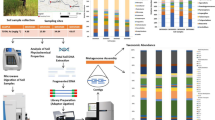
Abstract
In recent years, the role of microorganisms inhabiting rice rhizosphere in promoting arsenic contamination has emerged. However, little is known concerning the species and metabolic properties involved in this phenomenon. In this study, the influence of water management on the rhizosphere microbiota in relation to arsenic dissolution in soil solution was tested. Rice plants were cultivated in macrocosms under different water regimes: continuous flooding, continuous flooding with a 2-week period drainage before flowering, and dry soil watered every 10 days. The active bacterial communities in rhizosphere soil and in rhizoplane were characterized by 16S rRNA pyrosequencing. An in-depth analysis of microbial taxa with direct or indirect effects on arsenic speciation was performed and related contribution was evaluated. Continuous flooding promoted high diversity in the rhizosphere, with the plant strongly determining species richness and evenness. On the contrary, under watering the communities were uniform, with little differences between rhizosphere soil and rhizoplane. Arsenic-releasing and arsenite-methylating bacteria were selected by continuous flooding, where they represented 8% of the total. On the contrary, bacteria decreasing arsenic solubility were more abundant under watering, with relative abundance of 10%. These values reflected arsenic concentrations in soil solution: 135 μg L−1 and negligible in continuous flooding and under watering, respectively. When short-term drainage was applied before flowering, intermediate conditions were achieved. This evidence strongly indicates an active role of the rhizosphere microbiota in driving arsenic biogeochemistry in rice paddies, influenced by water management, explaining amounts and speciation of arsenic often found in rice grains.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Arsenic contamination of groundwater resources and soils represents an issue in many areas of the world (Singh et al. 2015; Heikens 2006). However, arsenic (As) speciation and the physicochemical characteristics of the environment determine its bioavailability more than its concentrations. In rice fields, the prolonged flooding usually adopted for cultivation leads to As release from soil minerals with the consequent accumulation of the metalloid in the grains (Zhu et al. 2014; Sun et al. 2014; Ma et al. 2014). Recent studies revealed that, on average, As content in rice from different countries exceeds the law limits established by the Commission regulation (EU) 2015/1006 (Ma et al. 2014; EFSA 2009, 2014).
The two inorganic As species mainly present in rice field soil, arsenate [As(V)] and arsenite [As(III)], have different biogeochemical properties. Nevertheless, both As species show high affinity for iron oxides/hydroxides (Meharg and Zhao 2012; Martin et al. 2014).
Continuous flooding in paddy soils leads to strongly reduced conditions, with the consequent rapid dissolution of these minerals and the release of As into the pore water (Zhu et al. 2014; Meharg and Zhao 2012). Furthermore, As(V) is reduced to As(III) abiotically by sulfide, ferrous iron [Fe(II)], H2 or reduced organic acids, or by As(V)-reducing bacteria (Cavalca et al. 2013; Meharg and Zhao 2012). Several studies reported that As(III) in flooded rice fields is the predominant As species (Zhu et al. 2014; Takahashi et al. 2004). At very low redox potentials, where microbial sulfate reduction is favored, sulfide produced by this activity can co-precipitate with As(III) forming a variety of minerals, such as orpiment (As2S3) (Zhu et al. 2014; Kocar and Fendorf 2009). If soil is aerated, for example after a drainage period, As(III) can be oxidized to As(V) by oxygen, manganese oxides, and H2O2 as well as by microbial As(III) oxidation (Meharg and Zhao 2012).
The genetic properties and the encoded enzymatic systems that allow several groups of microorganisms to resist to high As concentrations or to use As for metabolic purposes have been recently reviewed by various authors (Andres and Bertin 2016; Zhu et al. 2014; Yamamura and Amachi 2014; Cavalca et al. 2013; Zheng et al. 2012; Slyemi and Bonnefoy 2012). Interestingly, in rice paddy soils with low As concentration, a high diversity of microbial genes for As processing has been detected (Xiao et al. 2016), indicating the potential role of native communities on As transformations beyond abiotic factors. Among these processes, the microbial methylation of As(III) in rice rhizosphere is receiving great attention in the last few years. Recent studies indicated that rice roots microbiome is entirely responsible for the production of methylated As present in rice grains, which accounts for 50% of total As content (Lomax et al. 2012; Arao et al. 2011; Zhao et al. 2013). Furthermore, continuous flooding of rice fields has been demonstrated to increase the concentration of methylated As in rice grains (Ma et al. 2014; Li et al. 2009a, b).
In addition to direct As transformations, several metabolic activities of microorganisms could indirectly influence As speciation and bioavailability in the environment. Given the above-mentioned affinity of As for iron and sulfide minerals, microorganisms involved in iron and sulfur cycles could promote either the release or the sequestration of As from the pore water of rice paddies. Dissimilatory iron-reducing bacteria (DFeRB) use ferric iron [Fe(III)] as electron acceptor for anaerobic respiration, contributing to the release of As from iron minerals (Lee 2013). Conversely, iron-oxidizing bacteria (FeOB) are chemolithoautotrophic bacteria that couple the oxidation of Fe(II) to the reduction of a variety of electron acceptors (Emerson 2012; Hedrich et al. 2011; Emerson et al. 2010). With their activity, these bacteria promote the co-precipitation of As with iron minerals. As already stated, dissimilatory sulfate-reducing bacteria (DSRB) are strict anaerobes that reduce sulfate (SO4 2−) to sulfide (HS−) for their energy metabolism (Rabus et al. 2015; Ramel et al. 2015; Pester et al. 2012; Pereira et al. 2011), potentially contributing to the formation of As2S3 in anoxic compartments of rice fields soils. On the other hand, a variety of sulfur-oxidizing bacteria (SOB) can oxidize HS− to SO3 2−, and/or the latter to SO4 2−, leading to the release of As into the pore water in rice paddies (Stubner et al. 1998; Friedrich et al. 2005; Hamilton et al. 2015; Dahl et al. 2008).
The recent instructions established by the Commission regulation (EU) 2015/1006 concerning rice consumption in relation to As exposure have arisen great concern in the most important European rice-producing countries like Italy. Although the scientific community has often focused the attention on As contamination of rice, especially in Asia, very little is known about the role of different rhizospheric microbial populations and the microbial metabolic processes that drive As biogeochemistry. In this study, the bacterial community inhabiting the rhizosphere of rice plants cultivated with different water regimes in an unpolluted soil has been investigated in order to identify specific populations responsible for As contamination of rice grains.
