Abstract
Bacteriophages (phages), natural enemies of bacteria, can encode enzymes able to degrade polymeric substances. These substances can be found in the bacterial cell surface, such as polysaccharides, or are produced by bacteria when they are living in biofilm communities, the most common bacterial lifestyle. Consequently, phages with depolymerase activity have a facilitated access to the host receptors, by degrading the capsular polysaccharides, and are believed to have a better performance against bacterial biofilms, since the degradation of extracellular polymeric substances by depolymerases might facilitate the access of phages to the cells within different biofilm layers. Since the diversity of phage depolymerases is not yet fully explored, this is the first review gathering information about all the depolymerases encoded by fully sequenced phages. Overall, in this study, 160 putative depolymerases, including sialidases, levanases, xylosidases, dextranases, hyaluronidases, peptidases as well as pectate/pectin lyases, were found in 143 phages (43 Myoviridae, 47 Siphoviridae, 37 Podoviridae, and 16 unclassified) infecting 24 genera of bacteria. We further provide information about the main applications of phage depolymerases, which can comprise areas as diverse as medical, chemical, or food-processing industry.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phages are natural bacterial viruses and constitute the most abundant entities on Earth (Clokie et al. 2011). Phages have been used in clinical practice since the early 1920s (Kutter et al. 2010; Abedon et al. 2011), and although there was a period of declining interest on phage therapy after the antibiotics discovery (Kutter et al. 2010), the interest on phages re-emerged with the rise of antibiotic-resistant bacteria (Sulakvelidze 2005; Matsuzaki et al. 2005; Kutateladze and Adamia 2010). Therefore, many studies have reported the potential of phages as antimicrobial agents (McVay et al. 2007; Capparelli et al. 2007; Guenther et al. 2009; Oliveira et al. 2010; Morello et al. 2011; Trigo et al. 2013; Bertozzi Silva and Sauvageau 2014). However, the activity of phages is often hindered by the presence of polysaccharides at the bacterial surfaces and/or when bacteria are living in the biofilm form.
Biofilms are defined as microbial communities attached to a surface and encased in a matrix composed by extracellular polymeric substances (EPS) (Costerton et al. 1995; O’Toole et al. 2000), which represent over 90 % of the biofilm biomass (Flemming and Wingender 2010). The EPS matrix is constituted by a mixture of polysaccharides, proteins, nucleic acids, and lipids (Davey and O’toole 2000; Flemming et al. 2007; Flemming and Wingender 2010), and its composition varies with bacterial species, age of the biofilms, and environmental conditions such as temperature or nutrients availability (Flemming et al. 2007; Vu et al. 2009; Flemming and Wingender 2010; Harper et al. 2014). Biofilms are ubiquitous in nature (Davey and O’toole 2000; Donlan 2002), and on account of their architecture that protects bacteria against external harsh conditions, they constitute an important survival strategy (Mah and O’Toole 2001; Flemming et al. 2007; Glonti et al. 2010). Furthermore, biofilms are frequently associated as a common source of contamination especially in healthcare (Reid 1999; Donlan 2001a, b) and food-processing settings (Brooks and Flint 2008; Shi and Zhu 2009). This happens as a consequence of their inherent tolerance to antimicrobial agents due to diffusion limitations, which can potentially lead to severe persistent infections, particularly difficult to treat (Costerton et al. 1999; Donlan 2001b; Parsek and Singh 2003; Hall-Stoodley et al. 2004).
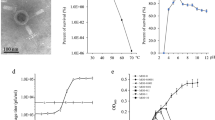
In the late 1990s, Hughes et al. partially purified enzymes capable of degrading bacterial polysaccharides (also known as depolymerases) from phage solutions (Hughes et al. 1998a). More recently, several works evidenced that phages can encode depolymerases to degrade polymers, either associated with the cell surface (capsule polysaccharides, e.g., K30 antigens) to facilitate phage adsorption, or EPS involved in biofilm matrix in order to promote phage diffusion through the bacterial slime (Glonti et al. 2010; Cornelissen et al. 2011, 2012; Hsu et al. 2013). The depolymerase activity is commonly identified by a constantly increasing halo surrounding the phage plaques (Cornelissen et al. 2011, 2012; Harper et al. 2014), as depicted in Fig. 1. Nonetheless, the formation of these halo zones is not so clear in some phages encoding depolymerases, such as the Salmonella phage phi PVP-SE1 that encodes a pectate-lyase in the gene 99 (Santos et al. 2009). It is believed that phages encoding depolymerases might have a better performance against bacterial biofilms (Hughes et al. 1998b; Yan et al. 2014). This is not completely true since there are phages deprived of depolymerases that have excellent anti-biofilm properties, which is the case of the Pseudomonas fluorescens phage phiIBB-PF7A (Sillankorva et al. 2008). Nonetheless, depolymerases may confer phages a great advantage in biofilm interaction (Lu and Collins 2007; Cornelissen et al. 2011, 2012).
As the presence and the distribution of depolymerases in the phage structures is not well understood, here, we report for the first time a complete study about EPS-degrading enzymes from all the phages completely sequenced to date.
Overview of phage-encoded depolymerases
The presence of depolymerases in phage genomes may act as an adjuvant for phage infection, since they degrade polymers either present in the bacterial surface, such as structural or capsular polysaccharides, or EPSs present in bacterial biofilms (Cornelissen et al. 2011; Yan et al. 2014). To get a better understanding of the diversity, structure, and function of phage depolymerases, we compiled information about all depolymerases present in fully sequenced dsDNA phages deposited in NCBI (Table S1).
Overall, 160 putative depolymerases were identified in 143 phages (43 Myoviridae, 47 Siphoviridae, 37 Podoviridae, and 16 unclassified) that infect 24 genera of bacteria, with lengths ranging from 197 to 2417 amino acids (Table S1). According to Drulis-Kawa et al., phage depolymerases appear in two forms: (i) as integral components of the phage particles or (ii) as soluble proteins generated during host cell lysis (Drulis-Kawa et al. 2015). Based on our search, the huge majority of phage depolymerases (126 proteins) are encoded in the same open reading frame of phage structural proteins (mostly on tail fibers, base plates, but sometimes also in the neck) or in close proximity to those genes (Fig. 2), and were thus considered as structural proteins (Table S1). Twenty other depolymerases found in this work might be soluble proteins since they are distant from any structural gene (Table S1). We were not able to classify 14 phage depolymerases, and thus, we considered them as unknown (Table S1).
