Abstract
Trihalomethanes such as chloroform and bromoform, although well-known as a prominent class of disinfection by-products, are ubiquitously distributed in the environment due to widespread industrial usage in the past decades. Chloroform and bromoform are particularly concerning, of high concentrations detected and with long half-lives up to several hundred days in soils and groundwater. In this study, we report a Dehalobacter- and Desulfovibrio-containing co-culture that exhibits dehalogenation of chloroform (~0.61 mM) to dichloromethane and bromoform (~0.67 mM) to dibromomethane within 10–15 days. This co-culture was further found to dechlorinate 1,1,1-trichloroethane (1,1,1-TCA) (~0.65 mM) to 1,1-dichloroethane within 12 days. The Dehalobacter species present in this co-culture, designated Dehalobacter sp. THM1, was found to couple growth with dehalogenation of chloroform, bromoform, and 1,1,1-TCA. Strain THM1 harbors a newly identified reductive dehalogenase (RDase), ThmA, which catalyzes chloroform, bromoform, and 1,1,1-TCA dehalogenation. Additionally, based on the sequences of thmA and other identified chloroform RDase genes, ctrA, cfrA, and tmrA, a pair of chloroform RDase gene-specific primers were designed and successfully applied to investigate the chloroform dechlorinating potential of microbial communities. The comparative analysis of chloroform RDases with tetrachloroethene RDases suggests a possible approach in predicting the substrate specificity of uncharacterized RDases in the future.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Trihalomethanes are the primary class of disinfection by-products in water treatment systems and have been used in large quantities as solvents and intermediates for chemical manufacturing (ATSDR 1989; ATSDR 1997; ATSDR 2005). A large amount of trihalomethanes have been released into the environment and subsequently distributed into groundwater and anoxic zones of aquifers through wastewater effluent discharge, extensive industrial utilization, and inappropriate disposal. The US Environmental Protection Agency has identified chloroform, bromoform, chlorodibromomethane, and dichlorobromomethane in at least 783, 136, 167, and 237 of 1322 National Priorities List sites, respectively (ATSDR 2013). This intensive environmental distribution has raised public concerns because trihalomethanes are known to depress the central nervous system and cause liver, kidney, or even fatal damage to mammals (ATSDR 1989; ATSDR 1997; ATSDR 2005). Additionally, chloroform is a potent inhibitor of many vital microbial processes, including some bioremediation processes which may be quenched at concentrations as low as 300 μg/L (Maymo-Gatell et al. 2001; Weathers and Parkin 2000). However, limited information is available about the fate of trihalomethanes in the polluted environment.
The detected maximum concentrations of trihalomethanes, especially for chloroform and bromoform, often exceed drinking water regulations set by the World Health Organization. For example, concentrations of chloroform as high as 21,800 μg/L have been detected in a leachate from a solid waste landfill, almost two orders of magnitude higher than the regulatory limit of 300 μg/L (ATSDR 1997). Similarly, bromoform was detected in water samples from the Southampton Water estuary at concentrations ranging from 10 to 2597 μg/L, significantly higher than the regulatory limit of 100 μg/L (ATSDR 1997; World Health Organisation 2004). Among available remediation strategies, microbial degradation has been shown to be an effective strategy to treat halogenated contaminants. Biological transformation of chloroform has been observed both co-metabolically, in methanogenic and sulfate-reducing environments, and metabolically, in proton-reducing environments, whereas little is known about the microbial transformation of bromoform (Bagley and Gossett 1995; McClay et al. 1996; Oldenhuis et al. 1991). However, the microbes which co-metabolically degrade chloroform usually show low tolerance to chloroform and require the addition of ancillary growth substrates (Oldenhuis et al. 1991). These severely restrict the application of a biological process in highly contaminated sites (Duhamel et al. 2002). Metabolic degradation can overcome the low tolerance of microbes capable of co-metabolic chloroform transformation. Metabolic degradation of chloroform can be achieved through stepwise reductive dechlorination, resulting in the production of dichloromethane. Only a few bacterial cultures, belonging to the genera Dehalobacter and Desulfitobacterium, have been found to be involved in such stepwise reductive dechlorination (Ding et al. 2014; Grostern et al. 2010; Lee et al. 2012).
In this study, we report the identification of a novel Dehalobacter sp. strain THM1 in a co-culture that metabolically dehalogenates chloroform, bromoform, and 1,1,1-trichloroethane (1,1,1-TCA). A reductive dehalogenase, ThmA, responsible for chloroform, 1,1,1-TCA, and bromoform dehalogenation, was identified in strain THM1. The obtainment of thmA and the comparative study conducted on thmA and other three chloroform RDase genes have assisted the design of a gene marker to indicate the chloroform dechlorinating potential of microbial communities and further demonstrated a possible approach in predicting the substrate specificity of uncharacterized RDases.
Materials and methods
Establishment of microcosms
A total of 20 anaerobic microcosms were established with sediments or sludge samples and spiked with trihalomethanes for dehalogenating activity tests, among which samples showing trihalomethane dehalogenating activities are indicated as the following: SN*, mangrove sediment, Singapore; CWRP, anaerobic digester in wastewater treatment plant, Singapore; IVP, anaerobic tank in sewage treatment plant, Singapore; and MR, Marina Bay sediment, Singapore. Another five environmental samples were collected from river sediment, tropical forest, e-recycling sites, and wastewater treatment plants from both China and Singapore, which were used to test the newly designed chloroform RDase gene primers. Microcosm studies were conducted in 60-mL serum bottles containing ~10 g of sediment/sludge and 25 mL autoclaved bicarbonate-buffered mineral salt medium, as previously described (He et al. 2007). Bottles were sealed with butyl rubber septa (Geo-Microbial Technologies, Inc., Ochelata, OK, USA) and secured with aluminum crimp caps. Microcosms were spiked with trihalomethane (CHCl3, CHBrCl2, CHBr2Cl, or CHBr3) as the electron acceptor (~24 μmol/bottle) as well as lactate (10 mM) as the carbon and electron donor source. Abiotic controls were established by autoclaving the microcosms prepared in the same way as above. All bottles were incubated in the dark at 30 °C without shaking.
