Abstract
This study describes the thiosulfate-supported respiratory electron transport activity of Thiomonas bhubaneswarensis strain S10 (DSM 18181T). Whole-genome sequence analysis revealed the presence of complete sox (sulfur oxidation) gene cluster (soxCDYZAXB) including the sulfur oxygenase reductase (SOR), sulfide quinone reductase (SQR), sulfide dehydrogenase (flavocytochrome c (fcc)), thiosulfate dehydrogenase (Tsd), sulfite dehydrogenase (SorAB), and intracellular sulfur oxidation protein (DsrE/DsrF). In addition, genes encoding respiratory electron transport chain components viz. complex I (NADH dehydrogenase), complex II (succinate dehydrogenase), complex III (ubiquinone-cytochrome c reductase), and various types of terminal oxidases (cytochrome c and quinol oxidase) were identified in the genome. Using site-specific electron donors and inhibitors and by analyzing the cytochrome spectra, we identified the shortest thiosulfate-dependent electron transport chain in T. bhubaneswarensis DSM 18181T. Our results showed that thiosulfate supports the electron transport activity in a bifurcated manner, donating electrons to quinol (bd) and cytochrome c (Caa 3 ) oxidase; these two sites (quinol oxidase and cytochrome c oxidase) also showed differences in their phosphate esterification potential (oxidative phosphorylation efficiency (P/O)). Further, it was evidenced that the substrate-level phosphorylation is the major contributor to the total energy budget in this bacterium.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chemolithotrophic sulfur-oxidizing bacteria utilize reduced inorganic sulfur compounds (RISCs) as electron donors (energy source) in their respiratory activity (Friedrich et al. 2005; Ghosh and Dam 2009). Among the chemolithotrophic sulfur oxidizers, biochemical differences exist for the oxidative enzymes including the respiratory electron transport components employed in oxidation of particular species of RISCs (Kelly et al. 1997; Friedrich et al. 2005; Ghosh and Dam 2009). Reports are available on multiple pathways for sulfur oxidation (Ghosh and Dam 2009). The Sox (sulfur oxidizing) pathway that uses a group of enzymes transcribed by sox gene cluster (Kelly et al. 1997; Friedrich et al. 2005; Meyer et al. 2007; Ghosh and Dam 2009) completely oxidizes thiosulfate to sulfate without any detectible intermediate sulfur compounds (Dam et al. 2007). In addition, sulfur oxidation mediated by the S4I pathway (involves thiosulfate dehydrogenase) has been reported from obligate chemolithoautotrophs of Gammaproteobacteria (Halothiobacillus) and Acidithiobacillia (Acidithiobacillus and Thermithiobacillus) and from facultative chemolithoautotrophic Betaproteobacterium Advenella kashmirensis (Pronk et al. 1990; Meulenberg et al. 1993; Hallberg et al. 1996; Kelly et al. 1997; Dam et al. 2007; Ghosh and Dam 2009; Kikumoto et al. 2013). Further, the thiosulfate: quinone oxidoreductase (TQO)-mediated thiosulfate oxidation is reported from thermo-acidophilic archaeon Acidianus ambivalens (Muller et al. 2004), in which TQO acts as an alternative to thiosulfate dehydrogenase (cytochrome c). Unlike eukaryotes, the electron transport components in the respiratory chain of bacteria are highly flexible, both at electron input and output sites including the intermediate redox components, that help to survive under changing ecological conditions (Richardson 2000).
Although several studies have described the physiology and biochemistry of sulfur oxidation in facultative chemolithotrophs belonging to Alphaproteobacteria, the mechanism of RISC oxidation and its energy-yielding process is poorly understood in obligate chemolithotrophs of Betaproteobacteria, Gammaproteobacteria, and Epsilonproteobacteria (Ghosh and Dam 2009).
Thiomonas bhubaneswarensis DSM 18181T belongs to the class Betaproteobacteria and is capable of oxidizing thiosulfate during mixotrophic growth on mineral salts-thiosulfate-yeast extract medium (Panda et al. 2009). This bacterium fails to grow when thiosulfate is replaced with other reduced sulfur compounds such as tetrathionate, sulfite, or elemental sulfur (Panda et al. 2009). The thiosulfate-dependent growth of this obligately mixotrophic bacterium leads us to investigate the pathway(s) and mechanism of electron transport during thiosulfate oxidation. Our results suggested the presence of two different electron transport pathways [e.g., cytochrome c to Caa 3 and quinone to quinol oxidase (bd)] with appreciable differences in the rate of thiosulfate oxidation and the ability to develop proton motive force. Sox pathway appeared to be the primary pathway of thiosulfate oxidation; besides, the existence of an alternative pathway mediated by thiosulfate dehydrogenase was postulated based on the genomic information and enzymatic activity. Finally, we have described the mechanism of electron transport during the oxidation of thiosulfate to sulfate in T. bhubaneswarensis.
Materials and methods
Bacterial growth and culture conditions
T. bhubaneswarensis DSM 18181T used in this study was reported by Panda et al. (2009). Growth, thiosulfate utilization, and sulfate production by this bacterium were examined by growing in a 500-ml culture flask containing 100 ml of liquid mineral salts-thiosulfate-yeast extract medium at 37 °C and 200 rpm for 72 h. The medium contained (l−1) the following: Na2HPO4, 4.0 g; KH2PO4, 1.5 g; CaCl2·5H2O, 0.01 g; NH4Cl, 1.0 g; MgSO4·7H2O, 0.5 g; Na2S2O3·5H2O, 5.0 g; and yeast extract powder, 2.0 g. The medium additionally contained phenol red (0.02 g l−1) to detect acid formation. The pH of the medium was adjusted to 7.5 with 10 M NaOH and autoclaved. Solid agar medium was prepared by adding agar powder (15.0 g l−1). Utilization of inorganic sulfur compounds was tested in mineral salts medium containing various concentrations of different inorganic sulfur sources (1.0, 2.5, or 5.0 g l−1). The sulfur compounds used were sodium thiosulfate, tetrathionate, sulfite, and elemental sulfur (Panda et al. 2009). The growth of cells was monitored by measuring the optical density at 600 nm using appropriate blank.