Distribution of the different depolymerase classes through the structure of tailed bacteriophages. Levanases, xylosidades, hyaluronidases, dextranases, and lipases represent rare depolymerases domains found in phages, and therefore, the location was not possible to identify. The Bacillus phage CampHawk represents the only phage to encode four depolymerases in different genes (one sialidase and three pectate lyases)
According to their mode of action, the putative phage depolymerases were distributed in two main classes: hydrolases and lyases, which were further divided into different subclasses (Table 1). Exolysins (structural lysins) or endolysins, enzymes that hydrolase the structural LPS and peptidoglycan encoded by all Caudovirales phages, were already previously identified and described (Oliveira et al. 2013). Therefore, in this review, we focus on less explored enzymes that degrade EPS.
Sixteen different domain combinations were found throughout this analysis, and one representative of each architecture is depicted in Fig. 3. Although most of domains were found individually represented in a phage protein, combinations of domains were also found. For instance, Pectate_lyase_3 was found alone (Shigella phage Sf6) and combined with Peptidase_S74 (Bacillus phage CampHawk), End_tail_spike-Peptidase_S74 complex (Enterobacteria phage K5), and with Pectin_lyase (Streptomyces phage phiSASD1).
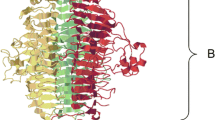
Different phage depolymerase architectures. For the 16 different architectures, a phage representative was selected. Accession numbers of phage proteins: (1) YP_003714746, (2) AAL15086, (3) AAQ12204, (4) AFX93507, (5) AGY47556, (6) CCE60816, (7) AFH14728, (8) ADA79897, (9) YP_654147, (10) CAJ29458, (11) YP_008242013, (12) AGO47740, (13) YP_008318429, (14) ACE96035, (15) AGY46940, and (16) YP_009056510
Most phages encode only one or two depolymerase motifs in the same gene (Table S1). Nevertheless, some phages can encode depolymerases in different genes. For example, phage ϕK1-5 is able to infect and replicate on both K1 and K5 Escherichia coli strains because it encodes two different enzymatic tail fiber proteins: an endosialidase that enables the phage to attach to and degrade K1 polysaccharide capsule, and a K5 lyase that cleaves the K5 capsular polysaccharides (Scholl et al. 2001). Another case is the Salmonella enterica serovar Enteritidis phage Marshall that encodes two tail proteins with beta helix/pectin lyase domains (Luna et al. 2013). A very interesting and rare example is the Bacillus subtilis phage CampHawk which encodes a putative peptidase and three pectate lyase domains in four different tail proteins (Table S1) (Ritz et al. 2013).
Diversity of phage depolymerases
Hydrolases
O-glycosyl hydrolases (EC 3.2.1.X) are enzymes that catalyze the hydrolysis of glycosidic bonds (Davies and Henrissat 1995). In this class of enzymes, six different groups of phage depolymerases were found: sialidases, levanases, xylosidases, dextranases, rhamnosidases, and peptidases (Table 1).
Sialidases or neuraminidases (EC 3.2.1.18) are a group of enzymes that hydrolyses the α-linkage of the terminal sialic acids of various glycans in diverse organisms (Kim et al. 2011b). Sialic acids represent a family of nine-carbon acidic sugars that are mostly found in the terminal position of oligosaccharide chains of glycan molecules present in cell surfaces (Traving and Schauer 1998; Kim et al. 2011b). Sialic acid plays an important role on bacterial pathogenesis since pathogenic bacteria such as E. coli K1, Haemophilus influenza, Pasteurella multocida, Neisseria spp., Campylobacter jejuni, and Streptococcus agalactiae can use this molecule as a source of nutrients or to increase resistance to host’s immune defenses by coating themselves in sialic acid (Severi et al. 2007). Nonetheless, it has also been shown that many bacterial species produce sialidases and that their presence has been associated with increased pathogenesis because they promote bacterial survival in mucosal niche environments and have important roles in the interaction with other organisms (Kim et al. 2011b; Lewis and Lewis 2012). So far, several phages have been reported to encode endosialidases as part of the tail structure, which enables the recognition and degradation of the capsular polysaccharides (Petter and Vimr 1993; Gerardy-Schahn et al. 1995; Long et al. 1995; Scholl et al. 2001; Scholl and Merril 2005; Schwarzer et al. 2012). Here, we found sialidase domains in a wide range of phages targeting different bacterial species. However, the majority of phages carrying sialidases are specific for Streptococcus spp. (Table S1). Sialic acid is reported to be one of the most important carbohydrates for Streptococcus pneumoniae because it plays a significant role as source of carbon and energy, receptor of adhesion, and invasion and promotion of biofilm formation (Gualdi et al. 2012). However, among Gram-positive bacteria, only S. agalactiae and Streptococcus suis are reported to produce sialylated polysaccharides in the form of capsular polysaccharides (Chaffin et al. 2005; Severi et al. 2007). It was already demonstrated that the intraperitoneal administration of a phage-derived endosialidase in E. coli-infected mice successfully prevented bacteremia and death from systemic infection (Mushtaq et al. 2004, 2005). Although the enzyme had no effect on the viability of E. coli cells, the degradation of capsular polysaccharides sensitized bacteria to the bactericidal action of the complement system (Mushtaq et al. 2004) and also enhanced phagocytosis by macrophages (Mushtaq et al. 2005).
Levanases (EC 3.2.1.65) are enzymes found in some microorganisms (e.g., Bacillus and Pseudomonas) that catalyze the hydrolysis of the β-2,6-linked main chain of levan, a natural product from the group of fructans (Murakami et al. 1992; Miasnikov 1997). Levan polysaccharide is known to play an ecological role on Bacillus biofilm formation and maintenance (Marvasi et al. 2010), while on some strains of Pseudomonas are suggested to be part of a protective capsule against phages (PATON 1960). Therefore, with no surprise, the only phage depolymerise domain with levan degradation (Glyco_hydro_32) activity found belongs to the Bacillus phage SP10 (Table S1).