Chemicals, culture, and growth conditions
All chemical reagents were purchased from Sigma-Aldrich (St. Louis, MO) or Merck (Darmstadt, Germany) at the highest purity available (>98%). Mixed and co-culture SN were grown in 60-mL serum bottles containing 30 mL mineral salt medium spiked with acetate as the sole carbon source and amended with hydrogen as an electron donor (0.33 atm) and trihalomethane, as indicated, as an electron acceptor as previously described unless otherwise specified (He et al. 2007). Culture dechlorination time course studies were conducted in 100 mL mineral salt medium in 160-mL serum bottles with all other conditions as described above.
For the substrate tests of co-culture SN, chloroethenes (~0.3 mM each tetrachloroethene, trichloroethene, trans-dichloroethene, cis-dichloroethene, 1,1-dichloroethene, and vinyl chloride), chloroethanes (~0.3 mM each of 1,1,1-trichloroethane, 1,1,2-trichloroethane, 1,2-dichloroethane, and 1,1-dichloroethane), chlorophenols (0.05 mM each of pentachlorophenol, 2,4,6-trichlrophenol, and 2,4,5-trichlorophenol), polybrominated diphenyl ethers (PBDEs) (octa-BDE technical mixture and penta-BDE technical mixture which include BDEs in the range from tetra-BDE to nona-BDE as previously described) (Ding et al. 2013), the polychlorinated biphenyl (PCB) mixture Aroclor 1260 (10 ppm), and trihalomethanes (~0.3 mM each of chloroform, bromoform, chlorodibromomethane, and dichlorobromomethane) were added in acetate/H2-amended medium. The culture medium in serum bottles was spiked with neat chloroethenes, chloroethanes, and trihalomethanes. Chlorophenols and PCBs were dissolved in hexane with stock concentrations of 100 mM and 50,000 ppm, respectively. The penta-BDE technical mixture was dissolved in ethyl acetate (500 ppm), and the octa-BDE technical mixture was dissolved in n-nonane (3100 ppm). All time course studies were performed in biological triplicate and with abiotic controls (without bacterial inocula or with autoclaved cultures). Cultures were incubated in the dark at 30 °C without shaking for 3 months before chemical analyses.
Molecular analyses
Clone library
Clone libraries of 16S ribosomal RNA (rRNA) genes of mixed cultures SN were established using the pGEMT Easy Vector Systems (Promega) according to the manufacturer’s instructions. The DNA for constructing a clone library of each culture was extracted in the late dehalogenation stage (~20% substrate left), and 72 clones in total were picked in each clone library. Plasmids were extracted using Plasmid Miniprep Kit (OMEGA). Restriction fragment length polymorphism (RFLP) analysis was performed on the extracted plasmid DNA using the restriction endonucleases HhaI and MspI (NEB, Ipswich, MA, USA). The 16S rRNA gene inserts representing distinct enzyme restriction patterns were subsequently sequenced by Integrated DNA Technology, Singapore. Sequences were aligned using the BioEdit assembly software package (http://www.mbio.ncsu.edu/BioEdit/bioedit.html) and were analyzed with BLASTn (http://www.ncbi.nlm.nih.gov/). The 16S rRNA gene sequences of interesting strains were aligned by ClustalW, and phylogenetic trees were constructed with Molecular Evolutionary Genetics Analysis 6 (MEGA6) using the Maximum Likelihood methods supported by 1000 bootstraps (Tamura et al. 2013).
PCR-denaturing gradient gel electrophoresis
Genomic DNA was extracted using the Qiagen DNeasy Blood and Tissue Kit (Qiagen® GmbH, Hilden, Germany) according to the manufacturer’s instructions. PCR of bacterial 16S rRNA genes for denaturing gradient gel electrophoresis (DGGE) analysis was carried out using the primer pair 341GC and 518R with a touchdown thermal program as previously described (Duhamel et al. 2004). PCR amplicons were separated by DGGE using a denaturing gradient ranging from 20 to 80% for 12 h at 120 V and 60 °C (U.K. Laemmli 1970).
Quantitative real-time PCR
qPCR was conducted on an ABI 7500 Fast System using the SYBR Green reporter (Lo-ROX Kit, SensiFAST™ SYBR, Bioline). The thermocycler program was initiated at 95 °C for 3 min, followed by 40 cycles of 15 s at 94 °C and 30s at 55–60 °C, which depends on the annealing temperature of primers, and then 30 s at 72 °C. Plasmids carrying targeted genes were used to serve as qPCR standards. These qPCR standards were constructed using a pGEM-T Easy Vector system (Promega) according to the manufacturer’s instruction and were later extracted using QIAprep Spin Miniprep Kit (Qiagen GmbH). Linear standard curves (amplification efficiencies from 90 to 110% and R 2 > 98%) were established with concentrations ranging from 1.0 × 101 to 1.0 × 109 copies/reaction. Dehalobacter-specific primers (Dhb441F and Dhb645R), Desulfovibrio-specific primers (Dsv691F and Dsv826R), and 16S rRNA gene-specific primers (338F and 518R) were utilized to monitor cell growth (Fite et al. 2004; Smits et al. 2004).
