Abstract
Cytochromes P450 (CYP) are attractive enzyme targets in biotechnology as they catalyze stereospecific C-hydroxylations of complex core skeletons at positions that typically are difficult to access by chemical synthesis. Membrane bound CYPs are involved in nearly all plant pathways leading to the formation of high-value compounds. In the present study, we systematically maximize the heterologous expression of six different plant-derived CYP genes in Escherichia coli, using a workflow based on C-terminal fusions to the green fluorescent protein. The six genes can be over-expressed in both K- and B-type E. coli strains using standard growth media. Furthermore, sequences encoding a small synthetic peptide and a small bacterial membrane anchor markedly enhance the expression of all six genes. For one of the CYPs, the length of the linker region between the predicted N-terminal transmembrane segment and the soluble domain is modified, in order to verify the importance of this region for enzymatic activity. The work describes how membrane bound CYPs are optimally produced in E. coli and thus adds this plant multi-membered key enzyme family to the toolbox for bacterial cell factory design.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Escherichia coli is a model organism and a popular workhorse for protein production with a comprehensive set of available genetic tools. An obvious extension of producing individual proteins for downstream applications is to produce a set of enzymes that catalyze an entire biosynthetic pathway. In our desire to advance from a petrochemical-based society towards a bio-based society, it is critical that we deepen our understanding of how such heterologous biosynthetic pathways should be designed and maintained for the production of high-value chemicals from renewable feedstock (Møller 2014). Plant cytochrome P450 enzymes (CYPs) are central to this approach, as they are involved in nearly all pathways that lead to the formation of high-value compounds such as terpenoids, alkaloids and phenylpropanoids (e.g., flavonoids, isoflavonoids, chalcones, aurones, and lignans). Many of these compounds are used as medicines, condiments, flavors, and fragrances (Morant et al. 2003), with well-known examples being the antimalarial diterpenoid artemisinin (Chang et al. 2007), the anti-cancer drug paclitaxel (Chau et al. 2004; Biggs et al. 2016), the flavor compound vanillin (Hansen et al. 2009; Gallage and Møller 2015) and the steviol glucoside-based sweeteners (Davies and Deroles 2014). Typically, the membrane anchored CYPs catalyze stereospecific hydroxylations of complex core carbon skeletons, at positions that are difficult to access by de novo chemical synthesis. CYPs often show high substrate specificity and although members of this ancient multigene family are found in all domains of life, plants seem to be particularly enriched with often more than 250 CYPs in a single species (Nelson 2011). Unfortunately, plant CYPs have the reputation of being notoriously difficult to produce in E. coli (Chang et al. 2007), possibly due to their hydrophobic nature combined with co-factor and posttranslational modification requirements. Furthermore, in order to work efficiently, CYPs often need co-production of other membrane-associated proteins such as reductases (Eugster et al. 1992) or cytochromes b5 (Zhang et al. 2007; Paddon et al. 2013).
Previous approaches to improve plant CYP production in E. coli includes N-terminal truncations (Haudenschild et al. 2000), adding an alanine/GCT in codon position 2 (Haudenschild et al. 2000; Biggs et al. 2016), exchanging parts of the native amino-terminal region with more hydrophilic amino acids (Morrone et al. 2010; Biggs et al. 2016) or with parts of the amino-terminal region of a codon-optimized bovine CYP17alpha, often referred to in the field as “Barnes sequence” (Barnes et al. 1991; Bak et al. 1997; Bak et al. 1998a; Leonard and Koffas 2007). Similarly, bacterial production of human CYPs have been aided by bacterial leader sequences such as the membrane translocation signals from pelB and ompA (Pritchard et al. 1997); for recent reviews on this topic see Zelasko et al. (2013) and Kitaoka et al. (2015).
Membrane protein production in E. coli has benefited greatly from the introduction of a streamlined green fluorescent protein-based pipeline for rapid and simple assessment of proper expression (Drew et al. 2001; Drew et al. 2005; Drew et al. 2006; Lee et al. 2014). Briefly, a carboxy-terminal GFP fusion has many advantages: without interfering with membrane targeting and integration, it acts as an expression reporter as fluorescence indicates that the mRNA was translated in full. Importantly, if the upstream protein is mislocalized and subsequently aggregated, GFP fluorescence is completely quenched. Further, once folded, GFP remains fluorescent even during otherwise denaturing applications such as SDS-PAGE. In summary, the carboxy-terminal GFP provides a cheap, reliable and fast readout of the proper production of membrane proteins. Moreover, the fluorescent properties of such chimeric proteins can be used for screening the optimal solubilization conditions for downstream applications, such as structural studies (Kawate and Gouaux 2006; Sonoda et al. 2011).
Currently, there is no established formula for the heterologous expression of CYPs in E. coli. The literature is scattered with a wealth of different genetic manipulations and use of various strains and growth conditions. To enable efficient heterologous CYP production, there is a need to systematically determine proper and generally useful expression conditions. Here, we demonstrate how production of six different plant CYPs in E. coli is achieved by very simple means and confirm that three of these retain their function. The carboxy-terminal GFP approach readily lends itself to high-throughput applications, as well as standard methods compatible with fluorescent readouts such as SDS-PAGE, flow cytometry, and size-exclusion chromatography. Thus, the experimental setup is bound to be highly useful for large scale expression screens.
Materials and methods
Bacterial strains
Escherichia coli strain NEB 5-alpha (New England BioLabs, Ipswich, USA) was used for cloning of PCR products and propagation of plasmids. The following E. coli strains were used for gene expression: Rosetta2(DE3) pLysS (Novagen, Merck Millipore, Germany), Bl21(DE3) pLysS (Promega, Madison, USA), C41(DE3) (Miroux and Walker 1996), KRX (Promega, Madison, USA), MC4100(DE3)pLysS, and MG1655(DE3)pLysS. MC4100(DE3) and MG1655(DE3) were made using a λDE3 Lysogenization kit (Novagen, Merck Millipore, Germany).