Xylosidases (EC 3.2.1.37), dextranases (EC 3.2.1.11), and rhamnosidases (EC 3.2.1.40) are the classes of enzymes representing the rarest depolymerase phage domains (Table S1). These enzymes catalyze the hydrolysis of sugars such as xylan (Sunna and Antranikian 1997), dextran (Jiménez 2009), and rhamnogalacturonan (Yadav et al. 2010), respectively. Such polymers are present in many bacterial species as described in Table 1. The xylosidase domain (Glyco_hydro_39) was found in the only Caulobacter phage present in Table S1, while the two dextranase domains (Glyco_hydro_66) were found in the only Lactobacillus phage. Although no rhamnosidases domains were found in our search of phage genomes, rhamnosidase activity was already reported in Enterobacteria phage P22 (Eriksson et al. 1979), Shigella phage SF6 (Chua et al. 1999), and Escherichia phage HK620 (Barbirz et al. 2008). The three Podoviridae phages cleave the respective host cell receptor polysaccharide by their homotrimeric tailspike proteins (Barbirz et al. 2008). Furthermore, after being identified, the endorhamnosidase of phage SF6 was cloned, and its activity was shown by the cleavage of O-antigen on LPS chain in endorhamnosidase-treated samples (Chua et al. 1999).
Among hydrolases, there are enzymes that act on peptide bonds known as peptidases (EC 3.4.X.X). In the present study, a poly-γ-glutamate (γ-PGA) hydrolase (EC 3.4.19.9) encoding a peptidase domain DUF867 (PF05908) was found in four Bacillus phages, namely Bobb, phiAGATE, SPP1, and phiNIT1. Kimura et al. identified and purified a γ-PGA hydrolase from phage phiNIT1 and demonstrated its enzymatic activity against capsular polypeptides of glutamate with a gamma-linkage, called poly-γ-glutamate (Kimura and Itoh 2003). This polypeptide is a natural polymer exclusively synthesized by Gram-positive bacteria, including some strains of Bacillus (Kimura and Itoh 2003; Candela and Fouet 2006).
Lyases
Polysaccharide lyases (EC 4.2.2.X) are a class of enzymes that cleave (1,4) glycosidic bonds by a β-elimination mechanism (Sutherland 1995; Michaud et al. 2003). This class of enzymes comprises three groups of phage depolymerases: hyaluronate, alginate, and pectin/pectate lyases.
Hyaluronidases or hyaluronate lyases (EC 4.2.2.1 or 4.2.99.1) are a class of enzymes capable of, but not exclusively, digesting the hyaluronate, a linear unsulfated glycosaminoglucan polymer found in several organisms, including bacteria (Staphylococcus, Clostridium, streptococci groups A, B, C as well as S. pneumoniae) (Hynes and Walton 2000). Many extracellular bacterial hyaluronidases are able to cleave the β-1-4 glycosidic linkage by β-elimination and are often suggested to be a virulence factor, by increasing tissue permeability, facilitating the pathogen invasion (Hynes and Walton 2000). The same type of hyaluronidases has been found on some prophages from Streptococcus pyogenes and Streptococcus equi strains (Timoney et al. 1982; Hynes et al. 1995; Baker et al. 2002; Singh et al. 2014), and are thought to play a role in phage penetration of streptococcal hyaluronan capsules, facilitating bacterial lysogenization (Singh et al. 2014). As no extracellular milieu hyaluronidase activity was detected, these enzymes were suggested to be part of the phage particle to degrade the host’s hyaluronic acid capsules, in order to gain access to their surface where phage receptors are located. In our search, we have identified two novel temperate siphoviruses that contain hyaluronidase-degrading activities, namely the group A streptococcal phage phiNIH1 and S. equi phage P9.
Alginate lyases are mannuronate (EC 4.2.2.3) or guluronate lyases (EC 4.2.2.11) that catalyze the degradation of alginate, a linear polysaccharide of α-L-guluronate and its C5 epimer β-D-mannuronate (Wong et al. 2000; Kim et al. 2011a). Alginates are abundant in nature as a structural component of brown algae (Phaeophyceae) (Wong et al. 2000; Kim et al. 2011a), and they are also synthesized by two families of bacteria, the Pseudomonadaceae (Monday and Schiller 1996; Albrecht and Schiller 2005) and the Azotobacteriaceae (Clementi 1997). Alginate lyases have been isolated from different sources including marine algae, marine molluscs, fungi, bacteria, and viruses (Wong et al. 2000), and can be used to enhance antibiotic killing efficacy against mucoid bacteria such as P. aeruginosa (Alkawash et al. 2006). To date, some Azobacter and Pseudomonas spp. phages were found to encode alginate lyases (Bartell et al. 1966; Davidson et al. 1977; Glonti et al. 2010), which help phages to penetrate through the acetylated poly(M)-rich EPSs produced by their bacterial hosts (Wong et al. 2000; Yan et al. 2014). An example of a phage with alginate-degrading activity is the P. aeruginosa phage PT-6 (Glonti et al. 2010). Glonti et al. showed that this phage was able to reduce the viscosity of four alginate polymers by 62 to 66 % within 15 min (Glonti et al. 2010).
Pectin or pectate lyases (EC 4.2.2.10 and EC 4.2.2.2) are pectolytic enzymes that degrade galacturonic acid, a major constituent of bacterial polysaccharides and plant cell walls (Frirdich and Whitfield 2005; Garron and Cygler 2010). Although these enzymes are mainly produced by plant-associated organisms, they are also commonly produced by bacteria (Hugouvieux-Cotte-Pattat et al. 2014). During this search, we found that the majority of phage depolymerases belong to the class of pectolytic enzymes (Table S1). The Pseudomonas putida phages φ15 and AF, which exhibit biofilm-degrading properties, encode both pectate lyases on their tailspike genes 17 (Cornelissen et al. 2011) and 19 (Cornelissen et al. 2012), respectively. The EPS-degrading activity of the tailspikes of phage φ15 was further confirmed after dropping the recombinant expressed and purified protein on a bacterial lawn followed by the formation of opaque halo zones (Cornelissen et al. 2011). However, biofilm growth was not influenced after addition of UV-inactivated φ15 phage particles, which are non-infective but keep the enzymatic activity of their tailspikes (Cornelissen et al. 2011). The same was observed with phage AF (Cornelissen et al. 2012). Another example of phages encoding these enzymes are the Staphylococcus epidermidis phages vB_SepiS_phiIPLA5 and vB_SepiS_phiIPLA7 (Gutiérrez et al. 2012). The depolymerase domains, located in the pre-neck appendage proteins of those phages, are responsible for the good biofilm-degrading properties of both phages (Gutiérrez et al. 2012; Melo et al. 2014).