Proteomic analyses and enzyme assays
Native polyacrylamide gels (native-PAGE) composed of 5% stacking gel and 10% resolving gel were prepared as previously described (U.K. Laemmli 1970), except that protein denaturation was prevented by running the native-PAGE in an ice box and removing sodium dodecyl sulfate (SDS) from all buffers. Cells were harvested by centrifugation (12,000 rpm, 20 min, 4 °C) and re-suspended in degassed Tris-HCl buffer (100 mM, pH 7.0). The cells were lysed by ultra-sonication using a VCX130 sonicator (Vibra-Cell, Sonics) (130 W; 20% duty cycle; for 3 min). The crude protein (supernatant partial) was harvested from the lysate after centrifugation at 12,000 rpm for 5 min. The membrane proteins were further extracted from crude protein via ultra-centrifugation at 120,000 rpm for 1 h at 4 °C. The pellets containing membrane proteins were solubilized in 10 mM CHAPS and concentrated by centrifugation through a membrane filter with a molecular cutoff of 5 kDa (Vivaspin 2, Sartorius Stedim Biotech). Membrane proteins of co-culture SN were obtained, and the native-PAGE electrophoresis was run as previously described, except that the gel was stained using the PlusOne Silver Staining Kit (Adrian et al. 2007). Briefly, the same protein samples were electrophoresed on two adjacent gel lanes, after which one lane was left unstained for further in vitro activity assays of RDase activity, while the other was silver stained to mark the position of protein bands in the unstained lane. The in vitro assays were carried out in an anaerobic chamber and initiated by addition of the gel pieces excised from the unstained lane corresponding to all the visible protein bands in the stained lanes, and the mixtures were incubated at 30 °C for 24 h prior to headspace analysis. In vitro activity assays were carried out in 4-mL vials containing 2 mL assay solution (2 mM methyl viologen; 1.5 mM titanium(III) citrate; 100 mM Tris-HCl buffer (pH 7.0)) and single halogenated substrate (chloroform, 1,1,1-TCA, and bromoform at 0.1 mM) as previously described (Adrian et al. 2007).
Prominent bands in the stained lanes corresponding to gel regions exhibiting chloroform, 1,1,1-TCA, and bromoform dehalogenating activity in the parallel non-stained bands were excised and sent for analysis at the Protein and Proteomics Centre, National University of Singapore. Briefly, after being washed, dehydrated, trypsin digested, extracted, and eluted, the samples were spotted on to the MALDI sample target plate. A MALDI TOF MS 4800 proteomics analyzer (AB Sciex) was used for mass spectrometric analysis. The membrane proteins collected from co-cultured SN spiked with 1,1,1-TCA or bromoform were trypsin digested and then analyzed by LC-MS/MS through TripleTOF 5600 System (AB Sciex). Briefly, the digested peptides were separated on Eksigent NanoLC Ultra and cHiPLC-nanoflex (Eksigent, Dublin, CA, USA) by a gradient formed by mobile phase A (2% acetonitrile, 0.1% formic acid) and mobile phase B (98% acetonitrile,0.1% formic acid): 5–12% mobile phase B in 20 min, 12–30% mobile phase B in 40 min, and 30–90% mobile phase B in 2 min, at a flow rate of 300 nL/min. The MS analysis was conducted on TripleTOF 5600 System (AB Sciex, Foster City, CA, USA). The MS/MS-based peptides and proteins were visualized using ProteinPilot software 4.5 (AB Sciex). Output signals of MALDI TOF MS and TripleTOF of 5600 MS were searched using the Mascot (v 2.1, Matrix Science Ltd., London, UK) in the National Center for Biotechnology Information (NCBInr) Protein Database using the parent ion mass with an error tolerance of 100 ppm and an MS/MS fragment mass tolerance of 0.2 Da for probable protein hits. Identification of peptides and proteins was validated if 95 and 99% probabilities were achieved by the Paragon Algorithm, respectively (Shilov et al. 2007).
Chemical analyses
Chloroethenes and chloroethanes concentrations were determined by headspace analyses using an Agilent gas chromatograph (GC7890) equipped with a flame ionization detector and a GS-GasPro column (30 m × 0.32 mm × 0.25 μm; J&W Scientific) as previously described (Zhao et al. 2015). Headspace samples of chloroethanes and chloroethenes were injected manually using a gas-tight glass syringe (Model Gastight ® #1725, Hamilton Co., Reno, Nevada). The temperature program was initially held at 50 °C for 2 min, increased at 30 °C/min to 220 °C, and held for 1 min. Trihalomethane concentrations were detected using an Agilent gas chromatograph (GC 6890) equipped with an electron capture detector and an HP-5 column (30 m × 0.32 mm × 0.25 μm, Agilent Technologies) by injecting headspace samples. The oven temperature was initially held at 35 °C for 1 min, increased to 150 °C at a rate of 30 °C/min, and held for 2 min. Chlorophenols, PCBs, and PBDEs were extracted by isooctane for further analyses. PCBs were tested on the same equipment used for trihalomethane detection. The oven program was initially set at 170 °C for 5 min, increased at 5 °C/min to 260 °C, and held for 5 min (Wang and He 2013). Chlorophenols and PBDEs were monitored on an Agilent gas chromatograph-mass spectrometer (GC 6890-MSD 5975) with an Rxi-5ms column (15 m × 0.25 mm × 0.25 μm; Restek, Bellefonte, PA, USA). The oven temperature of the GC-MS for PBDEs was initially set at 110 °C and was increased to 310 °C at a rate of 15 °C/min, and held at 310 °C for 3 min (Lee and He 2010). As for chlorophenol detection, the oven was programmed at an initial temperature of 40 °C, after which it was increased to 200 °C at a rate of 15 °C/min and held at 200 °C for 3 min. Helium was used as the carrier gas, with a column flow of 1.2 mL/min. Calibration curves were prepared using serum bottles containing defined mineral salt medium with known amounts of each compound, and the concentrations were calculated by total mass in both gas and liquid phases per bottle.
Nucleotide sequence accession number
The sequences were deposited in GenBank with accession numbers KX344912 for the 16S rRNA gene and KX344907 for the thmA gene of Dehalobacter sp. strain THM1.