Materials and methods
Experimental setup
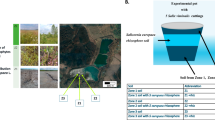
Nine rice paddy macrocosms containing 10–15 rice plants (Oryza sativa subsp. japonica, variety Loto) each were set up in tanks filled with rice field soil (sandy-loam texture; pH 6.0; 11.4 mg kg−1 of total As content and 33.1 g kg−1 of aqua regia extractable Fe). Three replicates for each macrocosm were managed either under continuous flooding (CF), under continuous flooding with a 2-week period of drainage before flowering (CF-D), or in dry soil with watering nearly every 7–10 days, or when the soil was excessively dry, always taking care to maintain aerobic conditions in soil all over the cropping season (D) (Fig. 1). After 12 days from germination, CF and CF-D macrocosms were flooded and D were watered. CF-D treatments were drained 47 days after germination for 14 days, followed by re-flooding. Then, they were re-flooded until sampling. All samplings were performed at the flowering stage. At this time point, the CF macrocosms were still flooded, whereas the CF-D and the D macrocosms were re-flooded the previous day. Before re-flooding, CF-D macrocosms underwent the 14-day period of drainage.
Chemical analyses of pore water
Three replicates of pore water samples were obtained from each macrocosm using Rhizon soil moisture samplers (Rhizosphere®, Rhizosphere Research Products, Wageningen, NL). In D macrocosms, pore water was sampled the day after watering, in order to allow the restoration of soil-solution equilibria and, at the same time, to have the possibility to obtain a solution sample. The Eh was measured in the soil right before sampling (portable pH/mV Measuring Instrument pH197i, WTW, Weinheim, Germany; equipped with a WTW SenTix® ORP electrode). The concentration of total As in the pore water samples was quantified with HG-AAS (Perkin-Elmer 4100 equipped with a FIAS 400 hydride generator; Perkin-Elmer Inc., Waltham, MA). For As speciation in pore water and in rice grain, refer to the treatments labeled as CF, 2IED, and AR in Zecchin et al. (2017), corresponding to CF, CF-D, and D, respectively. Fe(II) was determined with the orthophenantroline method (Loeppert and Inskeep 1996); N-nitrate, P-phosphate, and S-sulfate were determined with ion chromatography [Dionex DX-500 system, AS9 analytical column, with AG9 pre-column (Dionex, Sunnyvale, CA, USA)].
Rhizosphere soil and rhizoplane separation
Three plants from each macrocosm replicate, for a total of nine plants for each water management, were sampled after 60 days from germination, at flowering stage. This stage was chosen because previous studies indicated that, in rice, As is mainly translocated during flowering (Zheng et al. 2011).
Immediately after sampling, samples were pooled in one composite sample for each treatment, according to Somenahally et al. (2011), and rhizosphere soil and rhizoplane were collected according to Cavalca et al. (2010). Rhizosphere soil was defined as the soil removed from roots after shaking (180 rpm) in tetrasodium pyrophosphate buffer (0.2%, pH 8.0) for 1 h at 30 °C. The rhizoplane fraction was the biomass suspension resulting after 3 cycles of sonication (30 s each) of roots, previously washed thoroughly with sterile distilled water and submerged in 1:2 ratio (w/v) in 1× phosphate buffer saline (PBS) solution. Immediately after separation, samples were stored at −80 °C. An aliquot of the original soil used for the experiment was also sampled as the time zero control (T0).
RNA isolation
Total RNA was isolated using the RNA PowerSoil® Total RNA Isolation Kit (MO BIO, Carlsbad, USA), according to manufacturer’s instructions. To remove residual genomic DNA from isolated RNA, 1 U of DNaseI (Thermo Fisher Scientific, Waltham, MA, USA) was added to 1 μg of RNA and each reaction was incubated according to manufacturer’s instructions. The purity of RNA was tested both via agarose gel electrophoresis and with PCR amplification of bacterial 16S rRNA genes (see details in the following subsection). The purified RNA was reverse transcribed with iScript™ cDNA Synthesis Kit (BIO-RAD, Hercules, CA, USA) according to manufacturer’s instructions.
Barcoded pyrosequencing of 16S rRNA
Pyrosequencing of 16S rRNA was performed from reverse-transcribed RNA isolated from the T0 soil and from rhizosphere soil and rhizoplane sampled during the reproductive phase. Bacterial 16S rRNA was amplified with the universal bacterial primers 27F (5′ - GAG AGT TTG ATC CTG GCT CAG - 3′) and 1495R (5′ - CTA CGG CTA CCT TGT TAC GA - 3′) from each sample in triplicate in a 25 μL reaction volume containing 10 ng of cDNA, 0.3 μM primers, and 1× Taq PCR Mastermix kit (QIAGEN, Hilden, Germany). The thermal incubation included a first denaturation at 95 °C for 5 min followed by 35 cycles of denaturation at 95 °C for 1 min, annealing at 55 °C for 40 s, and elongation at 72 °C for 1 min and 40 s; the final elongation was performed at 72 °C for 10 min. Replicated amplicons were pooled and purified with MinElute PCR Purification kit (QIAGEN) to a final concentration of 20 ng μL−1. Pyrosequencing was performed at Molecular Research LP (MRDNA, Shallowater, TX, USA) by bacterial Tag-Encoded FLX Amplicon Pyrosequencing (bTEFAP), using the primer 27F. The sequences were processed and analyzed with the QIIME tools (Caporaso et al. 2010a). Sequences with less than 200 bases of with barcodes or primer biases, homopolymers, and chimeras were removed from the analysis. According to the quality scores, all sequences were trimmed at 400 bp. Operational taxonomic units (OTUs) were defined with a 97% similarity cutoff with the uclust method (Edgar 2010) using the last SILVA SSU Ref dataset (Quast et al. 2013) as reference database. Representative sequences for each OTU were aligned using the PyNAST method (Caporaso et al. 2010b). After taxonomic assignment, sequences belonging to chloroplasts were removed and OTU tables were generated for each sample. Phylogenetic analysis was performed using the FastTree method (Price et al. 2009). To measure the bacterial diversity within the samples, the OTU tables were rarefied and different indices of alpha diversity were calculated assuming a sample size of 2000. The OTU richness and the diversity within each sample was evaluated with different alpha diversity indices (observed species, Fisher’s alpha, ACE, Simpson evenness). To compare the bacterial diversity between the samples, principal coordinates analysis (PCoA) of the rarefied OTU tables was performed calculating unweighted and weighted UniFrac distances (Hamady and Knight 2009). The significance of the differences emerged with the beta diversity was tested with the multi-response permutation procedure (MRPP; Mielke et al. 1976). To distinguish between core and rare taxa, the rank abundance was plotted. Statistically significant differences of single OTUs among different water managements were evaluated with one-way analysis of variance (ANOVA) at p < 0.05 with Bonferroni’s correction.
Predictive microbial arsenic, iron and sulfur processing profiling
In order to point out the bacterial populations involved in arsenic, iron, and sulfur cycles, a reference database was specifically constructed on the bases of information available in the literature on bacterial strains with experimentally demonstrated metabolisms related to these elements. For taxa included in the database, the presence of genes related to arsenic, iron, and sulfur metabolisms was checked in the NCBI database. Genera were selected for their documented capacity to reduce As(V) as an electron acceptor [dissimilatory As(V)-reducing bacteria, DAsRB], to resist to arsenic with different mechanisms (arsenic resistant bacteria, AsRB), to oxidize As(III) [As(III)-oxidizing bacteria, AsOB], or to methylate As(III) [As(III)-methylating bacteria, AsMB]. The metabolisms that indirectly influence arsenic dynamics considered in this analysis were those of: dissimilatory Fe(III)-reducing bacteria (DFeRB), Fe(II)-oxidising bacteria (FeOB), dissimilatory SO4 2−-reducing bacteria (DSRB), and sulfur-oxidizing bacteria (SOB).
