and scales. Arch Mikrobiol 66(2):105–120. https://doi.org/10.1007/bf00410218
Lin Y-C, Campbell T, Chung C-C, Gong G-C, Chiang K-P, Worden AZ (2012) Distribution patterns and phylogeny of marine stramenopiles in the North Pacific Ocean. Appl Environ Microbiol 78(9):3387–3399. https://doi.org/10.1128/aem.06952-11
Logares R, Audic S, Santini S, Pernice MC, de Vargas C, Massana R (2012) Diversity patterns and activity of uncultured marine heterotrophic flagellates unveiled with pyrosequencing. ISME J 6(10):1823–1833. https://doi.org/10.1038/ismej.2012.36
López-García P, Philippe H, Gail F, Moreira D (2003) Autochthonous eukaryotic diversity in hydrothermal sediment and experimental microcolonizers at the mid-Atlantic ridge. Proc Natl Acad Sci U S A 100(2):697–702. https://doi.org/10.1073/pnas.0235779100
Machida RJ, Knowlton N (2012) PCR primers for metazoan nuclear 18S and 28S ribosomal DNA sequences. PLoS One 7(9):e46180. https://doi.org/10.1371/journal.pone.0046180
Massana R (2011) Eukaryotic picoplankton in surface oceans. Annu Rev Microbiol 65:91–110. https://doi.org/10.1146/annurev-micro-090110-102903
Moon-van der Staay SY, De Wachter R, Vaulot D (2001) Oceanic 18S rDNA sequences from picoplankton reveal unsuspected eukaryotic diversity. Nature 409(6820):607–610. https://doi.org/10.1038/35054541
O'Rorke R, Lavery S, Jeffs A (2012) PCR enrichment techniques to identify the diet of predators. Mol Ecol Resour 12(1):5–17. https://doi.org/10.1111/j.1755-0998.2011.03091.x
Orsi W, Song YC, Hallam S, Edgcomb V (2012) Effect of oxygen minimum zone formation on communities of marine protists. ISME J 6(8):1586–1601. https://doi.org/10.1038/ismej.2012.7
Parris DJ, Ganesh S, Edgcomb VP, DeLong EF, Stewart FJ (2014) Microbial eukaryote diversity in the marine oxygen minimum zone off northern Chile. Front Microbiol 5(543). https://doi.org/10.3389/fmicb.2014.00543
Peano C, Lesignoli F, Gulli M, Corradini R, Samson MC, Marchelli R, Marmiroli N (2005) Development of a peptide nucleic acid polymerase chain reaction clamping assay for semiquantitative evaluation of genetically modified organism content in food. Anal Biochem 344(2):174–182. https://doi.org/10.1016/j.ab.2005.04.009
Pernice MC, Giner CR, Logares R, Perera-Bel J, Acinas SG, Duarte CM, Gasol JM, Massana R (2016) Large variability of bathypelagic microbial eukaryotic communities across the world's oceans. ISME J 10(4):945–958. https://doi.org/10.1038/ismej.2015.170
Puchooa D (2004) A simple, rapid and efficient method for the extraction of genomic DNA from lychee (Litchi chinensis Sonn.) Afr J Biotechnol 3(4):253–255. https://doi.org/10.5897/AJB2004.000-2046
Reñé A, Camp J, Garcés E (2015) Diversity and phylogeny of Gymnodiniales (Dinophyceae) from the NW Mediterranean Sea revealed by a morphological and molecular approach. Protist 166(2):234–263. https://doi.org/10.1016/j.protis.2015.03.001
Sauvadet A-L, Gobet A, Guillou L (2010) Comparative analysis between protist communities from the deep-sea pelagic ecosystem and specific deep hydrothermal habitats. Environ Microbiol 12(11):2946–2964. https://doi.org/10.1111/j.1462-2920.2010.02272.x
Schloss PD, Gevers D, Westcott SL (2011) Reducing the effects of PCR amplification and sequencing artifacts on 16s rRNA-based studies. PLoS One 6(12):e27310. https://doi.org/10.1371/journal.pone.0027310
Seyama T, Ito T, Hayashi T, Mizuno T, Nakamura N, Akiyama M (1992) A novel blocker-PCR method for detection of rare mutant alleles in the presence of an excess amount of normal DNA. Nucleic Acids Res 20(10):2493–2496. https://doi.org/10.1093/nar/20.10.2493