Results
Dehalogenation of trihalomethanes by different microcosms
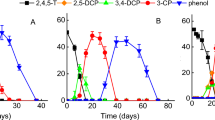
Among the 20 microcosms set up with inocula collected from different locations in Singapore, including wastewater treatment plants, mangroves, river sediments, and tropical forests, four showed dehalogenation of up to 50% of at least two types of trihalomethanes after 60 days of incubation. Only the microcosm established with mangrove sediment (SN*) was found to dechlorinate chloroform to dichloromethane (Table 1). More microcosms exhibited dehalogenation of trihalomethanes via single bromine removal, resulting in the formation of CH2Br2, CH2BrCl, and CH2Cl2 from CHBr3, CHBr2Cl, and CHBrCl2, respectively (Fig. 1). Additionally, CH3Br and CH3Cl were observed as minority products resulting from removal of two halogens from CHBr3 and CHBr2Cl, respectively. Although methane appeared in various amounts in each bottle, it is difficult to assign methane as the metabolite from dehalogenation of trihalomethanes regardless of the contribution from ubiquitous distribution of methanogens. Additionally, complete dehalogenation, producing methane as the sole end product, was not observed in the microcosms. In sterile controlled microcosms, less than 0.5% of abiotic degradation of chloroform occurred while all brominated trihalomethanes decreased from 8.8 to 11.8% throughout the study.
Enrichment and characterization of dehalogenation communities
Chloroform and bromoform are the more concerned pollutants among the four trihalomethanes (CHCl3, CHBrCl2, CHBr2Cl, or CHBr3) since they are produced and utilized in higher amounts. Among the 20 microcosms collected to test dehalogenation activity on trihalomethanes, microcosm SN was the only one showing dehalogenation on both chloroform and bromoform. Starting with microcosm SN, two subcultures, designated SN-CF and SN-BF, were enriched by more than 10 consecutive transfers in a mineral salt medium amended with acetate (10 mM) and hydrogen (0.33 ppmv) and spiked with either chloroform or bromoform (~0.4 mM). The clone library followed by RFLP of 16S rRNA genes of SN-CF showed three dominant populations, Clostridium, Dehalobacter, and Desulfovibrio, accounting for 86% of the total community (Fig. 2a). These same populations dominated the community of SN-BF, comprising 75% of the total. SN-BF also contained a fourth major population, Caulobacter, comprising 7% of the community. Dilution-to-extinction was performed to further enrich chloroform- and bromoform-dehalogenating bacteria in both cultures SN-CF and SN-BF. After three consecutive batches of serial dilutions, chloroform and bromoform dehalogenation activities were repeatedly detected in 10−7 dilutions. The 16S rRNA gene-based PCR-DGGE analysis showed the presence of only Dehalobacter and Desulfovibrio populations in both SN-CF and SN-BF (Fig. 2b). The roles of Clostridium and Caulobacter in dehalogenation were ruled out after the loss of the populations during serial dilution-to-extinction, showing no negative effect on chloroform dechlorination and bromoform debromination. Subsequently, both the SN-CF and SN-BF cultures were found to dehalogenate both chloroform and bromoform. The clone libraries yielded an identical restriction pattern for each culture SN-CF and SN-BF, each of which contained two distinct restriction patterns for the 72 clones after digestion with two enzymes, HhaI and MspI. Twenty clones from the clone library of each culture were randomly picked and sequenced. The sequencing results exhibited two distinct 16S rRNA gene sequences (with primer pair of 8F/1392R), and the bacterial populations were subjected to either Dehalobacter (1423 bp) or Desulfovibrio (1405 bp), which was in agreement with the microbial community analyses conducted by DGGE. This indicated that the culture contained two bacterial strains, and therefore, these identical co-cultures were designated SN for the following studies.
Characterization of chloroform/bromoform dehalogenating anaerobe
Co-culture SN dechlorinated chloroform (0.6 mM) to dichloromethane within 10 days at an average dechlorination rate of 61.0 ± 1.2 μmol L−1·day−1 (Fig. 3a) and also debrominated bromoform to dibromomethane within 15 days at an average debromination rate of 40.3 ± 2.1 μmol L−1 day−1 (Fig. 3b). The maximum dehalogenation rates reached as high as 123 μmol L−1 day−1 (as measured from day 6 to day 10) for chloroform and 43.6 μmol L−1 day−1 (as measured from day 3 to day 12) for bromoform. To discern whether Dehalobacter, Desulfovibrio, or both were responsible for dehalogenation of chloroform and bromoform, cell growth was monitored by qPCR during dehalogenation of each compound using genus-specific 16S rRNA gene primers. DNA samples from cultures without halogenated compound amendment were used as controls. During the dehalogenation process, the growth yields of Dehalobacter were estimated by monitoring cell numbers in late dehalogenation processes (>80% conversion) in different subcultures inoculated by various amounts of chloroform or bromoform. The growth yields of Dehalobacter were calculated to be (5.6 ± 0.7) × 107 16S rRNA gene copies per micromole of Cl− released from chloroform and (3.3 ± 0.8) × 107 16S rRNA gene copies per micromole of Br− released from bromoform. This chloroform-supported growth yield of Dehalobacter in co-culture SN is comparable with that of other Dehalobacter strains, ranging from (1.8 ± 0.1) × 107 to (12.3 ± 0.2) × 107 16S rRNA gene copies per micromole of Cl− released (Grostern et al. 2010; Sun et al. 2002). No cell growth of Dehalobacter was observed in the absence of halogenated compounds (Fig. 3d). Contrarily, growth of Desulfovibrio showed no correlation with chloroform or bromoform dehalogenation (Fig. 3). Therefore, the Dehalobacter in co-culture SN is responsible for dehalogenating chloroform and bromoform and was designated strain THM1.
