Abstract
This study aimed to compare the microbial community structures and compositions in composting and vermicomposting processes. We applied 454 high-throughput pyrosequencing to analyze the 16S rRNA gene of bacteria obtained from bio-stabilization of sewage sludge and cattle dung. Results demonstrated that vermicomposting process presented higher operational taxonomic units and bacterial diversity than the composting. Analysis using weighted UniFrac indicated that composting exhibited higher effects on shaping microbial community structure than the vermicomposting. The succession of dominant bacteria was also detected during composting. Firmicutes was the dominant bacteria in the thermophilic phase of composting and shifted to Actinomycetes in the maturing stage. By contrast, Proteobacteria accounted for the highest proportions in the whole process of the vermicomposting. Furthermore, vermicomposting contained more uncultured and unidentified bacteria at the taxonomy level of genus than the composting. In summary, the bacterial community during composting significantly differed from that during vermicomposting. These two techniques played different roles in changing the diversity and composition of microbial communities.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Composting and vermicomposting are used to bio-stabilize solid organic wastes (Lazcano et al. 2008; Vivas et al. 2009). Generally, two stages can be categorized in the composting process: (i) thermophilic stage, which is characterized by high pile temperature and intensive decomposition of organics; this stage constitutes the active phase of composting (Lazcano et al. 2008); and (ii) maturing stage, in which the temperature decreases to ambient range and the organic compounds continue to degrade slowly (Takaku et al. 2006). In comparison with composting, vermicomposting involves bio-oxidation and stabilization of organic materials through the joint action of earthworms and microorganisms (Vivas et al. 2009). It can also be divided into two phases: (i) active phase, in which earthworms grow fast, metabolize actively, and modify the physical state and microbial composition of substrate greatly; in this period, organic degradation significantly occurs because of the strong synergy effect between earthworms and microorganisms (Lores et al. 2006; Lazcano et al. 2008); and (ii) maturation phase, which is characterized by decreased earthworm activity or movement to fresh layers to obtain undigested wastes, and the modified microbes are responsible for the decomposition of remaining organic wastes (Gomez-Brandon et al. 2011).
Microorganisms are the mainly contributor for biochemical degradation of organic matter during composting and vermicomposting (de Gannes et al. 2013; Huang et al. 2013). Thus, a comprehensive understanding of microbial communities throughout the composting and vermicomposting processes cannot only elucidate the decomposition systems and enhance the quality of final products but also reveal the differences between these two processes for organic stabilization. Currently, the composition and dynamics of microbial communities during composting and vermicomposting have been examined through culture-dependent or independent techniques (Li et al. 2013; Huang et al. 2013). Nevertheless, early studies have focused only on the microbial communities in individual systems, composting, or vermicomposting (Fernandez-Gomez et al. 2010; He et al. 2013). Previous studies have also emphasized the comparison of microbial community in the final compost and vermicompost produced by single or mixed substrates (Fracchia et al. 2006; Vivas et al. 2009; Fernandez-Gomez et al. 2012). However, few studies have compared the microbial structure and community during the entire process of composting and vermicomposting.
Conventional molecular biological methods, including terminal restriction fragment length polymorphism, denaturing gradient gel electrophoresis (DGGE), and clone library (Vivas et al. 2009; Szekely et al. 2009; Partanen et al. 2010; Li et al. 2013), are used to specify the changes in microbial community during composting or vermicomposting process. These techniques can provide novel insights into microbial community in culture-independent manner compared with culture-dependent methods. However, the inherent limitations of these methods restrict the access of detailed information for microbial community structures (de Gannes et al. 2013). For example, the clone library method is expensive, time consuming, and have limited sequences generated, which made it difficult to provide comprehensive and systematic information about microbial community structures (DeSantis et al. 2007). In PCR–DGGE, one band may contain more than one species (Ma et al. 2013). Given these limitations, the new and advanced methods are needed to further investigate the dynamics and differences of microbial community structures and composition in composting and vermicomposting.
Pyrosequencing is a promising new tool which has 10 to 100 times higher output than the traditional Sanger techniques (Roesch et al. 2007; Acosta-Martínez et al. 2010). Thus, it can be used to completely and accurately elucidate the microbial communities compared with traditional molecular biological techniques (Acosta-Martínez et al. 2010). Pyrosequencing has been widely used to analyze the microbial communities in various environmental samples, such as drinking water distribution system biofilm (Luo et al. 2013), wastewater treatment plants (Sanapareddy et al. 2009), soils (Acosta-Martínez et al. 2010), and composts (de Gannes et al. 2013). However, to our knowledge, only a few studies have used pyrosequencing to analyze bacterial communities in vermicomposting and compare the bacterial communities between composting and vermicomposting. Thus, this study aimed to (i) simultaneously investigate the bacterial structure and composition during composting and vermicomposting of sewage sludge (SS) and cattle dung (CD) by 454 pyrosequencing and (ii) to compare the bacterial community characteristics between composting and vermicomposting processes.