Accession numbers
All the 16S rRNA sequences retrieved in this study are deposited in the NCBI Bioproject (https://www.ncbi.nlm.nih.gov/bioproject/) PRJNA353766.
Results
Effect of agronomic management on pore water chemistry and rice grain contamination
At the flowering stage, relatively high concentrations of As and Fe(II) were dissolved in the pore water in the CF macrocosms (Table 1), indicating reduced conditions induced by continuous flooding. The prevalent As form was As(V) comprising 88% of the total As in pore water. Organic arsenic forms were not detected (Zecchin et al. 2017). In CF-D and D, both dissolved As and Fe(II) were almost negligible at the considered sampling date, proving that the recent drainage of the CF-D macrocosms had been effective in restoring an oxidative environment in soil and that the watering of the D macrocosms the day before sampling did not induce any appreciable reductive dissolution of Fe and As. In the CF macrocosms, the average redox potential measured in the three replicates was below −200 mV, whereas in the CF-D and D macrocosms the values were above zero (data not shown). Nitrate and sulfate, which are reducible species, showed different patterns, being more abundant in the aerobic rice test D and in the just drained macrocosm CF-D, compared with the flooded ones. The concentration of dissolved phosphate, as expected, remained comparable in the different treatments. Total arsenic content in rice grains varied significantly according to the water regime: 237 ± 38 μg kg−1 in CF, 68 ± 4 μg kg−1 in CF-D, and 5 ± 3 μg kg−1 in D. Methylated As forms represented 46% of total As in CF, and 18% in CF-D and they were negligible in D (Zecchin et al. 2017).
Ecology of active microbiota in rice rhizosphere
Sequencing of 16S rRNA produced 230,791 reads. The average length of reads with quality score above 25 was 408 bp; therefore, the sequence region beyond the nucleotide position 400 was removed in all reads. The total number of sequences that passed the quality control for each sample and the related number of OTUs are listed in Table 2.
Different indexes for alpha diversity (ACE, Simpson evenness, observed species) were calculated and compared (Fig. 1). The expected number of species in all rhizosphere soil samples was similar to the T0 soil. The rhizoplane samples of CF and CF-D plants showed the highest species richness (Fig. 2a), whereas the rhizoplane of D was the sample with the lowest species richness, evidencing a plant effect induced by flooding. Simpson’s evenness in all samples was below 0.5, indicating the predominance of specific groups among the whole community. In the rhizoplane of CF plants, species were more evenly distributed with respect to all the other samples. In the rhizoplane of CF-D plants, although species richness was similar to CF plants, evenness was lower. The bacterial community of the rhizosphere soil of CF plants was the most heterogeneous. The rarefaction analysis performed on the observed species confirmed the ACE trend, with the diversity within the samples decreasing as follows: CF rhizoplane < CF-D rhizoplane < CF rhizosphere soil < T0 < D rhizosphere soil < CF-D rhizosphere soil < D rhizoplane (Fig. 2b).
Alpha diversity evaluated with ACE index and Simpson’s evenness measure, assuming a sample size of 2000 reads (a) and by observed species on rarefied samples (b). Values are shown for the original soil (T0), rhizosphere soil, and rhizoplane treated under continuous flooding (CF RS and CF RP), under CF with 14 days of drainage before flowering (CF-D RS and CF-D RP), and under watering every 10 days (D RS and D RP)
Driving forces of microbial community differentiation
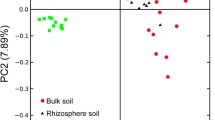
According to the PCoA analysis, the bacterial communities developed in both compartments of D were similar to each other, clustering in groups III and V according to unweighted and weighted UniFrac, respectively (Fig. 3a, d). On the contrary, the CF and CF-D treatments led to a significant differentiation between rhizosphere soil and the rhizoplane. Interestingly, the communities selected in CF and CF-D rhizoplanes were similar, clustering in groups I and IV according to unweighted and weighted UniFrac, respectively (Fig. 3a, d). In CF and in CF-D rhizosphere soils, the phylogenetic composition of the community was similar to the community in T0 (Fig. 3a, d). However, when considering the relative abundances of the taxa, CF-D rhizosphere soil samples were more similar to CF and CF-D rhizoplanes (Fig. 3d). The water regime and the compartment by themselves did not influence significantly the bacterial communities developed in the different samples (Fig. 3b, c, e and f, A: 0.08, 0.03, 0.26, and 0.11, respectively). More likely, a combination of factors drove the differentiation of the communities, which was best reflected by groups IV, V, and VI (A: 0.38, p < 0.001).
Beta diversity analysis of all treatments, included original soil (T0), rhizosphere soil, and rhizoplane treated with continuous flooding (CF RS and CF RP), with 14 days of drainage before flowering (CF-D RS and CF-D RP), and with watering every 10 days (D RS and D RP). The principal coordinate analysis was performed calculating unweighted (a, b, c) and weighted UniFrac distances (d, e, f) according to Hamady and Knight (2009). The significance of the groups determined by the sample identity (a, d), the water regime (b, e), and the root-soil compartment (c, f) was tested with multi-response permutation procedure (MRPP)
Phylogenetic composition of the different communities
In total 40,193 OTUs at genus level were found in all samples. Two opposite trends were observed concerning the number of OTUs exclusively present after each treatment in rhizosphere soil and in the rhizoplane (Fig. 4). In rhizosphere soil, 25.8, 30.5, and 30.9% of the total number of OTUs were exclusively present in CF, CF-D, and D, respectively (Fig. 4a). On the contrary, in rhizoplane they were 33.6, 30.6, and 22.6%, respectively (Fig. 4b). In rhizosphere soil, CF-D shared 4.6% of OTUs with D treatments, with respect to 3% with CF. In the rhizoplane, CF-D shared 7.2% of the OTUs with CF treatments, compared to 1.7% in common with D.
A total of 33 phyla were detected in the samples. The number of phyla in the T0 soil was higher with respect to all rhizosphere compartments of rice cultivated with different water regimes (Table 2). In both compartments, the number of phyla decreased from CF to CF-D to D. On the basis of the rank abundance plot, taxa with relative abundance below 0.01% were considered as part of the rare biosphere (Online Resource, Fig. S1a). According to this definition, the fraction of rare phyla was higher in the T0 soil, followed by both compartments of CF plants, both compartments of CF-D plants, and both compartments of D plants (Online Resource, Fig. S1b).