Others
Lipases (EC 3.1.1.3), or triacylglycerol hydrolases, are enzymes acting on the carboxyl ester bonds of triacylglycerols to liberate organic acids and glycerol (Jaeger et al. 1994; Gupta et al. 2004). Lipases are ubiquitous in nature, and they are produced by different types of organisms including bacteria. Bacillus, Pseudomonas, and Burkholderia are some of the most important lipase-producing bacterial genera (Gupta et al. 2004). In this work, only one phage depolymerase domain with lipids hydrolysis activity (Lipase_GDSL_3) was found in eight Cellulophaga phages and one Pseudomonas phage (Table S1).
Potential applications of phage depolymerases
Overall, phages with depolymerase activity have an improved performance against their target bacteria since these enzymes help phages in many important processes such as adsorption, invasion, and disintegration of host bacterial cell as well as biofilm disruption (Yan et al. 2014).
Phages encoding EPS depolymerases are expected to be more suitable for phage therapy purposes. In 2002, Tait el al. reported a complete eradication of Enterobacter cloacae biofilms after applying a cocktail of three phages with depolymerase activity (Tait et al. 2002), which revealed the potential of these enzymes to act as adjuvants for biofilm control. Nonetheless, most of the phages do not encode such enzymes, and therefore, phages can be engineered to express depolymerases during phage infection. For this purpose, Lu and Collins genetically engineered a T7 phage to express the biofilm-degrading enzyme dispersin B (Lu and Collins 2007). As a result, the enzymatic phage was able to efficiently kill biofilm cells and, at the same time, degrade the biofilm EPS matrix (Lu and Collins 2007).
Although not derived from phages, there are already EPS-degrading enzymes commercially available such as alginate lyase (Sigma-Aldrich) and dispersin B (Kane Biotech). This suggests that phage-based depolymerases may also have commercial potential.
Furthermore, phage depolymerases can be heterologous expressed having a wide range of other potential applications in medical, chemical, or food-processing industry as suggested in Table 2. Phage depolymerases can also be used in combination with antibiotics to act as adjuvants. These enzymes play an important role, for example, against alginate-producing strains that use alginate as a barrier to block the diffusion of antibiotics (Bayer et al. 1991; Hatch and Schiller 1998; Abdi-Ali et al. 2006). The ability of phages to effectively reduce the viscosity of alginates and EPS produced by P. aeruginosa cells was already demonstrated (Hanlon et al. 2001), and it was also shown that the efficacy of antibiotics against mucoid P. aeruginosa biofilms is enhanced by the administration of alginate lyase (Alkawash et al. 2006).
Other works have reported the potential of using phage depolymerases for multiple applications. It was shown that a capsular depolymerase encoded by a B. subtilis phage improved phagocytic killing of encapsulated Bacillus anthracis both in vitro (Scorpio et al. 2007) and in vivo (Scorpio et al. 2008). Another work also revealed the potential of a phage endosialidase to treat systemic infections in mice caused by E. coli K1 (Mushtaq et al. 2005). The enzyme degraded the capsular polysaccharide of pathogen cells rendering them sensitive to phagocytosis (Mushtaq et al. 2005). It was also shown that a depolymerase from an Erwinia amylovora phage effectively reduced colonization of fire blight host plants by E. amylovora (Kim and Geider 2000).
Another application of phage depolymerases is the detection or diagnosis of bacteria. Lin et al. successfully cloned and expressed a capsule depolymerase from a K. pneumoniae phage with specificity for K1 capsule suggesting a potential tool for the diagnosis of K1 K. pneumoniae infections (Lin et al. 2014). Furthermore, a work reported by Hsu et al. described the characterization of a K. pneumoniae phage depolymerase with specificity to KN2 capsular polysaccharides which can be successfully used for efficient Klebsiella capsular typing (Hsu et al. 2013).
Conclusions
The advances in omics and in silico techniques opened the way to new knowledge about the diversity of phage-encoded proteins. Phages are natural toolboxes that can offer an arsenal of proteins with a multitude of potential biotechnological applications. In this review, we demonstrated that phage genomes encode a diverse group of enzymes, called depolymerases, which are used to facilitate early steps of the phage infection cycle (adsorption and viral DNA injection). Depolymerases acting on many different substrates (including sialic acid, levan, xylan, dextran, rhamnogalacturonan, poly-γ-glutamante, hyaluronate, alginate, and triacylglycerol) were found in phage genomes, illustrating the complex phage-host interaction system that was refined during millions of years. The ability of phage depolymerases to degrade EPS makes them very attractive for a broad range of applications especially in clinical and industrial settings.