Materials and methods
Source materials
SS was obtained from a municipal wastewater treatment plant (Shanghai, China). CD was acquired from a farm in Pudong New District, Shanghai, China. The main physico-chemical characteristics of SS and CD are listed in Table 1. A mixture of fresh SS and CD at 2:3 (w/w dry weight) was used for composting and vermicomposting experiments.
Bio-stabilization processes
Composting was performed in cylindrical vessels (35 cm diameter × 50 cm height). Ambient temperature was maintained at 40 °C for the first 10 days, so the compost can rapidly enter the thermophilic period. The initial moisture content was adjusted to 65 %, and uniformly forced ventilation was used to supply oxygen. In the first 30 days of composting, the piles were manually turned every 5 days. In the last 30 days, the forced ventilation was terminated and the piles were stirred daily for stabilization. Vermicomposting was conducted in circular plastic containers (30 cm diameter × 20 cm depth) filled with 4 kg of mixed substrate (wet weight). The initial mixture was manually turned every 24 h in the first 7 days to reduce volatiles and other substances toxic to earthworms. Thereafter, 50 non-clitellated earthworms (Eisenia fetida) were inoculated into the containers. No new materials were added during whole vermicomposting. The containers with pierced cover lid were placed in dark with constant temperature (23 ± 2 °C). Deionized water was used to maintain the moisture content at the range of 70 to 80 %.
The composting and vermicomposting run were conducted in triplicate and performed for 100 days. Sampling was conducted on the day 10 and day 60 for the composting treatment, while the samples were collected on the day 20 and day 80 for the vermicomposting experiment. Approximately 30 g fresh compost/vermicompost sample was collected from three different locations and mixed thoroughly to provide a composite sample. Then the entire sample was divided into two parts: one part was immediately stored at −20 °C until DNA extraction, while the other part was air-dried, grounded in a stainless steel blender, and stored in a desiccator prior to physico-chemical properties analysis.
Physico-chemical analysis
The temperature was recorded at the center of piles every day during the composting process using a mercury thermometer. The moisture content of fresh samples was determined by oven-drying to a constant weight at 105 °C. The total organic carbon (TOC) was determined by TOC-VCPN analyzer (Shimadzu, Japan). The pH meter was used to measure the pH on aqueous suspensions of the dried samples (1:10, w/v, sample/water ratio). The concentrations of ammonia (N-NH3) were determined by modified sulfamic acid technique. Available of phosphate (AP) was determined in 0.5 M NaHCO3 extracts (1:10 w/v) followed by molybdenum-ascorbic colorimetric method.
DNA extraction and PCR amplification
The FastDNA® SPIN kit for soil (MP Biomedicals, Illkirch, France) was used to extract DNA from compost and vermicompost samples according to the manufacturer’s protocols. Concentration and purity of the extracted DNA were determined by analyzing its absorbance at 260 and 280 nm with a Nanodrop® ND-1000 spectrophotometer (Labtech International, UK). The qualified DNA samples were stored at −20 °C before subsequent analyses.
High-throughput 16S rRNA gene pyrosequencing
To construct 454 pyrosequencing gene libraries, we amplified the extracted DNA through PCR using the primer set 8 F (5ʹ-AGAGTTTGATCCTGGCTCAG-3ʹ) and 533R (5ʹ-TTACCGCGGCTGCTGGCAC-3ʹ) for the V1–V3 region of the 16S rRNA gene. In order to sort various DNA samples in a single 454 GS-FLX run, we inserted the fused forward primer, including a 10-nucleotide barcode, between the Life Sciences primer A and the 8 F primer. The following amplification cycling scheme was used: 95 °C for 2 min; followed by 25 cycles at 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s; and a final extension step at 72 °C for 5 min. The barcodes PCR products were mixed in equal concentrations and sequenced using a Roche 454 FLX Titanium sequencer (Roche, Nutley, NJ, USA) according to standard protocols (Margulies et al. 2005). The pyrosequencing results have been deposited into the NCBI short reads archive database (accession number: PRJNA272469).
Sequence analysis
The 16S rRNA raw sequence were optimized in terms of length, quality, primer and barcode mismatches, and chimera identification and removal by using the QIIME pipeline (Caporaso et al. 2010). The sequences were classified into operational taxonomic unit (OTU) by using a 97 % identity threshold, and the most abundant sequence of each OTU was considered as its representative sequence (Edgar 2010). Rarefaction curve, Shannon diversity index, and species richness estimator of Chao1 were calculated in QIIME (version 1.17). The RDP classifier (version 2.2) included in the QIIME (version 1.17) was used for taxonomic identities at a confidence threshold of 0.7 (Wang et al. 2007). OTUs were considered unclassified if a reliable match was not found for the representative sequence within the RDP database. The relative percent of a given phylogenetic group was set as the quotient of its sequence number to the total number of sequences per sample. A phylogenetic tree was built by FastTree using the aligned OTU representative sequences, and Fast UniFrac was used to compare bacterial community structures (Hamady et al. 2010; Price et al. 2010).
