Abstract
Burkholderia is an incredibly diverse and versatile Gram-negative genus, within which over 80 species have been formally named and multiple other genotypic groups likely represent new species. Phylogenetic analysis based on the 16S rRNA gene sequence and core genome ribosomal multilocus sequence typing analysis indicates the presence of at least three major clades within the genus. Biotechnologically, Burkholderia are well-known for their bioremediation and biopesticidal properties. Within this review, we explore the ability of Burkholderia to synthesise a wide range of antimicrobial compounds ranging from historically characterised antifungals to recently described antibacterial antibiotics with activity against multiresistant clinical pathogens. The production of multiple Burkholderia antibiotics is controlled by quorum sensing and examples of quorum sensing pathways found across the genus are discussed. The capacity for antibiotic biosynthesis and secondary metabolism encoded within Burkholderia genomes is also evaluated. Overall, Burkholderia demonstrate significant biotechnological potential as a source of novel antibiotics and bioactive secondary metabolites.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Burkholderia represents a rapidly expanding group of Gram-negative non-fermenting bacteria that occur worldwide in virtually all possible environments. Some species occur in plain soil or in planktonic form in fresh water, but most occur in association with an ever-increasing number of hosts including humans, animals (both vertebrates and invertebrates), plants and fungi. The type of interaction with these hosts is often not known, but a growing body of literature demonstrates that these interactions can be beneficial, harmful or both. Within the genus Burkholderia, a cluster of closely related species is known as the Burkholderia cepacia complex (Bcc) and presents particular challenges. Bcc bacteria are indeed well-known as rare but potentially life-threatening pathogens in patients with cystic fibrosis (CF) but simultaneously have been studied intensively for their biotechnological applications in plant growth promotion, biological control of plant pests and bioremediation.
The present mini-review provides an update on the taxonomy of these bacteria and addresses recent developments in terms of their capacity for secondary metabolism and antibiotic biosynthesis. The interested reader is referred to recent reviews on Burkholderia and Bcc that focused on extracellular products (Vial et al. 2007), relevance as contaminants in the pharmaceutical industry (Torbeck et al. 2011), conflicting life styles (Vial et al. 2011), potential for aromatic compound degradation (Pérez-Pantoja et al. 2012), common characteristics of plant-associated species (Suarez-Moreno et al. 2012), melioidosis (Currie 2015) and to the book entitled ‘Burkholderia: from genomes to function’ (2014).
Taxonomy
The genus Burkholderia belongs to the Betaproteobacteria class within the phylum of the Proteobacteria. When first described in 1992, it comprised seven species, two of which were subsequently reclassified into another novel genus, Ralstonia (Yabuuchi et al. 1992; Yabuuchi et al. 1995). During the past two decades, a large number of novel Burkholderia species have been reported and validly named. Yet, several species proved poorly characterised and needed further reclassification (Coenye et al. 1999; Coenye et al. 2000); at present (January 2016), the genus consists of 90 validly named species (Parte 2014) and a large number of uncultivated candidate species (van Oevelen et al. 2004; Verstraete et al. 2011; Lemaire et al. 2011; Lemaire et al. 2012). However, literature data and an analysis of publicly available 16S rRNA gene sequences suggest that many additional Burkholderia species await formal description. In addition, mining the Bcc PubMLST database (Jolley and Maiden 2010) using the 3 % threshold value of average concatenated allele sequence divergence for species delineation (Vanlaere et al. 2009; Peeters et al. 2013) demonstrated that also within the Bcc, a substantial number of additional Burkholderia species awaits formal naming (Vandamme and Peeters 2014).
The genus Burkholderia is phylogenetically diverse and consists of multiple deep-branching 16S rRNA lineages (Fig. 1). The first deep-branching Burkholderia clade comprises the type species, Burkholderia cepacia, and consists of all Bcc species, a group of plant-pathogenic species that includes Burkholderia gladioli, Burkholderia plantarii and Burkholderia glumae and a group of species closely related to the risk class 3 pathogens Burkholderia mallei and Burkholderia pseudomallei, the causative agents of glanders in Equidae and melioidosis in humans, respectively (Fig. 1). This first deep-branching Burkholderia clade comprises the majority of well-known pathogens in this genus but also includes many strains that have been used for plant growth promotion or biological control, such as Burkholderia vietnamiensis TVV74 and Burkholderia ambifaria AMMDT, respectively (Parke and Gurian-Sherman 2001). The former is a rice isolate which, when inoculated in field studies on rice, increased grain yield by 13 to 22 % (Tran Van et al. 2000). The latter was isolated from the rhizosphere of peas and has activity against Pythium aphanidermatum (responsible for pre- and post-emergence damping-off in peas) and Aphanomyces euteiches (responsible for root rot in peas) (Parke 1990; Bowers and Parke 1993; Heungens and Parke 2000; Heungens and Parke 2001; Parke and Gurian-Sherman 2001). In addition, although Burkholderia cenocepacia is generally considered the most problematic Bcc species in patients with CF (Lipuma 2010), recently, a genome sequence of a plant-beneficial endophytic B. cenocepacia strain with both biocontrol and plant growth-promoting characteristics was reported (Ho and Huang 2015).
Phylogenetic tree based on partial 16S rRNA gene sequences of Burkholderia species. Sequences (1125–1610 bp) were aligned against the SILVA SSU reference database using SINA v1.2.11 (http://www.arb-silva.de/aligner/) (Pruesse et al. 2012). Phylogenetic analysis was conducted using MEGA6 (Tamura et al. 2013). All positions containing gaps and missing data were eliminated, resulting in a total of 1087 positions in the final dataset. The optimal tree (highest log likelihood) was constructed using the maximum likelihood method and Tamura-Nei model (Tamura and Nei 1993). A discrete Gamma distribution was used to model evolutionary rate differences among sites (five categories [+G, parameter = 0.3498]) and allowed for some sites to be evolutionarily invariable ([+I], 68.6154 % sites). The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) are shown next to the branches if greater than 50 %. The sequence of Ralstonia solanacearum LMG 2299T was used as outgroup. The scale bar indicates the number of substitutions per site
The second deep-branching Burkholderia clade comprises Burkholderia glathei (Zolg and Ottow 1975) and 11 recently named Burkholderia species (Fig. 1). Several of these species were isolated from polluted soils. For instance, Burkholderia udeis comprises naphthalene-degrading isolates from a polycyclic aromatic hydrocarbon-contaminated hillside soil (Wilson et al. 2003; Vandamme et al. 2013), while Burkholderia jiangsuensis and Burkholderia zhejiangensis are methyl parathion-degrading bacteria isolated from methyl parathion-contaminated soil and a wastewater treatment system, respectively (Lu et al. 2012; Liu et al. 2014). Other species in this clade have been isolated from less studied sources such as fungal mycelia and mosses (Burkholderia sordidicola and Burkholderia grimmiae, respectively) (Lim et al. 2003; Tian et al. 2013). This clade also includes isolates from insect guts (Kikuchi et al. 2011; Shibata et al. 2013) and several candidate species with an endophytic lifestyle in plants (Carlier and Eberl 2012; Verstraete et al. 2013). Although these bacteria show a remarkable diversity in terms of ecological niches, to our knowledge, none of the present species within this group has been involved in human or animal infections. However, at least two novel B. glathei group species have been isolated from human sources including pleural fluid and blood, and await formal classification (own unpublished data).
The third deep-branching Burkholderia clade comprises more than 40 primarily environmental and plant-associated species, many of which are diazotrophic and have been documented as beneficial to their host (Suarez-Moreno et al. 2012) (Fig. 1). Among these species, Burkholderia fungorum is a most striking exception, as it has been isolated from a wide range of human and veterinary samples including human blood, cerebrospinal fluid, vaginal secretions, sputum and lavage samples of CF patients, the brain of a pig with neurological deficit, the brain stem of an injured deer and the nose of mice (Coenye et al. 2001b; Coenye et al. 2002; Gerrits et al. 2005) (own unpublished data). In addition, Burkholderia tropica has been isolated from a neonatal patient with necrotizing enterocolitis and bowel perforation, who developed septicaemia (Deris et al. 2010).
In addition to these three main clades, several Burkholderia species represent unique deep-branching 16S rRNA lineages (Fig. 1). These include Burkholderia rhizoxinica and Burkholderia endofungorum (two endosymbionts of the plant-pathogenic fungus Rhizopus microsporus) (Partida-Martinez et al. 2007) and a group consisting of Burkholderia caryophylli (a pathogen of carnations and onions) (Ballard et al. 1970), Burkholderia symbiotica (a root nodule endosymbiont of Mimosa species) (Sheu et al. 2012) and Burkholderia soli (a soil bacterium) (Yoo et al. 2007). These species do not cluster closely with any other Burkholderia species and their 16S rRNA sequence-based phylogenetic position appears variable and dependent on the other taxa included in a phylogenetic analysis. Finally, Burkholderia andropogonis, a pathogen causing stripe disease of sorghum and leaf spot of velvet bean (Coenye et al. 2001a), clustered within the B. glathei group clade in the present analysis (Fig. 1); yet it often occupies a distinct position in 16S rRNA-based phylogenetic trees as well (see e.g. Sawana et al. 2014; Estrada-de los Santos et al. 2015).