Genomic DNA preparation, sequencing, and assembly
The genomic DNA was isolated using standard methods (Sambrook et al. 1989). The draft genome of T. bhubaneswarensis DSM 18181T was generated at the DOE Joint Genome Institute (JGI) using the Illumina HiSeq 2000 platform (Bennett 2004). An Illumina standard shotgun library was constructed and sequenced using the Illumina HiSeq 2000 platform. The raw Illumina read sequences were filtered using DUK program developed by JGI, USA. The filtered Illumina reads were assembled using Velvet (Zerbino and Birney 2008), wgsim (https://github.com/lh3/wgsim), and Allpaths-LG (Gnerre et al. 2011) tools. Genes were identified using Prodigal (Hyatt et al. 2010) and GenePRIMP (Pati et al. 2010) programs. The predicted genes/candidate protein-coding sequences (CDSs) were annotated using the NCBI-nr, UniProt, TIGRFam, Pfam, KEGG, COG, and InterPro databases within the Integrated Microbial Genomes (IMG) platform (http://www.ncbi.nlm.nih.gov/pubmed/24165883) developed by the JGI (Markowitz et al. 2009).
Nucleotide sequence accession number
The draft genome sequence of T. bhubaneswarensis DSM 18181T is deposited at NCBI GenBank under the accession no. NZ_LIPV00000000.1. The version described in this paper is version NZ_LIPV00000000.1 and consists of sequences LIPV01000001–LIPV01000023.
Preparation of cell-free extract
Cells were harvested from logarithmic phase by centrifuging at 25 °C (10,000×g for 10 min). The pellet (wet cell mass ≈5–6 g) was washed twice with 10 ml of suspension buffer (100 mM Tris-HCl, pH 7.2 containing 0.1 mM EDTA) and finally resuspended in 40 ml of same buffer. The cell suspension was sonicated in two batches of 20 ml each (35% amplitude, 30-s pulse on/30-s pulse off) for 20 min on ice (Vibra-Cell sonicator). Unbroken cells were removed (20,000×g centrifugation for 60 min), and the supernatant (crude extract) was centrifuged at 144,000×g for 80 min at 4 °C to obtain the soluble (supernatant) and particulate (pellet) fractions. The particulate fraction was washed twice with 10 ml of suspension buffer. Both fractions were aliquoted in small volume of same buffer and stored at −80 °C until use.
Artificial electron donors and electron transport inhibitors
In addition to thiosulfate, artificial electron donors such as durohydroquinol (DQH2 reduces bc 1 complex and other complexes that utilize quinols) and reduced TMPD (N,N,N′,N′-tetramethyl-p-phenylediamine (TMPDH2) reduces cytochrome c and quinol oxidases) were used to identify the redox components of electron transport chain present in T. bhubaneswarensis DSM 18181T. Further, inhibitors like rotenone (an inhibitor of complex I), thenoyltrifluoroacetone (TTFA, an inhibitor of complex II), antimycin A (an inhibitor of complex III), 2-n-heptyl-4-hydroxyquinoline N-oxide (HQNO, inhibitor of complexes that oxidize quinols), potassium cyanide (KCN, inhibitor of heme-copper oxidase superfamily complexes viz., bo 3, aa 3, cbb 3, and bd), azide (an inhibitor of complex IV), and N-ethylmaleimide (NEM, a sulfhydryl-binding agent) were also used to decipher the route of thiosulfate-mediated electron flow in the T. bhubaneswarensis DSM 18181T.
Substrate-dependent O2 consumption
Oxygen consumption was measured polarographically with a Clark-type O2 electrode assembly (model DW1; Hansatech Instruments Ltd.) at 25 °C. Cells grown in mineral salts-thiosulfate-yeast extract medium were harvested from mid log phase by centrifugation at 10,000×g for 10 min, washed, and resuspended in 50 mM sodium phosphate buffer (pH 7.0) for thiosulfate-, tetrathionate-, elemental sulfur-, sulfite-, DQH2-, and TMPDH2-dependent O2 uptake assay. Substrate-dependent O2 uptake was measured in the whole cells, crude extract, and soluble and particulate fractions. The thiosulfate assay mixture contained 20 mM of Na2S2O3, whole cells, and 50 mM of sodium phosphate buffer (pH 7.0) in a final volume of 1.0 ml. A similar assay mixture was used to measure tetrathionate-, elemental sulfur-, DQH2-, and TMPDH2-dependent O2 uptake except that 20 mM of tetrathionate, 0.64 mg (≈20 mM) of elemental sulfur, 0.5 mM of DQH2, and 0.4 mM of TMPDH2 were used instead of thiosulfate. The sulfite-dependent O2 uptake assay contained 20 mM of Na2SO3, whole cells, and 50 mM of Tris-HCl buffer (pH 8.0) in a final volume of 1.0 ml. Whole cells equivalent to 50 μg of protein were used in all assay system. Similar procedure was followed for the crude extract and soluble and particulate fractions except that the protein concentration was maintained at a concentration of 1 mg ml−1. Substrate-dependent O2 uptake rate was expressed as nanomoles of O2 consumed per minute per milligram of protein.
Further, characterization of electron transport components of the respiratory chain was performed by using the site-specific electron donors and inhibitors. KCN and azide were used at concentrations of 1 μM to 10 mM whereas other inhibitors used at 1 to 100 μM. The membrane energization studies were performed using uncouplers at concentrations of 1 to 100 μM. DQH2 was prepared following the method of Izawa and Pan (1978). Unless otherwise mentioned, inhibitors and uncouplers were added to the reaction mixture 1 min after substrate addition. The effect of valinomycin (K+) was examined in the presence of 5 mM KCl in the reaction mixture.
Phosphate esterification
Phosphate esterification (phosphorylation) efficiency of the bacterium was monitored using whole cells. Quantification of ATP was made following the luciferin-luciferase reaction (Lu and Kelly 1983) using CellTiter-Glo Luminescent Cell Viability Assay kit (Promega) with minor modification. The reaction mixture consisted of 50 mM potassium phosphate buffer (pH 7.8), 6 mM MgCl2, 10 mM NaF, 2 mM ADP, and bacterial cell suspension (equivalent to 43 μg protein) in 1-ml volume. Under this assay condition, bacterial cells maintained linear O2 consumption up to 5 min. The reaction was allowed to proceed for 4 min, following the addition of the substrate (thiosulfate or DQH2). The reaction solution was centrifuged at 15,000×g for 5 min at 4 °C. The cell pellet was lysed with boiling distilled water (Yang et al. 2002). The lysed bacterial cells were used to measure the luminescence intensity after adding the luciferin-luciferase solution in 1:1 ratio. The quantity of ATP was determined by comparing luminescence from lysed cells and quantified using standard curve prepared with ATP in the range of 0.01–1.0 μM. Depending on the experimental approach, the effect of carbonyl cyanide m-cholorophenyl hydrazone (CCCP) was studied. The marginal luminescence obtained from ADP was deducted from the final estimation of ATP synthesized. All the solutions for phosphate esterification studies were prepared in sterile double-distilled water and kept in glass containers.