The percentage of sequences that could not be assigned to any known phylum ranged between 2.3 and 6.9% (Fig. 5). In the T0 soil, the phyla Acidobacteria, Actinobacteria, and Proteobacteria were the most abundant, accounting for 9.13, 30.69, and 36.72% of the bacterial community, respectively (Fig. 5a). In CF rhizosphere soil, the abundance of Proteobacteria decreased to 25.10%, with the concomitant increase of Acidobacteria and Actinobacteria, which accounted for 12.61 and 37.14% of the total, respectively. In the rhizoplane under CF, Actinobacteria represented only 2.52%, whereas Proteobacteria and Acidobacteria accounted for 54.43 and 20.83%, respectively. In CF rhizosphere soil, Actinobacteria belonging to an uncultured genus of the order Gaiellales were significantly more abundant (p < 0.05) with respect to the other treatments, whereas in the rhizoplane the same microorganisms were more abundant in D (Fig. 5b, Online Resource, Table S3). The genera Marmoricola and Nocardioides also contributed with 3.8 and 1.9%, respectively. In CF rhizoplane, Acidobacteria of the order DA023 and Candidatus Chloracidobacterium were represented by 7.81 and 4.21% of the sequencing library, respectively (Fig. 5b). Bacteria belonging to the class Deltaproteobacteria were selected by CF, accounting for 4.5 and 6% of the total community in CF rhizosphere soil and rhizoplane (Fig. 5a). Iron-reducing Deltaproteobacteria belonging to the genera Anaeromyxobacter and Geobacter were the most contributors for this class in CF rhizosphere soil and rhizoplane, respectively (Online Resource, Table S2).
Relative abundance (%) of phyla and proteobacterial classes (a) and genera above 1% of relative abundance (b) retrieved in the treatments: initial soil (T0), continuously flooded rhizosphere soil and rhizoplane (CF RS and CF RP), with 14 days of drainage before flowering (CF-D RS and CF-D RP) and with watering every 10 days (D RS and D RP)
In the rhizosphere soil and in the rhizoplane under CF-D, the phylum Proteobacteria accounted respectively for 82.34 and 68.10% of all bacterial phyla (Fig. 5a). In CF-D rhizosphere soil, this phylum was the only one with abundance above 10%, whereas in CF-D rhizoplane the phylum Acidobacteria contributed with 17.78%. The classes Alphaproteobacteria and Betaproteobacteria were mainly responsible for the dominance of the phylum Proteobacteria in both compartments. In CF-D rhizosphere soil, members of the class Betaproteobacteria accounted for more than 45%, with the Comamonadaceae family accounting for 38% of the total bacterial community (Fig. 5b, Online Resource, Table S2). Within this family, Ramlibacter, Piscinibacter, and other unknown genera represented 11, 3.3, and 21% of the whole community, respectively. In CF-D rhizoplane, on the other hand, several members of the class Alphaproteobacteria made up almost 50% of the total community (Fig. 5a). Within these members, the genus Sphingomonas accounted for 17% in both compartment.
In both compartments of D, the two most abundant phyla were Proteobacteria and Actinobacteria, respectively, accounting for 53.68 and 31.31% in D rhizosphere soil and 60.56 and 25.25% in D rhizoplane (Fig. 5a). Within Proteobacteria, the classes Alphaproteobacteria, Betaproteobacteria, and Gammaproteobacteria were the most abundant. In D rhizosphere soil, they accounted for 30.3, 6.8, and 15.6%, respectively, with Sphingomonas responsible for 15.2% of the total community. In the D rhizoplane, the three above-mentioned classes accounted for 39.2, 11.2, and 9.5%. In this compartment, Sphingomonas and Variovorax represented 28 and 6.8% of the total community, respectively. Within the phylum Actinobacteria, the genus Arthrobacter accounted for 14.3 and 5.9% in D rhizosphere soil and D rhizoplane, respectively. Different OTUs belonging to this genus were significantly more abundant (p < 0.05) in D with respect to the other water regimes (Online Resource, Table S3). In D rhizoplane, the genera Marmoricola and Nocardioides also contributed with 2.1 and 2.9%, respectively.
Bacterial populations potentially involved in arsenic, iron, and sulfur cycles
In order to predict the bacterial community functional profiles, although potential, related to As, Fe, and S processing, a survey was conducted in the literature to search for taxa that were experimentally demonstrated to be involved in those biogeochemical cycles or in whose genomes the presence of genetic markers was displayed (Online Resource, Table S1).
In the T0 soil, bacteria able to resist or to process As, either by reduction, oxidation, or methylation, accounted for 6.39% of the bacterial community (Online Resource, Fig. S2a). Within these, 58.96% were putative AsMB, 23.83% were AsOB, 12.35% were putatively ars/ACR operons-carrying species (AsRB), and 4.85% DAsRB. Bacteria involved in iron and sulfur cycle represented 1.4% of the total: within these, 22.34% were DFeRB, 35.55% were FeOB, 3.61% were DSRB, and 38.5% were SOB (Online Resource, Fig. S2b).
DAsRB, DFeRB, and DSRB were significantly more abundant in rhizosphere soil and in the rhizoplane of plants cultivated under CF with respect to CF-D and D (Fig. 6). In the rhizoplane, DAsRB and DFeRB accounted for more than 2% of the total community, whereas DSRB represented 0.1% of the community. Among DAsRB and DFeRB, Geobacter and Bacillus genera were significantly higher in CF and CF-D water regimes. The same trend was observed for DFeRB of the genus Anaeromyxobacter. Among DSRB, Desulfobacteraceae varied significantly being more abundant in the rhizosphere soil of CF plants (Table 3).
Relative abundance (% ± SD) of species potentially able to process arsenic directly (a) or indirectly as a consequence of their metabolism (b) in rhizosphere soil and rhizoplane of plants cultivated under continuous flooding (CF RS and CF RP), under continuous flooding with 14 days drainage before flowering (CF-D RS and CF-D RP) or with watering every 10 days (D RS and D RP). The metabolic groups considered in this analysis were dissimilatory As(V)-reducing bacteria (DAsRB), As-resistant bacteria (AsRB), As(III)-oxidizing bacteria (AsOB) and As(III)-methylating bacteria (AsMB), dissimilatory Fe(III)-reducing bacteria (DFeRB), Fe(II)-oxidizing bacteria (FeOB), dissimilatory SO4 2−-reducing bacteria (DSRB), and sulfur-oxidizing bacteria (SOB)
AsOB were significantly more abundant in D plants of both root compartments (Fig. 5). AOB accounted for 10% of the total community in D plants, whereas in CF and CF-D plants they were always overall below 3%. Rhizobium, Variovorax, Pseudomonas, and Stenotrophomonas significantly contributed to this variation. FeOB mirrored this trend in rhizospheric soil, whereas in the rhizoplane they were not affected by the agronomic managements and they were always below 1%. FeOB as Thermomonas and Pseudomonas were more abundant in D, whereas Leptothrix, Rhodobacter, and Aquabacterium were more abundant in CF (Table 3). Differently from As- and Fe-oxidizing bacteria, SOB as Rhodovolum and Anaeromyxibacter were significantly more abundant in CF plants in both root compartments, doubling from 1.49% in rhizosphere soil to 2.88% in the rhizoplane (Fig. 6), while Aurantimonas was the only SOB genus significantly higher in D (Table 3).