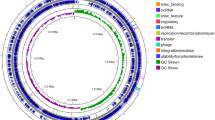
Today, the availability of whole genome sequences enables phylogenetic analyses based on the entire part of the genome that is shared between organisms, a discipline referred to as phylogenomics (Yutin et al. 2012; Wang and Wu 2013). Although the number of Burkholderia species for which whole genome sequences are available is still limited, phylogenomics only partially reveals the same major subdivisions as the 16S rRNA tree (Fig. 1). Analysing the diversity across the Burkholderia genus using complete genome comparison is difficult given their inherent genetic diversity. Nevertheless, comparison of the 53 ribosomal protein-encoding genes within the current ribosomal multilocus sequence typing (rMLST) scheme provides a robust and biologically meaningful representation of the core Burkholderia genome (Fig. 2) (Jolley et al. 2012). A first well-supported rMLST Burkholderia clade also comprises the Bcc, plant-pathogenic and B. pseudomallei species groups, though with greater support for their divergence (Fig. 2). The second 16S rRNA sequence-based branch including B. glathei group species is also distinct when analysed by rMLST (Fig. 2). However, it clusters among species belonging to the third 16S rRNA-based lineage. In the rMLST analysis too, B. rhizoxinica and B. symbiotica occupy very distinct positions each. In contrast to what is observed in the 16S rRNA-based phylogeny, B. andropogonis represents a unique, very deep-branching lineage in the rMLST tree, illustrating its isolated taxonomic position. It will be interesting to see how other Burkholderia species will group when additional whole genome sequences become available.
Burkholderia phylogeny reconstructed from concatenated ribosomal protein gene sequences. Aligned, concatenated gene sequences from defined ribosomal multilocus sequence typing (rMLST) loci were downloaded from the rMLST database at http://pubMLST.org (Jolley et al. 2012). Low confidence regions of the alignment were removed with Gblocks (Talavera and Castresana 2007), resulting in a total of 18,490 positions in the final dataset. A phylogeny was reconstructed with FastTree (Price et al. 2010) using the generalised time-reversible (GTR) model of nucleotide evolution, and the resulting tree was visualised with FigTree (http://tree.bio.ed.ac.uk/software/figtree). Type strains are indicated in bold type. Node confidence is shown if less than 80 %. The sequence of Ralstonia solanacearum PSI07 was used as outgroup. The scale bar indicates the number of substitutions per site
The phylogenetic heterogeneity described above inspired several researchers to suggest (Gyaneshwar et al. 2011; Suarez-Moreno et al. 2012; Estrada-de los Santos et al. 2013; Angus et al. 2014) and eventually propose (Sawana et al. 2014) a taxonomic subdivision of the genus Burkholderia. Although it is clear that one can distinguish beneficial and harmful interactions of Burkholderia strains, there is no such phylogenetic subdivision in this genus. In a study of whole genome sequences of 45 Burkholderia strains representing some 25 formally named species and several unclassified strains, species belonging to the first Burkholderia clade were characterised by a percentage guanine plus cytosine content in their genomes of 65 to 69 %, while all other Burkholderia strains examined had a percentage guanine plus cytosine content in their genomes of 61 to 65 %. In addition, species belonging to the first Burkholderia clade shared six conserved sequence indels. The remaining Burkholderia strains represented species belonging to 16S rRNA clades 2 and 3 described above and one of the ungrouped species (B. rhizoxinica), and shared two conserved sequence indels. However, the phylogenetic diversity among the clade 2 and 3 species and B. rhizoxinica as revealed by 16S rRNA-based divergence and by differences in the distribution of 22 additional conserved sequence indels was ignored, as the authors proposed to restrict the name Burkholderia to 16S rRNA clade 1 species while reclassifying all other species into a single novel genus, Paraburkholderia (Sawana et al. 2014). These novel names were subsequently validated (Garrity and Oren 2015). Clearly, rMLST analysis is also supportive of greater evolutionary divergence between Burkholderia senso strictu (i.e. 16S rRNA clade 1 species) and Paraburkholderia but excludes B. rhizoxinica, B. symbiotica and B. andropogonis from the latter (Figs. 1 and 2). It will be up to the scientific community to adopt these novel names or not. According to the International Code of Nomenclature of Bacteria, researchers who are convinced that these name changes are ill-founded can continue to work with the original species names as these all were validly published.
Capacity for secondary metabolism: quorum sensing and antibiotic biosynthesis
Burkholderia bacteria are incredibly versatile organisms, with a phenomenal capacity for secondary metabolite production (Cimermancic et al. 2014), among which many with antifungal, antibacterial, herbicidal or insecticidal properties. Strains of Burkholderia are known to produce pyrrolnitrin (El-Banna and Winkelmann 1998), xylocandins (Meyers et al. 1987), cepafungins/glidobactins (Schellenberg et al. 2007), altericidins (Kirinuki et al. 1984), cepacins (Parker et al. 1984), cepaciamides (Jiao et al. 1996), phenazines (Cartwright et al. 1995) and quinoline derivatives (Moon et al. 1996). Although historical interest in Burkholderia secondary metabolites was largely focused on these primarily antifungal compounds, there is now growing evidence that Burkholderia also produce a range of potent antibacterial antibiotics, such as enacyloxin IIa (Mahenthiralingam et al. 2011). The following discussion will provide an overview of the state of the art on antimicrobial products from Burkholderia, as an update to the literature review by Vial et al. (2007) and genome-driven analysis of Liu and Cheng (2014). Biosynthesis of multiple Burkholderia antibiotics is controlled by quorum sensing (QS). For example, production of enacyloxin IIa and the resultant bioactivity of B. ambifaria AMMDT against B. multivorans is lost when the QS system is genetically disrupted (Mahenthiralingam et al. 2011) and the production of multiple other Burkholderia antibiotics is regulated in a similar way (Schmidt et al. 2009; Seyedsayamdost et al. 2010). Since QS and the signalling molecules produced by this process represent a major class of Burkholderia secondary metabolites, this process will be discussed first.
Quorum sensing in the genus Burkholderia
The expression of extracellular products is tightly regulated in bacteria. The immediate environment of the organism has a major influence on the production and secretion of extracellular products, but bacteria themselves also have a global regulation system in place to coordinate their behaviour. This QS system, as it is known, allows bacteria to alter their gene expression according to population density, as a form of cell-to-cell communication. In Gram-negative bacteria, N-acyl homoserine lactones (AHLs) are the most commonly used signal molecules, usually produced by an autoinducer synthase of the LuxI protein family and perceived by a transcriptional regulator belonging to the LuxR family (Whitehead et al. 2001). Bcc bacteria contain a LuxI/R type QS system, known as CepI/R, which was first discovered in B. cenocepacia K56-2 (Lewenza et al. 1999). The CepI AHL synthase is responsible for the production of N-octanoyl homoserine lactone and, as a minor by-product, N-hexanoyl homoserine lactone, whereas CepR acts as a transcriptional regulator. This CepI/R QS system is highly conserved among members of the Bcc (Sokol et al. 2007) and has been shown to regulate the production of a variety of extracellular products, including siderophores, fungicides and proteases (Lewenza et al. 1999; Zhou et al. 2003; Malott et al. 2005), as well numerous antibiotics, as discussed below.
Certain Bcc members harbour additional QS systems, such as BviI/R in B. vietnamiensis (Conway and Greenberg 2002) and CciI/R in B. cenocepacia strains belonging to the epidemic ET12 lineage (Malott et al. 2005). In addition, Boon et al. (2008) described another QS system in B. cenocepacia, which uses cis-2-dodecenoic acid (also known as Burkholderia diffusible signal factor or BDSF) as a signal molecule. BDSF is structurally similar to the diffusible signal factor (DSF) produced by the plant pathogen Xanthomonas campestris pv. campestris in order to regulate virulence. A homologue of the rpfF gene, the key enzyme in DSF biosynthesis, has been identified in B. cenocepacia and appears to be conserved throughout the Bcc (Deng et al. 2010). RpfF deletion mutants show reduced motility and adherence to porcine mucin, decreased extracellular polysaccharide production and diminished biofilm formation (Ryan et al. 2009). BDSF thus acts as an intraspecies signal in B. cenocepacia, yet it is also involved in interspecies and interkingdom communication, as antagonistic effects on Candida albicans have been observed (Boon et al. 2008).