Cytochrome spectral analysis
The cytochrome(s) spectral analysis was performed with whole cells by measuring the difference spectrum of reduced-minus-air-oxidized samples, in the wavelength of 350–750 nm at 25 °C using a Varian Cary 300 spectrophotometer. Reductants used were thiosulfate, TMPDH2, and DQH2. As and when required, ammonium persulfate and dithionite were used as effective oxidizing and reducing agents, respectively. The reaction was performed in 1 ml of 100 mM Tris-HCl buffer (pH 7.2). Similar procedure was followed to analyze the cytochrome(s) in soluble and particulate fractions.
Thiosulfate dehydrogenase assay
The thiosulfate dehydrogenase (Tsd) activity was assayed in soluble and particulate fractions as described by Hensen et al.(2006). The activity was measured in 1 ml of reaction buffer (100 mM Tris-HCl, pH 7.2) containing 1 mM ferricyanide [K3Fe(CN)6], 5 mM thiosulfate, and crude enzyme. Enzyme activity was determined spectrophotometrically at 25 °C by measuring the rates of reduction of ferricyanide at 420 nm with an extinction coefficient of (ε = 1.09 mM−1 cm−1) and was expressed as nanomoles of ferricyanide reduced per minute per milligram of protein. The reaction was started by addition of thiosulfate, and appropriate corrections were made for substrate and enzyme blanks.
Analytical methods
Protein concentration was determined by dye binding method (Bradford 1976) with bovine serum albumin as standard. Concentrations of thiosulfate and tetrathionate in culture filtrates were measured according to Kelly et al. (1969). The sulfate content was determined by following the methods of Berglund and Sorbo (1960) and Gleen and Quastel (1953). The statistical significance of inhibitor effect on thiosulfate oxidation was determined by using ANOVA on PRISM 5.01. The standard deviation from mean value was shown as “vertical bars” and “±” values in figures and tables, respectively.
Results
Genomic properties
Whole-genome sequencing of T. bhubaneswarensis DSM 18181T revealed the presence of sox genes including the components of electron transport chain responsible for sulfur oxidation. The final draft assembly of the genome of T. bhubaneswarensis DSM 18181T consisted of 25 contigs in 23 scaffolds; the largest contig was 542.4 kb long, and N50 contig size was 288.4 kb. The total genome size was 3.2 Mb with a G+C content of 65.0%. The draft genome sequence had 3057 CDSs, 43 tRNAs, and 3 rRNA operons.
Genetic basis of sulfur oxidation and electron transport system
The analysis of whole-genome sequence of T. bhubaneswarensis DSM 18181T revealed the presence of complete sox gene cluster (consisting of soxCDYZAXB and an ArsR family regulator). In addition, enzymes involved in sulfur oxidation pathway such as sulfur oxygenase reductase (SOR), sulfide-quinone oxidoreductase (SQR), flavocytochrome c (fcc), thiosulfate dehydrogenase (Tsd), sulfite dehydrogenase (SorAB), and intracellular sulfur oxidation protein (DsrE/DsrF) were present (Table S1). The genome also possessed essential subunits of respiratory electron transport chain viz. complex I (NADH dehydrogenase), complex II (succinate dehydrogenase), and complex III (ubiquinone-cytochrome c reductase). Further, genes encoding various types of terminal oxidases such as cytochrome c-type aa 3 oxidase (cox A, B, and C), cytochrome cbb 3 oxidase (ccoN, O, P, and Q), a bo 3 quinol type oxidase (cyoA, B, C, D, E), cytochrome bd (cyd A, B) quinol oxidase, and ba 3 oxidase were identified (Table S1).
Physiology of sulfur oxidation
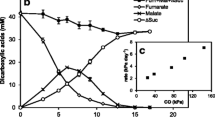
Different RISCs viz., thiosulfate, tetrathionate, sulfite, or elemental sulfur were supplemented in mineral salts-yeast extract medium to check the growth of T. bhubaneswarensis DSM 18181T. However, except thiosulfate, no other sulfur compounds supported the growth (data not shown). Growth was not observed in the absence of yeast extract. Further, no autotrophic growth was observed in the presence of thiosulfate. The growth rate and the change in pH of the medium with time have been shown in Fig. 1a. In mineral salts-thiosulfate-yeast extract medium, thiosulfate was consumed entirely within 36 h producing sulfate in 1 (S2O3 −2): 2 (SO4 −2) mole stoichiometry (Fig. 1b). However, tetrathionate was not detected in the growth medium during thiosulfate utilization. The specificity of thiosulfate oxidation was further examined in the respiratory activity of the organism.
Growth, change in medium pH, and the stoichiometric relation of substrate consumed to product formation in T. bhubaneswarensis DSM 18181T during mixotrophic growth on mineral salts-thiosulfate-yeast extract medium. a Time-dependent alteration in growth (squares) behavior of the bacterium and the associated decline in pH (circles). b Stoichiometry of thiosulfate (triangles) oxidation to sulfate (circles) production during the 72 h of growth. Each data point is the average of two independent determinations. The vertical bars represent the ± SD
Substrate-dependent respiratory activity
Reduced sulfur compounds like thiosulfate, tetrathionate, sulfite, or elemental sulfur were used for substrate-dependent O2 consumption activity in T. bhubaneswarensis. Addition of thiosulfate substantially enhanced the O2 consumption activity (referred as electron transport activity) in whole-cell assays. O2 consumption was not observed, when thiosulfate was replaced with tetrathionate, sulfite, or elemental sulfur. Besides thiosulfate, this bacterium also oxidized TMPDH2 and DQH2 (Table 1). The thiosulfate-supported electron transport rate was enhanced with increasing the concentration of thiosulfate (Fig. 2a). Reciprocal plot analysis (i.e., 1/S versus 1/V) suggested the presence of a high and a low affinity site (Fig. 2b–d) of electron donation by thiosulfate having K m 0.011 ± 0.006 and 0.54 ± 0.13 mM, respectively. Further, to know the site of electron donation by thiosulfate, O2 consumption activity was measured in the presence of HQNO. Thiosulfate concentration-dependent electron transport activity in the presence of HQNO and its reciprocal analysis (Fig. 2e, f, respectively) indicated that the low affinity site of electron donation by thiosulfate was HQNO insensitive.