PCR and uracil excision
All DNA manipulations were performed using uracil excision technology as previously described (Nour-Eldin et al. 2006; Nørholm 2010) and all oligonucleotides are listed in Table S1. PCR products were amplified in 50-μL reactions containing: 1-μL PfuX7 DNA polymerase, 0.2 mM dNTPs (Thermo Scientific, Waltham, USA), 1.5 mM MgCl2, 0.5 μM forward oligonucleotide, 0.5 μM reverse oligonucleotide, Phusion® HF Reaction Buffer (New England BioLabs, Ipswich, USA) and 50-ng plasmid template. A touch-down PCR program was used for amplification: step 1: 2 min 98 °C; step 2: 15 s 98 °C, 20 s 65 °C (−1 °C per cycle), 45 s per kb at 72 °C (step 2 repeated nine times until 55 °C, then repeated 20 cycles at the annealing temperature 55 °C); step 3: 5 min 72 °C; step 4 hold at 10 °C. PCR products were gel purified from 1% (w/V) agarose gel using NucleoSpin Gel and PCR Clean-up (Macherey-Nagel, Düren, Germany) and eluted in 10% TE buffer. Purified PCR products were incubated with 1-μL USER™ enzyme (New England BioLabs, Ipswich, USA) for 30 min at 37 °C and subsequently mixed with linearized vector backbone in a molar ratio 3:1 and incubated at 18 °C for 1 h. A Nanodrop spectrophotometer 2000 (Thermo Scientific, Waltham, USA) was used for estimation of PCR product and vector concentration. Approx. 5 μL of the assembled PCR product:vector solution was transformed into NEB 5-alpha chemically competent cells according to the manufacturer’s protocol. Transformants were selected on Luria Bertoni (LB) agar plates supplemented with 50 μg/mL kanamycin, 25 μg/mL chloramphenicol, or 10 μg/mL gentamycin. Colonies were screened for gene insert by colony PCR using OneTaq 2X Master mix (New England BioLabs, Ipswich, USA). Vectors were extracted and purified using QIAprep Spin Kit (Qiagen) and verified by DNA sequencing (Eurofins Genomics, Ebersberg, Germany).
DNA constructs
All DNA constructs were made with uracil excision as previously described (Nour-Eldin et al. 2006; Nørholm 2010). Details of oligonucleotides, template DNA, and references can be found in Tables S1 and S2 and below. Briefly, AsiSI restriction enzyme recognition sites, present in the kanamycin resistance cassettes, were deleted from all pET28a(+)-derived construct by uracil excision-based site-directed mutagenesis. AsiSI uracil excision compatible cloning cassettes were inserted between an upstream sequence encoding either the 28-tag (Nørholm et al. 2013) (Fig. 1a), a Barnes-like N-terminal sequence MALLLAVF, SohB (residues 1-48, KDT39511) or YafU (residues 1-88, KEN61237) and a downstream cassette encoding a TEV protease site, GFP and a polyhistidine tag. Six genes encoding different plant CYPs were generously supplied from different laboratories at The Department of Plant and Environmental Sciences (Sorghum bicolor CYP51G1 (U74319); Avena strigosa CYP51H10 (DQ680849), Sorghum bicolor CYP71E1 (O48958), Sorghum bicolor CYP79A1 (U32624), Arabidopsis thaliana CYP79B2 (NM_120158), and Picea sitchensis CYP720B4 (HM245403)) were PCR amplified with sequence specific oligonucleotides and cloned into the different pET28a(+)-derived vectors either treated with the restriction enzyme AsiSI (Thermo Scientific, Waltham, USA) and the nicking enzyme Nb. BbvCI (New England Biolabs, Ipswich, USA) or PCR amplified. The sohB- and yafU-based E. coli membrane anchors have their C-termini on the cytoplasmic side of the E. coli inner membrane (Daley et al. 2005) and therefore the plant CYPs cloned into the corresponding backbones had their predicted N-terminal anchors truncated to avoid translocation of the CYP into the periplasmic space. Truncations were designed using TMHMM v. 2.0 (Sonnhammer et al. 1998). The E. coli lepB-membrane anchor has its C-termini in the periplasm and was therefore fused directly to the N-terminal end of the full-length CYP. A Strep-HRV3C tagged, codon optimized Sorghum bicolor CPR2b (Wadsäter et al. 2012) was cloned into pET28a(+)-tev-gfp-his8 and gfp-his8 was subsequently deleted and the origin of replication was swapped with the corresponding parts from pSEVA63 (Silva-Rocha et al. 2013) by amplifying pSEVA63 with the oligo nucleotides 5′-ATCCGCTUTAATTAAAGGCATCAAATAAAAC-3′ and 5′-ACTAGTCTUGGACTCCTGTTGATAGATC-3′ and the pET28-based cpr2b construct with the oligo nucleotides 5′-AAGCGGAUCTACGAGTTGCATGATAAAGAAGACAGTC-3′ and 5′-AAGACTAGUCAATCCGGATATAGTTCCTCCTTTCAG-3′. A truncated version of E. coli lepB was amplified from a previously described pGem1-Lep construct (Hessa et al. 2005) using the oligonucleotides 5′-ACTCGAGGAUGGCGAATATGTTTGCCCTGATTC-3′ and 5′-ATCGCTGCUTCCAGGACCACCACTAGTCTCG-3′ and combined with the pET28-based full-length CYP constructs amplified with 5′-ATCCTCGAGUCTCCTTCTTAAAG-3′ in combination with the gene-specific forward oligonucleotides.