A phylogenetic comparison of a nearly full-length 16S rRNA gene from Dehalobacter sp. strain THM1 with previously identified Dehalobacter strains and trihalomethyl group-dehalogenating anaerobes showed 99% identity (1418/1422 bp) between strain THM1 and its closet relative, a 2,4,6-trichlorophenol-dechlorinating species, Dehalobacter sp. AD14-TCP (Wang and He 2013). However, strain THM1 only showed 97% identity (1385/1424 bp) with other chloroform-dechlorinating strains, such as Dehalobacter CF and Dehalobacter UNSWDHB (Fig. 4) (Grostern et al. 2010; Wong et al. 2016).
Co-culture SN exhibited a narrow substrate utilization profile after a 3-month incubation with all tested electron acceptors, which included chloroethenes as well as specific members of chloroethanes, chlorophenols, PBDEs, and PCBs. In addition to the above described chloroform and bromoform, only 1,1,1-TCA was dechlorinated (to 1,1-DCA) by co-culture SN (Fig. 3c).
Identification of chloroform reductive dehalogenase gene in strain THM1
Membrane proteins from chloroform-fed co-culture SN were harvested during the exponential growth phase and separated by native-PAGE. An in vitro assay of chloroform dechlorination activity found dichloromethane production from only one band (Fig. 5a), with no activity from other fragments. This suggests that components of this band should be responsible for chloroform dechlorination. Additionally, this single band was found to dehalogenate 1,1,1-TCA to 1,1-DCA and bromoform to dibromomethane. The silver-stained counterpart of the gel fragment showing dehalogenating activity was analyzed by MALDI-TOF-MS. The comparison of high-accuracy MALDI peptide mass fingerprinting against the NCBInr database provided the most likely protein hits. The results demonstrated a protein sequence coverage with the chloroform RDases in Dehalobacter sp. strain CF (CfrA) and Desulfitobacterium sp. strain PR (CtrA) of 13.2 and 12.7%, respectively. Specific primers were designed based on the detected peptide sequences, and the amplified gene fragments were obtained by the clone library. From the clones, a partial sequence of the RDase gene was captured and the full sequence was obtained through genome walking. Given that the gel band containing chloroform RDase in strain THM1 was also found to dehalogenate bromoform and 1,1,1-TCA, the role of this RDase in bromoform debromination and 1,1,1-TCA dechlorination was investigated through proteomic analyses. A proteomic analysis of membrane proteins harvested from co-culture SN fed with bromoform or 1,1,1-TCA detected the chloroform RDase as the most abundant protein in both samples, sharing 55.7 and 51.5% coverage with chloroform RDase sequence in strain THM1. This further implicates this newly identified RDase’s multi-functional roles for chloroform, bromoform, and 1,1,1-TCA dehalogenation; thus, this trihalomethyl-dehalogenating RDase was designated ThmA.
a Native polyacrylamide gel electrophoresis profile of membrane proteins extracted from co-culture SN fed with chloroform. b Detection of chloroform RDase genes in microcosms using specific primer set cf159F/cf387R. Note: 1, 100 bp DNA ladder; 2, river sediment, Singapore; 3, tropical forest, Singapore; 4, e-recycling sites, China; 5, anaerobic digester, Singapore; 6, river sediment proximal to the outfalls of a chemical factory, China; 7, co-culture SN; 8, Desulfitobacterium sp. strain PR; 9, H2O
Chloroform RDase gene-specific primers and their application
Chloroform RDase gene-specific primers (cf159F 5′-GGAACGGTCAAGGAACTGGA-3′ and cf387R 5′-ATAATCAACGGCAAAGCTTG-3′) were designed in the conserved regions by alignment of known chloroform RDase genes (ctrA from Desulfitobacterium sp. strain PR, cfrA from Dehalobacter sp. strain CF, tmrA from Dehalobacter sp. strain UNSWDHB, and thmA from Dehalobacter sp. strain THM1). The specificity of the selected primers was primarily examined by comparison with two highly similar homologs (dcrA, of 97–98% similarity of the identified chloroform RDase genes, mediating 1,1-DCA to CA in Dehalobacter sp. strain DCA and an uncharacterized rdhA of 75% similarity with the identified chloroform RDase genes from Dehalobacter restrictus DSM5455) that catalyze dehalogenation of different halogenated substrates and later checked on NCBI Primer BLAST (Table 2). The newly designed chloroform RDase gene-specific primers were applied to five microcosms with inocula from different locations including river sediment, tropical forest, e-recycling sites, and anaerobic digester in Singapore and China. Among the five microcosms, only two, one from e-recycling sites, China, and the other from river sediment, China, showed chloroform dechlorination activity of 78 and 63% conversion, respectively, with an initial chloroform concentration of ~0.6 mM after a 2-month incubation. PCR amplification of DNA extracted from all five microcosms using cf159F/cf387R primers yielded amplicons of expected size and sequences in the two microcosms showing chloroform dechlorination activity (Fig. 5b). No amplicon was obtained in the three microcosms without chloroform dechlorination.