Respiratory O2 consumption activity in whole-cell assays of T. bhubaneswarensis DSM 18181T . The O2 consumption activity with the increasing concentrations of thiosulfate is shown in a. The reciprocal analysis of rate (1/V) versus thiosulfate concentration (1/[S]) has been shown in 2(b). c, d Reciprocal plotting in an extended scale showing the presence of a high- (c) and low- (d) affinity sites of thiosulfate-dependent electron donation to the electron transport chain. e, f Electron transport rate with increasing concentrations of thiosulfate in presence of HQNO and the reciprocal analysis of the same, respectively. Each data point is the average of two independent determinations
Inhibitor and uncoupler sensitivity of electron transport in whole cells
Different inhibitors and uncouplers were used to know the thiosulfate-dependent electron donation site in the electron transport chain of T. bhubaneswarensis DSM 18181T. Thiosulfate oxidation in T. bhubaneswarensis DSM 18181T showed varied sensitivity towards three inhibitors HQNO, KCN, and azide (Table 2). Thiosulfate oxidation was inhibited by about 30–35% in the presence of HQNO (Table 2, Fig. 2e, f); the degree of inhibition in the presence of azide was much lower than reported for other bacterial species (Beffa et al. 1992; Meulenberg et al. 1992; Suzuki 1999; Masau et al. 2001). Oxidation of DQH2 was inhibited by nearly 80–85% in the presence of HQNO. Unlike thiosulfate and DQH2, the TMPDH2-supported electron flow was HQNO insensitive (Table 2).
KCN that interacts with the heme-copper moiety of O2 reductase complex was highly effective in inhibiting the electron transport activity (Fig. S1). A clear variation in KCN sensitivity was noticed with respect to the type of electron donor used to initiate the reaction (Fig. 3). The I 50 value of KCN (Fig. 3, inset table) with respect to donor DQH2 was 360 ± 20 μM > TMPDH2 (75 ± 12 μM) = thiosulfate (70 ± 14 μM). A significant difference was found in the KCN concentration to induce a nearly complete cessation of electron flow to O2 in the presence of thiosulfate or DQH2. In this regard, 2 mM KCN completely inhibited the reaction rate supported by thiosulfate, whereas addition of 5 mM KCN inhibited about 65% when DQH2 was used as electron donor. The lipophilic electron donor (TMPDH2)-supported activity was inhibited by nearly 95% at 5 mM KCN (Fig. 3). A marginal inhibition of the thiosulfate-supported electron flow was observed in the presence of azide and remained completely insensitive to inhibitors like TTFA, antimycin A, NEM, and rotenone (Table 2). The maximum inhibitory effect in the presence of high KCN (2 mM) suggested involvement of the bd complex, which was also suggested by the inhibitory effect of HQNO (Table 2). This observation was also supported by the requirement of high concentration of KCN to inhibit DQH2-supported O2 consumption rate (Table 3).
Relative inhibition of donor-dependent electron transport activity with increasing concentration of KCN. The respective donor-dependent I 50 values of KCN inhibition are shown in the inset table. The 100% activity for thiosulfate, TMPDH2, and DQH2 refers to 1338, 692, and 570 (nmol O2 consumed mg protein−1 min−1), respectively
The thiosulfate- and TMPDH2-supported electron flow was increased by 2-fold in the presence of CCCP, a known proton-conducting uncoupler that inhibits formation of proton motif force (Ikonomidis et al. 2008). However, this phenomenon was not discernible in DQH2-supported electron transport activity. Instead, in the presence of CCCP, the oxidation of DQH2 was reduced nearly 45–50% as compared to the control (thiosulfate) activity (Table 2). The effect of 2,4-dinitrophenol (2–4 DNP) was marginal. No alteration in electron transport rate was evident with ion-conducting uncoupler like valinomycin (K+).
Electron transport activity in crude extract and soluble and particulate fractions
The thiosulfate-, TMPDH2-, or DQH2-supported respiratory electron transport activity of T. bhubaneswarensis DSM 18181T was further measured in crude extract and particulate and soluble fractions to assess the oxidation of substrate and O2 reduction. None of these preparations supported the oxidation of thiosulfate, although they retained electron flow from TMPDH2 and DQH2 to O2. Reduced TMPD supported KCN-sensitive but HQNO-insensitive electron flow to O2 in the soluble and particulate fractions. The TMPDH2-dependent electron transport rate was nearly 3-fold higher in the particulate fraction as compared to the soluble fraction. The DQH2-supported O2 consumption was restricted to the particulate fraction, and nearly 80–85% of this activity was sensitive to HQNO inhibition. Further, the DQH2-supported electron flow was less sensitive to KCN as compared to TMPDH2 and thiosulfate (Table 3). CCCP failed to stimulate the TMPDH2-dependent O2 consumption in the particulate fraction, suggesting that the particulate fraction contains non-sealed membrane.
Cytochromes involved in electron transport activity
Since cytochromes are active redox components of electron transport chain and have wide variation in types and occurrence, substrate-dependent differential spectral analysis was performed to assess the type of cytochrome(s) present in T. bhubaneswarensis DSM 18181T. The redox difference spectral analysis (reduced-minus-air oxidized) in whole cells showed the presence of asymmetric electronic transitions having spectral signatures for cytochrome c (α peak 552, 554 nm, β peak 520, 522 nm, and γ peak 420–423 nm), b (α peak 590–595 nm, β peak 556–562 nm, and γ peak 427–428 nm) aa 3 (α peak 603–610 nm and γ peak 442–445 nm), and d types (α peak 630–632 nm) (Fig. 4). The signature for cytochrome b was discernible mostly as humps. No difference was observed using ammonium persulfate as oxidant replacing air in thiosulfate reduced-minus-persulfate oxidized spectra. The addition of dithionite over and above thiosulfate neither affected the intensity nor the characteristic features of the spectra induced by thiosulfate (thiosulfate + dithionite reduced-minus-air oxidized).