The abundance of AsRB in rhizosphere soil was included between 2.6 and 3.4%, without significant variations due to the water regimes (Fig. 6). This was reflected by the opposite trends of relative abundance of different genera, like Pseudomonas (stimulated in D) and Anaeromyxibacter (higher in CF) (Table 3). In the rhizoplane, on the contrary, AsRB accounted for 3.91% of the total in CF plants, with respect to 1.14 and 0.72% in CF-D and D plants.
Similarly, AsMB were present in rhizosphere soil with abundances between 1.93 and 4.97%, with no significant variations among the three water managements. In the rhizoplane, AsMB were significantly more abundant in CF plants (4.64%) than in CF-D (2.66%) and D (1.11%) plants (Fig. 6).
Discussion
Continuous flooding lead to the dissolution of As and Fe in the CF macrocosms, as expected according to previous studies (Yamaguchi et al. 2011; Borch et al. 2010; Takahashi et al. 2004), following the well-assessed reductive dissolution of Fe and Mn oxides and the reduction of As(V) to the more soluble As(III). The temporary establishment of an oxic environment in the CF-D soils was followed by very low As and Fe concentrations in solution, comparable to those encountered in the D ones, assessing a fast chemical response of the system to the water management. The mobilization of As in pore water in CF agronomic schemes has been recently proven to lead to As enrichment of rice grains, if compared to the CF-D and D ones (Zecchin et al. 2017; Spanu et al. 2012).
The species richness and evenness of the samples, the PCoA analysis, and the core microbiome analysis reflected the chemical characteristics determined by the different water regimes and indicated a strong influence of the presence of the plant in CF and CF-D agronomic conditions. The rhizosphere of CF and CF-D plants reflected an anoxic environment, where strictly anaerobic species were selected. By contrast, in the rhizoplane of these plants, the release of organic matter and oxygen by the roots likely promoted a higher diversity. In D rhizosphere, the proximity to the plant did not lead to a sharp differentiation in the bacterial community, possibly linking the lower development of the roots and the aerenchyma in such condition to a lower turn-over of oxygen and carbon sources, and consequently the lack of a chemical gradient (Suralta and Yamauchi 2008). In rhizosphere soil of CF and CF-D treatments, populations with similar phylogenetic affiliation were selected during the vegetative phase, whereas the 2-week drainage period led to a differentiation in the abundances but not in the phylogenetic structure of the community (Fig. 3). In rhizosphere soil of CF-D plants, the stress induced by sharp short-term changes in the redox conditions could have selected for more versatile species in common with D. In the rhizosphere of D, less species where stimulated, probably indicating a lower degree of electron acceptors restoration given by either the absence of a redox interface or by the fact that these plants do not grow under optimal conditions (i.e., continuous flooding).
The three dominant phyla found in all the samples, i.e., Actinobacteria, Acidobacteria, and Proteobacteria, resembled what commonly found in plants rhizosphere (Bulgarelli et al. 2013). Actinobacteria represented a significant part of the original community of the rice paddy soil used for this experiment, contrary to what seen in previous studies carried out on rice paddy soils from different locations (Edwards et al. 2015; Wörner et al. 2016). These microorganisms are common soil inhabitants and plant commensals (Ventura et al. 2007). Most of them, including members of the genera Marmoricola and Nocardioides and of the order Gaiellales, are aerobic and degrade a variety of complex polysaccharides deriving from the plant (Barka et al. 2016; Kügler et al. 2015; Urzì et al. 2000; Lee 2007; Dastager et al. 2008; Lee and Lee 2010; Lee et al. 2011; Yoon and Park 2006; Albuquerque et al. 2011). The genus Arthrobacter, quite abundant in rhizosphere soil of D plants, is commonly found in soils with neutral pH. Members of this genus are versatile concerning carbon source and highly resistant to environmental stress like aridity (Jones and Keddie 2006). The high abundance of members of the order Gaiellales together with Fe(III)- and SO4 2−-reducing genera in the rhizosphere of CF plants indicates the presence of microhabitats with different levels of oxygen and electron acceptors. Members of the phylum Acidobacteria are commonly found in soils as well as in rhizosphere soil (Bulgarelli et al. 2013; Ward et al. 2009). As a confirmation of these outcomes, in previous studies these organisms were found to be more abundant in bulk and rhizosphere soil with respect to the rhizoplane (Edwards et al. 2015). They are usually aerobic, capable of nitrate and nitrite reduction, heterotrophs, able to degrade complex substrates, and tolerant to variation of soil humidity (Ward et al. 2009).
Members of the Proteobacteria phylum were favored by the proximity to the plant, where the release of C compounds is higher and determines the prevalence of r-strategist bacteria (Edwards et al. 2015). These microorganisms have often been found more abundant in the rhizoplane and in the endosphere (Edwards et al. 2015) with respect to the bulk of rice field soil. The high abundance observed in CF-D for Alphaproteobacteria and Betaproteobacteria might be due to the decrease of members of the other taxa, less resistant to the alternation of wet/dry periods. Shrestha et al. (2009) also observed that members belonging to these two classes were more active in oxic paddy soil. The genus Ramlibacter sp., including aerobic heterotrophs, often isolated from soils, has been demonstrated to be resistant to dryness stress (Lee et al. 2014). The genus Sphingomonas, strictly aerobic and facultative photoorganotroph (Yabuuchi and Kosako 2005), found its ideal habitat in the rhizoplane of D. Piscinibacter are described as chemoheterotrophs and facultative aerobic (Song and Cho 2007; Stackebrandt et al. 2009). Deltaproteobacteria are more frequently detected in anoxic rice paddies (Shrestha et al. 2009; Pester et al. 2012; Lu et al. 2006). According to these observations, in this study members of this class were more abundant in CF rhizosphere compartments.
Most of the above-mentioned genera, with high relative abundance in the different treatments, are not known to have arsenic-processing capacities. This aspect probably reflects the fact that the soil used for this experiment did not contain high As concentration. Therefore, As was not the main factor shaping the bacterial communities in this environment.
Nowadays, apart from metagenomic analysis, tools to predict functional profiling of microbial systems relay on 16S rRNA data associated to databases using marker gene data and reference genomes (Langille et al. 2013). The central idea is that despite various important forms of microbial genome plasticity (gene loss, duplication, or gene transfer), the genes present in microbial genomes are much more similar among related bacteria or archaea than distant relatives. In line with this approach, the database built in the presence study was used to assess potential functionalities related to arsenic/iron/sulfur biogeochemical cycles. Such data might be useful in future studies to focus the attention on specific microbial taxa when considering metagenomic libraries. Although this approach can indicate only potential functions in a microbial community, the trends observed were consistent with the soil physical-chemical conditions in the different water managements and with previous data based on marker gene quantification (Zecchin et al. 2017).