Production of six different CYP enzymes in standard growth media. a Schematic representation of the cloning procedure used. Genes encoding six different CYPs were PCR amplified with specific uracil-containing tailed oligonucleotides and assembled with a compatible AsiSI/Nb NbvcI treated vector. This way, the individual CYPs were fused with a 28 codon tag in their 5′end and a cassette encoding a TEV protease site, GFP and a His tag at their 3′end. b Effect of media on production of CYP51G1, CYP51H10, CYP71E1, CYP79A1, CYP79B2, and CYP720B4 using the 28-tag:CYP:tev-gfp-his8 expression construct in the KRX strain. Four media types were tested: Luria Bertoni (LB), Terrific Broth (TB), minimal media M9, and autoinducing media (AI). As cell cultures reached the exponential growth phase, cultures were induced for 3 h. Cell cultures were subsequently harvested by centrifugation and resuspended in loading buffer for in-gel fluorescence detection. c Strain-dependent production of CYP enzymes. Three K-strains and three B-strains were transformed with 28-tag:CYP:tev-gfp-his8 expression constructs. Cells were grown in TB media and induced for 3 h, as cell cultures reached the exponential growing phase. Overall expression was monitored by in-gel fluorescence detection in the three B- and the three K-strains, but significant levels of expression were only obtained in Rosetta2(DE3)pLysS and KRX
Culture media and expression conditions
All strains were grown aerobically in liquid cultures. For plasmid propagation, single colonies were grown over night in 2xYeast/Tryptone medium at 37 °C, 250 rpm. For single plasmid transformations, chemically competent cells were transformed with 20-ng plasmid according to the manufacturer’s protocol. For co-transformation of plasmids for expression assays, electrocompetent cells were transformed with 20 ng of each plasmid. Four different media were tested for expression, lysogeny broth (LB) (1% tryptone, 0.5% yeast extract, 1% NaCl), Terrific Broth (TB) (1.2% tryptone, 2.4% yeast extract, 0.4% glycerol, 17 mM potassium phosphate (monobasic), 72 mM potassium phosphate (dibasic)), the defined rich medium PA-5052 (Studier 2005) (50 mM Na2HPO4, 50 mM KH2PO4, 25 mM (NH4) 2SO4, 2 mM MgSO4, 10 μM metals, 0.5% glycerol, 0.05% glucose, 0.2% alpha-lactose, 200 μg/mL amino acids E, D, K, R, H, A, P, G, T, S, Q, N, V, L, I, F, W, and M) and minimal M9 medium (M9 salts, 2 mM MgSO4, 0.1 mM CaCl2, 0.2% glycerol, 0.2% glucose, 10 μM Fe3+ (due to the heme requirement; 200 μg/mL amino acids E, D, K, R, H, A, P, G, T, S, Q, N, V, L, I, F, W, and M). For media tests, over night cultures were prepared with each of the four media supplemented with 0.5% (w/V) d-glucose and appropriate antibiotics. The pre-cultures were subsequently inoculated into each of the corresponding media for expression. For other expression assays, over night cultures were made in 96 deep well plates (CR1296, sealed with metal sandwich covers, Enzyscreen B.V. Harlem, Netherlands), inoculating from single colonies into 800-μL TB supplemented with 0.5% (w/V) d-glucose and appropriate antibiotics and grown at 30 °C, 250 rpm in Innova®44R incubator shaker system (5 cm orbital shaking) (New Brunswick Scientific, Eppendorf, USA). The optical density (OD) of the overnight cultures was measured at Abs600nm on Plate Reader SynergyMx (SMATLD) (BioTek, Winooski, USA). Overnight cultures were inoculated into 5-mL fresh TB medium in 24 deep well plates (CR1224, sealed with metal sandwich covers, Enzyscreen B.V. Harlem, Netherlands) to a final OD of 0.05. Cells were incubated for approx. 2 h at 37 °C 250 rpm to an OD of 0.3–0.5. All strains were induced with a final conc. of 0.4 mM isopropyl ß-D-1-thiogalactopyranoside (IPTG, dioxane free, Thermo Scientific, Waltham, USA). Despite the autoinduction capacity of the PA-5052 medium, as described previously (Lee et al. 2014), IPTG was use for induction in this medium as well. The strain KRX was furthermore induced with a final conc. of 5 mM L-rhamnose (Sigma-Aldrich, St. Louis, USA). Cultures were subsequently incubated at 25 °C 150 rpm for 3 or 22 h.
Whole-cell fluorescence measurements
Whole-cell fluorescence was measured using 2-mL of induced culture. The cells were harvested (2,500×g, 20 min, 4 °C) and resuspended in a total of 100-μL PBS buffer. The GFP fluorophore was allowed to form for 1 h at room temperature and then fluorescence was detected using excitation at 485 nm and emission at 512 nm with a window of ±9 nm, gain value 50, using plate reader SynergyMx SMATLD (BioTek, Winooski, USA).
SDS-page
Whole-cells were harvested (2,500×g, 20 min, 4 °C), resuspended to an OD of 10, and lysed for 1.5 h at room temperature in TRIS buffer (50 mM Tris HCl pH 7.5, 150 mM NaCl, 2 mM MgCl2) with 250 U/mL Benzonase ® nuclease (Sigma-Aldrich, St. Louis, USA), 5 mg/mL lysozyme egg white powder (Amresco, Ohio, USA) and the cOmplete ULTRA EDTA-free protease inhibitor cocktail (Roche, Basel, Switzerland). 10-μL cell lysate (OD600nm 0.1) was analyzed in parallel with PageRuler™ Prestained Protein Ladder 10-170K (Thermo Scientific, Waltham, USA) by standard SDS-page using Mini-PROTEAN® TGX™ 4-15% gels (Bio-Rad, Hercules, USA). In-gel fluorescence was detected on a G:BOX UV-table (Syngene, Cambridge, UK).
Enzyme activity measurements
For CYP79A1 enzymatic assays, cells were harvested by centrifugation 2,500×g 4 °C for 10 min, washed once in 50 mM potassium phosphate (KPi) buffer pH 7.5 and resuspended to 0.03 OD units per microliter in 50 mM KPi buffer. The enzyme reaction was carried out in 30-μL volume consisting of 5 mM NADPH, 0.5 mM L-Tyrosine (Sigma-Aldrich, St. Louis, USA), 50 mM KPi buffer, and 20-μL cell suspension. Cells were incubated at 30 °C 400 rpm for 60 min. The product (E)-p-hydroxyphenylacetaldoxime (oxime) was extracted by adding 150-μL methanol followed by incubation at room temperature for 10 min. Cells debris was discarded twice by centrifugation (20,000×g, 10 min) and the supernatant was transferred into HPLC vials and stored at −20 °C prior to analysis. The oxime was also extracted directly from cultures co-expressing SbCYP79A1 and SbCPR2b for 22 h. One hundred-microliter culture was extracted with 100-μL methanol, as described above.
LCMS detection
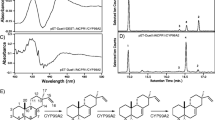
The CYP79A1 product (E)-p-hydroxyphenylacetaldoxime (Mw 151.17) was detected and quantified by LC-MS. A chemically synthesized (Z)-p-hydroxyphenylacetaldoxime standard was kindly provided by Mohammed Saddik Motawie (University of Copenhagen, Department of Plant and Environmental Sciences). The two geometrical isomers (Z)-p-hydroxyphenylacetaldoxime and (E)-p-hydroxyphenylacetaldoxmie were detected in both the chemical oxime standard and the oxime sample product due to instability and chemical equilibrium. Thus, for the purpose of total oxime quantification, the chromatogram area of the both peaks were summed. LC-MS data was collected on a Bruker Evoq triple quadropole mass spectrometer equipped with an Advance UHPLC pumping system. Samples were held in the CTC PAL autosampler at a temperature of 10 °C during the analysis. Injections (2 μL) of the sample were made onto a Supellco Discovery HS F5-3 HPLC column (3-μm particle size, 2.1-mm i.d., and 150 mm long). The column was held at a temperature of 30 °C. The solvent system (flow rate 1.0 mL/min) was milli-Q water with (A) 100 mM ammonium formate and (B) acetonitrile using the following elution profile: 0.5 min 95% A/5% B, linear gradient to 50% A/50% B for 3.0 min, 1.5 min 50% A/50% B and re-equilibration for 2 min 95% A/5% B. The column eluent flowed directly into the heated ESI probe of the MS, which was held at 350 °C and a voltage of 4500 V. SRM data was collected in centroid at unit mass resolution. Positive ion mode with Q1 set to monitor 152.70 m/z, Q3 set to monitor 136 m/z, and Q2 set to a collision energy of 10.0 eV, with an Argon pressure of 1.5 mTorr. The other MS settings were as follows, sheath gas flow rate of 40 units, aux gas flow rate of 40 units, sweep gas flow rate of 20 units, ion transfer tube temp was 350 °C.