Difference spectral analysis of whole-cell assays with thiosulfate (a), TMPDH2 (b), and durohydroquinol (c) as reductant against air-oxidized samples. Presence of cytochrome c, b, a, and d types was shown with corresponding α, β, and γ electronic transitions with respective marked wavelengths (numbers). For clear resolution, all the spectra have been shown at an extended scale. To obtain a clear picture of the transitions, the spectra (c2) was subjected to its second derivative (d 2 A / dλ 2, c2a) inset). The cell mass considered for the spectral characterization was 300 μg equivalent protein
The soluble fraction when subjected to difference spectral analysis with thiosulfate as reductant induced the cytochrome c (α peak 552 nm, β peak 520 nm, and γ peak 420 nm) as the prominent species without appearance of electronic transitions associated with cytochrome b (α peak 590–595 nm, β peak 556–562 nm, and γ peak 427–428 nm) or aa 3 (α peak 603–610 nm and γ peak 442–445 nm) (Fig. 5 (panel 1: a1, a2)). However, no O2 consumption activity was noticed in the presence of thiosulfate in soluble fraction. The reduction of aa 3 (α peak 610 nm and γ peak 443 nm)-type cytochrome with measurable O2 consumption activity was clearly evident in the presence of TMPDH2 (Fig. 5 (panel 1: b1, b2)). The thiosulfate-mediated reduced spectra remain insensitive to HQNO. Further, addition of DQH2 induced the reduction of c-type cytochrome (α peak 554 nm, β peak 523 nm, and γ peak 420 nm) (Fig. 5 (panel 1: c1, c2)) but did not supported O2 consumption.
Difference cytochrome spectral analysis of soluble (panel 1) and particulate (panel 2) fractions prepared from T. bhubaneswarensis DSM 18181T . The cytochrome reduction was achieved using thiosulfate, TMPDH2, and DQH2 against air-oxidized samples. Mark the specific spectral resolution (443 and 610 nm) induced by TMPDH2 in soluble fraction (panel 1: b1, b2). Also note the appearance of 427 and 595–610 nm absorption in presence of DQH2 in particulate fraction (panel 2: b, c). The protein concentration was maintained at 220 μg ml−1
Unlike the soluble fraction, thiosulfate failed to show any spectral resolution in particulate fraction (Fig. 5 (panel 2: a)). Signatures for the presence of a 3 - (α peak 603–610 nm and γ peak 443 nm) and b- (α peak 590–595 nm and γ peak 427 nm) type cytochromes appeared as humps in presence of DQH2 (Fig. 5 (panel 2: c)), which were further amplified (443, 576, and 608 nm) by the addition of TMPDH2 (Fig. 5 (panel 2: b)). However, the inclusion of HQNO with DQH2 completely suppressed the reduction of cytochrome.
KCN effect on the spectral characteristics of soluble fraction
As KCN showed varied inhibitory effect towards thiosulfate-dependent O2 consumption (Table 2 and Fig. 3) in whole cells, further studies were conducted with soluble and particulate fractions. The measurement of difference spectra in presence of cyanide (KCN-minus-air) induced electronic transition, having similar signature as found for c-type cytochrome. However, the intensity of absorption increased in a time-dependent manner achieving the maximal value in 45 min. A discernible hump at 595-nm region was clearly evident in the samples supplemented with KCN (Fig. S2).
Phosphorylation efficiency
Phosphorylation activities were measured to assess the efficiency of functional terminal oxidases in terms of energy conservation in T. bhubaneswarensis DSM 18181T during oxidation of natural (thiosulfate) and artificial electron donor (DQH2). Esterification efficiency showed significant differences with respect to specific electron donors. Thiosulfate-dependent esterification potential was 1.4–1.6-fold higher in comparison to DQH2-supported esterification activity (Table 4, a). In order to pull out the fraction of phosphate esterified by electron flow through oxidase complex (oxidative phosphorylation) from non-oxidative (substrate level including others), the experiment was further repeated without electron donor. The difference in phosphorylation activity between total (Table 4, a) and without electron donor (Table 4, d) represented the magnitude of oxidative phosphorylation (Table 4, a–d). Thiosulfate-supported oxidative phosphorylation activity was several folds higher (nearly 4–5) as compared to DQH2, which was also reflected in their respective oxidative phosphorylation efficiencies (i.e., P/O). An interesting observation was recorded with the selective effect of CCCP towards thiosulfate- and DQH2-supported O2 consumption and esterification activities. While the CCCP enhanced the thiosulfate-dependent O2 consumption rate about 1.44-fold, it inhibited DQH2-dependent O2 consumption about 2.0-fold. CCCP also showed an inhibitory effect on the non-oxidative phosphorylation efficiency to a higher extent with thiosulfate than DQH2 (compare c and d, Table 4).
Thiosulfate dehydrogenase activity
Genomic analysis predicted that the genes encoding thiosulfate dehydrogenase (Tsd) were present (Table S1) in T. bhubaneswarensis DSM 18181T. In this regard, the activity of Tsd enzyme was measured in particulate and soluble fractions. The Tsd activity was 5-fold higher in soluble fraction than particulate fraction (Fig. S3 and inset therein).
Discussion
In view of the substrate specificity (only thiosulfate-dependent growth), T. bhubaneswarensis DSM 18181T is unique among sulfur-oxidizing bacteria (Loya et al. 1982; Lu and Kelly 1983; Ghosh and Dam 2009). The presence of complete sox gene cluster (sox CDYZAXB) in the genome and stoichiometric conversion of thiosulfate to sulfate with concomitant decline in the pH of the growth medium (Fig. 1a, b) suggests that T. bhubaneswarensis DSM 181811T has a functional Sox pathway (Dam et al. 2007; Houghton et al. 2016). Further, presence of genetic signatures for proteins such as SOR, SQR, fcc, and SorAB in the genome indicates that T. bhubaneswarensis DSM181811T can grow on or oxidize other RISCs. However, this bacterium neither grows in presence of elemental sulfur, sulfite, and tetrathionate nor oxidizes these compounds (Table 1), suggesting that these proteins may not function efficiently if expressed during thiosulfate oxidation (Yin et al. 2014). The presence of the genetic signature of Tsd and its activity in the soluble and particulate fractions (Fig. S3) further suggest that T. bhubaneswarensis DSM 18181T can oxidize thiosulfate to tetrathionate. However, tetrathionate formation was not detected in the culture filtrates during thiosulfate oxidation. This can be explained by the fact that under normal physiological conditions (neutral pH), this bacterium uses the Sox pathway for the oxidation of thiosulfate to form sulfate, while under acidic condition (pH <6), thiosulfate oxidation proceeds via tetrathionate as an intermediate compound. In fact, thiosulfate is acid labile and there are a number of bacteria that oxidize thiosulfate to tetrathionate under acidic condition (Houghton et al. 2016).