Results
Physico-chemical changes
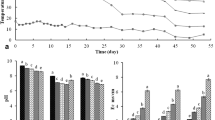
Figure 1a shows the variation of temperature profile during 100 days of composting. The primary indicator of microbiological activity in the composting process is temperature. In this study, the evolution of temperature had three periods, which are the initial heating phase, thermophilic phase, and cooling phase. It rose very quickly during the initial heating phase and achieved 50 °C on day 4. Troy et al. (2012) suggested that a quick increase in temperature during the heating phase was due to the rapid breakdown of easily degradable organic matter in raw materials by microbial activity. The thermophilic period was maintained for 14 consecutive days. Then the temperature dropped gradually which indicated the end of thermophilic phase, and the composting turned to cooling and maturation stages. On the other hand, TOC was also reduced rapidly in the thermophilic phase of composting due to the intense activity of microbes (Fig. 1b). As composting entered the cooling and maturation phases, TOC began to decrease slowly. Therefore, the samples from day 10 and day 60 of the composting can be considered as the active and mature stage (Com-A and Com-M), respectively. As for vermicomposting, the individual live weight of earthworm increased greatly from 0.25 to 0.71 g during the first 40 days of the vermicomposting, and the earthworm weight remained stable for the next 2 weeks. Moreover, the earthworm was also very active in the first 40 days. After 60 days, the biomass of earthworm decrease significantly and the activity of earthworm was reduced significantly as well due to the shortage of feedstock (Gomez-Brandon et al., 2011). In addition, TOC decreased rapidly in the first 60 days as compared to the latter stage. Thus, the samples collected from day 20 and day 80 of the vermicomposting could also be considered as the active and mature stage (Vom-A and Vom-M), respectively.
Microbial diversity and richness of bacterial phylotypes
Five 16S rRNA gene libraries were constructed by pyrosequencing the amplified DNA from the initial mixture (IM), Com-A, Vom-A, Com-M, Vom-M; these libraries contained 6818, 9635, 7800, 10,140, and 9230 high-quality reads (average length of 420 bp), respectively. By aligning at a uniform length of 420 bp, 1153 (IM), 1083 (Com-A), 1580 (Vom-A), 656 (Com-M), and 1511 (Vom-M) OTUs were clustered at a 3 % distance (Table 2). The rarefaction curves of the five samples at 3 % distance threshold are shown in Fig. 2. No rarefaction curves reached a plateau, and new OTUs still continued to appear in spite of the 10,000 sequences generated by pyrosequencing. This result indicated that the microbial community was abundant in the composting and vermicomposting systems. Moreover, pyrosequencing successfully revealed a higher diversity of bacterial communities during composting and vermicomposting than other conventional molecular biological methods, such as DGGE and clone library (Vivas et al., 2009; Huang et al., 2013). As shown in Table 2, the maximum numbers of OTUs estimated using the Chao1 estimator in infinite sampling were 2079 (IM), 2139 (Com-A), 2661 (Vom-A), 1057 (Com-M), and 2465 (Vom-M). This finding indicated that vermicomposting exhibited higher bacterial community richness than the composting, with the maximum OTUs detected in Vom-A. Ace estimators also proved this observation.
The Shannon diversity index provides not only the species richness (the species number) but also how the distribution of each species (evenness of the species) among all species in the community (Shannon and Weaver, 1963; Lu et al. 2012). In this study, Vom-A exhibited the highest diversity (Shannon = 6.23) among the five communities. Moreover, the Shannon index in vermicomposting process was significantly higher than that in the composting process.
Comparative analysis of bacterial community
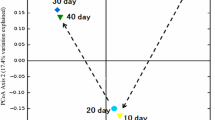
The similarity/dissimilarity of the 16S sequences among different samples was measured using the weighted UniFrac clustering method (Hamady et al. 2010). As depicted in Fig. 3, the compost groups (including Com-A and Com-M) were separated from other groups, and Com-A exhibited the lowest similarity, while the two samples from vermicomposting (including Vom-A and Vom-M) clustered together and presented a relatively close distance with IM. However, Com-A and Com-M were separated from IM, and a clear distinction was detected between the samples obtained from composting and vermicomposting. Moreover, the community structure significantly differed between Com-A and Com-M even if these samples were obtained from the composting process.
Taxonomic complexity of bacterial community
Figure 4 shows the relative bacterial community abundance at the phylum level in composting and vermicomposting. A total of 33 phyla were identified, and Proteobacteria, Bacteroidetes, Actinobacteria, Planctomycetes, and Firmicutes were identified as the dominant phyla.
