Abstract
The presence of metal resistance determinants in bacteria usually is attributed to geological or anthropogenic metal contamination in different environments or associated with the use of antimicrobial metals in human healthcare or in agriculture. While this is certainly true, we hypothesize that protozoan predation and macrophage killing are also responsible for selection of copper/zinc resistance genes in bacteria. In this review, we outline evidence supporting this hypothesis, as well as highlight the correlation between metal resistance and pathogenicity in bacteria. In addition, we introduce and characterize the “copper pathogenicity island” identified in Escherichia coli and Salmonella strains isolated from copper- and zinc-fed Danish pigs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Essential metals such as iron and copper can cycle between different oxidation states and are used in metalloenzymes that catalyze electron transport reactions. Zinc also plays a major structural and catalytic role in metalloenzymes and has been reported to counter oxidative stress. But in excess, all of these metals are deleterious to cells. To ensure their own survival, prokaryotes have developed mechanisms of maintaining cellular Zn2+ and Cu+ homeostasis, while eukaryotes invented very original Zn- and Cu-binding structures not present in prokaryotes. Such structures allow accumulation of Zn2+ in the intracellular organelles followed by its utilization in biological processes specific for a given cell type. In particular, macrophages employ Zn2+ and Cu+ to attack Fe–S clusters essential for bacterial survival (Braymer and Giedroc 2014; Dupont et al. 2011; Festa and Thiele 2012; Kashyap et al. 2014; Macomber and Imlay 2009; Neyrolles et al. 2015; Subashchandrabose et al. 2014; Xu and Imlay 2012). We hypothesize that such a mechanism, where bacterial killing occurs through accumulation of Zn2+ and Cu+ in the phagosome/vacuole, originated in protozoa long before multicellular life arose and that it later evolved in eukaryotic phagocytes. Our hypothesis is supported by the presence of the homologous copper transporter 1 (Ctr1) in macrophages and P80 in Dictyostelium discoideum and Acanthamoeba castellanii—proteins, both of which are involved in Cu+ uptake upon phagocytosis. In addition, amoebae are known to contain P-type ATPases (German et al. 2013; Burlando et al. 2002), and similar to macrophages, at least one of these P-type ATPases in A. castellanii could be pumping Zn2+ or Cu+ into the phagosome of amoeba (Fig. 1). Importantly, our hypothesis explains selection of genes involved in conferring copper and zinc resistance not only by the presence of these metals in the environment but also by protozoan predation as well. Since these determinants would aid survival in both protozoans and macrophages, one could expect a higher occurrence of additional copper and zinc resistance determinants in virulent bacteria.
A schematic overview of Zn and Cu involvement in phagosomal killing of bacteria. Macrophages and amoeba can exploit similar molecules for Zn2+ and Cu+ trafficking. ZIP family transporters allow Zn2+ uptake into the cytoplasm, and cation diffusion facilitator proteins (CDF) could deliver Zn2+ to the phagosome and other organelles, like mitochondria, Golgi, and endoplasmic reticulum (ER). Cu+ uptake and delivery to phagosomes occur due to copper transporter 1 (Ctr1, in amoeba known as P80), antioxidant 1 copper chaperone (Atox1), and in human macrophages, the P-type ATPase ATP7A. H+–ATPase causes acidification of the phagosomal milieu, while natural resistance-associated macrophage protein 1 (NRAMP1) removes Fe2+ and Mn2+, which are needed to protect (Mn2+) and rebuild degraded Fe–S clusters of bacteria. In addition, Cu+ amplifies toxicity of reactive oxygen species (hydroxyl radical (·OH) and hydroxide anion (OH−)). E. coli express genes encoding ZntA for Zn2+ efflux, CopA for Cu+ efflux, and the CusCBA complex for periplasmic Cu+ efflux, but virulent strains have additional copper resistance systems
A copper/silver resistance cluster or an ancestral defense to phagosomal killing using copper?
Genome sequencing projects have revealed that several strains of Salmonella enterica subspecies enterica harbor a ca. 12-kb copper resistance locus. This cluster is shown to either form part of a Tn7-like transposon inserted at the 3′ end of the gene that encodes a NAD-utilizing dehydrogenase on the chromosome (Peters et al. 2014), as found in isolates from serovars Heidelberg, Montevideo, Senftenberg, or Tennessee, or to be a part of a larger integrating conjugative element inserted at the pheV phenylalanine tRNA, as present in strains of the serovars Senftenberg, Ohio, or Cubana. Although there is a history of copper- and zinc-resistant bacteria being isolated from feces of animals fed with metal-supplement-containing diets, the presence of this resistance cluster has just recently been recognized.