References
Abdi-Ali A, Mohammadi-Mehr M, Agha Alaei Y (2006) Bactericidal activity of various antibiotics against biofilm-producing Pseudomonas aeruginosa. Int J Antimicrob Agents 27:196–200. doi:10.1016/j.ijantimicag.2005.10.007
Abedon ST, Kuhl SJ, Blasdel BG, Kutter EM (2011) Phage treatment of human infections. Bacteriophage 1:66–85. doi:10.4161/bact.1.2.15845
Albrecht MT, Schiller NL (2005) Alginate lyase (AlgL) activity is required for alginate biosynthesis in Pseudomonas aeruginosa. J Bacteriol 187:3869–3872. doi:10.1128/JB.187.11.3869-3872.2005
Alkawash MA, Soothill JS, Schiller NL (2006) Alginate lyase enhances antibiotic killing of mucoid Pseudomonas aeruginosa in biofilms. APMIS 114:131–138. doi:10.1111/j.1600-0463.2006.apm_356.x
Andres D, Hanke C, Baxa U, Seul A, Barbirz S, Seckler R (2010) Tailspike interactions with lipopolysaccharide effect DNA ejection from phage P22 particles in vitro. J Biol Chem 285:36768–36775. doi:10.1074/jbc.M110.169003
Baker JR, Dong S, Pritchard DG (2002) The hyaluronan lyase of Streptococcus pyogenes bacteriophage H4489A. Biochem J 365:317. doi:10.1042/BJ20020149
Barbirz S, Müller JJ, Uetrecht C, Clark AJ, Heinemann U, Seckler R (2008) Crystal structure of Escherichia coli phage HK620 tailspike: podoviral tailspike endoglycosidase modules are evolutionarily related. Mol Microbiol 69:303–316. doi:10.1111/j.1365-2958.2008.06311.x
Bartell PF, Orr TE, Lam GK (1966) Polysaccharide depolymerase associated with bacteriophage infection. J Bacteriol 92:56–62
Bayer AS, Speert DP, Park S, Tu J, Witt M, Nast CC, Norman DC (1991) Functional role of mucoid exopolysaccharide (alginate) in antibiotic-induced and polymorphonuclear leukocyte-mediated killing of Pseudomonas aeruginosa. Infect Immun 59:302–308
Beg QK, Kapoor M, Mahajan L, Hoondal GS (2001) Microbial xylanases and their industrial applications: a review. Appl Microbiol Biotechnol 56:326–338. doi:10.1007/s002530100704
Bertozzi Silva J, Sauvageau D (2014) Bacteriophages as antimicrobial agents against bacterial contaminants in yeast fermentation processes. Biotechnol Biofuels 7:123. doi:10.1186/s13068-014-0123-9
Brooks JD, Flint SH (2008) Biofilms in the food industry: problems and potential solutions. Int J Food Sci Technol 43:2163–2176. doi:10.1111/j.1365-2621.2008.01839.x
Candela T, Fouet A (2006) Poly-gamma-glutamate in bacteria. Mol Microbiol 60:1091–1098. doi:10.1111/j.1365-2958.2006.05179.x
Capparelli R, Parlato M, Borriello G, Salvatore P, Iannelli D (2007) Experimental phage therapy against Staphylococcus aureus in mice. Antimicrob Agents Chemother 51:2765–2773. doi:10.1128/AAC.01513-06
Chaffin DO, Mentele LM, Rubens CE (2005) Sialylation of group B streptococcal capsular polysaccharide is mediated by cpsK and is required for optimal capsule polymerization and expression. J Bacteriol 187:4615–4626. doi:10.1128/JB.187.13.4615-4626.2005
Chua JE, Manning PA, Morona R (1999) The Shigella flexneri bacteriophage Sf6 tailspike protein (TSP)/endorhamnosidase is related to the bacteriophage P22 TSP and has a motif common to exo- and endoglycanases, and C-5 epimerases. Microbiology 145:1649–1659. doi:10.1099/13500872-145-7-1649
Clark LE, Mellette JR (1994) The use of hyaluronidase as an adjunct to surgical procedures. J Dermatol Surg Oncol 20:842–844. doi:10.1111/j.1524-4725.1994.tb03718.x
Clementi F (1997) Alginate production by Azotobacter vinelandii. Crit Rev Biotechnol 17:327–361. doi:10.3109/07388559709146618
Clokie MR, Millard AD, Letarov AV, Heaphy S (2011) Phages in nature. Bacteriophage 1:31–45. doi:10.4161/bact.1.1.14942
Cornelissen A, Ceyssens P-J, T’Syen J, Van Praet H, Noben J-P, Shaburova OV, Krylov VN, Volckaert G, Lavigne R (2011) The T7-related Pseudomonas putida phage φ15 displays virion-associated biofilm degradation properties. PLoS One 6:e18597. doi:10.1371/journal.pone.0018597
Cornelissen A, Ceyssens P-J, Krylov VN, Noben J-P, Volckaert G, Lavigne R (2012) Identification of EPS-degrading activity within the tail spikes of the novel Pseudomonas putida phage AF. Virology 434:251–256. doi:10.1016/j.virol.2012.09.030
Costerton JW, Lewandowski Z, Caldwell DE, Korber DR, Lappin-Scott HM (1995) Microbial biofilms. Annu Rev Microbiol 49:711–745. doi:10.1146/annurev.mi.49.100195.003431
Costerton JW, Stewart PS, Greenberg EP (1999) Bacterial biofilms: a common cause of persistent infections. Science (80-) 284:1318–1322. doi:10.1126/science.284.5418.1318
Dahech I, Ben Ayed H, Belghith KS, Belghith H, Mejdoub H (2013) Microbial production of levanase for specific hydrolysis of levan. Int J Biol Macromol 60:128–133. doi:10.1016/j.ijbiomac.2013.05.002
Davey ME, O’toole GA (2000) Microbial biofilms: from ecology to molecular genetics. Microbiol Mol Biol Rev 64:847–867. doi:10.1128/MMBR.64.4.847-867.2000
Davidson IW, Lawson CJ, Sutherland IW (1977) An alginate lysate from Azotobacter vinelandii phage. J Gen Microbiol 98:223–229
Davies G, Henrissat B (1995) Structures and mechanisms of glycosyl hydrolases. Structure 3:853–859. doi:10.1016/S0969-2126(01)00220-9
Donlan RM (2001a) Biofilm formation: a clinically relevant microbiological process. Clin Infect Dis 33:1387–1392. doi:10.1086/322972
Donlan RM (2001b) Biofilms and device-associated infections. Emerg Infect Dis 7:277–281. doi:10.3201/eid0702.700277
Donlan RM (2002) Biofilms: microbial life on surfaces. Emerg Infect Dis 8:881–890. doi:10.3201/eid0809.020063