In this study, we observed that DSRB were more abundant in rhizosphere soil of CF if compared to rhizoplane, whereas DAsRB and DFeRB in the same treatment were closer to the roots. The observed pattern, with DFeRB more abundant than DAsRB, probably reflects the redox scale predicted by Kocar and Fendorf (2009), who demonstrated that at pH < 6.5 Fe(III)-reduction is favored over dissimilatory As(V)-reduction. Members of the genera Geobacter and Anaeromyxobacter were confirmed to be common Fe(III)-reducing inhabitants of anoxic paddy fields, promoted by a CF water regime (Hori et al. 2010; Shrestha et al. 2009; Treude et al. 2003).
The best habitat for FeOB was the rhizosphere of D plants. This could be due to a sharp redox interface in CF and CF-D, with only little areas with the optimal concentration of Fe(II) and O2, and a wider microoxic area in D rhizosphere. Similarly, populations of AsOB were more abundant in D rhizosphere, confirming what observed in previous studies (Das et al. 2016). The presence of both FeOB and AsOB might contribute to the conversion of As(III) to As(V) and its co-precipitation with Fe oxides. Conversely, in CF DFeRB predominate over FeOB. Therefore, the process of dissolution of Fe oxides and consequent release of As is promoted over its precipitation.
Genera of SOB were more abundant in the rhizoplane of CF plants, in contrast with what observed by Das et al. (2016). The rhizoplane of CF plants could be an optimal habitat for SOB since reduced sulfur compounds are produced by DSRB but, at the same time, little amounts of oxygen needed by these organisms are released by the roots. It has often been reported that SOB are related to ecosystems characterized by sharp opposing gradients of O2 and reduced sulfur compound (Dahl et al. 2008). Considering that SOB potentially promote sulfide minerals dissolution, their presence in CF rhizosphere might contribute to As release from sulfide minerals. Microbial mineral weathering for nutrient acquisition has already been reported as a cause of As release to soil solution from apatite (Mailloux et al. 2009). These evidences strongly suggest that these microorganism-mediated processes affect As translocation to rice grains in accordance with previous studies (Zecchin et al. 2017; Somenahally et al. 2011), thus posing a health issue.
Although in the rhizosphere soil of all the treatments AsRB were equally represented, in the rhizoplane these organisms were more abundant in CF plants. This was probably a consequence of the fact that arsenic was strongly released in that region, where iron oxides and sulfide minerals are present and probably dissolved by DFeRB and SOB. In the rhizoplane, the bacterial diversity was generally lower than in the rhizosphere soil. In the latter, the higher diversity included microorganisms generally present in soil characterized by an average distribution of As resistance.
Arsenic-methylating bacteria were present in rice rhizosphere. Particularly, in CF plants this activity was selected and probably enhanced by the presence of As(III) and organic matter, which are the substrates for this reaction. This is in accordance with recent evidence that under CF more organic As is produced with respect to sprinkler irrigation and aerobic rice (Moreno-Jiménez et al. 2014; Li et al. 2009a, b). Organic As constituted almost 50% of total As in rice grains in CF condition in the same experimental set up (Zecchin et al. 2017). Furthermore, recent evidence indicates that organic As accumulated in rice grains derives from microbial methylation carried out in the rhizosphere (Lomax et al. 2012). Among versatile bacteria related to arsenic reduction or methylation and to Fe dissimilative reduction, Geobacter, rare in the T0 soil according to rank abundance analysis (Online Resource, Fig. S1), significantly increased in CF and CF-D rhizosphere soils and rhizoplanes, but not in D water regime. Previous data of Geobacter-specific gene quantification (Zecchin et al. 2017) are in accordance with the barcoded sequence data obtained in the present work, thus evidencing a putative role of Geobacter in As release from soil mineral to pore water and in As methylation. In accordance with the increase of DAsRB and AsMB in CF and CF-D water regimes, higher As(III) concentrations were detected in the corresponding soil pore water and higher methylated As was detected in the corresponding rice grains.
Together, these outcomes indicate a dramatic effect of water management of rice paddies in shaping the rhizosphere microbiota. Continuous flooding promotes the proliferation of As-releasing bacterial taxa, whereas in aerobic rice microorganisms that reduce the solubility of As in the soil solution are favored. Introducing a 2-week drainage period before the flowering stage within a continuous flooding regime leads to intermediate relative abundances of As-affecting bacteria. A decrease in the flooding intensity might be helpful for the selection of an As-stabilizing microbial community with the reduction of bioavailable As concentration in the soil solution, thus decreasing As contamination in rice grains.
References
Albuquerque L, França L, Rainey FA, Schumann P, Nobre MF, da Costa MS (2011) Gaiella occulta gen. nov., sp. nov, a novel representative of a deep branching phylogenetic lineage within the class Actinobacteria and proposal of Gaiellaceae fam. nov. and Gaiellales ord. nov. System Appl Microbiol 34:595–599
Andres J, Bertin PN (2016) The microbial genomics of arsenic. FEMS Microbiol Reviews 39:1–24
Arao T, Kawasaki A, Baba K, Matsumoto S (2011) Effects of arsenic compound amendment on arsenic speciation in rice grain. Environ Sci Technol 45:1291–1297
Barka EA, Vatsa P, Sanchez L, Gaveau-Vallant N, Jacquard C, Klenk HP, Clément C, Ouhdouch Y, van Wezel GP (2016) Taxonomy, physiology, and natural products of Actinobacteria. Microbiol Mol Biol Reviews 80:1–43
Borch T, Kretzschmar R, Kappler A, Van Cappellen P, Ginder-Vogel M, Voegelin A, Campbell K (2010) Biogeochemical redox processes and their impact on contaminant dynamics. Environ Sci Technol 44:15–23
Bulgarelli D, Schlaeppi K, Spaepen S, van Themaat EVL, Schulze-Lefert P (2013) Structure and functions of the bacterial microbiota of plants. Annu Rev Plant Biol 64:807–838
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Gonzalez Peña A, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010a) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010b) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26(2):266–267