The five communities evidently differed in terms of the distribution of phyla Bacteroidetes, Firmicutes, and Proteobacteria in the total community composition. As delineated in Fig. 4, Firmicutes was the dominant phylum in Com-A with a proportion of 71.1 %. Previous investigations have identified Firmicutes as a prominent member in the thermophilic stage (Yamada et al. 2008; Adams and Frostick 2009). Nevertheless, Firmicutes significantly reduced in Com-M and accounted for 3.3 % of all OTUs only. Proteobacteria was the second most abundant phylum in Com-A, and it increased to 33.1 % in Com-M. Actinomycetes, the most dominant phylum in Com-M, presented an increasing trend during composting. In addition, Bacteroidetes significantly increased during composting and accounted for 0.5 and 10.3 % in Com-A and Com-M, respectively.
The microbial community structure during vermicomposting distinctly differed from that in composting. As described in Fig. 4, Proteobacteria was the single largest group that accounted for more than 50 % of the total sequences in vermicomposting and its proportion minimally changed between Vom-A and Vom-M. Moreover, Bacteroidetes was another dominant phylum that occupied 15.1 and 14.0 % in Vom-A and Vom-M, respectively. In addition, Actinomycetes, Acidobacteria, and Planctomycetes demonstrated a large proportion during vermicomposting. The vermicomposting also presented a significantly higher proportion of Acidobacteria than the composting.
Analysis on the genus level allows us to infer the functions of the community (Fig. 5). On the one hand, the dominant populations in Com-A include Ureibacillus (53.9 %), Tepidimicrobium (8.1 %), Symbiobacterium (4.3 %), and Pseudoxanthomonas (4.4 %). These bacteria, particularly Ureibacillus, can strongly decompose proteins and cellulose. Similarly, several studies have reported that Ureibacillus and Tepidimicrobium, which belong to the phylum Firmicutes, are typical genera in the thermophilic stages of composting (Takaku et al. 2006; Nakasaki et al. 2009). On the other hand, Kribbella (22.6 %) was the most dominant genus in Com-M, indicating that composting reached the maturation stage. Bordetella (10.2 %), Olivibacter (5.3 %), Streptomyces (8.2 %), Parapedobacter (3.8 %), Mycobacterium (2.7 %), and Bacillus (2.5 %) were also detected in Com-M.
The dominant genus in vermicomposting clearly differed from that in the composting. As shown in Fig. 5, the prevailing populations during vermicomposting included Rhodanobacter, Altererythrobacter, Lysobacter, Paenibacillus, Rhizomicrobium, and Luteimonas, which are usually detected in various environmental samples (Yasir et al. 2009; Acosta-Martínez et al. 2010; de Gannes et al. 2013). Moreover, Acidobacterium was only detected in vermicomposting, and its content considerably increased from Vom-A to Vom-M. This finding could be attributed to the lower pH of vermicomposting relative to that of the composting (Table 1). In addition, pyrosequencing demonstrated that the ranges of 14.3 to 15.5 % (composting) and 24.8 to 31.2 % (vermicomposting) of the total OTUs were not classified at the genus level; hence, these bacteria are considered unknown.
Discussion
Comparison of the effects of composting and vermicomposting on the structure of microbial community
Chao1 estimators and Shannon indices showed that the bacteria in vermicomposting were richer and higher in terms of diversity than those in the composting. Moreover, the lower Shannon diversity in the composting samples than that in the initial materials suggested that some microbes cannot adapt to the compost environment and were eliminated in the process of aerobic composting (He et al. 2013). Furthermore, previous studies have demonstrated that the high temperatures in the thermophilic stage of composting can severely inhibit the growth of mesophilic bacteria and reduce the bacterial population diversity (Tiquia 2005; de Gannes et al. 2013). Although the temperatures declined in the maturation stage of composting, the residue is mainly dominated by refractory organics. Moisture content also decreased (Table 1); thus, the growth and reproduction of microbes were significantly inhibited, resulting in low community structure diversity (Cahyani et al. 2003; Tiquia 2005; Partanen et al. 2010).
By contrast to composting, vermicomposting considerably enriched the bacterial community diversity in initial feedstock. Our results were in accordance with several studies in which vermicompost exhibited more bacterial composition than the feedstock and compost (Vivas et al. 2009; Huang et al. 2013). The higher microbe diversity may be attributed to the mesophilic conditions or favorable action of earthworms (Tiunov and Scheu 2000; Sheehan et al. 2008) in the vermicomposting, which are beneficial to various types of bacteria (Domínguez et al. 2010). The abundant bacterial community in earthworm guts and casts may also contribute to high bacterial community during vermicomposting (Aira et al. 2009; Sen and Chandra 2009; Liu et al. 2012). Additionally, Toyota and Kimura (2000) demonstrated that E. fetida have an indigenous gut-associated microflora that can be beneficial to the microbial community in the mature vermicompost. Furthermore, several studies have concluded that earthworms can change the diversity and abundance of microorganisms directly by predating and/or stimulating special bacteria (Byzov et al. 2007; Yasir et al. 2009).