Nucleotide sequence accession numbers
Sequences used for design of the bacterial anchors of SohB (residues 1-48), YafU (residues 1-88), and LepB (residue 1-228) can be found under the accession numbers KDT39511, KEN61237 and 947040, respectively. The six plant P450 enzymes are derived from Avena strigosa CYP51H10 (DQ680849), A. thaliana CYP79B2 (NM_120158), Picea sitchensis CYP720B4 (HM245403) and Sorghum bicolor CYP51G1 (U74319), CYP71E1 (O48958), CYP79A1 (U32624). Furthermore the cytochrome P450 reductase is derived from S. bicolor CPR2b (XP_002444097.1).
Results
Full-length production of six plant CYPs in E. coli grown in standard media
Previously, it was shown that simple addition of an 84 nucleotide/28 amino acid tag (28-tag) to the amino-termini greatly enhanced the expression of challenging membrane protein encoding genes in E. coli (Nørholm et al. 2013). Here, we used a similar design principle to sandwich plant CYPs between an amino-terminal 28-tag and a carboxy-terminal TEV-GFP-His cassette (Fig. 1a). Six native CYP gene sequences from four different mono- and dicotyledonous plants were cloned into the pET28-28-tag-tev-gfp-his8 plasmid: S. bicolor CYP51G1 (Bak et al. 1997), CYP71E1 (Bak et al. 1998a) and CYP79A1 (Koch et al. 1995), Avena strigosa CYP51H10 (Qi et al. 2006), A. thaliana CYP79B2 (Bak et al. 1998b) and Picea sitchensis CYP720B4 (Hamberger et al. 2011) and transformed into E. coli KRX. Following 3-h induction in four different standard growth media (LB, TB, M9 supplemented with Fe3+, and the synthetic rich medium PA-5052), expression and protein integrity was analyzed by measuring whole-cell- and in-gel fluorescence. M9 medium performed poorly compared to the other media, with significantly lower activity detected—in most cases, only just above background (Fig. 1b) and the highest cell densities were achieved on rich TB medium (data not shown). Moreover, analysis of the samples by in-gel fluorescence indicated that the majority of the constructs were expressed as full-length proteins (Fig. 1c), except for CYP51H10 for which a significant proportion seemed to be truncated, possibly due to the presence of internal ribosome binding sites, or partly degraded. Thus, whole-cell fluorescence can be a poor proxy for CYP-gfp expression, but combined with in-gel fluorescence, it is a very convenient and informative assay enabling further optimization efforts. Addition of 5-aminolevulinic acid suggested to aid the functional production of heme-containing proteins (Sudhamsu et al. 2010) did not have any apparent effect on expression levels (data not shown). Based on the highest expression level and cell density, we chose TB medium for further expression optimization.
E. coli strains KRX and Rosetta excel in plant CYP production
Next, we compared expression levels of the six different CYP constructs in the six different E. coli strains MC4100(DE3) pLysS, MG1655(DE3) pLysS, KRX, BL21(DE3) pLysS, Rosetta2(DE3) pLysS and C41(DE3) that all vary in the way they control expression of the T7 RNA polymerase. In all strains except for KRX and C41, expression from the T7 promoter is catalyzed by the T7 phage derived RNA polymerase controlled by the PlacUV5 IPTG-inducible promoter. In three of the strains, potential toxic effects of over-expression are counteracted by expression of the natural T7 DNA polymerase inhibitor T7 lysozyme from the pLysS or the pLysSRARE2 plasmid (Studier 1991), whereas a similar effect is obtained in C41 by mutations in PlacUV5 that make the promoter less strong (Wagner et al. 2008). In the commercial strain KRX, expression of T7 RNA polymerase is tightly controlled by a rhamnose inducible promoter. The six pET28-28tag-CYP-tev-gfp-his8 constructs were transformed into the six different strains and expression was monitored by in-gel fluorescence after 3-h induction. As judged from the whole-cell fluorescence (data not shown) and presence of full-length protein on SDS-gels, strains KRX and Rosetta2(DE3) pLysS clearly outperformed the other four strains (Fig. 1c), although again, the truncation/degradation of CYP51H10 was evident in most strains.
A transmembrane domain encoded by E. coli sohB normalizes expression of plant CYP genes to high levels
Most eukaryotic CYPs utilize N-terminal hydrophobic peptides to localize to the endoplasmic reticulum, and sequence modifications in this region have previously proven essential for heterologous expression (Barnes et al. 1991). We hypothesized that it would be beneficial to exchange the plant ER/membrane localization signals from the CYPs with membrane anchors that are derived from the heterologous expression host and that have previously been shown to express at high levels from the T7 promoter. To find suitable candidates, we mined a previously published library of membrane protein encoding genes expressed in E. coli (Daley et al. 2005). We chose three of the most highly expressed genes (Fig. S1.) that encoded membrane proteins with different topologies: sohB, encoding a single-pass membrane protein with the C-terminal end localized inside the bacterial cytoplasm (Cin); yafU, encoding two membrane-spanning regions with a Cin topology; and lepB, encoding two membrane-spanning regions, but with the opposite (Nout) topology (Fig. 2a). To mimic the overall topology of the microsomal plant CYPs and ensure localization of the CYP catalytic domain to the cytoplasm, we replaced the hydrophobic sequences of the CYPs with the transmembrane parts of SohB and YafU, whereas the full-length CYP was fused to the periplasmic carboxy-terminus of LepB. For benchmarking purposes, we added a Barnes-like MALLLAVF-encoding sequence to the six CYP genes and comparisons were made to unmodified (native) plant CYP sequences. The native controls, 28-tag-, MALLLAVF-, sohB-, yafU-, and lepB-tagged CYP constructs were transformed into KRX and Rosetta2(DE3) pLysS and expression was monitored by in-gel fluorescence (Fig. 2b). None of the native N-terminally unmodified sequences expressed to any significant levels. In contrast, both the 28-tag and the SohB-domain normalized the expression of the CYPs to high levels and in particular, the combination of the KRX strain with the SohB-tagged CYPs gave high levels of full-length protein in five out of six cases. Furthermore, compared to the 28-tag constructs, a higher proportion of SohB-tagged CYP79A appeared as full-length, both 3 and 22-h postinduction (Fig. S2). “Free” GFP (Fig. S2) has previously been observed using similar membrane protein GFP fusions and likely originates from degradation of the intact fusion protein as they can form after solubilization of membrane fractions (Sonoda et al. 2010).