Cavalca L, Zanchi R, Corsini A, Colombo M, Romagnoli C, Canzi E, Andreoni V (2010) Arsenic-resistant bacteria associated with roots of the wild Cirsium arvense (L.) plant from an arsenic polluted soil, and screening of potential plant growth-promoting characteristics. Syst Appl Microbiol 33:154–164
Cavalca L, Corsini A, Zaccheo P, Andreoni V, Muyzer G (2013) Microbial transformations of arsenic: perspectives for biological removal of arsenic from water. Future Microbiol 86:753–768
Commission E (2015) Commission regulation (EC) No 2015/1006 of 25 June 2015 amending Regulation (EC) No 1881/2006 as regard maximum levels of inorganic arsenic in foodstuffs. OJ L 161:14–16
Dahl C, Friedrich C, Kletzin A (2008) Sulfur oxidation in prokaryotes. In: Encyclopedia of life sciences (ELS). Wiley, Chichester
Das S, Chou ML, Jean JS, Liu CC, Yang HJ (2016) Water management impacts on arsenic behavior and rhizosphere bacterial communities and activities in a rice agro-ecosystem. Sci Total Environ 542:642–652
Dastager SG, Lee J, Ju Y, Park D, Kim C (2008) Marmoricola bigeumensis sp. nov., a member of the family Nocardioidaceae. Int J Syst Evol Micr 58:1060–1063
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2480–2461
Edwards J, Johnson C, Santos-Medellín C, Lurie E, Podishetty NK, Bhatnagar S, Eisen JA, Sundaresan V (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. PNAS 112:911–920
Emerson D (2012) Biogeochemistry and microbiology of microaerobic Fe(II) oxidation. Biochem Soc Trans 40:1211–1216
Emerson D, Fleming EJ, McBeth JM (2010) Iron-oxidizing bacteria: an environmental and genomic perspective. Ann Rev Microbiol 64:561–583
European Food Safety Authority (2009) Scientific opinion on arsenic in food. EFSA J 7:1–1351
European Food Safety Authority (2014) Dietary exposure to inorganic arsenic in the European population. EFSA J 12:3597
Friedrich CG, Bardischewsky F, Rother D, Quentmeier A, Fischer J (2005) Prokaryotic sulfur oxidation. Curr Opin Microbiol 8:253–259
Hamady M, Knight R (2009) Microbial community profiling for human microbiome projects: tools, techniques and challenges. Genome Res 19:1141–1152
Hamilton TL, Jones DS, Schaperdoth I, Macalady JL (2015) Metagenomic insight into S(0) precipitation in a terrestrial subsurface lithoautotrophic ecosystem. Front Microbiol 5:1–16
Hedrich S, Schlömann M, Johnson DB (2011) The iron-oxidizing proteobacteria. Microbiology 157:1551–1564
Heikens A (2006) Arsenic contamination of irrigation water, soil and crops in Bangladesh: risk implications for sustainable agriculture and food safety in Asia. RAP Publication (FAO), Bangkok
Hori T, Müller A, Igarashi Y, Conrad R, Friedrich MW (2010) Identification of iron-reducing microorganisms in anoxic rice paddy soil by 13C-acetate probing. ISME J 4:267–278
Jones D, Keddie RM (2006) The genus Arthrobacter. Prokaryotes 3:945–960
Kocar BD, Fendorf S (2009) Thermodynamic constraints on reductive reactions influencing the biogeochemistry of arsenic in soils and sediments. Environ Sci Technol 43:4871–4877
Kügler JH, Le Roes-Hill M, Syldatk C, Hausmann R (2015) Surfactants tailored by the class Actinobacteria. Frontier Microbiol 6:1–23
Langille MGI, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, Knight R, Beiko RG, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31(9):814–821
Lee SD (2007) Marmoricola aequoreus sp. nov., a novel actinobacterium isolated from a marine sediment. Int J Syst Evol Microbiol 57:1391–1395
Lee S (2013) Enhancement of arsenic mobility by Fe(III)-reducing bacteria from iron oxide minerals. J Mater Cycles Waste 15:362–369
Lee DW, Lee SD (2010) Marmoricola scoriae sp. nov., isolated from volcanic ash. Int J Syst Evol Microbiol 60:2135–2139
Lee SD, Lee DW, Ko Y (2011) Marmoricola korecus sp. nov. Int J Syst Evol Microbiol 61:1628–1631
Lee HJ, Lee SH, Lee S, Lee JS, Kim Y, Kim S, Joen CO (2014) Ramlibacter solisilvae sp. nov., isolated from forest soil, and emended description of the genus Ramlibacter. Int J Syst Evol Microbiol 64:1317–1322
Li R, Ago Y, Liu W, Mitani N, Feldmann J, McGrath SP, Ma JF, Zhao F (2009a) The rice aquaporin Lsi1 mediates uptake of methylated arsenic species. Plant Physiol 150:2071–2080
Li RY, Stroud JL, Ma JF, McGrath SP, Zhao FJ (2009b) Mitigation of arsenic accumulation in rice with water management and silicon fertilization. Environ Sci Technol 43:3778–3783
Loeppert RH, Inskeep WP (1996) Iron. In: Sparks DL, Page AL, Helmke PA, Loeppert RH, Soltanpour PN, Tabatabai MA, Summer ME (eds) Methods of soil analysis. Part 3. Chemical methods. Soil Science Society of America, Madison, pp 639–663
Lomax C, Liu WJ, Wu L, Xue K, Xiong J, Zhou J, McGrath SP, Meharg AA, Miller AJ, Zhao FJ (2012) Methylated arsenic species in plants originate from soil microorganisms. New Phytol 193:665–672
Lu YH, Rosencrantz D, Liesack W, Conrad R (2006) Structure and activity of bacterial community inhabiting rice roots and the rhizosphere. Environ Microbiol 8:1351–1360
Ma R, Shen J, Wu J, Tang Z, Shen Q, Zhao FJ (2014) Impact of agronomic practices on arsenic accumulation and speciation in rice grain. Environ Pollut 194:217–223
Mailloux BJ, Alexandrova E, Keimowitz AR, Wovkulich K, Freyer GA, Herron M, Stolz JF, Kenna TC, Pichler T, Polizzotto ML, Dong H, Bishop M, Knappett PSK (2009) Microbial mineral weathering for nutrient acquisition releases arsenic. Appl Environ Microbiol 75:2558–2565
Martin M, Violante A, Ajmone-Marsan F, Barberis E (2014) Surface interactions of arsenite and arsenate on soil colloids. Soil Sci Soc Am J 78:157–170
Meharg AA, Zhao F (2012) Biogeochemistry of arsenic in paddy environments. In: Arsenic & Rice. Springer, Berlin, pp 71–101
Mielke PW, Berry KJ, Johnson ES (1976) Multi-response permutation procedures for a priori classifications. Commun Stat Theor Methods 5:1409–1424
Moreno-Jiménez E, Meharg AA, Smolders E, Manzano R, Becerra D, Sánchez-Llerena J, Albarrán Á, López-Piñero A (2014) Sprinkler irrigation of rice fields reduces grain arsenic but enhances cadmium. Sci Total Environ 485-486:468–473
Pereira IAC, Ramos AR, Grein F, Marques MC, da Silva SM, Venceslau SS (2011) A comparative genomic analysis of energy metabolism in sulfate reducing bacteria and archaea. Front Microbiol 2:1–22
Pester M, Knorr K, Friedrich MW, Wagner M, Loy A (2012) Sulfate-reducing microorganisms in wetlands—fameless actors in carbon cycling and climate change. Front Microbiol 3:1–19
Price MN, Dehal PS, Arkin AP (2009) FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol 26:1641–1650