Although QS has been less explored outside the Bcc, it appears to be widespread in the genus Burkholderia. For example, the plant pathogens B. glumae and B. plantarii are known to have QS systems similar to CepI/R, known as TofI/R and PlaI/R, respectively (Kim et al. 2004; Solis et al. 2006). Another distinct AHL-based QS system, designated BraI/R, is present in Burkholderia kururiensis and other members of 16S rRNA clade 3 (Suarez-Moreno et al. 2008). This QS system is similar to the LasI/R system found in Pseudomonas aeruginosa and relies on N-3-oxo-dodecanoyl homoserine lactone as main signal molecule. Despite the high conservation of the BraI/R system in this Burkholderia clade, most phenotypes (including biofilm formation, plant colonisation and degradation of aromatic compounds) seem to be regulated in a species-specific manner, suggesting that its role has evolved to suit the niche-specific needs of each species (Coutinho et al. 2013). Finally, several highly conserved and complex QS systems are present in members of the B. pseudomallei group. Both B. pseudomallei and Burkholderia thailandensis contain three complete QS circuits (QS-1, QS-2 and QS-3) and at least two orphan luxR homologues, whereas B. mallei has lost a large genomic region containing the QS-2 system through reductive evolution (Majerczyk et al. 2014).
Antibiotic biosynthesis by members of the Bcc
Members of the Bcc are known to produce multiple antimicrobial products, as described below. Pyrrolnitrin is a potent antifungal and antibacterial metabolite produced by strains of Burkholderia, Pseudomonas, Myxococcus, Serratia and Enterobacter (El-Banna and Winkelmann 1998), and it plays a role in the biocontrol activity of Bcc strains against phytopathogenic fungi such as Rhizoctonia solani and Fusarium spp. (Burkhead et al. 1994; Hwang et al. 2002). Pyrrolnitrin also inhibits growth of C. albicans and several Gram-positive bacteria, whereas Gram-negative organisms, except Proteus vulgaris, are not affected (El-Banna and Winkelmann 1998). A recent study determined the distribution of prnD, the gene responsible for the last step of pyrrolnitrin biosynthesis, within the genus Burkholderia (Schmidt et al. 2009). The pyrrolnitrin operon was found in strains belonging to eight Bcc species (Burkholderia pyrrocinia, B. cepacia, B. cenocepacia, B. ambifaria, Burkholderia ubonensis, Burkholderia lata and two novel Bcc groups) and in three species from the B. pseudomallei group (B. pseudomallei, B. thailandensis and Burkholderia oklahomensis). In addition, pyrrolnitrin production was shown to be under the control of the CepI/R QS system in B. lata 383T, as both cepI and cepR mutants lost inhibitory activity against R. solani, which could be restored in the cepI mutant through the addition of exogenous AHLs.
El-Banna and Winkelmann (1998) previously reported that glycerol strongly enhanced the production of pyrrolnitrin by B. cepacia NB-1. This finding was later confirmed in Burkholderia sp. O33, which produced increased amounts of pyrrolnitrin and polyhydroxyalkanoates in the presence of glycerol (Keum 2009). Based on these observations, Mahenthiralingam and colleagues used a basal salt medium, supplemented with glycerol as the only carbon source, to screen members of the Bcc for antimicrobial production (Mahenthiralingam et al. 2011). This led to the discovery that several B. ambifaria isolates show strong antimicrobial activity against pan-resistant Gram-negative pathogens, including Acinetobacter baumannii and two closely related Bcc species, B. multivorans and Burkholderia dolosa. The compounds responsible for this activity, enacyloxin IIa and the novel isomer cis-enacyloxin IIa, are produced by an unusual hybrid modular polyketide synthase (PKS) gene cluster. The fact that this cluster contains two orphan luxR-type homologues, disruption of which abolishes enacyloxin production, combined with the observation that cepI mutants no longer produce the enacyloxins, also indicates that QS plays a key role in regulation of this antibiotic biosynthesis cluster.
Another group of potent antifungals, named occidiofungins, was recently isolated from cultures of Burkholderia contaminans MS14 (Lu et al. 2009). These compounds display potent antifungal activity against a range of animal- and plant-pathogenic fungi, including Alternaria alternata, Aspergillus fumigatus, R. solani and several Phytium species. Occidiofungins are cyclic glycosylated oligopeptides, synthesised by a nonribosomal peptide synthetase (NRPS) and are structurally similar to the fungicidal xylocandins (Meyers et al. 1987) and the newly described burkholdines from B. ambifaria 2.2N (Tawfik et al. 2010). An NRPS gene cluster with close homology to the occidiofungin biosynthetic cluster was recently identified in B. ambifaria AMMDT and B. vietnamiensis DBO1. This gene cluster was responsible for hemolytic and insecticidal activity in B. vietnamiensis DBO1, suggesting that occidiofungins/burkholdines are not strictly fungicidal, but rather cytotoxic (Thomson and Dennis 2012). The synthesis of this group of bioactive oligopeptides is likely also QS-regulated, as the occidiofungin/burkholdine gene cluster was identified among several QS-controlled loci in B. ambifaria (Chapalain et al. 2013).
Finally, B. ambifaria is known to produce a range of bioactive volatile compounds that inhibit growth of the phytopathogenic fungi A. alternata and R. solani (Groenhagen et al. 2013). In addition to exhibiting fungicidal activity, the volatiles also increased biomass in the model plant Arabidopsis thaliana and were able to induce increased levels of antibiotic resistance in Escherichia coli.
An unidentified Burkholderia species related to the Bcc produces cepafungins, also known as glidobactins, which exhibit broad-spectrum antifungal and antitumor activities (Schellenberg et al. 2007). Similar gene clusters were found in B. pseudomallei, and glidobactins were recently identified as strong inhibitors of the eukaryotic proteasome (Groll et al. 2008).
Antibiotic biosynthesis by non-Bcc Burkholderia
Although a wide variety of antimicrobial compounds have been isolated from Bcc bacteria, several non-Bcc members of the genus Burkholderia are also known to exhibit antimicrobial activity. Species belonging to the B. pseudomallei group in particular appear to be excellent sources of antimicrobial natural products such as betulinans (Biggins et al. 2011), malleilactone/burkholderic acid (Franke et al. 2012; Biggins et al. 2012), thailandamides (Nguyen et al. 2008), thailandepsins (Wang et al. 2011), capistruin (Knappe et al. 2008) and bactobolins (Seyedsayamdost et al. 2010). Three of these bioactive compounds, capistruin, bactobolin and thailandamide, were discovered through genome mining of B. thailandensis E264T, which indicated the presence of a gene cluster involved in the synthesis of a lasso peptide, a type of ribosomally assembled bioactive peptide frequently isolated from Actinobacteria. The predicted molecule, capistruin, could be isolated from culture supernatant and demonstrated antibacterial activity against closely related Burkholderia and Pseudomonas strains (Knappe et al. 2008). Indication for the production of another antibacterial molecule came from a study investigating the role of the second QS system (QS-2) in B. thailandensis, encoded by BtaI2/R2, as these genes were found in clusters predicted to be involved in antibiotic biosynthesis. Supernatants from stationary-phase cultures of B. thailandensis E264T, but not a btaR2 mutant or a strain defective in AHL production, showed inhibitory activity against Gram-positive bacteria such as Bacillus subtilis, Staphylococcus aureus and Streptococcus pyogenes (Duerkop et al. 2009). This inhibitory activity was later attributed to a mixture of polar antibiotic compounds, known as bactobolins A–D, synthesised by a NRPS/PKS hybrid gene cluster (Seyedsayamdost et al. 2010).
The link between Burkholderia antibiotic production and QS was confirmed by Ishida et al. (2010). In this study, induction of thailandamide lactone A production in B. thailandensis was achieved by genetic manipulation of QS. Bioinformatic mining of the B. thailandensis E264T genome identified biosynthetic gene clusters which were congruent with the biosynthesis of polyketide metabolites. One of the metabolites, thailandamide A, was inconsistently detected in minute quantities within the supernatants of early-stage B. thailandensis cultures. Examination of the putative thailandamide biosynthetic cluster demonstrated the presence of an orphan luxR gene, encoding a transcriptional regulator (designated pthaA), which was phylogenetically distinct from other Burkholderia LuxR homologues. Disruption of the pthaA transcription factor binding region by mutagenesis resulted in a strain that accumulated large quantities of yellow pigment. This pigment was subsequently identified as thailandamide A and displayed antiproliferative, cytotoxic and anticancer activities (Ishida et al. 2010). Contrastingly, production of the cytotoxin malleilactone is regulated by an orphan luxR homologue, known as malR, which is not responsive to AHLs (Truong et al. 2015).