Effect of different N-terminal peptides and small bacterial membrane anchors on the expression of six different CYPs. a Illustration of the hypothetical membrane topology of native CYPs and recombinant bacterial anchor constructs: (from left) reference CYP with native N-terminal sequence, SohB membrane domain (1TM, highlighted in red) fused with a truncated CYP, YafU anchor (2TM, Cin, highlighted in orange) fused with a truncated CYP, and LepB anchor (2TM, Cout, highlighted in yellow) fused to a native CYP. All constructs have TEV:GFP:His8 fusions at their C-terminal. b Six CYPs were tested with different N-terminal modifications: (from left) native sequence as reference, 28-tag fused to native CYP sequence, Barnes-like MALLAVF peptide fused to native CYP sequence, SohB membrane domain fused to a truncated CYP sequence, YafU membrane domain fused to a truncated CYP sequence and the LepB membrane domain fused to a native sequence. Constructs were expressed in two strains (KRX and Rosetta2(DE3)pLysS). Cell cultures were induced for 3 h before harvest. Cell lysates were analyzed by in-gel fluorescence to determine the level of full-length protein present
Activity is preserved in all engineered versions of CYP79A1 except for those truncated in a proline-rich region that precedes the soluble catalytic domain
To obtain detectable expression levels of CYPs, we substantially re-engineered plant CYP proteins with truncations and/or heterologous protein domains such as the five different amino-terminal peptides and the large carboxy-terminal GFP domain described above. To test that a CYP is able to retain activity with these major modifications, we functionally assayed all the modified versions of CYP79A1, catalyzing the formation of (Z)-p-dihydroxyphenylacetaldoxime from L-tyrosine (Fig. 3a) while monitoring expression levels by fluorescence. Moreover, we functionally assayed the best expressing CYP71E1 and CYP79B2 versions (Fig. S3) and checked the effect of truncating CYP79A1 at the residue Y36, likely placed at the interface between the membrane and the aqueous face, at the residue P69, placed at a proline-rich, proposed hinge region observed in many P450s (Chen et al. 1998; Williams et al. 2000; Leonard and Koffas 2007) and at two positions in between these two extremes (P44 and P60, Fig. 3b). CYP79A1 and the reductase S. bicolor CPR2b were co-expressed in the KRX strain and (E)-p-hydroxyphenylacetaldoxime production was detected by separation using high-performance liquid chromatography and mass detection (LCMS, Fig. 3c). Whole-cell fluorescence confirmed expression of all constructs except the construct with the native, unmodified CYP79A1 sequence, as described above, and most of the expressed constructs were catalytically active except for those with the most heavily truncated CYP79A1 sequence at position P69 (Fig. 3d). Cells expressing the YafU and LepB fusions were less active than those expressing the 28- and the SohB-fusions (data not shown). In order to see if the distance to the membrane is important, we increased the linker between the membrane anchor and the soluble domain by duplicating the sequence Y36-P44 in the SohB-CYP79A1 fusion construct. The artificially increased size of the linker resulted in a seemingly small increase in enzyme activity (Fig. 3d). Under the short-term expression conditions, different constructs with the full-length CYP79A1 sequence conserved or with the shortest truncation, showed good correlations between whole-cell fluorescence and enzyme activity (Fig. S4). Finally, particularly because the CYP needs to functionally interact with its reductase partner, we tested if the GFP fusion had a negative impact on the proper interactions, but only observed minor improvements in the activity when GFP was removed (Fig. S5).
Functional assaying of engineered variants of CYP79A1. a Illustration of the chemical reaction catalyzed by CYP79A1. b Schematic overview of CYP79A1 modified expression constructs used in this study. Four truncated expression constructs were designed to remove various parts of the N-terminal sequence: Δ1-35: at the end of the predicted transmembrane segment between residues 1 and 35, and at three proline residues at positions 44, 60, and 70 (Δ1-43, Δ1-59, Δ1-69) preceding the catalytic domain. In addition, an artificially extended linker region was engineered by inserting a repeat of the amino acids Y36-P44. c High-performance liquid chromatography chromatograms of (i) chemical oxime standard low conc. (0.1 μg/mL), (ii) chemical oxime standard high conc. (10 μg/mL), (iii) sample low product conc., (iv) sample high product conc., and (v) blank illustrated with a fixed scale of signal intensity on the Y-axes. d CYP79A1-catalyzed conversion of L-tyrosine to (E)-p-hydroxyphenylacetaldoxime in 22-h induced cultures. Differently truncated and engineered CYP79A1 constructs were co-produced with the compatible reductase electron donor CPR2b in the KRX strain. Cells were induced for 22 h, before oxime was extracted from the culture. Error bars indicate standard error of the mean (n = 3)
Discussion
The multitude of available genetic tools, growth media, and genotypes for the model organism E. coli is a blessing, but can also be a curse. Optimization of the physical parameters for heterologous gene expression is a multifactorial problem and the details—from transcription to translation and posttranslational processing—can be endlessly tinkered with. To this end, a fast reliable and inexpensive assay to optimize expression conditions in a high-throughput fashion is of great importance, especially for the design of cell factories that are based on expression of multiple challenging enzymes like CYPs.
Here, we have explored the use of GFP fusions to report on the functional production of a commercially attractive group of single spanning membrane proteins, the multigene encoded cytochrome P450 enzyme family (Morant et al. 2003). Our ambition was to replace the commonly used, but very laborious, CO-spectral analysis with the more convenient and high-throughput compatible fluorescence readouts. In doing this, we discovered that the two expression strains KRX and Rosetta2(DE3) pLysS consistently outperform the other tested expression strains with respect to the amount of functionally active P450 enzyme produced under short-term induction conditions. KRX is a K strain and Rosetta2 is a B strain, suggesting that there is no obvious restriction in the use of K vs. B strains for CYP expression. The Rosetta2 strain provides additional copies of transfer RNAs that are present in low concentration in E. coli strains, whereas KRX does not, suggesting that this complementation is not the major causative effect on CYP expression in Rosetta2. Rather, a likely similarity between the two strains is the carefully balanced expression of the T7 RNA polymerase—in KRX by controlling expression directly from the Prha promoter and in Rosetta2 indirectly by expression of lysS from the pLysSRARE2 plasmid. Several of the other strains express lysS from the pLysS plasmid, but these do not perform as well, and we speculate that the difference in lysS carrying plasmid may change the production of lysozyme in a way that affects CYP expression. In line with this, upon removal of lysS from pLysS, we obtained similar expression levels in Rosetta and BL21 with all six P450s (Fig. S6) and we have previously shown that lysozyme is produced at lower levels from pLysSRARE2 compared to pLysS (Søgaard and Nørholm 2016). It is likely that other factors such as the expression vector copy number and the antibiotic selection will have similar effects on protein production levels as shown recently for a difficult to express membrane protein (Kim et al. 2016). Our comparison of growth media suggests that there is no “magic” medium for CYP expression, but that the minimal medium M9 without additional supplements is suboptimal.