Sherr EB, Sherr BF (1994) Bacterivory and herbivory: key roles of phagotrophic protists in pelagic food webs. Microb Ecol 28(2):223–235. https://doi.org/10.1007/BF00166812
Stoeck T, Behnke A, Christen R, Amaral-Zettler L, Rodriguez-Mora MJ, Chistoserdov A, Orsi W, Edgcomb VP (2009) Massively parallel tag sequencing reveals the complexity of anaerobic marine protistan communities. BMC Biol 7(1):1–20. https://doi.org/10.1186/1741-7007-7-72
Team RC (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Vaulot D, Eikrem W, Viprey M, Moreau H (2008) The diversity of small eukaryotic phytoplankton (<= 3 mu m) in marine ecosystems. FEMS Microbiol Rev 32(5):795–820. https://doi.org/10.1111/j.1574-6976.2008.00121.x
Vestheim H, Deagle B, Jarman S (2011) Application of blocking oligonucleotides to improve signal-to-noise ratio in a PCR. In: Park DJ (ed) PCR Protocols. Methods in molecular biology, 687. Humana Press, New York, pp 265–274
Vestheim H, Jarman SN (2008) Blocking primers to enhance PCR amplification of rare sequences in mixed samples—a case study on prey DNA in Antarctic krill stomachs. Front Zool 5:12. https://doi.org/10.1186/1742-9994-5-12
von Wintzingerode F, Landt O, Ehrlich A, Göbel UB (2000) Peptide nucleic acid-mediated PCR clamping as a useful supplement in the determination of microbial diversity. Appl Environ Microbiol 66(2):549–557. https://doi.org/10.1128/aem.66.2.549-557.2000
Watanabe MM, Suda S, Inouya I, Sawaguchi T, Chihara M (1990) Lepidodinium viride gen. et sp. nov. (Gymnodinaiales, Dinophyta), a green dinoflagellate with a chlorophyll a- and b-containing endosymbiont. J Phycol 26(4):741–751. https://doi.org/10.1111/j.0022-3646.1990.00741.x
Yi XY, Zhang H, Liu GX (2014) Primers to block the amplification of symbiotic apostome ciliate 18S rRNA gene in a PCR-based copepod diet study. Chin J Oceanol Limnol 32(3):515–521. https://doi.org/10.1007/s00343-014-3129-z
Yu D, Mukai M, Liu Q, Steinman CR (1997) Specific inhibition of PCR by non-extendable oligonucleotides using a 5′ to 3′ exonuclease-deficient DNA polymerase. BioTechniques 23(4):714–716 718-20
Zubkov MV, Tarran GA (2008) High bacterivory by the smallest phytoplankton in the North Atlantic Ocean. Nature 455(7210):224–226. https://doi.org/10.1038/nature07236
Acknowledgements
We thank Dr. Wenxiong Wang for providing Tigriopus japonicus, Jian Wang for Apocyclops royi, Kailin Liu for Thalassiosira weissflogii, Ying Ke for Bodoridae sp. and Caecitellas sp., Chongping Li for the help with DNA extraction of the pure cultures and Shun Yan Cheung for collecting seawater samples and extracting the DNA.
Funding
This work was financially supported by the National Basic Research Program (“973” Program) of China through the grant number 2009CB421203, the Natural Science Foundation of China (41330961), and the Research Grants Council of Hong Kong RGF grants (661912, 661813 and 16128416).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
No specific permissions were required for collecting the samples. This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Tan, S., Liu, H. Unravel the hidden protistan diversity: application of blocking primers to suppress PCR amplification of metazoan DNA. Appl Microbiol Biotechnol 102, 389–401 (2018). https://doi.org/10.1007/s00253-017-8565-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8565-1