Discussion
In this study, a Dehalobacter sp. strain THM1 was identified to be able to dehalogenate trihalomethanes (i.e., chloroform and bromoform). Though only a few Dehalobacter strains have been identified, they exhibit versatile dehalogenation capabilities for a variety of halogenated compounds. Dehalobacter strains are known to dehalogenate chloroethenes (Holliger et al. 1998; Wild et al. 1997), chloroethanes (Grostern and Edwards 2009; Sun et al. 2002), chloroform (Grostern et al. 2010; Lee et al. 2012), chlorophenols (Wang et al. 2014), chlorobenzenes (Nelson et al. 2011), polybrominated diphenyl ethers (Robrock et al. 2008), and 4,5,6,7-tetrachlorophthalide (Yoshida et al. 2009) via organohalide respiration and ferment dichloromethane to acetate, hydrogen, and carbon dioxide (Justicia-Leon et al. 2012; Lee et al. 2012). Here, a co-culture containing Dehalobacter sp. strain THM1 was enriched from mangrove sediment, which exhibits distinct physiological and phylogenetic properties compared with previous identified Dehalobacter strains. Strain THM1 is the first known organohalide respiring bacterium to couple growth with reductive debromination of bromoform. The rapid debromination of bromoform to a less toxic daughter compound, dibromomethane, makes strain THM1 a promising candidate for remediation of bromoform in anaerobic environments. Additionally, strain THM1 is able to remove single chlorine moieties from both chloroform and 1,1,1-TCA. So far, strain THM1 and three previous known chloroform-respiring bacteria—Dehalobacter sp. strain CF (Grostern et al. 2010), Dehalobacter sp. strain UNSWDHB (Lee et al. 2012; Wong et al. 2016), and Desulfitobacterium sp. strain PR (Ding et al. 2014)—can not only couple reductive dechlorination of chloroform to cell growth, but also dechlorinate 1,1,1-TCA with cell growth, suggesting a probable alternative electron acceptor for either chloroform- or 1,1,1-TCA-dechlorinating bacteria.
Dehalobacter sp. strain THM1 possesses a RDase, ThmA, responsible for dehalogenation of chloroform, 1,1,1-TCA, and bromoform. ThmA shares 96 to 98% amino acid identity over 456 residues with other identified chloroform RDases, CtrA from Desulfitobacterium sp. strain PR, CfrA from Dehalobacter sp. strain CF, and TmrA from Dehalobacter sp. strain UNSWDHB. The high degree of identity among the chloroform RDases allowed for the design of specific primers which can be used to detect chloroform-respiring bacteria, so as to predict the chloroform dechlorinating potential of endogenous microbial communities at contaminated sites. Targeting the genomic DNA extracted from five microcosms using the newly designed chloroform RDase gene-specific primers revealed the presence of the chloroform RDase gene in two of the five microcosms, which is consistent with the observed chloroform dechlorination activity in these two samples but not in the other three microcosms. The chloroform reductive dehalogenases—ThmA, CtrA, CfrA, and TmrA—also showed capability to catalyze the dechlorination of 1,1,1-TCA (Ding et al. 2014; Tang and Edwards 2013; Wong et al. 2016), suggesting that such multi-functionality is a feature of the chloroform RDase. This explains the versatile dechlorinating phenotypes exhibited by single bacterial strains containing chloroform RDases.
Identical functionality has been reported in many RDases despite low homology in nucleic acid or amino acid sequence (Hug et al. 2013). The mechanisms that mediate substrate preference in this diverse group of enzymes remain largely unknown due to a dearth of structural information. Thanks to recently published efforts describing the crystal structure of the tetrachloroethene (PCE) RDase, PceA, from S ulfurospirillum multivorans which identified specific amino acid residues that confer substrate selectivity and specificity, it is possible to make comparisons of primary and secondary amino acid structures of functionally similar RDases at these critical residues. While not necessarily predictive of function, such comparisons may provide insight into how enzymes with such low overall homology perform such similar tasks.
Substrate specificity in PceA is mediated by a substrate channel gated by a stereoselective “letter-box” comprising five amino acid residues (Thr39, Phe44, Phe57, Leu186, Glu189) that are conserved among PCE RDases (Bommer et al. 2014). Alignment of all known chloroform RDases shows that these critical residues exhibit a single highly conserved difference to PceA, with the exception of Leu186 which is also variable among the PCE RDases, at Phe57, which is replaced by Ala57 (Fig. 6). The other residues responsible for stereoselectivity, Thr39, Phe44, and Glu189, are highly conserved among the chloroform RDases. The high degree of similarity shared between PceA and chloroform RDases in this stereoselective region reflects the comparable shape and volume of PCE and chloroform.
Sequence alignment of PceA and chloroform RDases by Clustal Omega. PceA_Sm, Sulfurospirillum multivorans PCE/TCE RDase (AF022812); PceA_SmSL2, Sulfurospirillum sp. SL2 PCE-only RDase (AGW23615); PceA_Dres, Dehalobacter restrictus DSM 9455 PCE/TCE RDase (AHF10423); CprA_Ddh, Desulfitobacterium sp. PCE-1/o-chlorophenol reductive RDase (AF259790); TmrA, Dehalobacter sp. UNSWDHB chloroform/TCA RDase (EQB22800); CfrA, Dehalobacter sp. CF chloroform/1,1,1-TCA RDase (AFQ20272); CtrA, Desulfitobacterium sp. PR chloroform/TCA RDase (AGO27983); ThmA, Dehalobacter sp. THM1 chloroform/1,1,1-TCA/bromoform RDase (KX344907). Green box indicates the amino acid residues lined for entrance of the substrate channel, and red box indicates the predominant amino acid residues lined for the activity site pocket (Bommer et al. 2014) (color figure online).
Substrate specificity at the active site of PceA is mediated by three conserved adjacent amino acids, Tyr246, Arg305, and Asn276, which line a cavity containing a corrin-binding site created by two amino acids, Phe38 and Trp376 (Bommer et al. 2014). Same as in all PCE RDases, both Phe38 and Trp376 are invariant in the chloroform RDases. However, the chloroform RDases again exhibit conserved differences to PceA at residue positions corresponding to substrate specificity (Fig. 6). While Arg305 is the same in both PCE and chloroform RDases, the Tyr 246 and Asn272 in PceA are replaced by cysteine and either serine or alanine residue in the chloroform RDase subset, respectively, suggesting that Cys246, Ser (Ala)272, and Arg305 define the unique specificity of the active site in the chloroform RDase. The highly conserved sequence at the active site of both PCE and chloroform RDases may suggest an approach to predicting the substrate range of uncharacterized RDases in the future.