Identification and characterisation of bioactive natural products outside of the Bcc and B. pseudomallei group has not been as extensive. The plant pathogen B. plantarii produces tropolone, a compound with antibacterial, antifungal and phytotoxic properties (Azegami et al. 1987). The observation that a sesquiterpene signal molecule from Trichoderma virens PS1–7, a biocontrol agent of B. plantarii, represses tropolone production via transcriptional suppression of the AHL synthase plaI suggests that tropolone production is at least partially QS-regulated (Wang et al. 2013). Another plant pathogen, B. glumae, is known for the production of the phytotoxin toxoflavin (Jeong et al. 2003), which is regulated by the TofI/R QS system (Kim et al. 2004). Recently, another TofI/R-independent regulatory mechanism for toxoflavin production was discovered. Deletion mutants of this regulatory factor, tofM, produced lower levels of toxoflavin and showed reduced virulence, suggesting that tofM is a positive regulator of toxoflavin production (Chen et al. 2012). Besides this well-known phytotoxin, B. glumae also produces a bioactive pyrazole with antibacterial activity against Erwinia amylovora and several other Erwinia and Pseudomonas species (Mitchell et al. 2008). B. gladioli, a third plant pathogen in the genus Burkholderia, is known to produce several toxic and antimicrobial metabolites. The respiratory toxin, bongkrekic acid, associated with the fermented coconut-based Indonesian food, tempe bongkrek, was identified as the product of a polyketide biosynthesis cluster in B. gladioli pv. cocovenenans (Moebius et al. 2012). Tempe bongkrek is produced via fermentation by the mould Rhizopus oligosporus, which was initially thought to be the source of bongkrekic acid. Later, it was shown that B. gladioli pv. cocovenans, which was found as a contaminant of the fungal cultures and fermentations, was responsible for the production of this toxin (Moebius et al. 2012). Further evidence that B. gladioli can secrete bioactive molecules has been observed by Bharti et al. (2012), who noted that strain OR1 produces a range of, as yet uncharacterised, antimicrobial compounds with activity against Staphylococcus and Candida species. Finally, B. gladioli was shown to produce a polyketide of the enacyloxin family with antibacterial and antifungal activities when grown in co-culture with R. microsporus (Ross et al. 2014). B. rhizoxinica, an endosymbiont of the plant-pathogenic fungus R. microsporus, was also shown to be the source of a polyketide toxin, rhizoxin (Partida-Martinez and Hertweck 2005; Scherlach et al. 2012). Interkingdom interactions, in particular with plants and fungi, are key aspects of the natural biology of Burkholderia and it will be interesting to see if further bioactive natural products will be discovered as these complex environmental lifestyles are explored.
A genomic perspective on the capacity of Burkholderia for antibiotic biosynthesis
Greater access to complete microbial genome sequences facilitates the discovery of novel antibiotics via genome mining (Zerikly and Challis 2009). Several genomic approaches have been used to identify multiple antibiotics within Burkholderia, and particularly within the B. pseudomallei group (Fig. 1), as recently reviewed by Liu and Cheng (2014). In the last decade, hundreds of genomes were obtained for this group of potential bioterrorism agents, facilitating the application of genome mining for nondefence-related research. With the recent availability of genome sequences of other B. pseudomallei-related Burkholderia strains (Figs. 1 and 2), spanning the phylogenetic diversity of this group, it appears that the genomic capacity for antibiotic biosynthesis is an intrinsic feature of this group of organisms. Several bioinformatics tools are available for identifying secondary metabolite pathways within microbial genomes, the antibiotics Secondary Metabolite Analysis Shell (antiSMASH) being among the most advanced and well-curated (Blin et al. 2013; Weber et al. 2015). To provide a perspective on the genomic capacity for antibiotic biosynthesis within the genus Burkholderia, 15 complete genomes of strains representative of current diversity (Fig. 2) were analysed using antiSMASH 2.2.1 (Blin et al. 2013). The metrics for overall secondary metabolism and for the encoded PKS, NRPS, terpene and homoserine lactone biosynthetic capacities encoded in these 15 genomes are summarised in Table 1. The use of predictive biology software such as antiSMASH greatly accelerates the potential for novel secondary metabolite pathway discovery and was able to identify known PKS and NRPS pathways within the Burkholderia genomes analysed (Table 1). However, structure-pathway correlation and conventional chemical characterisation will still be required to characterise the wealth of potential bioactive molecules encoded by Burkholderia.
A large genome size is a well-recognised feature of Burkholderia bacteria, which stand out in terms of their general functional versatility (Suarez-Moreno et al. 2012): the mean genome size of the strains included in the analysis is nearly 7 Mb (Table 1). The number of secondary metabolite clusters encoded within Burkholderia genomes varies greatly: for the 15 genomes analysed, a mean of 13 clusters was observed (range 7 to 22). On average, this equates to more than 7 % of the Burkholderia genome (>450 kb) being devoted to secondary metabolism. Another key feature of this genomic capacity for secondary metabolism is the substantial size of the encoded pathways, at averages of 49.5, 58.7, 21.8 and 18.3 kb for PKS, NRPS, terpene and homoserine lactone biosynthetic loci, respectively (Table 1). Given the size of the identified clusters and the current knowledge about large multimodular pathways such as those found in PKS and NRPS operons, the majority of pathways are likely to be complete, functional and able to synthesise complex compounds, provided their expression is activated. With complex regulatory controls involving QS (Schmidt et al. 2009; Ishida et al. 2010), inducing carbon sources (El-Banna and Winkelmann 1998; Mahenthiralingam et al. 2011) and as yet uncharacterised environmental interactions as potential stimulants, understanding how to activate cryptic antibiotic biosynthetic pathways within Burkholderia will be key to unlocking their biotechnological potential as a source of fine pharmaceuticals.
Liu and Cheng (2014) suggested that B. thailandensis E264T could be considered a champion Burkholderia in terms of its encoded capacity for natural product biosynthesis. From our preliminary analysis, the species within the B. pseudomallei group collectively encode the largest capacity for secondary metabolite biosynthesis (>11 % of their genomes). Evidence for significant antibiotic biosynthetic capacity is also present within the Bcc, with B. ambifaria currently leading the group, devoting over 9 % of its genome to secondary metabolism (Table 1). B. gladioli and B. glumae also dedicate 10 % or more of their genomes to antibiotic biosynthesis (Table 1). The potential of Burkholderia species belonging to the B. glathei and B. xenovorans clades (Fig. 1 and 2) appears less impressive, with fewer pathways and less than 5 % of their genomes in general being dedicated to antibiotic production (Table 1). However, within the latter group, fewer genomes have been characterised to date and the genomic distance between species is substantial as demonstrated by their deep-branching phylogenies, suggesting that greater diversity within these groups still remains to be characterised. B. rhizoxinica is an outlier, both in terms of its phylogenetic position (Figs. 1 and 2), as well as its capacity for antibiotic biosynthesis, which is substantial at 12.9 % of its relatively small endosymbiotic-adapted genome (Table 1).
In addition to the potential for the biosynthesis of antimicrobial compounds such as polyketide antibiotics and NRPS products, another highly conserved feature, observed across all Burkholderia genomes, is the significant potential for terpene production (Table 1). The well-known interactions of Burkholderia with plants (Suarez-Moreno et al. 2012) and the intrinsic ability of these bacteria to colonise the rhizosphere (Vidal-Quist et al. 2014) suggest that terpene biosynthesis and the potential interplay of these molecules during bacteria-plant interactions are areas worthy of future biotechnological exploration. Finally, another interesting feature of Burkholderia genomes is their organisation in a multireplicon structure. Species within the Bcc have a three replicon genome (Table 1) and can tolerate deletion of the smallest replicon (Agnoli et al. 2012). This results in strains which are highly attenuated in virulence as well as antibiotic production, especially in the case of B. ambifaria AMMDT, where this deletion results in the loss of the enacyloxin pathway, encoded on the third replicon (Mahenthiralingam et al. 2011). The ability to colonise the rhizosphere is not affected by deletion of this third chromosome in Bcc strains (Vidal-Quist et al. 2014), suggesting that in the future, it may be possible to engineer biological control strains that contain the antimicrobial pathways required for biopesticidal activity but lack the virulence pathways associated with Burkholderia pathogenicity.
Conclusion
Burkholderia continue to fascinate as a diverse group of Gram-negative bacteria. They are arguably better known as primary pathogens such as B. pseudomallei, opportunistic pathogens such as members of the B. cepacia complex and plant pathogens such as B. glumae. However, the diverse environmental interactions of these bacteria are now pointing towards multiple beneficial properties extending beyond their known capacities for bioremediation and biological control, towards the significant biotechnological potential as antibiotic producers, as illustrated within this review. For most antimicrobial metabolites, the function in the natural environment is still not known. However, with insights into the ecological relevance of antibiotics, we could take advantage of the multifunctionality of these natural products, although positive exploitation of Burkholderia as biotechnological agents will have to balance against their potential pathogenicity. With the ability to rapidly define the entire functional content of Burkholderia strains using genomics, future exploitation of these organisms for biotechnological purposes will be greatly accelerated.