Another finding is that expression levels can be normalized to high levels using small peptides such as the 28-tag, which to our knowledge provides no association to membranes, or membrane-spanning domains like SohB. Similar effects have been observed before with tags such as polyhistidine, the maltose binding protein (MBP), and the small ubiquitin-related modifier (SUMO, for a recent review see Costa et al. 2014). The frequently observed positive effect of adding sequences to the 5′-end is possibly due to an incompability between the native plant 5′sequences with high-level expression, as previously reported for membrane protein encoding genes like araH and narK (Nørholm et al. 2013). An alternative solution that previously has proven successful is to introduce synonymous changes in the first couple of codons downstream from the start codon (Nørholm et al. 2013) (Zelasko et al. 2013). Indeed, we have successfully expressed the native CYP79A1 and CYP71E1 genes by introducing just a few codon changes in the pET expression system (data not shown).
We hypothesized that the inherent membrane targeting of, e.g., SohB, would have a positive impact on expression and activity of an ER-derived protein targeted to the E. coli inner membrane but found no significant enhancement in activity when comparing SohB with, e.g., 28-tagged CYPs. Membrane association in the absence of N-terminal hydrophobic sequence has been observed before (Doray et al. 2001), suggesting that some CYPs are sufficiently hydrophobic to localize to the membrane in the absence of an N-terminal localization signal (Jensen et al. 2011). This is also supported by molecular dynamics studies suggesting interactions between helices located far downstream from the N-terminus and biological membranes (Denisov et al. 2012). The identification of SohB as a generic tool to enhance expression, while being native to the E. coli membrane insertional machinery, may become useful when engineering higher order membrane assemblies such as the suggested multi-CYP metabolons (Moller 2010; Laursen et al. 2015). Also, even though the lepB fusions expressed to a lower level than those based on, e.g., SohB, the chimeric proteins may become useful because they maintain the native plant membrane anchor and (presumably) topology. This is particularly relevant because our results clearly demonstrate that care should be taken when truncating CYPs, as exemplified with the inactive P69-truncated CYP79A1.
The presented work demonstrates the usefulness of the GFP-reporter approach for de-bugging and maximizing production of a group of related enzymes and suggests that there are no inherent limitations in using different standard E. coli strains and expression conditions for exploiting CYPs for biotechnological applications. Further, the peptides and membrane domains used in this study add new biobricks to a toolbox for engineering pathways of higher complexity such as the taxol biosynthetic pathway consisting of eight steps catalyzed by CYPs (Chau et al. 2004) (Biggs et al. 2016).
References
Bak S, Kahn RA, Nielsen HL, Moller BL, Halkier BA (1998a) Cloning of three A-type cytochromes P450, CYP71E1, CYP98, and CYP99 from Sorghum bicolor (L.) Moench by a PCR approach and identification by expression in Escherichia coli of CYP71E1 as a multifunctional cytochrome P450 in the biosynthesis of the cyanogenic glucoside dhurrin. Plant Mol Biol 36:393–405
Bak S, Kahn RA, Olsen CE, Halkier BA (1997) Cloning and expression in Escherichia coli of the obtusifoliol 14 alpha-demethylase of Sorghum bicolor (L.) Moench, a cytochrome P450 orthologous to the sterol 14 alpha-demethylases (CYP51) from fungi and mammals. Plant J 11:191–201
Bak S, Nielsen HL, Halkier BA (1998b) The presence of CYP79 homologues in glucosinolate-producing plants shows evolutionary conservation of the enzymes in the conversion of amino acid to aldoxime in the biosynthesis of cyanogenic glucosides and glucosinolates. Plant Mol Biol 38:725–734
Barnes HJ, Arlotto MP, Waterman MR (1991) Expression and enzymatic activity of recombinant cytochrome P450 17 alpha-hydroxylase in Escherichia coli. Proc Natl Acad Sci U S A 88:5597–5601
Biggs BW, Lim CG, Sagliani K, Shankar S, Stephanopoulos G, De Mey M, Ajikumar PK (2016) Overcoming heterologous protein interdependency to optimize P450-mediated Taxol precursor synthesis in Escherichia coli. Proc Natl Acad Sci U S A 113:3209–3214. doi:10.1073/pnas.1515826113
Chang MCY, Eachus RA, Trieu W, Ro D-K, Keasling JD (2007) Engineering Escherichia coli for production of functionalized terpenoids using plant P450s. Nat Chem Biol 3:274–277. doi:10.1038/nchembio875
Chau M, Jennewein S, Walker K, Croteau R (2004) Taxol biosynthesis: molecular cloning and of a cytochrome p450 characterization taxoid 7 beta-hydroxylase. Chemistry & Biology 11:663–672. doi:10.1016/j.chembiol.2004.02.025
Chen CD, Doray B, Kemper B (1998) A conserved proline-rich sequence between the N-terminal signal-anchor and catalytic domains is required for assembly of functional cytochrome P450 2C2. Arch Biochem Biophys 350:233–238. doi:10.1006/abbi.1997.0524
Costa S, Almeida A, Castro A, Domingues L (2014) Fusion tags for protein solubility, purification and immunogenicity in Escherichia coli: the novel Fh8 system. Front Microbiol 5:63. doi:10.3389/fmicb.2014.00063
Daley DO, Rapp M, Granseth E, Melen K, Drew D, Heijne von G (2005) Global topology analysis of the Escherichia coli inner membrane proteome. Science 308:1321–1323. doi:10.1126/science.1109730
Davies KM, Deroles SC (2014) Prospects for the use of plant cell cultures in food biotechnology. Curr Opin Biotechnol 26:133–140. doi:10.1016/j.copbio.2013.12.010