Inhibitor studies indicated that the complex I (NADH dehydrogenase) and complex II (succinate dehydrogenase) that are highly sensitive to rotenone and TTFA, respectively, do not participate as electron carriers during thiosulfate oxidation. Partial inhibition of thiosulfate-dependent electron flow by HQNO (Table 2) suggests that electrons released by the oxidation of thiosulfate are received at multiple sites in T. bhubaneswarensis DSM 18181T electron transport chain. This was further supported by the measurement of O2 consumption activity with increasing concentration of thiosulfate, which yields a low- (cytochrome c) and a high- (quinone pool) affinity site of electron donation (Fig. 2). The electron donation of thiosulfate to cytochrome c and quinone pool was identified to be an independent process without involving cytochrome bc 1 since there was no inhibition of thiosulfate oxidation in presence of antimycin A (Table 2).
Unlike HQNO, antimycin A is a specific inhibitor of high potential site (b H) of cytochrome bc 1 complex. Although HQNO inhibits electron transport by binding to high (b H) and low (b L ) potential quinone binding sites of the bc 1 complex, it has other non-specific sites of inhibition, especially in bacterial electron transport system having multiple complexes of O2 reductases that include both cytochrome c and quinol oxidases (Esposti 1989; Brasseur et al. 2004). Evidence for presence of quinol oxidase in T. bhubaneswarensis DSM 18181T is obtained from whole-genome sequence analysis (Table S1) and from DQH2-supported electron transport activity and its high sensitivity to HQNO (Tables 2 and 3). The DQH2-supported electron flow to O2 was identified in the whole cells, crude, and particulate fractions but not in the soluble fraction (Tables 1 and 3), suggesting that the quinol oxidase in T. bhubaneswarensis DSM 18181T is membrane bound. Therefore, it is advocated that thiosulfate-dependent electron donation at the high-affinity site with low V max is restricted to quinol oxidase. The low affinity site with high V max was unaffected by HQNO, suggesting a preference for another type of oxidase, probably cytochrome c oxidase (Fig. 6). The argument for the presence of a quinol type of oxidase activity is supported by KCN inhibition studies. In agreement with previous reports, the I 50 value of KCN inhibition of DQH2-supported electron flow to O2 is nearly 5-fold high as compared to TMPDH2- or thiosulfate-dependent electron donation activity (Fig. 3, inset table), suggesting that T. bhubaneswarensis DSM 18181T prefers to donate electrons to cytochrome c when grown on thiosulfate. As reported earlier (Junemann 1997), the O2 reduction activity by TMPDH2 in a KCN-sensitive but HQNO-insensitive manner suggests that the site of electron donation by the compound lies in the terminal oxidase complexes.
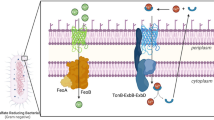
A hypothetical model showing overall mechanism of thiosulfate oxidation and electron transport in T. bhubaneswarensis DSM 18181T. a Thiosulfate oxidation by Sox pathway: SoxY thiosulfate-binding protein (WP_055450809.1), SoxZ sulfur compound chelating protein (WP_055450810.1), SoxA diheme cytochrome (WP_055450811.1), SoxX monoheme cytochrome (WP_055450812.1), SoxC sulfane dehydrogenase subunit (WP_055450807.1), and SoxD sulfur dehydrogenase subunit (WP_055450808.1). b Thiosulfate oxidation by thiosulfate quinone oxidoreductase (TQO) to quinol oxidase (bd) in a high-affinity and HQNO-sensitive manner. c Different routes of electron flow from donors like thiosulfate (blue arrow) and DQH2 (dashed green arrow) in T. bhubaneswarensis DSM 18181T bacterium. The inhibitory effect of HQNO for DQH2 and thiosulfate-supported electron flow has been depicted in red arrow. The diagram also shows the low- (solid blue arrow) and high-affinity (dotted blue arrow) sites of electron donation by thiosulfate including the low- and high-concentration-dependent KCN interactions (dotted black arrows) (Color figure online)
The cytochrome spectral analysis with whole cells provides further insight on the type of cytochrome participating in electron flow in this bacterium. The major form of cytochrome was c type (423, 520, 552–554 nm), although signature pertaining to b (427, 524, 558–560 nm), aa 3 (443–445 nm, 603–610 nm), and d (α peak 630 nm) was also evident as discernible humps (Fig. 4). This phenomenon may be due to the presence of high concentration of cytochrome c (Gel’man et al. 1967; Loya et al. 1982; Sorokin et al. 1999). The characteristic of oxidation of DQH2 in particulate fraction in a HQNO-sensitive manner (Table 3) with low affinity to KCN inhibition (Fig. 3, inset table) suggests the presence of quinol-type oxidase. The appearance of b, a 3 , and d types of cytochrome signatures in whole cells (Fig. 4) further suggests the involvement of bd-type quinol oxidase and a ba 3 oxidase as well (Brasseur et al. 2004). However, the major O2 reductase in T. bhubaneswarensis DSM 18181T is suggested to be Caa 3 type as evidenced from both cytochrome spectral analysis (Fig. 5 (panel 1)) and inhibition effect by KCN in soluble fraction (Table 3). KCN-supported increase in the intensity of c-type cytochrome signature in a time-dependent manner (Fig. S2) and the presence of aa 3 type of cytochrome signature (Fig. 4) corroborate this suggestion. A similar phenomenon was reported in Bacillus subtilis (Assempour and Hill 1997). However, still, there is some kind of reverse electron flow from a 3 to a and eventually to cytochrome c/CuA complex, thus inducing a time-dependent increase in cytochrome c absorption. A similar phenomenon is found in Thermus thermophilus (Siletsky et al. 2013). The reverse electron flow in T. bhubaneswarensis DSM 18181T may be one of the strategies to increase proton pumping capacity, thus increasing the efficiency of ATP synthesis (Siletsky et al. 2013).