Drulis-Kawa Z, Majkowska-Skrobek G, Maciejewska B (2015) Bacteriophages and phage-derived proteins—application approaches. Curr Med Chem 22:1757–1773. doi:10.2174/0929867322666150209152851
Eriksson U, Svenson SB, Lönngren J, Lindberg AA (1979) Salmonella phage glycanases: substrate specificity of the phage P22 endo-rhamnosidase. J Gen Virol 43:503–511. doi:10.1099/0022-1317-43-3-503
Flemming H-C, Wingender J (2010) The biofilm matrix. Nat Rev Microbiol 8:623–633. doi:10.1038/nrmicro2415
Flemming H-C, Neu TR, Wozniak DJ (2007) The EPS matrix: the “house of biofilm cells”. J Bacteriol 189:7945–7947. doi:10.1128/JB.00858-07
Frirdich E, Whitfield C (2005) Characterization of Gla(KP), a UDP-galacturonic acid C4-epimerase from Klebsiella pneumoniae with extended substrate specificity. J Bacteriol 187:4104–4115. doi:10.1128/JB.187.12.4104-4115.2005
Garron M-L, Cygler M (2010) Structural and mechanistic classification of uronic acid-containing polysaccharide lyases. Glycobiology 20:1547–1573. doi:10.1093/glycob/cwq122
Gerardy-Schahn R, Bethe A, Brennecke T, Mühlenhoff M, Eckhardt M, Ziesing S, Lottspeich F, Frosch M (1995) Molecular cloning and functional expression of bacteriophage PK1E-encoded endoneuraminidase Endo NE. Mol Microbiol 16:441–450. doi:10.1111/j.1365-2958.1995.tb02409.x
Glonti T, Chanishvili N, Taylor PW (2010) Bacteriophage-derived enzyme that depolymerizes the alginic acid capsule associated with cystic fibrosis isolates of Pseudomonas aeruginosa. J Appl Microbiol 108:695–702. doi:10.1111/j.1365-2672.2009.04469.x
Gualdi L, Hayre JK, Gerlini A, Bidossi A, Colomba L, Trappetti C, Pozzi G, Docquier J-D, Andrew P, Ricci S, Oggioni MR (2012) Regulation of neuraminidase expression in Streptococcus pneumoniae. BMC Microbiol 12:200. doi:10.1186/1471-2180-12-200
Guenther S, Huwyler D, Richard S, Loessner MJ (2009) Virulent bacteriophage for efficient biocontrol of Listeria monocytogenes in ready-to-eat foods. Appl Environ Microbiol 75:93–100. doi:10.1128/AEM.01711-08
Gupta R, Gupta N, Rathi P (2004) Bacterial lipases: an overview of production, purification and biochemical properties. Appl Microbiol Biotechnol 64:763–781. doi:10.1007/s00253-004-1568-8
Gutiérrez D, Martínez B, Rodríguez A, García P (2012) Genomic characterization of two Staphylococcus epidermidis bacteriophages with anti-biofilm potential. BMC Genomics 13:228. doi:10.1186/1471-2164-13-228
Hall-Stoodley L, Costerton JW, Stoodley P (2004) Bacterial biofilms: from the natural environment to infectious diseases. Nat Rev Microbiol 2:95–108. doi:10.1038/nrmicro821
Hanlon GW, Denyer SP, Olliff CJ, Ibrahim LJ (2001) Reduction in exopolysaccharide viscosity as an aid to bacteriophage penetration through Pseudomonas aeruginosa biofilms. Appl Environ Microbiol 67:2746–2753. doi:10.1128/AEM.67.6.2746-2753.2001
Harper D, Parracho H, Walker J, Sharp R, Hughes G, Werthén M, Lehman S, Morales S (2014) Bacteriophages and biofilms. Antibiotics 3:270–284. doi:10.3390/antibiotics3030270
Hatch RA, Schiller NL (1998) Alginate lyase promotes diffusion of aminoglycosides through the extracellular polysaccharide of mucoid Pseudomonas aeruginosa. Antimicrob Agents Chemother 42:974–977
Hsu C-R, Lin T-L, Pan Y-J, Hsieh P-F, Wang J-T (2013) Isolation of a bacteriophage specific for a new capsular type of Klebsiella pneumoniae and characterization of its polysaccharide depolymerase. PLoS One 8:e70092. doi:10.1371/journal.pone.0070092
Hughes KA, Sutherland IW, Clark J, Jones MV (1998a) Bacteriophage and associated polysaccharide depolymerases-novel tools for study of bacterial biofilms. J Appl Microbiol 85:583–590. doi:10.1046/j.1365-2672.1998.853541.x
Hughes KA, Sutherland IW, Jones MV (1998b) Biofilm susceptibility to bacteriophage attack: the role of phage-borne polysaccharide depolymerase. Microbiology 144:3039–3047. doi:10.1099/00221287-144-11-3039
Hugouvieux-Cotte-Pattat N, Condemine G, Shevchik VE (2014) Bacterial pectate lyases, structural and functional diversity. Environ Microbiol Rep 6:427–440. doi:10.1111/1758-2229.12166
Hynes WL, Walton SL (2000) Hyaluronidases of gram-positive bacteria. FEMS Microbiol Lett 183:201–207. doi:10.1111/j.1574-6968.2000.tb08958.x
Hynes WL, Hancock L, Ferretti JJ (1995) Analysis of a second bacteriophage hyaluronidase gene from Streptococcus pyogenes: evidence for a third hyaluronidase involved in extracellular enzymatic activity. Infect Immun 63:3015–3020
Jaeger KE, Ransac S, Dijkstra BW, Colson C, van Heuvel M, Misset O (1994) Bacterial lipases. FEMS Microbiol Rev 15:29–63. doi:10.1111/j.1574-6976.1994.tb00121.x
Jiménez ER (2009) Dextranase in sugar industry: a review. Sugar Tech 11:124–134. doi:10.1007/s12355-009-0019-3
Juturu V, Wu JC (2012) Microbial xylanases: engineering, production and industrial applications. Biotechnol Adv 30:1219–1227. doi:10.1016/j.biotechadv.2011.11.006
Kim WS, Geider K (2000) Characterization of a viral EPS-depolymerase, a potential tool for control of fire blight. Phytopathology 90:1263–1268. doi:10.1094/PHYTO.2000.90.11.1263
Kim HS, Lee C-G, Lee EY (2011a) Alginate lyase: structure, property, and application. Biotechnol Bioprocess Eng 16:843–851. doi:10.1007/s12257-011-0352-8
Kim S, Oh D-B, Kang HA, Kwon O (2011b) Features and applications of bacterial sialidases. Appl Microbiol Biotechnol 91:1–15. doi:10.1007/s00253-011-3307-2
Kimura K, Itoh Y (2003) Characterization of poly-gamma-glutamate hydrolase encoded by a bacteriophage genome: possible role in phage infection of Bacillus subtilis encapsulated with poly-gamma-glutamate. Appl Environ Microbiol 69:2491–2497. doi:10.1128/AEM.69.5.2491-2497.2003