In the present study, strong shifts in the bacterial structure were observed in different composting stages, indicating that the bacterial community structure changed dynamically during composting. Self-heating composting can significantly alter the microbial community composition and structure because of pile temperature and substrates nutrient level to bacteria (Vivas et al. 2009; Adams and Frostick 2009). By contrast, vermicomposting can also affect the microbial community structure to some extent. Gomez–Brandon et al. (2011) suggested that earthworms strongly alter the physical and chemical states of initial wastes during vermicomposting then resulting in significant changes of bacterial communities within the initial feedstock. Moreover, the activity of earthworms might shape the microbial composition by selective ingestion and/or stimulation of specific taxa. Although vermicomposting can also change the microbial community in the initial material, composting evidently provided higher effects on the variation of microbial community structure as compared to the vermicomposting.
Bacterial phylum distribution and function as affected by composting and vermicomposting
In the present study, the most dominant bacteria in the thermophilic stages of composting were Firmicutes (genera Bacillus, Paenibacillus, Symbiobacterium, Tepidimicrobium, and Ureibacillus). Previous works on composting suggested that an increase of low G + C gram-positive bacteria from Firmicutes is often found in the hot stage of composting, even in different composting systems feed with various organic wastes (Takaku et al. 2006; de Gannes et al. 2013). Other studies have also stated that Bacillus and Ureibacillus genera are the most dominant bacterial taxon in thermophilic and the entire stage of composting (Dees and Ghiorse 2001; Juteau et al. 2004; Fracchia et al. 2006). Notably, previous studies on thermophilic composting have identified Proteobacteria as a minor group through PCR–DGGE or clone library (Ishii et al. 2000; Partanen et al. 2010). However, in this study, pyrosequencing revealed the prevalence of Proteobacteria (genera Bordetella and Pseudoxanthomonas), thus, advancing knowledge on microbes. In addition, Actinobacteria (genus Streptomyces) was also predominant in the active stage of composting. These bacteria can form spores and allow them to tolerate the high temperature during the hot stage of composting (Tian et al. 2013). The succession of microbial communities evidently occurred during composting, with Actinobacteria, Proteobacteria, and Bacteroidetes more dominant than Firmicutes in the maturing stage of composting. Takaku et al. (2006) found that the dominant bacterial group changed from phylum Firmicutes (thermophilic stage) to the phyla Actinobacteria, Bacteroidetes, and Proteobacteria (maturing stage) during composting of garbage. Actinobacteria is considered important during composting because of its ability in the decomposition of cellulose and chitin; hence, these bacteria play a crucial role in the bio-stabilization of refractory organics (Vivas et al. 2009; Partanen et al. 2010; Federici et al. 2011). The abundance of Proteobacteria (genera Bordetella, Candidatus Liberibacter, and Filomicrobium) was also significantly higher in the mature phase than that in the thermophilic phase; this finding confirmed that some mesophilic bacteria could enter dormancy during the thermophilic period and recover to grow at the cooling phase (Nakasaki et al. 1985; Yamada et al. 2008). Bacteroidetes at the maturing stage mainly included the genera Olivibacter and Parapedobacter. The Bacteroidetes often includes various bacteria that have strong ability to hydrolyze many macromolecules, such as starch, cellulose, proteins, and chitin (Takaku et al. 2006). Besides, members of this phylum have also been detected in different composts (Takaku et al. 2006; Danon et al. 2008; Li et al. 2013). Therefore, we can conclude that the bacteria in active stage of composting could tolerate the high temperature, while the dominant bacteria in the maturing stage of composting were characterized by the ability to degrade residual refractory organic matter.
As compared to the composting, Proteobacteria was the most dominant phylum in the entire vermicomposting process and it minimally changed between the active and maturing stages. However, its genera, such as Altererythrobacter, Brevundimonas, Pseudoxanthomonas, Pusillimonas, and Rhizomicrobium, significantly changed during vermicomposting. This finding indicated that the variation in bacterial community was also detected in the two steps of vermicomposting. Early studies have indicated that earthworms can digest some microorganisms, whereas others may survive or/and proliferate in the gut and then colonize into the vermicompost (Fracchia et al. 2006). The phylum Bacteroidetes was detected in all samples but was more abundant in Vom-A than in Vom-M. The anoxic environment and enrichment of nutrients in the gut of earthworm could favor these anaerobic microbes (Karsten and Drake 1995; Huang et al. 2013). Moreover, the genus of Bacteroidetes found in the present study was also detected in the earthworm’s digestive tract (Byzov et al. 2009). Furthermore, the members of Bacteroidetes demonstrated an essential role in the synergistic action with earthworms to stabilize refractory organic matter during vermicomposting (Danon et al. 2008; Liu et al. 2012). Actinobacteria (genera Arthrobacter, Demequina, Microbacterium, Mycobacterium, Nocardioides, and Streptomyces) was another dominant phylum during vermicomposting whose proportion increased. This finding suggested that the bacterial communities exhibited strong and specific changes in vermibed microbiota because of the degradation and stabilization of organic matter. Although the genus Bacilli was also detected during vermicomposting, its presence was very limited compare to that during composting. Thus, as compared to composting, the bacteria in the vermicomposting might be depended both on the activity of earthworm and the stability or variation of substrates.