The respiratory electron transport activity in bacteria is coupled to ATP synthesis by the formation of proton gradient. Protonophores like CCCP (anionic proton conducting uncouplers) and 2–4 DNP are known to collapse the formation of this proton gradient due to their own proton carrying nature (Grogan 1984; Guerrero and Makemson 1989). Protonophores while eliminate the ATP synthesis also induce a stimulation in electron transport rate. In this regard, only CCCP was able to show uncoupling activity on thiosulfate- and TMPDH2-supported electron flow by stimulating the rate of O2 consumption (Table 2) with concomitant termination of ATP synthesis (Table 4). 2–4 DNP, which is known to act as a potent inhibitor for thiosulfate-dependent electron transport activity in thiosulfate-oxidizing bacteria (Suzuki 1999; Masau et al. 2001), failed to exhibit any inhibition in T. bhubaneswarensis DSM 18181T. Instead, a marginal stimulation in O2 consumption activity was found during thiosulfate oxidation. These results suggest that the thiosulfate-dependent electron flow is coupled to ATP synthesis, which is apparent for the artificial electron donor TMPDH2 as well. With regard to the sites of H+ translocation in T. bhubaneswarensis DSM 18181T during electron flow from thiosulfate to O2, it could be assumed that the sites of proton translocation are associated with the activity of cytochrome oxidase. In many sulfur-oxidizing bacteria, the P/O ratio around 1.0 suggests the presence of a single proton-translocating site corresponding to oxidase complexes (Lu and Kelly 1983). We observe that a similar P/O ratio around 1.0 obtained from phosphate esterification studies during electron flow from thiosulfate to O2 in T. bhubaneswarensis DSM 18181T suggests the presence of only one proton-translocating site. Moreover, the presence of a single site of proton translocation and the extent of oxidative ATP synthesis may not be sufficient to support the energy requirement for growth of the bacterium. Our examination on phosphate esterification potential in T. bhubaneswarensis DSM 18181T revealed the operation of an alternative energy-yielding mechanism in the form of high substrate-level phosphorylation activity (Table 4). The oxidative phosphorylation potential in terms of P/O ratio obtained with DQH2 was nearly 6–6.5-fold lower than thiosulfate-supported P/O ratio, indicating that the quinol oxidase has poor contribution towards oxidative phosphorylation. The quinol oxidase may be an additional advantage for the growth of the bacterium during low concentration of thiosulfate and high O2 availability.
Genomic analysis predicted the presence of cytochrome c-type Tsd (Table S1) in this bacterium. However, partial inhibition of thiosulfate-supported electron transport activity in the presence of HQNO (Fig. 2e, f) suggests that the Tsd is the quinone type (TQO) (Wentzien and Sand 1999). HQNO sensitivity of TQO-mediated thiosulfate oxidation via the reduction of quinol oxidase has been proposed previously (Brasseur et al. 2004). Based on this observation, it can be postulated that two pathways of electron transport function during thiosulfate oxidation with different electron donation affinities (Fig. 6). The low-affinity site appears to operate in response to Sox complex-mediated thiosulfate oxidation, while high-affinity site acts during Tsd-dependent thiosulfate oxidation (Fig. 6).
In conclusion, T. bhubaneswarensis DSM 18181T oxidized thiosulfate in a bifurcated manner and used two different pathways. The primary pathway is Sox (Kelly-Friedrich) pathway, which operates under normal physiological condition and oxidizes thiosulfate to sulfate and transfers electron to cytochrome c to cytochrome c oxidase (Caa 3). However, under acidic condition (pH <6) and low concentration of thiosulfate, this bacterium may use TQO for thiosulfate oxidation by transferring electron through quinone pool to bd oxidase.
References
Assempour M, Hill BC (1997) Cyanide binding to different redox states of the cytochrome caa3 complex from Bacillus subtilis; a member of the cytochrome oxidase super-family of enzymes. Biochim Biophys Acta-Bioenergetics 1320:175–187
Beffa T, Berczy M, Aragno M (1992) Inhibition of respiratory oxidation of elemental sulfur (S0) and thiosulfate in Thiobacillus versutus and another sulfur-oxidizing bacterium. FEMS Microbiol Lett 90:123–127
Bennett S (2004) Solexa Ltd. Pharmacogenomics 5:433–438
Berglund F, Sorbo B (1960) Turbidimetric analysis of inorganic sulfate in serum, plasma and urine. Scand J Clin Lab Invest 12:147–153
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Brasseur G, Levican G, Bonnefoy V, Holmes D, Jedlicki E, Lemesle-Meunier D (2004) Apparent redundancy of electron transfer pathways via bc(1) complexes and terminal oxidases in the extremophilic chemolithoautotrophic Acidithiobacillus ferrooxidans. Biochim Biophys Acta 1656:114–126
Dam B, Mandal S, Ghosh W, Das Gupta SK, Roy P (2007) The S4-intermediate pathway for the oxidation of thiosulfate by the chemolithoautotroph Tetrathiobacter kashmirensis and inhibition of tetrathionate oxidation by sulfite. Res Microbiol 158:330–338
Esposti MD (1989) Prediction and comparison of the haem-binding sites in membrane haemoproteins. Biochim Biophys Acta 977:249–265
Friedrich CG, Bardischewsky F, Rother D, Quentmeier A, Fischer J (2005) Prokaryotic sulfur oxidation. Curr Opin Microbiol 8:253–259
Gel’man NS, Lukoyanova MA, Ostrovskii DN (1967) The respiratory chain of bacteria. pp 71–159. In respiration and phosphorylation of bacteria. Springer. doi:10.1007/978-1-4899-5526-5
Ghosh W, Dam B (2009) Biochemistry and molecular biology of lithotrophic sulfur oxidation by taxonomically and ecologically diverse bacteria and archaea. FEMS Microbiol Rev 33:999–1043
Gleen H, Quastel JH (1953) Sulphur metabolism in soil. Appl Microbiol 1:70–77
Gnerre S, Maccallum I, Przybylski D, Ribeiro FJ, Burton JN, Walker BJ, Sharpe T, Hall G, Shea TP, Sykes S, Berlin AM, Aird D, Costello M, Daza R, Williams L, Nicol R, Gnirke A, Nusbaum C, Lander ES, Jaffe DB (2011) High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proc Natl Acad Sci U S A 108:1513–1518