Although the five amino acids that form the letter-box and the three amino acids lining the corrin-binding pocket at the active site are the major selective barriers dictating substrate specificity in RDases, the entire substrate range of any RDase arises from a combinatorial effect of these core regions and from other domains. For example, variability in the amino acids that form the channel linking the letter-box to the active site also plays a role in selectivity. Such differences in non-core regions of sequence are likely responsible for the observed differences in substrate range among the chloroform RDases, such as 1,1,2-TCA dechlorination by TmrA and CtrA or bromoform debromination by ThmA. It is also notable that the stereoselectivity of RDases can be highly specific. For example, ThmA can dehalogenate neither CHBr2Cl nor CHBrCl2, highlighting the importance of the arrangement of halogen substituents on substrates. These results shed light on future exploration of the substrate diversity of organohalide-respiring bacteria.
References
Adrian L, Rahnenführer J, Gobom J, Hölscher T (2007) Identification of a chlorobenzene reductive dehalogenase in Dehalococcoides sp. strain CBDB1. Appl Environ Microbiol 73(23):7717–7724. doi:10.1128/aem.01649-07
ATSDR (1989) Toxicological profile for bromodichloromethane [WWW document]. URL https://www.atsdr.cdc.gov/toxprofiles/tp129.pdf
ATSDR (1997) Toxicological profile for chloroform [WWW document]. URL https://www.atsdr.cdc.gov/toxprofiles/tp6.pdf
ATSDR (2005) Toxicological profile for bromoform and dibromochloromethane [WWW document]. URL https://www.atsdr.cdc.gov/toxprofiles/tp130.pdf
ATSDR (2013) Summary data for 2013 priority list of hazardous substances [WWW document]. URL https://www.atsdr.cdc.gov/SPL/resources/ATSDR_2013_SPL_Detailed_Data_Table.pdf
Bagley DM, Gossett JM (1995) Chloroform degradation in methanogenic methanol enrichment cultures and by Methanosarcina barkeri 227. Appl Environ Microbiol 61(9):3195–3201
Bommer M, Kunze C, Fesseler J, Schubert T, Diekert G, Dobbek H (2014) Structural basis for organohalide respiration. Science 346(6208):455–458. doi:10.1126/science.1258118
Ding C, Chow WL, He J (2013) Isolation of Acetobacterium sp. strain AG, which reductively debrominates octa- and pentabrominated diphenyl ether technical mixtures. Appl Environ Microbiol 79(4):1110–1117. doi:10.1128/aem.02919-12
Ding C, Zhao S, He J (2014) A Desulfitobacterium sp. strain PR reductively dechlorinates both 1,1,1-trichloroethane and chloroform. Environ Microbiol 16(11):3387–3397. doi:10.1111/1462-2920.12387
Duhamel M, Mo K, Edwards EA (2004) Characterization of a highly enriched Dehalococcoides-containing culture that grows on vinyl chloride and trichloroethene. Appl Environ Microbiol 70(9):5538–5545. doi:10.1128/aem.70.9.5538-5545.2004
Duhamel M, Wehr SD, Yu L, Rizvi H, Seepersad D, Dworatzek S, Cox EE, Edwards EA (2002) Comparison of anaerobic dechlorinating enrichment cultures maintained on tetrachloroethene, trichloroethene, cis-dichloroethene and vinyl chloride. Water Res 36(17):4193–4202. doi:10.1016/s0043-1354(02)00151-3
Fite A, Macfarlane GT, Cummings JH, Hopkins MJ, Kong SC, Furrie E, Macfarlane S (2004) Identification and quantitation of mucosal and faecal Desulfovibrios using real time polymerase chain reaction. Gut 53(4):523–529. doi:10.1136/gut.2003.031245
Grostern A, Duhamel M, Dworatzek S, Edwards EA (2010) Chloroform respiration to dichloromethane by a Dehalobacter population. Environ Microbiol 12(4):1053–1060. doi:10.1111/j.1462-2920.2009.02150.x
Grostern A, Edwards EA (2009) Characterization of a Dehalobacter coculture that dechlorinates 1,2-dichloroethane to ethene and identification of the putative reductive dehalogenase Gene. Appl Environ Microbiol 75(9):2684–2693. doi:10.1128/aem.02037-08
He J, Holmes VF, Lee PKH, Alvarez-Cohen L (2007) Influence of vitamin B12 and cocultures on the growth of Dehalococcoides isolates in defined medium. Appl Environ Microbiol 73(9):2847–2853. doi:10.1128/aem.02574-06
Holliger C, Hahn D, Harmsen H, Ludwig W, Schumacher W, Tindall B, Vazquez F, Weiss N, Zehnder AJB (1998) Dehalobacter restrictus gen. nov. and sp. nov., a strictly anaerobic bacterium that reductively dechlorinates tetra- and trichloroethene in an anaerobic respiration. Arch Microbiol 169(4):313–321. doi:10.1007/s002030050577
Hug LA, Maphosa F, Leys D, Löffler FE, Smidt H, Edwards EA, Adrian L (2013) Overview of organohalide-respiring bacteria and a proposal for a classification system for reductive dehalogenases. Philosophical Transactions of the Royal Society of London, Series B: Biological Sciences 368(1616):19
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. doi:10.1038/227680a0
Justicia-Leon SD, Ritalahti KM, Mack EE, Löffler FE (2012) Dichloromethane fermentation by a Dehalobacter sp in an enrichment culture derived from pristine river sediment. Appl Environ Microbiol 78(4):1288–1291. doi:10.1128/aem.07325-11
Lee LK, He J (2010) Reductive debromination of polybrominated diphenyl ethers by anaerobic bacteria from soils and sediments. Appl Environ Microbiol 76(3):794–802. doi:10.1128/aem.01872-09