The most striking difference during composting and vermicomposting processes was the presence of Acidobacteria in the latter, and this phylum increased significantly during vermicomposting. Several studies also found that bacteria from the phylum Acidobacteria are exclusively detected in vermicompost or vermifilter (Vivas et al. 2009; Fernandez-Gomez et al. 2012; Liu et al. 2012). As shown in Table 1, the pH in vermicomposting was significantly lower than that in the composting. Therefore, the acidification of microbial activity that resulted in low pH value may be the reason for prevalence of Acidobacteria and Planctomycetes in vermicompost (Fracchia et al. 2006; Liu et al. 2012). In addition, the 16S rRNA sequences from the vermicompost at the genus level were mainly related to yet uncultured or unclassified bacteria (37.2 to 49.3 % of the total sequences), while this percentage was lower (10.3 to 15.4 %) in the compost. Therefore, more studies should focus on bacterial phylogenetic and functional diversity during vermicomposting.
Significance of the research
To the best of our knowledge, this study is the first attempt to use pyrosequencing technology for comparing microbial communities involved in the composting and vermicomposting processes. Pyrosequencing is an effective method used to elucidate the diversity of microorganisms in the bio-stabilization process, and some interesting results were acquired by this new technique. Although both the composting and vermicomposting can stabilize initial waste materials, the succession of bacterial community structure in the latter significantly differed from that in the former. The substrates initially used were similar in both treatments, but different species of bacterial microbes participated during vermicomposting and composting. Therefore, the treatment process may be an important factor that affects the abundance and diversity of microbial communities. In the composting process, the self-heating phase with its subsequent maturation phase is the major biological conversion step. The bacteria in active stage of composting could stand the high temperature, while the bacteria in the maturing stage of composting had the ability to degrade residual refractory organic matter. In the vermicomposting process, the activity of earthworms can modify the microbial diversity and composition of the vermibeds by ingesting or stimulating specific taxa. Moreover, the different physico-chemical properties of the substrate caused by composting and vermicomposting treatment also played an important role on bacterial community and composition.
The succession of dominant bacteria distinctly occurred during composting, and unique dominant species were observed at different stages, while the succession of predominant genera remained ambiguous in vermicomposting. Moreover, vermicomposting also showed significantly higher bacterial diversity than the composting. This finding suggested that the degradation and stabilization processes can be clearly distinguished during composting. Hence, the dominant populations played the most important role in the transformation of organic wastes at each stage of composting. By contrast, more functional gene compositions and microbial populations were involved in the vermicomposting process. Besides, the stabilization and treatment process presented more metabolic diversity and complexity in vermicomposting than that in aerobic composting. Nevertheless, divergence in microbial community structures, particularly the high abundance of unclassified and uncultured bacteria in vermicomposting, could be attributed to different factors. Therefore, further studies must be performed to elucidate the role of functional microorganisms in composting and vermicomposting processes.
References
Acosta-Martínez V, Dowd SE, Sun Y, Wester D, Allen V (2010) Pyrosequencing analysis for characterization of soil bacterial populations as affected by an integrated livestock-cotton production system. Appl Soil Ecol 45(1):13–25
Adams JD, Frostick LE (2009) Analysis of bacterial activity, biomass and diversity during windrow composting. Waste Manage 29(2):598–605
Aira M, Monroy F, Dominguez J (2009) Changes in bacterial numbers and microbial activity of pig slurry during gut transit of epigeic and anecic earthworms. J Hazard Mater 162(2–3):1404–1407
Byzov BA, Khomyakov NV, Kharin SA, Kurakov AV (2007) Fate of soil bacteria and fungi in the gut of earthworms. Eur J Soil Biol 43:S149–S156
Byzov BA, Nechitaylo TY, Bumazhkin BK, Kurakov AV, Golyshin PN, Zvyagintsev DG (2009) Culturable microorganisms from the earthworm digestive tract. Microbiology 78(3):360–368
Cahyani VR, Matsuya K, Asakawa S, Kimura M (2003) Succession and phylogenetic composition of bacterial communities responsible for the composting process of rice straw estimated by PCR-DGGE analysis. Soil Sci Plant Nutr 49(4):619–630