References
Agnoli K, Schwager S, Uehlinger S, Vergunst A, Viteri DF, Nguyen DT, Sokol PA, Carlier A, Eberl L (2012) Exposing the third chromosome of Burkholderia cepacia complex strains as a virulence plasmid. Mol Microbiol 83:362–378. doi:10.1111/j.1365-2958.2011.07937.x
Angus AA, Agapakis CM, Fong S, Yerrapragada S, Estrada-de los Santos P, Yang P, Song N, Kano S, Caballero-Mellado J, de Faria SM, Dakora FD, Weinstock G, Hirsch AM (2014) Plant-associated symbiotic Burkholderia species lack hallmark strategies required in mammalian pathogenesis. PLoS One 9:e83779. doi:10.1371/journal.pone.0083779
Azegami K, Nishiyama K, Watanabe Y, Kadota I, Ohuchi A, Fukazawa C (1987) Pseudomonas plantarii sp. nov., the causal agent of rice seedling blight. Int J Syst Bacteriol 37:144–152. doi:10.1099/00207713-37-2-144
Ballard RW, Palleroni NJ, Doudoroff M, Stanier RY, Mandel M (1970) Taxonomy of the aerobic pseudomonads: Pseudomonas cepacia, P. marginata, P. alliicola and P. caryophylli. J Gen Microbiol 60:199–214. doi:10.1099/00221287-60-2-199
Bharti P, Anand V, Chander J, Singh IP, Singh TV, Tewari R (2012) Heat stable antimicrobial activity of Burkholderia gladioli OR1 against clinical drug resistant isolates. Indian J Med Res 135:666–671
Biggins JB, Liu X, Feng Z, Brady SF (2011) Metabolites from the induced expression of cryptic single operons found in the genome of Burkholderia pseudomallei. J Am Chem Soc 133:1638–1641. doi:10.1021/ja1087369
Biggins JB, Ternei MA, Brady SF (2012) Malleilactone, a polyketide synthase-derived virulence factor encoded by the cryptic secondary metabolome of Burkholderia pseudomallei group pathogens. J Am Chem Soc 134:13192–13195. doi:10.1021/ja3052156
Blin K, Medema MH, Kazempour D, Fischbach MA, Breitling R, Takano E, Weber T (2013) antiSMASH 2.0—a versatile platform for genome mining of secondary metabolite producers. Nucleic Acids Res 41:W204–W212. doi:10.1093/nar/gkt449
Boon C, Deng Y, Wang L-H, He Y, Xu J-L, Fan Y, Pan SQ, Zhang L-H (2008) A novel DSF-like signal from Burkholderia cenocepacia interferes with Candida albicans morphological transition. ISME J 2:27–36. doi:10.1038/ismej.2007.76
Bowers JH, Parke JL (1993) Epidemiology of Pythium damping-off and Aphanomyces root rot of peas after seed treatment with bacterial agents for biological control. Phytopathology 83:1466. doi:10.1094/Phyto-83-1466
Burkhead KD, Schisler DA, Slininger PJ (1994) Pyrrolnitrin production by biological control agent Pseudomonas cepacia B37w in culture and in colonized wounds of potatoes. Appl Environ Microbiol 60:2031–2039
Carlier AL, Eberl L (2012) The eroded genome of a Psychotria leaf symbiont: hypotheses about lifestyle and interactions with its plant host. Environ Microbiol 14:2757–2769. doi:10.1111/j.1462-2920.2012.02763.x
Cartwright DK, Chilton WS, Benson DM (1995) Pyrrolnitrin and phenazine production by Pseudomonas cepacia, strain 5.5B, a biocontrol agent of Rhizoctonia solani. Appl Microbiol Biotechnol 43:211–216. doi:10.1007/BF00172814
Chapalain A, Vial L, Laprade N, Dekimpe V, Perreault J, Déziel E (2013) Identification of quorum sensing-controlled genes in Burkholderia ambifaria. Microbiologyopen 2:226–242. doi:10.1002/mbo3.67
Chen R, Barphagha IK, Karki HS, Ham JH (2012) Dissection of quorum-sensing genes in Burkholderia glumae reveals non-canonical regulation and the new regulatory gene tofM for toxoflavin production. PLoS One 7:e52150. doi:10.1371/journal.pone.0052150
Cimermancic P, Medema MH, Claesen J, Kurita K, Wieland Brown LC, Mavrommatis K, Pati A, Godfrey PA, Koehrsen M, Clardy J, Birren BW, Takano E, Sali A, Linington RG, Fischbach MA (2014) Insights into secondary metabolism from a global analysis of prokaryotic biosynthetic gene clusters. Cell 158:412–421. doi:10.1016/j.cell.2014.06.034
Coenye T, Holmes B, Kersters K, Govan JRW, Vandamme P (1999) Burkholderia cocovenenans (van Damme et al. 1960) Gillis et al. 1995 and Burkholderia vandii Urakami et al. 1994 are junior synonyms of Burkholderia gladioli (Severini 1913) Yabuuchi et al. 1993 and Burkholderia plantarii (Azegami et al. 1987) Urakami et al. 1994, respectively. Int J Syst Bacteriol 49:37–42. doi:10.1099/00207713-49-1-37
Coenye T, Falsen E, Hoste B, Ohlen M, Goris J, Govan J, Gillis M, Vandamme P (2000) Description of Pandoraea gen. nov. with Pandoraea apista sp. nov., Pandoraea pulmonicola sp. nov., Pandoraea pnomenusa sp. nov., Pandoraea sputorum sp. nov. and Pandoraea norimbergensis comb. nov. Int J Syst Evol Microbiol 50:887–899. doi:10.1099/00207713-50-2-887
Coenye T, Laevens S, Gillis M, Vandamme P (2001a) Genotypic and chemotaxonomic evidence for the reclassification of Pseudomonas woodsii (Smith 1911) Stevens 1925 as Burkholderia andropogonis (Smith 1911) Gillis et al. 1995. Int J Syst Evol Microbiol 51:183–185. doi:10.1099/00207713-51-1-183
Coenye T, Laevens S, Willems A, Ohlen M, Hannant W, Govan J, Gillis M, Falsen E, Vandamme P (2001b) Burkholderia fungorum sp. nov. and Burkholderia caledonica sp. nov., two new species isolated from the environment, animals and human clinical samples. Int J Syst Evol Microbiol 51:1099–1107. doi:10.1099/00207713-51-3-1099
Coenye T, Goris J, Spilker T, Lipuma JJ, Vandamme P (2002) Characterization of unusual bacteria isolated from respiratory secretions of cystic fibrosis patients and description of Inquilinus limosus gen. nov., sp. nov. J Clin Microbiol 40:2062–2069. doi:10.1128/JCM.40.6.2062
Conway B-A, Greenberg EP (2002) Quorum-sensing signals and quorum-sensing genes in Burkholderia vietnamiensis. J Bacteriol 184:1187–1191. doi:10.1128/jb.184.4.1187-1191.2002
Coutinho BG, Mitter B, Talbi C, Sessitsch A, Bedmar EJ, Halliday N, James EK, Camara M, Venturi V (2013) Regulon studies and in planta role of the BraI/R quorum-sensing system in the plant-beneficial Burkholderia cluster. Appl Environ Microbiol 79:4421–4432. doi:10.1128/AEM.00635-13
Currie B (2015) Melioidosis: evolving concepts in epidemiology, pathogenesis, and treatment. Semin Respir Crit Care Med 36:111–125. doi:10.1055/s-0034-1398389
Deng Y, Wu J, Eberl L, Zhang L-H (2010) Structural and functional characterization of diffusible signal factor family quorum-sensing signals produced by members of the Burkholderia cepacia complex. Appl Environ Microbiol 76:4675–4683. doi:10.1128/AEM.00480-10
Deris ZZ, Van Rostenberghe H, Habsah H, Noraida R, Tan GC, Chan YY, Rosliza AR, Ravichandran M (2010) First isolation of Burkholderia tropica from a neonatal patient successfully treated with imipenem. Int J Infect Dis 14:e73–e74. doi:10.1016/j.ijid.2009.03.005
Duerkop BA, Varga J, Chandler JR, Peterson SB, Herman JP, Churchill ME, Parsek MR, Nierman WC, Greenberg EP (2009) Quorum-sensing control of antibiotic synthesis in Burkholderia thailandensis. J Bacteriol 191:3909–3918. doi:10.1128/JB.00200-09
El-Banna N, Winkelmann G (1998) Pyrrolnitrin from Burkholderia cepacia: antibiotic activity against fungi and novel activities against streptomycetes. J Appl Microbiol 85:69–78. doi:10.1046/j.1365-2672.1998.00473.x
Estrada-de los Santos P, Vinuesa P, Martínez-Aguilar L, Hirsch AM, Caballero-Mellado J (2013) Phylogenetic analysis of Burkholderia species by multilocus sequence analysis. Curr Microbiol 67:51–60. doi:10.1007/s00284-013-0330-9