Denisov IG, Shih AY, Sligar SG (2012) Structural differences between soluble and membrane bound cytochrome P450s. J Inorg Biochem 108:150–158. doi:10.1016/j.jinorgbio.2011.11.026
Doray B, Chen CD, Kemper B (2001) N-terminal deletions and His-tag fusions dramatically affect expression of cytochrome p450 2C2 in bacteria. Arch Biochem Biophys 393:143–153. doi:10.1006/abbi.2001.2473
Drew D, Lerch M, Kunji E, Slotboom DJ, de Gier JW (2006) Optimization of membrane protein overexpression and purification using GFP fusions. Nat Methods 3:303–313. doi:10.1038/nmeth0406-303
Drew D, Slotboom D-J, Friso G, Reda T, Genevaux P, Rapp M, Meindl-Beinker NM, Lambert W, Lerch M, Daley DO, Van Wijk K-J, Hirst J, Kunji E, De Gier J-W (2005) A scalable, GFP-based pipeline for membrane protein overexpression screening and purification. Protein Sci 14:2011–2017. doi:10.1110/ps.051466205
Drew DE, Heijne von G, Nordlund P, de Gier JW (2001) Green fluorescent protein as an indicator to monitor membrane protein overexpression in Escherichia coli. FEBS Lett 507:220–224
Eugster HP, Bärtsch S, Würgler FE, Sengstag C (1992) Functional co-expression of human oxidoreductase and cytochrome P450 1A1 in Saccharomyces cerevisiae results in increased EROD activity. Biochem Biophys Res Commun 185:641–647
Gallage NJ, Møller BL (2015) Vanillin-bioconversion and bioengineering of the most popular plant flavor and its de novo biosynthesis in the vanilla orchid. Mol Plant 8:40–57. doi:10.1016/j.molp.2014.11.008
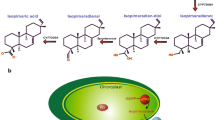
Hamberger B, Ohnishi T, Hamberger B, Séguin A, Bohlmann J (2011) Evolution of diterpene metabolism: Sitka spruce CYP720B4 catalyzes multiple oxidations in resin acid biosynthesis of conifer defense against insects. Plant Physiol 157:1677–1695. doi:10.1104/pp.111.185843
Hansen EH, Møller BL, Kock GR, Bünner CM, Kristensen C, Jensen OR, Okkels FT, Olsen CE, Motawia MS, Hansen J (2009) De novo biosynthesis of vanillin in fission yeast (Schizosaccharomyces pombe) and baker’s yeast (Saccharomyces cerevisiae). Appl Environ Microbiol 75:2765–2774. doi:10.1128/AEM.02681-08
Haudenschild C, Schalk M, Karp F, Croteau R (2000) Functional expression of regiospecific cytochrome P450 limonene hydroxylases from mint (Mentha spp.) in Escherichia coli and Saccharomyces cerevisiae. Arch Biochem Biophys 379:127–136. doi:10.1006/abbi.2000.1864
Hessa T, Kim H, Bihlmaier K, Lundin C, Boekel J, Andersson H, Nilsson I, White SH, Heijne von G (2005) Recognition of transmembrane helices by the endoplasmic reticulum translocon. Nature 433:377–381. doi:10.1038/nature03216
Jensen K, Osmani SA, Hamann T, Naur P, Møller BL (2011) Homology modeling of the three membrane proteins of the dhurrin metabolon: catalytic sites, membrane surface association and protein-protein interactions. Phytochemistry 72:2113–2123. doi:10.1016/j.phytochem.2011.05.001
Kawate T, Gouaux E (2006) Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14:673–681. doi:10.1016/j.str.2006.01.013
Kim SH, Cavaleiro AM, Rennig M, Nørholm MHH (2016) SEVA linkers: a versatile and automatable DNA backbone exchange standard for synthetic biology. ACS Synth Biol 5:1177–1181. doi:10.1021/acssynbio.5b00257
Kitaoka N, Lu X, Yang B, Peters RJ (2015) The application of synthetic biology to elucidation of plant mono-, sesqui-, and diterpenoid metabolism. Plant Metabolism and Synthetic Biology 8:6–16. doi:10.1016/j.molp.2014.12.002
Koch BM, Sibbesen O, Halkier BA, Svendsen I, Moller BL (1995) The primary sequence of cytochrome P450tyr, the multifunctional N-hydroxylase catalyzing the conversion of L-tyrosine to p-hydroxyphenylacetaldehyde oxime in the biosynthesis of the cyanogenic glucoside dhurrin in Sorghum bicolor (L.) Moench. Arch Biochem Biophys 323:177–186
Laursen T, Møller BL, Bassard J-E (2015) Plasticity of specialized metabolism as mediated by dynamic metabolons. Trends Plant Sci 20:20–32. doi:10.1016/j.tplants.2014.11.002
Lee C, Kang HJ, Hjelm A, Qureshi AA, Nji E, Choudhury H, Beis K, De Gier J-W, Drew D (2014) MemStar: a one-shot Escherichia coli-based approach for high-level bacterial membrane protein production. FEBS Lett 588:3761–3769. doi:10.1016/j.febslet.2014.08.025
Leonard E, Koffas MAG (2007) Engineering of artificial plant cytochrome p450 enzymes for synthesis of isoflavones by Escherichia coli. Appl Environ Microbiol 73:7246–7251. doi:10.1128/AEM.01411-07
Miroux B, Walker JE (1996) Over-production of proteins in Escherichia coli: mutant hosts that allow synthesis of some membrane proteins and globular proteins at high levels. J Mol Biol 260:289–298. doi:10.1006/jmbi.1996.0399
Moller BL (2010) Dynamic metabolons. Science 330:1328–1329
Morant M, Bak S, Møller BL, Werck-Reichhart D (2003) Plant cytochromes P450: tools for pharmacology, plant protection and phytoremediation. Curr Opin Biotechnol 14:151–162
Morrone D, Chen X, Coates RM, Peters RJ (2010) Characterization of the kaurene oxidase CYP701A3, a multifunctional cytochrome P450 from gibberelin biosynthesis. Biochem J 431:337–347. doi:10.1042/BJ20100597
Møller BL (2014) Disruptive innovation: channeling photosynthetic electron flow into light-driven synthesis of high-value products. Synth Biol 1:330–359
Nelson DR (2011) Progress in tracing the evolutionary paths of cytochrome P450. Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics 1814(1):14–18. doi:10.1016/j.bbapap.2010.08.008
Nour-Eldin HH, Hansen BG, Nørholm MHH, Jensen JK, Halkier BA (2006) Advancing uracil-excision based cloning towards an ideal technique for cloning PCR fragments. Nucleic Acids Res 34:e122–e122. doi:10.1093/nar/gkl635
Nørholm MHH (2010) A mutant Pfu DNA polymerase designed for advanced uracil-excision DNA engineering. BMC Biotechnol 10:21. doi:10.1186/1472-6750-10-21