Quast C, Pruesse E, Ylmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:590–596
Rabus R, Venceslau SS, Wöhlbrand L, Voordouw G, Wall JD, Pereira IAC (2015) A post-genomic view of the ecophysiology, catabolism and biotechnological relevance of sulphate-reducing prokaryotes. Adv Microb Physiol 66:55–321
Ramel F, Brasseur G, Pieulle L, Valette O, Hirschler-Réa A, Fardeau ML, Dolla A (2015) Growth of the obligate anaerobe Desulfovibrio vulgaris Hildenborough under continuous low oxygen concentration sparging: impact of the membrane-bound oxygen reductases. PLoS One 10:1–17
Shrestha PM, Kube M, Reinhardt R, Liesack W (2009) Transcriptional activity of paddy soil bacterial communities. Environ Microbiol 11:960–970
Singh R, Singh S, Parihar P, Singh VP, Prasad SM (2015) Arsenic contamination, consequences and remediation techniques: a review. Ecotoxicol Environ Saf 112:247–270
Slyemi D, Bonnefoy V (2012) How prokaryotes deal with arsenic. Environ Microbiol Rep 4:571–586
Somenahally AC, Hollister EB, Loeppert RH, Yan W, Gentry TJ (2011) Microbial communities in rice rhizosphere altered by intermittent and continuous flooding in fields with long-term arsenic application. Soil Biol Biochem 43:1220–1228
Song J, Cho J (2007) Methylibium aquaticum sp. nov., a betaproteobacterium isolated from a eutrophic freshwater pond. Int J Syst Evol Microbiol 57:2125–2128
Spanu A, Daga L, Orlandoni AM, Sanna G (2012) The role of irrigation techniques in arsenic bioaccumulation in rice (Oryza sativa L.) Environ Sci Technol 42:8333–8340
Stackebrandt E, Verbarg S, Frühling A, Busse H, Tindall BJ (2009) Dissection of the genus Methylibium: reclassification of Methylibium aquaticum as Piscinibacter aquaticus gen. nov., comb. nov. and Methylibium subsaxonicum as Rivibacter subsaxonicus gen. nov., comb. nov. and emended descriptions of the genera Rhizobacter and Methylibium. Int J Syst Evol Microbiol 59:2552–2560
Stubner S, Wind T, Conrad R (1998) Sulfur oxidation in rice field soil: activity, enumeration, isolation and characterization of thiosulfate-oxidizing bacteria. System Appl Microbiol 21:569–578
Sun L, Zheng M, Liu H, Peng S, Huang J, Cui K, Nie L (2014) Water management practices affect arsenic and cadmium accumulation in rice grains. Scientific World J 2014:Article ID 596438
Suralta RR, Yamauchi A (2008) Root growth, aerenchyma development, and oxygen transport in rice genotype subjected to drought and waterlogging. Environ Exp Bot 64:75–82
Takahashi Y, Minamikawa R, Hattori KH, Kurishima K, Kihou N, Yuita K (2004) Arsenic behavior in paddy fields during the cycle of flooded and non-flooded periods. Environ Sci Technol 38:1038–1044
Treude N, Rosencrantz D, Liesack W, Schnell S (2003) Strain FAc12, a dissimilatory iron-reducing member of the Anaeromyxobacter subgroup of Myxococcales. FEMS Microbiol Ecol 44:261–269
Urzì C, Salamone P, Schumann P, Stackebrandt E (2000) Marmoricola aurantiacus gen. nov., sp. nov., a coccoid member of the family Nocardioidaceae isolated from a marble statue. Int J Syst Evol Microbiol 50:529–536
Ventura M, Canchaya C, Tauch A, Chandra G, Fitzgerald GF, Chater KF, van Sinderen D (2007) Genomics of Actinobacteria: tracing the evolutionary history of an ancient phylum. Microbiol Mol Biol R 71:495–548
Ward NL, Challacombe JF, Janssen PH, Henrissat B, Coutinho PM, Wu M, Xie G, Haft DH, Sait M, Badger J, Barabote RD, Bradley B, Brettin TS, Brinkac LM, Bruce D, Creasy T, Daugherty SC, Davidsen TM, DeBoy RT, Detter JC, Dodson RJ, Durkin AS, Ganapathy A, Gwinn-Giglio M, Han CS, Khouri H, Paulsen I, Penn K, Ren Q, Rosovitz MJ, Selengut JD, Shrivastava S, Sullivan SA, Tapia R, Thompson LS, Watkins KL, Yang Q, Yu C, Zafar N, Zhou L, Kuske CR (2009) Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microbiol 75:2046–2056
Wörner S, Zecchin S, Dan J, Hristova Todorova N, Loy A, Conrad R, Pester M (2016) Gypsum amendment to rice paddy soil stimulated bacteria involved in sulfur cycling but largely preserved the phylogenetic composition of the total bacterial community. Environ Microbiol Rep 8:413–423
Xiao K, Li L, Ma L, Zhang S, Bao P, Zhang T, Zhu Y (2016) Metagenomic analysis revealed highly diverse microbial arsenic metabolism genes in paddy soils with low-arsenic contents. Environ Poll 211:1–8
Yabuuchi E, Kosako Y (2005) Sphingomonas. Bergey’s manual of systematics of archaea and bacteria. Wiley, Chichester
Yamaguchi N, Nakamura T, Dong D, Takahashi Y, Amachi S, Makino T (2011) Arsenic release from flooded paddy soils is influenced by speciation, Eh, pH, and iron dissolution. Chemosphere 83:25–932
Yamamura S, Amachi S (2014) Microbiology of inorganic arsenic: from metabolism to bioremediation. J Biosci Bioeng 1:1–9
Yoon J, Park Y (2006) The genus Nocardioides. PRO 3:099–1113
Zecchin S, Corsini A, Martin M, Romani M, Beone GM, Zanchi R, Zanzo E, Tenni D, Fontanella MC, Cavalca L (2017) Rhizospheric iron and arsenic bacteria affected by water regime: implications for metalloid uptake by rice. Soil Biol Biochem 106:129–137
Zhao FJ, Harris E, Yan J, Ma J, Wu L, Liu W, McGrath SP, Zhou J, Zhu YG (2013) Arsenic methylation in soils and its relationship with microbial arsM abundance and diversity, and As speciation in rice. Environ Sci Technol 47:7147–7154
Zheng MZ, Cai C, Hu Y, Sun GX, Williams PN, Cui HJ, Li G, Zhao FJ, Zhu YG (2011) Spatial distribution of arsenic and temporal variation of its concentration in rice. New Phytol 189:200–209
Zheng R, Sun G, Zhu Y (2012) Effects of microbial process on the fate of arsenic in paddy soil. Chinese Sci Bull 58:186–193
Zhu YG, Yoshinaga M, Zhao FJ, Rosen BP (2014) Earth abides arsenic biotransformation. Annu Rev Earth Planet Sci 42:443–467
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
The research was supported by Ministry of University and Research program PRIN (2010JBNLJ7-004). Sarah Zecchin was awarded of a PhD fellowship from the University of Milan, Doctorate School of Food Systems.
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any study with human participants or animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Electronic supplementary material
ESM 1
(PDF 460 kb)
Rights and permissions
About this article
Cite this article
Zecchin, S., Corsini, A., Martin, M. et al. Influence of water management on the active root-associated microbiota involved in arsenic, iron, and sulfur cycles in rice paddies. Appl Microbiol Biotechnol 101, 6725–6738 (2017). https://doi.org/10.1007/s00253-017-8382-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8382-6