Estrada-de los Santos P, Rojas-Rojas FU, Tapia-García EY, Vásquez-Murrieta MS, Hirsch AM (2015) To split or not to split: an opinion on dividing the genus Burkholderia. Ann Microbiol. doi:10.1007/s13213-015-1183-1
Franke J, Ishida K, Hertweck C (2012) Genomics-driven discovery of burkholderic acid, a noncanonical, cryptic polyketide from human pathogenic Burkholderia species. Angew Chemie - Int Ed 51:11611–11615. doi:10.1002/anie.201205566
Garrity GM, Oren A (2015) List of new names and new combinations previously effectively, but not validly, published. Int J Syst Evol Microbiol 65:2017–2025. doi:10.1099/ijs.0.000317
Gerrits GP, Klaassen C, Coenye T, Vandamme P, Meis JF (2005) Burkholderia fungorum septicemia. Emerg Infect Dis 11:1115–1117. doi:10.3201/eid1107.041290
Groenhagen U, Baumgartner R, Bailly A, Gardiner A, Eberl L, Schulz S, Weisskopf L (2013) Production of bioactive volatiles by different Burkholderia ambifaria strains. J Chem Ecol 39:892–906. doi:10.1007/s10886-013-0315-y
Groll M, Schellenberg B, Bachmann AS, Archer CR, Huber R, Powell TK, Lindow S, Kaiser M, Dudler R (2008) A plant pathogen virulence factor inhibits the eukaryotic proteasome by a novel mechanism. Nature 452:755–758. doi:10.1038/nature06782
Gyaneshwar P, Hirsch AM, Moulin L, Chen W-M, Elliott GN, Bontemps C, Estrada-de los Santos P, Gross E, dos Reis FB, Sprent JI, Young JPW, James EK (2011) Legume-nodulating Betaproteobacteria: diversity, host range, and future prospects. Mol Plant-Microbe Interact 24:1276–1288. doi:10.1094/MPMI-06-11-0172
Heungens K, Parke JL (2000) Zoospore homing and infection events: effects of the biocontrol bacterium Burkholderia cepacia AMMDR1 on two oomycete pathogens of pea (Pisum sativum L.). Appl Environ Microbiol 66:5192–5200. doi:10.1128/AEM.66.12.5192-5200.2000
Heungens K, Parke JL (2001) Postinfection biological control of oomycete pathogens of pea by Burkholderia cepacia AMMDR1. Phytopathology 91:383–391. doi:10.1094/PHYTO.2001.91.4.383
Ho Y-N, Huang C-C (2015) Draft genome sequence of Burkholderia cenocepacia strain 869T2, a plant-beneficial endophytic bacterium. Genome Announc 3:e01327–15. doi:10.1128/genomeA.01327-15
Hwang J, Chilton W, Benson D (2002) Pyrrolnitrin production by Burkholderia cepacia and biocontrol of Rhizoctonia stem rot of poinsettia. Biol Control 25:56–63. doi:10.1016/S1049-9644(02)00044-0
Ishida K, Lincke T, Behnken S, Hertweck C (2010) Induced biosynthesis of cryptic polyketide metabolites in a Burkholderia thailandensis quorum sensing mutant. J Am Chem Soc 132:13966–13968. doi:10.1021/ja105003g
Jeong Y, Kim J, Kim S, Kang Y, Nagamatsu T, Hwang I (2003) Toxoflavin produced by Burkholderia glumae causing rice grain rot is responsible for inducing bacterial wilt in many field crops. Plant Dis 87:890–895. doi:10.1094/PDIS.2003.87.8.890
Jiao Y, Yoshihara T, Ishikuri S, Uchino H, Ichihara A (1996) Structural identification of cepaciamide A, a novel fungitoxic compound from Pseudomonas cepacia D-202. Tetrahedron Lett 37:1039–1042. doi:10.1016/0040-4039(95)02342-9
Jolley KA, Maiden MC (2010) BIGSdb: scalable analysis of bacterial genome variation at the population level. BMC Bioinformatics 11:595. doi:10.1186/1471-2105-11-595
Jolley KA, Bliss CM, Bennett JS, Bratcher HB, Brehony C, Colles FM, Wimalarathna H, Harrison OB, Sheppard SK, Cody AJ, Maiden MCJ (2012) Ribosomal multilocus sequence typing: universal characterization of bacteria from domain to strain. Microbiology 158:1005–1015. doi:10.1099/mic.0.055459-0
Keum YS (2009) Effects of nutrients on quorum signals and secondary metabolite productions of Burkholderia sp. O33. J Microbiol Biotechnol. doi:10.4014/jmb.0901.465
Kikuchi Y, Hosokawa T, Fukatsu T (2011) An ancient but promiscuous host–symbiont association between Burkholderia gut symbionts and their heteropteran hosts. ISME J 5:446–460. doi:10.1038/ismej.2010.150
Kim J, Kim JG, Kang Y, Jang JY, Jog GJ, Lim JY, Kim S, Suga H, Nagamatsu T, Hwang I (2004) Quorum sensing and the LysR-type transcriptional activator ToxR regulate toxoflavin biosynthesis and transport in Burkholderia glumae. Mol Microbiol 54:921–934. doi:10.1111/j.1365-2958.2004.04338.x
Kirinuki T, Ichiba T, Katayama K (1984) General survey of action site of altericidins on metabolism of Alternaria kikuchiana and Ustilago maydis. J Pestic Sci 9:601–610. doi:10.1584/jpestics.9.601
Knappe TA, Linne U, Zirah S, Rebuffat S, Xie X, Marahiel MA (2008) Isolation and structural characterization of capistruin, a lasso peptide predicted from the genome sequence of Burkholderia thailandensis E264. J Am Chem Soc 130:11446–11454. doi:10.1021/ja802966g
Lemaire B, Robbrecht E, van Wyk B, Van Oevelen S, Verstraete B, Prinsen E, Smets E, Dessein S (2011) Identification, origin, and evolution of leaf nodulating symbionts of Sericanthe (Rubiaceae). J Microbiol 49:935–941. doi:10.1007/s12275-011-1163-5
Lemaire B, van Oevelen S, de Block P, Verstraete B, Smets E, Prinsen E, Dessein S (2012) Identification of the bacterial endosymbionts in leaf nodules of Pavetta (Rubiaceae). Int J Syst Evol Microbiol 62:202–209. doi:10.1099/ijs.0.028019-0
Lewenza S, Conway B, Greenberg EP, Sokol PA (1999) Quorum sensing in Burkholderia cepacia: identification of the LuxRI homologs CepRI. J Bacteriol 181:748–756
Lim YW, Baik KS, Han SK, Kim SB, Bae KS (2003) Burkholderia sordidicola sp. nov., isolated from the white-rot fungus Phanerochaete sordida. Int J Syst Evol Microbiol 53:1631–1636. doi:10.1099/ijs.0.02456-0
Lipuma JJ (2010) The changing microbial epidemiology in cystic fibrosis. Clin Microbiol Rev 23:299–323. doi:10.1128/CMR.00068-09
Liu X, Cheng Y-QQ (2014) Genome-guided discovery of diverse natural products from Burkholderia sp. J Ind Microbiol Biotechnol 41:275–284. doi:10.1007/s10295-013-1376-1
Liu X-Y, Li C-X, Luo X-J, Lai Q-L, Xu J-H (2014) Burkholderia jiangsuensis sp. nov., a methyl parathion degrading bacterium, isolated from methyl parathion contaminated soil. Int J Syst Evol Microbiol 64:3247–3253. doi:10.1099/ijs.0.064444-0
Lu S-E, Novak J, Austin FW, Gu G, Ellis D, Kirk M, Wilson-Stanford S, Tonelli M, Smith L (2009) Occidiofungin, a unique antifungal glycopeptide produced by a strain of Burkholderia contaminans. Biochemistry 48:8312–8321. doi:10.1021/bi900814c
Lu P, Zheng L-Q, Sun J-J, Liu H-M, Li S-P, Hong Q, Li W-J (2012) Burkholderia zhejiangensis sp. nov., a methyl-parathion-degrading bacterium isolated from a wastewater-treatment system. Int J Syst Evol Microbiol 62:1337–1341. doi:10.1099/ijs.0.035428-0
Mahenthiralingam E, Song L, Sass A, White J, Wilmot C, Marchbank A, Boaisha O, Paine J, Knight D, Challis GL (2011) Enacyloxins are products of an unusual hybrid modular polyketide synthase encoded by a cryptic Burkholderia ambifaria genomic island. Chem Biol 18:665–677. doi:10.1016/j.chembiol.2011.01.020
Majerczyk CD, Brittnacher MJ, Jacobs MA, Armour CD, Radey MC, Bunt R, Hayden HS, Bydalek R, Greenberg EP (2014) Cross-species comparison of the Burkholderia pseudomallei, Burkholderia thailandensis, and Burkholderia mallei quorum-sensing regulons. J Bacteriol 196:3862–3871. doi:10.1128/JB.01974-14