Kulkarni N, Shendye A, Rao M (1999) Molecular and biotechnological aspects of xylanases. FEMS Microbiol Rev 23:411–456. doi:10.1111/j.1574-6976.1999.tb00407.x
Kutateladze M, Adamia R (2010) Bacteriophages as potential new therapeutics to replace or supplement antibiotics. Trends Biotechnol 28:591–595. doi:10.1016/j.tibtech.2010.08.001
Kutter E, De Vos D, Gvasalia G, Alavidze Z, Gogokhia L, Kuhl S, Abedon ST (2010) Phage therapy in clinical practice: treatment of human infections. Curr Pharm Biotechnol 11:69–86. doi:10.2174/138920110790725401
Lewis AL, Lewis WG (2012) Host sialoglycans and bacterial sialidases: a mucosal perspective. Cell Microbiol 14:1174–1182. doi:10.1111/j.1462-5822.2012.01807.x
Lin T-L, Hsieh P-F, Huang Y-T, Lee W-C, Tsai Y-T, Su P-A, Pan Y-J, Hsu C-R, Wu M-C, Wang J-T (2014) Isolation of a bacteriophage and its depolymerase specific for K1 capsule of Klebsiella pneumoniae: implication in typing and treatment. J Infect Dis 210:1734–1744. doi:10.1093/infdis/jiu332
Long GS, Bryant JM, Taylor PW, Luzio JP (1995) Complete nucleotide sequence of the gene encoding bacteriophage E endosialidase: implications for K1E endosialidase structure and function. Biochem J 309(Pt 2):543–550. doi:10.1042/bj3090543
Lu TK, Collins JJ (2007) Dispersing biofilms with engineered enzymatic bacteriophage. Proc Natl Acad Sci U S A 104:11197–11202. doi:10.1073/pnas.0704624104
Luna AJ, Wood TL, Chamakura KR, Kuty Everett GF (2013) Complete genome of Salmonella enterica serovar enteritidis myophage Marshall. Genome Announc. doi:10.1128/genomeA.00867-13
Mah TF, O’Toole GA (2001) Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol 9:34–39. doi:10.1016/S0966-842X(00)01913-2
Manzanares P, Vallés S, Ramòn D, Orejas M (2007) α-L-rhamnosidases: old and new insights. In: Polaina J, MacCabe AP (eds) Industrial enzymes. Springer Netherlands, Dordrecht, pp 117–140
Marvasi M, Visscher PT, Casillas Martinez L (2010) Exopolymeric substances (EPS) from Bacillus subtilis: polymers and genes encoding their synthesis. FEMS Microbiol Lett 313:1–9. doi:10.1111/j.1574-6968.2010.02085.x
Matsuzaki S, Rashel M, Uchiyama J, Sakurai S, Ujihara T, Kuroda M, Ikeuchi M, Tani T, Fujieda M, Wakiguchi H, Imai S (2005) Bacteriophage therapy: a revitalized therapy against bacterial infectious diseases. J Infect Chemother 11:211–219. doi:10.1007/s10156-005-0408-9
McVay CS, Velásquez M, Fralick JA (2007) Phage therapy of Pseudomonas aeruginosa infection in a mouse burn wound model. Antimicrob Agents Chemother 51:1934–1938. doi:10.1128/AAC.01028-06
Melo LDR, Sillankorva S, Ackermann H-W, Kropinski AM, Azeredo J, Cerca N (2014) Characterization of Staphylococcus epidermidis phage vB_SepS_SEP9—a unique member of the Siphoviridae family. Res Microbiol 165:679–685. doi:10.1016/j.resmic.2014.09.012
Miasnikov AN (1997) Characterization of a novel endo-levanase and its gene from Bacillus sp. L7. FEMS Microbiol Lett 154:23–28. doi:10.1111/j.1574-6968.1997.tb12619.x
Michaud P, Da Costa A, Courtois B, Courtois J (2003) Polysaccharide lyases: recent developments as biotechnological tools. Crit Rev Biotechnol 23:233–266. doi:10.1080/07388550390447043
Monday SR, Schiller NL (1996) Alginate synthesis in Pseudomonas aeruginosa: the role of AlgL (alginate lyase) and AlgX. J Bacteriol 178:625–632
Morello E, Saussereau E, Maura D, Huerre M, Touqui L, Debarbieux L (2011) Pulmonary bacteriophage therapy on Pseudomonas aeruginosa cystic fibrosis strains: first steps towards treatment and prevention. PLoS One 6:e16963. doi:10.1371/journal.pone.0016963
Murakami H, Kuramoto T, Mizutani K, Nakano H, Kitahata S (1992) Purification and some properties of a new levanase from Bacillus sp. No. 71. Biosci Biotechnol Biochem 56:608–613. doi:10.1271/bbb.56.608
Mushtaq N, Redpath MB, Luzio JP, Taylor PW (2004) Prevention and cure of systemic Escherichia coli K1 infection by modification of the bacterial phenotype. Antimicrob Agents Chemother 48:1503–1508. doi:10.1128/AAC.48.5.1503-1508.2004
Mushtaq N, Redpath MB, Luzio JP, Taylor PW (2005) Treatment of experimental Escherichia coli infection with recombinant bacteriophage-derived capsule depolymerase. J Antimicrob Chemother 56:160–165. doi:10.1093/jac/dki177
O’Toole G, Kaplan HB, Kolter R (2000) Biofilm formation as microbial development. Annu Rev Microbiol 54:49–79. doi:10.1146/annurev.micro.54.1.49
Oliveira A, Sereno R, Azeredo J (2010) In vivo efficiency evaluation of a phage cocktail in controlling severe colibacillosis in confined conditions and experimental poultry houses. Vet Microbiol 146:303–308. doi:10.1016/j.vetmic.2010.05.015
Oliveira H, Melo LDR, Santos SB, Nóbrega FL, Ferreira EC, Cerca N, Azeredo J, Kluskens LD (2013) Molecular aspects and comparative genomics of bacteriophage endolysins. J Virol 87:4558–4570. doi:10.1128/JVI.03277-12
Parsek MR, Singh PK (2003) Bacterial biofilms: an emerging link to disease pathogenesis. Annu Rev Microbiol 57:677–701. doi:10.1146/annurev.micro.57.030502.090720
Paton AM (1960) The role of Pseudomonas in plant disease. J Appl Bacteriol 23:526–532. doi:10.1111/j.1365-2672.1960.tb00224.x
Petter JG, Vimr ER (1993) Complete nucleotide sequence of the bacteriophage K1F tail gene encoding endo-N-acylneuraminidase (endo-N) and comparison to an endo-N homolog in bacteriophage PK1E. J Bacteriol 175:4354–4363