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Tumbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336
Danon M, Franke-Whittle IH, Insam H, Chen Y, Hadar Y (2008) Molecular analysis of bacterial community succession during prolonged compost curing. Fems Microbiol Ecol 65(1):133–144
de Gannes V, Eudoxie G, Hickey WJ (2013) Prokaryotic successions and diversity in composts as revealed by 454-pyrosequencing. Bioresour Technol 133:573–580
Dees PM, Ghiorse WC (2001) Microbial diversity in hot synthetic compost as revealed by PCR-amplified rRNA sequences from cultivated isolates and extracted DNA. Fems Microbiol Ecol 35(2):207–216
DeSantis TZ, Brodie EL, Moberg JP, Zubieta IX, Piceno YM, Andersen GL (2007) High-density universal 16S rRNA microarray analysis reveals broader diversity than typical clone library when sampling the environment. Microb Ecol 53(3):371–383
Domínguez J, Aira M, Gómez-Brandón M (2010) Vermicomposting: earthworms enhance the work of microbes. In: Insam H, Franke-Whittle I, Goberna M (eds) Microbes at work: from wastes to resources. Springer, Berlin Heidelberg, pp 93–114
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19):2460–2461
Federici E, Pepi M, Esposito A, Scargetta S, Fidati L, Gasperini S, Cenci G, Altieri R (2011) Two-phase olive mill waste composting: community dynamics and functional role of the resident microbiota. Bioresour Technol 102(23):10965–10972
Fernandez-Gomez MJ, Nogales R, Insam H, Romero E, Goberna M (2010) Continuous-feeding vermicomposting as a recycling management method to revalue tomato-fruit wastes from greenhouse crops. Waste Manage 30(12):2461–2468
Fernandez-Gomez MJ, Nogales R, Insam H, Romero E, Goberna M (2012) Use of DGGE and COMPOCHIP for investigating bacterial communities of various vermicomposts produced from different wastes under dissimilar conditions. Sci Total Environ 414:664–671
Fracchia L, Dohrmann AB, Martinotti MG, Tebbe CC (2006) Bacterial diversity in a finished compost and vermicompost: differences revealed by cultivation-independent analyses of PCR-amplified 16S rRNA genes. Appl Microbiol Biot 71(6):942–952
Gomez-Brandon M, Aira M, Lores M, Dominguez J (2011) Changes in microbial community structure and function during vermicomposting of pig slurry. Bioresour Technol 102(5):4171–4178
Hamady M, Lozupone C, Knight R (2010) Fast UniFrac: facilitating high-throughput phylogenetic analyses of microbial communities including analysis of pyrosequencing and PhyloChip data. ISME J 4(1):17–27
He Y, Xie K, Xu P, Huang X, Gu W, Zhang F, Tang S (2013) Evolution of microbial community diversity and enzymatic activity during composting. Res Microbiol 164(2):189–198
Huang K, Li F, Wei Y, Chen X, Fu X (2013) Changes of bacterial and fungal community compositions during vermicomposting of vegetable wastes by Eisenia foetida. Bioresour Technol 150:235–241
Ishii K, Fukui M, Takii S (2000) Microbial succession during a composting process as evaluated by denaturing gradient gel electrophoresis analysis. J Appl Microbiol 89(5):768–777
Juteau P, Tremblay D, Villemur R, Bisaillon JG, Beaudet R (2004) Analysis of the bacterial community inhabiting an aerobic thermophilic sequencing batch reactor (AT-SBR) treating swine waste. Appl Microbiol Biot 66(1):115–122
Karsten GR, Drake HL (1995) Comparative assessment of the aerobic and anaerobic microfloras of earthworm guts and forest soils. Appl Environ Microb 61(3):1039–1044
Lazcano C, Gomez-Brandon M, Dominguez J (2008) Comparison of the effectiveness of composting and vermicomposting for the biological stabilization of cattle manure. Chemosphere 72(7):1013–1019
Li Q, Wang XC, Zhang HH, Shi HL, Hu T, Ngo HH (2013) Characteristics of nitrogen transformation and microbial community in an aerobic composting reactor under two typical temperatures. Bioresour Technol 137:270–277
Liu J, Lu Z, Yang J, Xing M, Yu F, Guo M (2012) Effect of earthworms on the performance and microbial communities of excess sludge treatment process in vermifilter. Bioresour Technol 117:214–221
Lores M, Gomez-Brandon M, Perez-Diaz D, Dominguez J (2006) Using FAME profiles for the characterization of animal wastes and vermicomposts. Soil Biol Biochem 38(9):2993–2996
Lu L, Xing D, Ren N (2012) Pyrosequencing reveals highly diverse microbial communities in microbial electrolysis cells involved in enhanced H2 production from waste activated sludge. Water Res 46(7):2425–2434
Luo J, Liang H, Yan L, Ma J, Yang Y, Li G (2013) Microbial community structures in a closed raw water distribution system biofilm as revealed by 454-pyrosequencing analysis and the effect of microbial biofilm communities on raw water quality. Bioresour Technol 148:189–195
Ma J, Wang Z, Yang Y, Mei X, Wu Z (2013) Correlating microbial community structure and composition with aeration intensity in submerged membrane bioreactors by 454 high-throughput pyrosequencing. Water Res 47(2):859–869