Malott RJ, Baldwin A, Mahenthiralingam E, Sokol PA (2005) Characterization of the cciIR quorum-sensing system in Burkholderia cenocepacia. Infect Immun 73:4982–4992. doi:10.1128/IAI.73.8.4982
Meyers E, Bisacchi GS, Dean L, Liu WC, Minassian B, Slusarchyk DS, Sykes RB, Tanaka SK, Trejo W (1987) Xylocandin: a new complex of antifungal peptides. I. Taxonomy, isolation and biological activity. J Antibiot (Tokyo) 40:1515–1519. doi:10.7164/antibiotics.40.1515
Mitchell RE, Greenwood DR, Sarojini V (2008) An antibacterial pyrazole derivative from Burkholderia glumae, a bacterial pathogen of rice. Phytochemistry 69:2704–2707. doi:10.1016/j.phytochem.2008.08.013
Moebius N, Ross C, Scherlach K, Rohm B, Roth M, Hertweck C (2012) Biosynthesis of the respiratory toxin bongkrekic acid in the pathogenic bacterium Burkholderia gladioli. Chem Biol 19:1164–1174. doi:10.1016/j.chembiol.2012.07.022
Moon SS, Kang PM, Park KS, Kim CH (1996) Plant growth promoting and fungicidal 4-quinolinones from Pseudomonas cepacia. Phytochemistry 42:365–368. doi:10.1016/0031-9422(95)00897-7
Nguyen T, Ishida K, Jenke-Kodama H, Dittmann E, Gurgui C, Hochmuth T, Taudien S, Platzer M, Hertweck C, Piel J (2008) Exploiting the mosaic structure of trans-acyltransferase polyketide synthases for natural product discovery and pathway dissection. Nat Biotechnol 26:225–233. doi:10.1038/nbt1379
Parke JL (1990) Population dynamics of Pseudomonas cepacia in the pea spermosphere in relation to biocontrol of Pythium. Phytopathology 80:1307. doi:10.1094/Phyto-80-1307
Parke JL, Gurian-sherman D (2001) Diversity of the Burkholderia cepacia complex and implications for risk assessment of biological control strains. Annu Rev Phytopathol 39:225–258. doi:10.1146/annurev.phyto.39.1.225
Parker WL, Rathnum ML, Seiner V, Trejo WH, Principe PA, Sykes RB (1984) Cepacin A and cepacin B, two new antibiotics produced by Pseudomonas cepacia. J Antibiot (Tokyo) 37:431–440
Parte AC (2014) LPSN—list of prokaryotic names with standing in nomenclature. Nucleic Acids Res 42:D613–D616. doi:10.1093/nar/gkt1111
Partida-Martinez LP, Hertweck C (2005) Pathogenic fungus harbours endosymbiotic bacteria for toxin production. Nature 437:884–888. doi:10.1038/nature03997
Partida-Martinez LP, Groth I, Schmitt I, Richter W, Roth M, Hertweck C (2007) Burkholderia rhizoxinica sp. nov. and Burkholderia endofungorum sp. nov., bacterial endosymbionts of the plant-pathogenic fungus Rhizopus microsporus. Int J Syst Evol Microbiol 57:2583–2590. doi:10.1099/ijs.0.64660-0
Peeters C, Zlosnik JEA, Spilker T, Hird TJ, LiPuma JJ, Vandamme P (2013) Burkholderia pseudomultivorans sp. nov., a novel Burkholderia cepacia complex species from human respiratory samples and the rhizosphere. Syst Appl Microbiol 36:483–489. doi:10.1016/j.syapm.2013.06.003
Pérez-Pantoja D, Donoso R, Agulló L, Córdova M, Seeger M, Pieper DH, González B (2012) Genomic analysis of the potential for aromatic compounds biodegradation in Burkholderiales. Environ Microbiol 14:1091–1117. doi:10.1111/j.1462-2920.2011.02613.x
Price MN, Dehal PS, Arkin AP (2010) FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS One 5:e9490. doi:10.1371/journal.pone.0009490
Pruesse E, Peplies J, Glockner FO (2012) SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28:1823–1829. doi:10.1093/bioinformatics/bts252
Ross C, Opel V, Scherlach K, Hertweck C (2014) Biosynthesis of antifungal and antibacterial polyketides by Burkholderia gladioli in coculture with Rhizopus microsporus. Mycoses 57:48–55. doi:10.1111/myc.12246
Ryan RP, McCarthy Y, Watt SA, Niehaus K, Dow JM (2009) Intraspecies signaling involving the diffusible signal factor BDSF (cis-2-dodecenoic acid) influences virulence in Burkholderia cenocepacia. J Bacteriol 191:5013–5019. doi:10.1128/JB.00473-09
Sawana A, Adeolu M, Gupta RS (2014) Molecular signatures and phylogenomic analysis of the genus Burkholderia: proposal for division of this genus into the emended genus Burkholderia containing pathogenic organisms and a new genus Paraburkholderia gen. nov. harboring environmental species. Front Genet 5:1–22. doi:10.3389/fgene.2014.00429
Schellenberg B, Bigler L, Dudler R (2007) Identification of genes involved in the biosynthesis of the cytotoxic compound glidobactin from a soil bacterium. Environ Microbiol 9:1640–1650. doi:10.1111/j.1462-2920.2007.01278.x
Scherlach K, Busch B, Lackner G, Paszkowski U, Hertweck C (2012) Symbiotic cooperation in the biosynthesis of a phytotoxin. Angew Chemie Int Ed 51:9615–9618. doi:10.1002/anie.201204540
Schmidt S, Blom JF, Pernthaler J, Berg G, Baldwin A, Mahenthiralingam E, Eberl L (2009) Production of the antifungal compound pyrrolnitrin is quorum sensing-regulated in members of the Burkholderia cepacia complex. Environ Microbiol 11:1422–1437. doi:10.1111/j.1462-2920.2009.01870.x
Seyedsayamdost MR, Chandler JR, Blodgett JAV, Lima PS, Duerkop BA, Oinuma K-I, Greenberg EP, Clardy J (2010) Quorum-sensing-regulated bactobolin production by Burkholderia thailandensis E264. Org Lett 12:716–719. doi:10.1021/ol902751x
Sheu S-Y, Chou J-H, Bontemps C, Elliott GN, Gross E, James EK, Sprent JI, Young JPW, Chen W-M (2012) Burkholderia symbiotica sp. nov., isolated from root nodules of Mimosa spp. native to north-east Brazil. Int J Syst Evol Microbiol 62:2272–2278. doi:10.1099/ijs.0.037408-0
Shibata TF, Maeda T, Nikoh N, Yamaguchi K, Oshima K, Hattori M, Nishiyama T, Hasebe M, Fukatsu T, Kikuchi Y, Shigenobu S (2013) Complete genome sequence of Burkholderia sp. strain RPE64, bacterial symbiont of the bean bug Riptortus pedestris. Genome Announc 1:e00441–13. doi:10.1128/genomeA.00441-13
Sokol PA, Malott RJ, Riedel K, Eberl L (2007) Communication systems in the genus Burkholderia: global regulators and targets for novel antipathogenic drugs. Future Microbiol 2:555–563. doi:10.2217/17460913.2.5.555
Solis R, Bertani I, Degrassi G, Devescovi G, Venturi V (2006) Involvement of quorum sensing and RpoS in rice seedling blight caused by Burkholderia plantarii. FEMS Microbiol Lett 259:106–112. doi:10.1111/j.1574-6968.2006.00254.x
Suarez-Moreno ZR, Caballero-Mellado J, Venturi V (2008) The new group of non-pathogenic plant-associated nitrogen-fixing Burkholderia spp. shares a conserved quorum-sensing system, which is tightly regulated by the RsaL repressor. Microbiology 154:2048–2059. doi:10.1099/mic.0.2008/017780-0
Suarez-Moreno ZR, Caballero-Mellado J, Coutinho BG, Mendonça-Previato L, James EK, Venturi V (2012) Common features of environmental and potentially beneficial plant-associated Burkholderia. Microb Ecol 63:249–266. doi:10.1007/s00248-011-9929-1
Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56:564–577. doi:10.1080/10635150701472164
Tamura K, Nei M (1993) Estimation of the number of base nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. doi:10.1093/molbev/mst197
Tawfik KA, Jeffs P, Bray B, Dubay G, Falkinham JO, Mesbah M, Youssef D, Khalifa S, Schmidt EW (2010) Burkholdines 1097 and 1229, potent antifungal peptides from Burkholderia ambifaria 2.2N. Org Lett 12:664–666. doi:10.1021/ol9029269
Thomson ELS, Dennis JJ (2012) A Burkholderia cepacia complex non-ribosomal peptide-synthesized toxin is hemolytic and required for full virulence. Virulence 3:286–298. doi:10.4161/viru.19355