Lee M, Low A, Zemb O, Koenig J, Michaelsen A, Manefield M (2012) Complete chloroform dechlorination by organochlorine respiration and fermentation. Environ Microbiol 14(4):883–894. doi:10.1111/j.1462-2920.2011.02656.x
Maphosa F, van Passel MWJ, de Vos WM, Smidt H (2012) Metagenome analysis reveals yet unexplored reductive dechlorinating potential of Dehalobacter sp E1 growing in co-culture with Sedimentibacter sp. Environ Microbiol Reports 4(6):604–616 doi:10.1111/j.1758-2229.2012.00376.x
Maymo-Gatell X, Nijenhuis I, Zinder SH (2001) Reductive dechlorination of cis-1,2-dichloroethene and vinyl chloride by “Dehalococcoides ethenogenes”. Environ Sci Technol 35(3):516–521. doi:10.1021/es001285i
McClay K, Fox BG, Steffan RJ (1996) Chloroform mineralization by toluene-oxidizing bacteria. Appl Environ Microbiol 62(8):2716–2722
Nelson JL, Jiang JD, Zinder SH (2014) Dehalogenation of chlorobenzenes, dichlorotoluenes, and tetrachloroethene by three Dehalobacter spp. Environl Sci Technol 48(7):3776–3782 doi:10.1021/es4044769
Nelson JL, Fung JM, Cadillo-Quiroz H, Cheng X, Zinder SH (2011) A role for Dehalobacter spp. in the reductive dehalogenation of dichlorobenzenes and monochlorobenzene. Environ Sci Technol 45(16):6806–6813. doi:10.1021/es200480k
Oldenhuis R, Oedzes JY, van der Waarde JJ, Janssen DB (1991) Kinetics of chlorinated hydrocarbon degradation by Methylosinus trichosporium OB3b and toxicity of trichloroethylene. Appl Environ Microbiol 57(1):7–14
Robrock KR, Korytar P, Alvarez-Cohen L (2008) Pathways for the anaerobic microbial debromination of polybrominated diphenyl ethers. Environ Sci Technol 42(8):2845–2852. doi:10.1021/es0720917
Shilov IV, Seymour SL, Patel AA, Loboda A, Tang WH, Keating SP, Hunter CL, Nuwaysir LM, Schaeffer DA (2007) The Paragon Algorithm, a next generation search engine that uses sequence temperature values and feature probabilities to identify peptides from tandem mass spectra. Molecular & cellular proteomics : MCP 6(9):1638–1655. doi:10.1074/mcp.T600050-MCP200
Smits THM, Devenoges C, Szynalski K, Maillard J, Holliger C (2004) Development of a real-time PCR method for quantification of the three genera Dehalobacter, Dehalococcoides, and Desulfitobacterium in microbial communities. Environ Sci Technol 57(3):369–378. doi:10.1016/j.mimet.2004.02.003
Sun BL, Griffin BM, Ayala-del-Rio HL, Hashsham SA, Tiedje JM (2002) Microbial dehalorespiration with 1,1,1-trichloroethane. Science 298(5595):1023–1025. doi:10.1126/science.1074675
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol Biol Evol 30(12):2725–2729. doi:10.1093/molbev/mst197
Tang S, Edwards EA (2013) Identification of Dehalobacter reductive dehalogenases that catalyse dechlorination of chloroform, 1,1,1-trichloroethane and 1,1-dichloroethane. Philoso Trans R Soc Lon B Biol Sci 368(1616):20120318. doi:10.1098/rstb.2012.0318
van der Zaan B, de Weert J, Rijnaarts H, de Vos WM, Smidt H, Gerritse J (2009) Degradation of 1,2-dichloroethane by microbial communities from river sediment at various redox conditions. Water Research 43(13):3207–3216 doi:10.1016/j.watres.2009.04.042
Wang S, He J (2013) Dechlorination of commercial PCBs and other multiple halogenated compounds by a sediment-free culture containing Dehalococcoides and Dehalobacter. Environ Sci Technol 47(18):10526–10534. doi:10.1021/es4017624
Wang S, Zhang W, Yang KL, He J (2014) Isolation and characterization of a novel Dehalobacter species strain TCP1 that reductively dechlorinates 2,4,6-trichlorophenol. Biodegradation 25(2):313–323. doi:10.1007/s10532-013-9662-1
Weathers LJ, Parkin GF (2000) Toxicity of chloroform biotransformation to methanogenic bacteria. Environ Sci Technol 34(13):2764–2767. doi:10.1021/es990948x
Wild A, Hermann R, Leisinger T (1997) Isolation of an anaerobic bacterium which reductively dechlorinates tetrachloroethene and trichloroethene. Biodegradation 7(6):507–511. doi:10.1007/bf00115297
Wong YK, Holland SI, Ertan H, Manefield M, Lee M (2016) Isolation and characterization of Dehalobacter sp. strain UNSWDHB capable of chloroform and chlorinated ethane respiration. Environ Microbiol. doi:10.1111/1462-2920.13287
World Health Organisation (2004) Trihalomethanes in drinking water
Yoshida N, Ye L, Baba D, Katayama A (2009) A novel Dehalobacter species is involved in extensive 4,5,6,7-tetrachlorophthalide dechlorination. Appl Environ Microbiol 75(8):2400–2405. doi:10.1128/aem.02112-08
Zhao S, Ding C, He J (2015) Detoxification of 1,1,2-trichloroethane to ethene by Desulfitobacterium and identification of its functional reductase gene. PLoS One 10(3):e0119507. doi:10.1371/journal.pone.0119507
Acknowledgments
This study was supported by the Singapore Agency for Science, Technology and Research (A*STAR) of the Science and Engineering Research Council under project No.: 102 101 0025 and the National Research Foundation, Prime Minister’s Office, Singapore, under the Competitive Research Programme, Project No.: NRF-CRP5-2009-05.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Zhao, S., Rogers, M.J. & He, J. Microbial reductive dehalogenation of trihalomethanes by a Dehalobacter-containing co-culture. Appl Microbiol Biotechnol 101, 5481–5492 (2017). https://doi.org/10.1007/s00253-017-8236-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8236-2