Grogan DW (1984) Interaction of respiration and luminescence in a common marine bacterium. Arch Microbiol 137:159–162
Guerrero MA, Makemson JC (1989) The cytochromes of luminous bacteria and their coupling to bioluminescence. Curr Microbiol 18:67–73
Hallberg KB, Dopson M, Lindstrom EB (1996) Reduced sulfur compound oxidation by Thiobacillus caldus. J Bacteriol 178:6–11
Hensen D, Sperling D, Truper HG, Brune DC, Dahl C (2006) Thiosulphate oxidation in the phototrophic sulphur bacterium Allochromatium vinosum. Mol Microbiol 62:794–810
Houghton JL, Foustoukos D, Flynn TM, Vetriani C, Bradley AS, Fike DA (2016) Thiosulfate oxidation by Thiomicrospira thermophila: metabolic flexibility in response to ambient geochemistry. Environ Microbiol. doi:10.1111/1462-2990.13232
Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11:119
Ikonomidis A, Tsakris A, Kanellopoulou M, Maniatis AN, Pournaras S (2008) Effect of the proton motive force inhibitor carbonyl cyanide-m-chlorophenylhydrazone (CCCP) on Pseudomonas aeruginosa biofilm development. Lett Appl Microbiol 47:298–302
Izawa S, Pan RL (1978) Photosystem I electron transport and phosphorylation supported by electron donation to the plastoquinone region. Biochem Biophys Res Commun 83:1171–1177
Junemann S (1997) Cytochrome bd terminal oxidase. Biochim Biophys Acta 1321:107–127
Kelly DP, Chambers LA, Trudinger PA (1969) Cyanolysis and spectrophotometric estimation of trithionate in mixture with thiosulfate and tetrathionate. Anal Chem 41:898–901
Kelly DP, Shergill JK, Lu WP, Wood AP (1997) Oxidative metabolism of inorganic sulfur compounds by bacteria. Antonie Van Leeuwenhoek 71:95–107
Kikumoto M, Nogami S, Kanao T, Takada J, Kamimura K (2013) Tetrathionate-forming thiosulfate dehydrogenase from the acidophilic, chemolithoautotrophic bacterium Acidithiobacillus ferrooxidans. Appl Environ Microbiol 79:113–120
Loya S, Yankofsky SA, Epel BL (1982) Lithotrophy to organotrophy conversion in Thiobacillus A2. Microbiology 128:865–874
Lu W-P, Kelly DP (1983) Thiosulphate oxidation, electron transport and phosphorylation in cell-free systems from Thiobacillus A2. Microbiology 129:1661–1671
Markowitz VM, Mavromatis K, Ivanova NN, Chen IM, Chu K, Kyrpides NC (2009) IMG ER: a system for microbial genome annotation expert review and curation. Bioinformatics 25:2271–2278
Masau RJ, Oh JK, Suzuki I (2001) Mechanism of oxidation of inorganic sulfur compounds by thiosulfate-grown Thiobacillus thiooxidans. Can J Microbiol 47:348–358
Meulenberg R, Pronk JT, Hazeu W, Bos P, Kuenen JG (1992) Oxidation of reduced sulphur compounds by intact cells of Thiobacillus acidophilus. Arch Microbiol 157:161–168
Meulenberg R, Scheer EJ, Pronk JT, Hazeu W, Bos P, Kuenen JG (1993) Metabolism of tetrathionate in Thiobacillus acidophilus. FEMS Microbiol Lett 112:167–172
Meyer B, Imhoff JF, Kuever J (2007) Molecular analysis of the distribution and phylogeny of the soxB gene among sulfur-oxidizing bacteria—evolution of the Sox sulfur oxidation enzyme system. Environ Microbiol 9:2957–2977
Muller FH, Bandeiras TM, Urich T, Teixeira M, Gomes CM, Kletzin A (2004) Coupling of the pathway of sulphur oxidation to dioxygen reduction: characterization of a novel membrane-bound thiosulphate:quinone oxidoreductase. Mol Microbiol 53:1147–1160
Panda SK, Jyoti V, Bhadra B, Nayak KC, Shivaji S, Rainey FA, Das SK (2009) Thiomonas bhubaneswarensis sp. nov., an obligately mixotrophic, moderately thermophilic, thiosulfate-oxidizing bacterium. Int J Syst Evol Microbiol 59:2171–2175
Pati A, Ivanova NN, Mikhailova N, Ovchinnikova G, Hooper SD, Lykidis A, Kyrpides NC (2010) GenePRIMP: a gene prediction improvement pipeline for prokaryotic genomes. Nat Methods 7:455–457
Pronk JT, Meulenberg R, Hazeu W, Bos P, Kuenen JG (1990) Oxidation of reduced inorganic sulphur compounds by acidophilic thiobacilli. FEMS Microbiol Lett 75:293–306
Richardson DJ (2000) Bacterial respiration: a flexible process for a changing environment. Microbiology 146:551–571
Sambrook J, Fritsch EF, Maniatis TA (1989) In molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Siletsky SA, Belevich I, Soulimane T, Verkhovsky MI, Wikstrom M (2013) The fifth electron in the fully reduced caa(3) from Thermus thermophilus is competent in proton pumping. Biochim Biophys Acta 1827:1–9
Sorokin DY, Cherepanov A, De Vries S, Kuenen GJ (1999) Identification of cytochrome c oxidase in the alkaliphilic, obligately chemolithoautotrophic, sulfur-oxidizing bacterium ‘Thioalcalomicrobium aerophilum’ strain AL 3. FEMS Microbiol Lett 179:91–99
Suzuki I (1999) Oxidation of inorganic sulfur compounds: chemical and enzymatic reactions. Can J Microbiol 45:97–105
Wentzien S, Sand W (1999) Polythionate metabolism in Thiomonas intermedia K12. Process Metallurgy 9:787–797
Yang NC, Ho WM, Chen YH, Hu ML (2002) A convenient one-step extraction of cellular ATP using boiling water for the luciferin-luciferase assay of ATP. Anal Biochem 306:323–327
Yin H, Zhang X, Li X, He Z, Liang Y, Guo X, Hu Q, Xiao Y, Cong J, Ma L, Niu J, Liu X (2014) Whole-genome sequencing reveals novel insights into sulfur oxidation in the extremophile Acidithiobacillus thiooxidans. BMC Microbiol 14:179
Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18:821–829
Acknowledgements
This work was supported in part by the funding received from the Department of Biotechnology, Government of India (D.O. no. BT/PR9712/NBD/52/91/2007), to SKD. Author KDN acknowledges the Council of Scientific and Industrial Research, Government of India, for providing the research fellowship. Thanks are due to Director, Institute of Life Sciences, for offering an adjunct faculty to SCS. The sequence data used in the study was produced by the US Department of Energy Joint Genome Institute (http://www.jgi.doe.gov/).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 645 kb)
Rights and permissions
About this article
Cite this article
Narayan, K.D., Sabat, S.C. & Das, S.K. Mechanism of electron transport during thiosulfate oxidation in an obligately mixotrophic bacterium Thiomonas bhubaneswarensis strain S10 (DSM 18181T). Appl Microbiol Biotechnol 101, 1239–1252 (2017). https://doi.org/10.1007/s00253-016-7958-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7958-x