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen Z, Dewell SB, Du L, Fierro JM, Gomes XV, Godwin BC, He W, Helgesen S, Ho CH, Irzyk GP, Jando SC, Alenquer ML, Jarvie TP, Jirage KB, Kim JB, Knight JR, Lanza JR, Leamon JH, Lefkowitz SM, Lei M, Li J, Lohman KL, Lu H, Makhijani VB, McDade KE, McKenna MP, Myers EW, Nickerson E, Nobile JR, Plant R, Puc BP, Ronan MT, Roth GT, Sarkis GJ, Simons JF, Simpson JW, Srinivasan M, Tartaro KR, Tomasz A, Vogt KA, Volkmer GA, Wang SH, Wang Y, Weiner MP, Yu P, Begley RF, Rothberg JM (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437(7057):376–380
Nakasaki K, Sasaki M, Shoda M, Kubota H (1985) Characteristics of mesophilic bacteria isolated during thermophilic composting of sewage sludge. Appl Environ Microb 49:42–45
Nakasaki K, le Tran TH, Idemoto Y, Abe M, Rollon AP (2009) Comparison of organic matter degradation and microbial community during thermophilic composting of two different types of anaerobic sludge. Bioresour Technol 100(2):676–682
Partanen P, Hultman J, Paulin L, Auvinen P, Romantschuk M (2010) Bacterial diversity at different stages of the composting process. BMC Microbiol 10:94
Price MN, Dehal PS, Arkin AP (2010) FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS One 5(3), e9490
Roesch LF, Fulthorpe RR, Riva A, Casella G, Hadwin AK, Kent AD, Daroub SH, Camargo FA, Farmerie WG, Triplett EW (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1(4):283–290
Sanapareddy N, Hamp TJ, Gonzalez LC, Hilger HA, Fodor AA, Clinton SM (2009) Molecular diversity of a North Carolina wastewater treatment plant as revealed by pyrosequencing. Appl Environ Microb 75(6):1688–1696
Sen B, Chandra TS (2009) Do earthworms affect dynamics of functional response and genetic structure of microbial community in a lab-scale composting system? Bioresour Technol 100(2):804–811
Shannon CE, Weaver W (1963) The mathematical theory of communication. The University of Illinois Press, Urbana
Sheehan C, Kirwan L, Connolly J, Bolger T (2008) The effects of earthworm functional diversity on microbial biomass and the microbial community level physiological profile of soils. Eur J Soil Biol 44(1):65–70
Szekely AJ, Sipos R, Berta B, Vajna B, Hajdu C, Marialigeti K (2009) DGGE and T-RFLP analysis of bacterial succession during mushroom compost production and sequence-aided T-RFLP profile of mature compost. Microb Ecol 57(3):522–533
Takaku H, Kodaira S, Kimoto A, Nashimoto M, Takagi M (2006) Microbial communities in the garbage composting with rice hull as an amendment revealed by culture-dependent and -independent approaches. J Biosci Bioeng 101(1):42–50
Tian W, Sun Q, Xu DB, Zhang ZH, Chen D, Li CY, Shen QR, Shen B (2013) Succession of bacterial communities during composting process as detected by 16S rRNA clone libraries analysis. Int Biodeter Biodegr 78:58–66
Tiquia SM (2005) Microbial community dynamics in manure composts based on 16S and 18S rDNA T-RFLP profiles. Environ Technol 26(10):1101–1113
Tiunov AV, Scheu S (2000) Microbial biomass, biovolume and respiration in Lumbricus terrestris L. cast material of different age. Soil Biol Biochem 32(2):265–275
Toyota K, Kimura M (2000) Microbial community indigenous to the earthworm Eisenia foetida. Biol Fert Soils 31(3–4):187–190
Troy SM, Nolan T, Kwapinski W, Leahy JJ, Healy MG, Lawlor PG (2012) Effect of sawdust addition on composting of separated raw and anaerobically digested pig manure. J Environ Manag 111:70–77
Vivas A, Moreno B, Garcia-Rodriguez S, Benitez E (2009) Assessing the impact of composting and vermicomposting on bacterial community size and structure, and microbial functional diversity of an olive-mill waste. Bioresour Technol 100(3):1319–1326
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb 73(16):5261–5267
Yamada T, Suzuki A, Ueda H, Ueda Y, Miyauchi K, Endo G (2008) Successions of bacterial community in composting cow dung wastes with or without hyperthermophilic pre-treatment. Appl Microbiol Biot 81(4):771–781
Yasir M, Aslam Z, Kim SW, Lee SW, Jeon CO, Chung YR (2009) Bacterial community composition and chitinase gene diversity of vermicompost with antifungal activity. Bioresour Technol 100(19):4396–4403
Acknowledgments
The research was funded by the National Natural Science Foundation of China (51109161), the PhD Programs Foundation of Ministry of Education of China (20110072120029), the Fundamental Research Funds for The Central Universities (0400219187), and the National Spark Program of China (2010GA680004). We would like to thank the editor and anonymous reviewers to improve the quality of the manuscript.
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lv, B., Xing, M., Yang, J. et al. Pyrosequencing reveals bacterial community differences in composting and vermicomposting on the stabilization of mixed sewage sludge and cattle dung. Appl Microbiol Biotechnol 99, 10703–10712 (2015). https://doi.org/10.1007/s00253-015-6884-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6884-7