Nørholm MHH, Toddo S, Virkki MTI, Light S, Heijne von G, Daley DO (2013) Improved production of membrane proteins in Escherichia coli by selective codon substitutions. FEBS Lett 587:2352–2358. doi:10.1016/j.febslet.2013.05.063
Paddon CJ, Westfall PJ, Pitera DJ, Benjamin K, Fisher K, McPhee D, Leavell MD, Tai A, Main A, Eng D, Polichuk DR, Teoh KH, Reed DW, Treynor T, Lenihan J, Fleck M, Bajad S, Dang G, Dengrove D, Diola D, Dorin G, Ellens KW, Fickes S, Galazzo J, Gaucher SP, Geistlinger T, Henry R, Hepp M, Horning T, Iqbal T, Jiang H, Kizer L, Lieu B, Melis D, Moss N, Regentin R, Secrest S, Tsuruta H, Vazquez R, Westblade LF, Xu L, Yu M, Zhang Y, Zhao L, Lievense J, Covello PS, Keasling JD, Reiling KK, Renninger NS, Newman JD (2013) High-level semi-synthetic production of the potent antimalarial artemisinin. Nature:496–528. doi:10.1038/nature12051
Pritchard MP, Ossetian R, Li DN, Henderson CJ, Burchell B, Wolf CR, Friedberg T (1997) A general strategy for the expression of recombinant human cytochrome P450s in Escherichia coli using bacterial signal peptides: expression of CYP3A4, CYP2A6, and CYP2E1. Arch Biochem Biophys 345:342–354. doi:10.1006/abbi.1997.0265
Qi X, Bakht S, Qin B, Leggett M, Hemmings A, Mellon F, Eagles J, Werck-Reichhart D, Schaller H, Lesot A, Melton R, Osbourn A (2006) A different function for a member of an ancient and highly conserved cytochrome P450 family: from essential sterols to plant defense. Proc Natl Acad Sci U S A 103:18848–18853. doi:10.1073/pnas.0607849103
Silva-Rocha R, Martínez-García E, Calles B, Chavarría M, Arce-Rodríguez A, de Las Heras A, Páez-Espino AD, Durante-Rodríguez G, Kim J, Nikel PI, Platero R, De Lorenzo V (2013) The Standard European Vector Architecture (SEVA): a coherent platform for the analysis and deployment of complex prokaryotic phenotypes. Nucleic Acids Res 41:D666–D675. doi:10.1093/nar/gks1119
Sonnhammer EL, Heijne von G, Krogh A (1998) A hidden Markov model for predicting transmembrane helices in protein sequences. Proc Int Conf Intell Syst Mol Biol 6:175–182
Sonoda Y, Cameron A, Newstead S, Omote H, Moriyama Y, Kasahara M, Iwata S, Drew D (2010) Tricks of the trade used to accelerate high-resolution structure determination of membrane proteins. FEBS Lett 12:2539–2547. doi:10.1016/j.febslet.2010.04.015
Sonoda Y, Newstead S, Hu N-J, Alguel Y, Nji E, Beis K, Yashiro S, Lee C, Leung J, Cameron AD, Byrne B, Iwata S, Drew D (2011) Benchmarking membrane protein detergent stability for improving throughput of high-resolution X-ray structures. Structure 19:17–25. doi:10.1016/j.str.2010.12.001
Studier FW (1991) Use of bacteriophage T7 lysozyme to improve an inducible T7 expression system. J Mol Biol 219:37–44
Studier FW (2005) Protein production by auto-induction in high-density shaking cultures. Protein Expr Purif 41:207–234. doi:10.1016/j.pep.2005.01.016
Sudhamsu J, Kabir M, Airola MV, Patel BA, Yeh S-R, Rousseau DL, Crane BR (2010) Co-expression of ferrochelatase allows for complete heme incorporation into recombinant proteins produced in E. coli. Protein Expr Purif 73:78–82. doi:10.1016/j.pep.2010.03.010
Søgaard KM, Nørholm MHH (2016) Side effects of extra tRNA supplied in a typical bacterial protein production scenario. Protein Sci 25:2102–2108. doi:10.1002/pro.3011
Wadsäter M, Laursen T, Singha A, Hatzakis NS, Stamou D, Barker R, Mortensen K, Feidenhans’l R, Møller BL, Cárdenas M (2012) Monitoring shifts in the conformation equilibrium of the membrane protein cytochrome P450 reductase (POR) in nanodiscs. J Biol Chem 287:34596–34603. doi:10.1074/jbc.M112.400085
Wagner S, Klepsch MM, Schlegel S, Appel A, Draheim R, Tarry M, Hogbom M, Van Wijk KJ, Slotboom DJ, Persson JO, de Gier JW (2008) Tuning Escherichia coli for membrane protein overexpression. Proc Natl Acad Sci U S A 105:14371–14376. doi:10.1073/pnas.0804090105
Williams PA, Cosme J, Sridhar V, Johnson EF, McRee DE (2000) Mammalian microsomal cytochrome P450 monooxygenase: structural adaptations for membrane binding and functional diversity. Mol Cell 5:121–131
Zelasko S, Palaria A, Das A (2013) Optimizations to achieve high-level expression of cytochrome P450 proteins using Escherichia coli expression systems. Protein Expr Purif 92:77–87. doi:10.1016/j.pep.2013.07.017
Zhang H, Im S-C, Waskell L (2007) Cytochrome b(5) increases the rate of product formation by cytochrome p450 2B4 and competes with cytochrome p450 reductase for a binding site on cytochrome p450 2B4. J Biol Chem 282:29766–29776. doi:10.1074/jbc.M703845200
Acknowledgements
We thank Victor de Lorenzo and the members of his laboratory for generously providing the pSEVA collection. We thank Tomas Laursen, Peter Naur, Søren Bak, Björn Hamberger, Johan Andersen-Ranberg, and Britta Hamberger for advice on CYPs and reductases. We thank David Drew, Daniel Daley, and Jan. Willem de Gier for discussions on E. coli gene expression and the GFP-based expression platform. SS is the recipient of VILLUM Foundation’s Young Investigator Programme grant VKR023128. This work was supported by the Novo Nordisk Foundation, from the VILLUM research center of excellence “Plant Plasticity,” from the UCPH Excellence Program for Interdisciplinary Research to Center of Synthetic Biology “bioSYNergy,” and by a European Research Council Advanced Grant Project No. 323034: LightdrivenP450s.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Ulla Christensen and Dario Vazquez-Albacete contributed equally to this work.
Electronic supplementary material
ESM 1
(PDF 2837 kb)
Rights and permissions
About this article
Cite this article
Christensen, U., Vazquez-Albacete, D., Søgaard, K.M. et al. De-bugging and maximizing plant cytochrome P450 production in Escherichia coli with C-terminal GFP fusions. Appl Microbiol Biotechnol 101, 4103–4113 (2017). https://doi.org/10.1007/s00253-016-8076-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-8076-5