Reid G (1999) Biofilms in infectious disease and on medical devices. Int J Antimicrob Agents 11:223–226. doi:10.1016/S0924-8579(99)00020-5
Ritz MP, Perl AL, Colquhoun JM, Chamakura KR, Kuty Everett GF (2013) Complete genome of bacillus subtilis myophage CampHawk. Genome Announc. doi:10.1128/genomeA.00984-13
Santos SB, Carvalho CM, Sillankorva S, Nicolau A, Ferreira EC, Azeredo J (2009) The use of antibiotics to improve phage detection and enumeration by the double-layer agar technique. BMC Microbiol 9:148. doi:10.1186/1471-2180-9-148
Scholl D, Merril C (2005) The genome of bacteriophage K1F, a T7-like phage that has acquired the ability to replicate on K1 strains of Escherichia coli. J Bacteriol 187:8499–8503. doi:10.1128/JB.187.24.8499-8503.2005
Scholl D, Rogers S, Adhya S, Merril CR (2001) Bacteriophage K1-5 encodes two different tail fiber proteins, allowing it to infect and replicate on both K1 and K5 strains of Escherichia coli. J Virol 75:2509–2515. doi:10.1128/JVI.75.6.2509-2515.2001
Schwarzer D, Buettner FFR, Browning C, Nazarov S, Rabsch W, Bethe A, Oberbeck A, Bowman VD, Stummeyer K, Muhlenhoff M, Leiman PG, Gerardy-Schahn R (2012) A multivalent adsorption apparatus explains the broad host range of phage phi92: a comprehensive genomic and structural analysis. J Virol 86:10384–10398. doi:10.1128/JVI.00801-12
Scorpio A, Chabot DJ, Day WA, O’brien DK, Vietri NJ, Itoh Y, Mohamadzadeh M, Friedlander AM (2007) Poly-gamma-glutamate capsule-degrading enzyme treatment enhances phagocytosis and killing of encapsulated Bacillus anthracis. Antimicrob Agents Chemother 51:215–222. doi:10.1128/AAC.00706-06
Scorpio A, Tobery SA, Ribot WJ, Friedlander AM (2008) Treatment of experimental anthrax with recombinant capsule depolymerase. Antimicrob Agents Chemother 52:1014–1020. doi:10.1128/AAC.00741-07
Severi E, Hood DW, Thomas GH (2007) Sialic acid utilization by bacterial pathogens. Microbiology 153:2817–2822. doi:10.1099/mic.0.2007/009480-0
Shi X, Zhu X (2009) Biofilm formation and food safety in food industries. Trends Food Sci Technol 20:407–413. doi:10.1016/j.tifs.2009.01.054
Sieiro C, García-Fraga B, López-Seijas J, da Silva AF, Villa TG (2012) Microbial Pectic Enzymes in the Food and Wine Industry. In: Valdez B (ed) Food Industrial Processes-Methods and Equipment. InTech, pp 201–218
Sillankorva S, Neubauer P, Azeredo J (2008) Pseudomonas fluorescens biofilms subjected to phage phiIBB-PF7A. BMC Biotechnol 8:79. doi:10.1186/1472-6750-8-79
Singh SK, Bharati AP, Singh N, Pandey P, Joshi P, Singh K, Mitra K, Gayen JR, Sarkar J, Akhtar MS (2014) The prophage-encoded hyaluronate lyase has broad substrate specificity and is regulated by the N-terminal domain. J Biol Chem 289:35225–35236. doi:10.1074/jbc.M113.507673
Sulakvelidze A (2005) Phage therapy: an attractive option for dealing with antibiotic-resistant bacterial infections. Drug Discov Today 10:807–809. doi:10.1016/S1359-6446(05)03441-0
Sunna A, Antranikian G (1997) Xylanolytic enzymes from fungi and bacteria. Crit Rev Biotechnol 17:39–67. doi:10.3109/07388559709146606
Sutherland IW (1995) Polysaccharide lyases. FEMS Microbiol Rev 16:323–347
Tait K, Skillman LC, Sutherland IW (2002) The efficacy of bacteriophage as a method of biofilm eradication. Biofouling 18:305–311. doi:10.1080/0892701021000034418
Timoney JF, Pesante L, Ernst C (1982) Hyaluronidase associated with a temperate bacteriophage of Streptococcus equi. In: Schlessinger D (ed) Microbiology. American Society for Microbiology, Washington, DC, pp 145–146
Traving C, Schauer R (1998) Structure, function and metabolism of sialic acids. Cell Mol Life Sci 54:1330–1349. doi:10.1007/s000180050258
Trigo G, Martins TG, Fraga AG, Longatto-Filho A, Castro AG, Azeredo J, Pedrosa J (2013) Phage therapy is effective against infection by Mycobacterium ulcerans in a murine footpad model. PLoS Negl Trop Dis 7:e2183. doi:10.1371/journal.pntd.0002183
Vu B, Chen M, Crawford RJ, Ivanova EP (2009) Bacterial extracellular polysaccharides involved in biofilm formation. Molecules 14:2535–2554. doi:10.3390/molecules14072535
Wong TY, Preston LA, Schiller NL (2000) Alginate lyase: review of major sources and enzyme characteristics, structure-function analysis, biological roles, and applications. Annu Rev Microbiol 54:289–340. doi:10.1146/annurev.micro.54.1.289
Yadav V, Yadav PK, Yadav S, Yadav KDS (2010) α-l-Rhamnosidase: a review. Process Biochem 45:1226–1235. doi:10.1016/j.procbio.2010.05.025
Yan J, Mao J, Mao J, Xie J (2014) Bacteriophage polysaccharide depolymerases and biomedical applications. BioDrugs 28:265–274. doi:10.1007/s40259-013-0081-y
Acknowledgments
DPP acknowledges the financial support from the Portuguese Foundation for Science and Technology (FCT) through the grant SFRH/BD/76440/2011. SS is an FCT investigator (IF/01413/2013). The authors also thank FCT for the Strategic Project of the UID/BIO/04469/2013 unit, FCT and European Union funds (FEDER/COMPETE) for the project RECI/BBB-EBI/0179/2012 (FCOMP-01-0124-FEDER-027462).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
(PDF 382 kb)
Rights and permissions
About this article
Cite this article
Pires, D.P., Oliveira, H., Melo, L.D.R. et al. Bacteriophage-encoded depolymerases: their diversity and biotechnological applications. Appl Microbiol Biotechnol 100, 2141–2151 (2016). https://doi.org/10.1007/s00253-015-7247-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-7247-0