Tian Y, Kong BH, Liu SL, Li CL, Yu R, Liu L, Li YH (2013) Burkholderia grimmiae sp. nov., isolated from a xerophilous moss (Grimmia montana). Int J Syst Evol Microbiol 63:2108–3113. doi:10.1099/ijs.0.045492-0
Torbeck L, Raccasi D, Guilfoyle DE, Friedman RL, Hussong D (2011) Burkholderia cepacia: this decision is overdue. PDA J Pharm Sci Technol 65:535–543. doi:10.5731/pdajpst.2011.00793
Tran Van V, Berge O, Ngo Ke S, Balandreau J, Heulin T (2000) Repeated beneficial effects of rice inoculation with a strain of Burkholderia vietnamiensis on early and late yield components in low fertility sulphate acid soils of Vietnam. Plant Soil 218(2):273–284. doi:10.1023/A:1014986916913
Truong TT, Seyedsayamdost M, Greenberg EP, Chandler JR (2015) A Burkholderia thailandensis acyl-homoserine lactone-independent orphan LuxR homolog that activates production of the cytotoxin malleilactone. J Bacteriol 197:3456–3462. doi:10.1128/JB.00425-15
van Oevelen S, de Wachter R, Vandamme P, Robbrecht E, Prinsen E (2004) “Candidatus Burkholderia calva” and “Candidatus Burkholderia nigropunctata” as leaf gall endosymbionts of African Psychotria. Int J Syst Evol Microbiol 54:2237–2239. doi:10.1099/ijs.0.63188-0
Vandamme P, Peeters C (2014) Time to revisit polyphasic taxonomy. Antonie Van Leeuwenhoek 106:57–65. doi:10.1007/s10482-014-0148-x
Vandamme P, De Brandt E, Houf K, Salles JF, Dirk van Elsas J, Spilker T, LiPuma JJ (2013) Burkholderia humi sp. nov., Burkholderia choica sp. nov., Burkholderia telluris sp. nov., Burkholderia terrestris sp. nov. and Burkholderia udeis sp. nov.: Burkholderia glathei-like bacteria from soil and rhizosphere soil. Int J Syst Evol Microbiol 63:4707–4718. doi:10.1099/ijs.0.048900-0
Vanlaere E, Baldwin A, Gevers D, Henry D, De Brandt E, LiPuma JJ, Mahenthiralingam E, Speert DP, Dowson C, Vandamme P (2009) Taxon K, a complex within the Burkholderia cepacia complex, comprises at least two novel species, Burkholderia contaminans sp. nov. and Burkholderia lata sp. nov. Int J Syst Evol Microbiol 59:102–111. doi:10.1099/ijs.0.001123-0
Verstraete B, Van Elst D, Steyn H, Van Wyk B, Lemaire B, Smets E, Dessein S (2011) Endophytic bacteria in toxic south African plants: identification, phylogeny and possible involvement in gousiekte. PLoS One 6:e19265. doi:10.1371/journal.pone.0019265
Verstraete B, Janssens S, Smets E, Dessein S (2013) Symbiotic ß-proteobacteria beyond legumes: Burkholderia in Rubiaceae. PLoS One 8:e55260. doi:10.1371/journal.pone.0055260
Vial L, Groleau M-C, Dekimpe V, Déziel E (2007) Burkholderia diversity and versatility: an inventory of the extracellular products. J Microbiol Biotechnol 17:1407–1429
Vial L, Chapalain A, Groleau MC, Déziel E (2011) The various lifestyles of the Burkholderia cepacia complex species: a tribute to adaptation. Environ Microbiol 13:1–12. doi:10.1111/j.1462-2920.2010.02343.x
Vidal-Quist JC, O’Sullivan LA, Desert A, Fivian-Hughes AS, Millet C, Jones TH, Weightman AJ, Rogers HJ, Berry C, Mahenthiralingam E (2014) Arabidopsis thaliana and Pisum sativum models demonstrate that root colonization is an intrinsic trait of Burkholderia cepacia complex bacteria. Microbiology 160:373–384. doi:10.1099/mic.0.074351-0
Wang Z, Wu M (2013) A phylum-level bacterial phylogenetic marker database. Mol Biol Evol 30:1258–1262. doi:10.1093/molbev/mst059
Wang C, Henkes LM, Doughty LB, He M, Wang D, Meyer-Almes F-J, Cheng Y-Q (2011) Thailandepsins: bacterial products with potent histone deacetylase inhibitory activities and broad-spectrum antiproliferative activities. J Nat Prod 74:2031–2038. doi:10.1021/np200324x
Wang M, Hashimoto M, Hashidoko Y (2013) Repression of tropolone production and induction of a Burkholderia plantarii pseudo-biofilm by carot-4-en-9,10-diol, a cell-to-cell signaling disrupter produced by Trichoderma virens. PLoS One 8:e78024. doi:10.1371/journal.pone.0078024
Weber T, Blin K, Duddela S, Krug D, Kim HU, Bruccoleri R, Lee SY, Fischbach MA, Muller R, Wohlleben W, Breitling R, Takano E, Medema MH (2015) antiSMASH 3.0—a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res 1–7. doi: 10.1093/nar/gkv437
Whitehead NA, Barnard AML, Slater H, Simpson NJL, Salmond GPC (2001) Quorum-sensing in Gram-negative bacteria. FEMS Microbiol Rev 25:365–404. doi:10.1111/j.1574-6976.2001.tb00583.x
Wilson MS, Herrick JB, Jeon CO, Hinman DE, Madsen EL (2003) Horizontal transfer of phnAc dioxygenase genes within one of two phenotypically and genotypically distinctive naphthalene-degrading guilds from adjacent soil environments. Appl Environ Microbiol 69:2172–2181. doi:10.1128/AEM.69.4.2172-2181.2003
Yabuuchi E, Kosako Y, Oyaizu H, Yano I, Hotta H, Hashimoto Y, Ezaki T, Arakawa M (1992) Proposal of Burkholderia gen. nov. and transfer of seven species of the genus Pseudomonas homology group II to the new genus, with the type species Burkholderia cepacia (Palleroni and Holmes 1981) comb. nov. Microbiol Immunol 36:1251–1275. doi:10.1111/j.1348-0421.1992.tb02129.x
Yabuuchi E, Kosako Y, Yano I, Hotta H, Nishiuchi Y (1995) Transfer of two Burkholderia and an Alcaligenes species to Ralstonia gen. nov.: proposal of Ralstonia pickettii (Ralston, Palleroni and Doudoroff 1973) comb. nov., Ralstonia solanacearum (Smith 1896) comb. nov. and Ralstonia eutropha (Davis 1969) comb. nov. Microbiol Immunol 39:897–904. doi:10.1111/j.1348-0421.1995.tb03275.x
Yoo S-H, Kim B-Y, Weon H-Y, Kwon S-W, Go S-J, Stackebrandt E (2007) Burkholderia soli sp. nov., isolated from soil cultivated with Korean ginseng. Int J Syst Evol Microbiol 57:122–125. doi:10.1099/ijs.0.64471-0
Yutin N, Puigbò P, Koonin EV, Wolf YI (2012) Phylogenomics of prokaryotic ribosomal proteins. PLoS One 7:e36972. doi:10.1371/journal.pone.0036972
Zerikly M, Challis GL (2009) Strategies for the discovery of new natural products by genome mining. ChemBioChem 10:625–633. doi:10.1002/cbic.200800389
Zhou H, Yao F, Roberts DP, Lessie TG (2003) AHL-deficient mutants of Burkholderia ambifaria BC-F have decreased antifungal activity. Curr Microbiol 47:174–179. doi:10.1007/s00284-002-3926-z
Zolg W, Ottow JCG (1975) Pseudomonas glathei sp. nov., a new nitrogen-scavenging rod isolated from acid lateritic relicts in Germany. Z Allg Mikrobiol 15:287–299. doi:10.1002/jobm.19750150410
Acknowledgments
MB acknowledges funding from the Cardiff University Synthetic Biology initiative. CP is indebted to the Special Research Council of Ghent University. TC acknowledges funding from the Fund for Scientific Research-Flanders and the Interuniversity Attraction Poles Programme initiated by the Belgian Science Policy Office. EM is a recipient of funding from the Biotechnology and Biological Sciences Research Council (Grant BB/L021692/1). PV and TC acknowledge funding from the Industrial Research Fund of Ghent University (Grant F2015/IOF-ConcepTT/142).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Depoorter, E., Bull, M.J., Peeters, C. et al. Burkholderia: an update on taxonomy and biotechnological potential as antibiotic producers. Appl Microbiol Biotechnol 100, 5215–5229 (2016). https://doi.org/10.1007/s00253-016-7520-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7520-x