Recently, we have sequenced the genomes of two Escherichia coli and three S. enterica serovar Typhimurium strains isolated from copper- and zinc-fed Danish pigs, hence displaying high-level copper resistance (Lüthje et al. 2014; Qin et al. 2014). One of the E. coli strains and all three of the Salmonella strains contained a specific 19-gene mobile genetic element that we have named as the “copper pathogenicity island.” In the E. coli isolate, we have identified this island as a part of aTn7-like transposon, while in the S. Typhimurium strains, it forms part of an approximately 80-kbp chromosomal element inserted at the pheU phenylalanine tRNA, similar to that identified in Heidelberg, Montevideo, Senftenberg, or Tennessee isolates (Qin et al. 2014). This genetic cluster is comprised of two previously reported metal ion resistance determinants, neither of which was realized until recently to be part of a single contiguous gene cluster (Crossman et al. 2010; Hobman and Crossman 2015). One, the pco determinant was first isolated from plasmid pRJ1004 from an Australian pig E. coli isolate (Brown et al. 1995) and confers copper resistance. The other, the sil determinant originally located on Salmonella Typhimurium plasmid pMG101, but later shown to have transferred into the chromosome of the host E. coli K-12 J53 strain (Randall et al. 2015), is associated with silver resistance (McHugh et al. 1975; Gupta et al. 1999). Later sequencing of pRJ1004 (NCBI accession no. KC146966) has identified two new genes among the entire 19-gene cluster—pcoF encoding a putative copper-binding protein and pcoG encoding a putative M23B metallopeptidase, an enzyme that has been implicated in pathogenicity (Bonis et al. 2010) (Fig. 2). We have identified this arrangement of pco/sil genes in a number of different genome and plasmid sequences.
Pco and Sil mechanisms in action. Proposed genes and protein products forming the molecular mechanisms of Pco- and Sil-mediated copper and silver detoxification and control in the cell. The bottom line indicates the genes and their transcriptional and translational directions, with the open circles representing potential promoter regions/transcript start sites. The illustrated function of each sil and pco gene product within the operon (Gram-negative) is deduced from homology modeling. The transcription of Pco proteins PcoABCDEFG appears to be regulated by PcoRS (left). The roles of PcoFG have not been elucidated. In addition to the oxidized catechol siderophores, copper may be detoxified from Cu+ to Cu2+ by the suggested multicopper oxidase PcoA. PcoB possibly functions as the outer membrane transporter, while, sitting in the inner membrane, PcoD drives the transport of Cu+ from the periplasm to the cytoplasm, with periplasmic PcoC chaperoning/delivering the Cu+ to PcoD. PcoE is an additional chaperone that binds copper in the periplasm and probably shuttles it to PcoA and/or PcoS. Similarly, the Sil system (right) contains a homolog to PcoE—periplasmic protein SilE. SilE is predicted to bind and chaperone Ag+, Cu+, and Cu2+ to the three-polypeptide, transmembrane, chemiosmotic RND exchange system (SilCBA), exporting the metal ions out of the cell. Likewise, SilF acts as a chaperone to SilCBA too. The other putative efflux pump mediating the mechanism is a P-type ATPase—SilP. Although conserved within the sil determinant, a role of SilG has not yet been determined. While the expression of silCFBAGP is thought to be governed by the two-component membrane sensor and transcriptional responder SilRS; the expression of silE, just like homolog pcoE, is thought to be regulated/co-regulated by the Cus system (Zimmerman et al. 2012), and therefore, SilE perhaps is involved in the activation of regulators SilRS (proposed by dotted arrow)
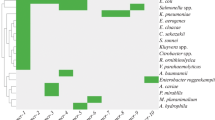
Similarly, the sil determinant has been associated with pathogenicity in the Enterobacter cloacae complex, where the presence of the genes conferring silver resistance was increased in isolates from hospital settings vs. strains associated with plants (Kremer and Hoffmann 2012). Although identification of the pco genes was not part of that study, their presence within the isolates harboring the full sil determinant is very likely based on the high rate of co-representation (Mourão et al. 2015) (Table 1).
Previous studies and genomic analysis have shown that the copper pathogenicity island is often plasmid associated. Transfer of such plasmids has resulted in a nosocomial outbreak of Klebsiella pneumoniae (Sandegren et al. 2012). Moreover, the pco/sil cluster has been identified on pAPEC-O2-R plasmids from avian pathogenic E. coli (Johnson et al. 2005), R478 from Serratia marcescens (Gilmour et al. 2004), plasmids pK2044 and pLVPK from K. pneumoniae strains (Chen et al. 2004; Wu et al. 2009), and on the chromosome or plasmids of many pathogenic enteric bacteria such as ETEC H10407 (Crossman et al. 2010) and EHEC O104:H4 (Hobman and Crossman 2015) and Enterobacter cloacae subsp. cloacae strain ATCC 13047 (Ren et al. 2010). For E. coli and Enterobacter strains, the copper pathogenicity island was identified in close vicinity to Tn7-like transposons with tnsABCD being present (Peters et al. 2014). At the same time, in Klebsiella strains, it was often associated with IS4-related elements and genes encoding a HNH endonuclease. These data together with the fact of a similar arrangement of a copper/silver resistance island found in Salmonella (Moreno Switt et al. 2012) indicate that the gene cluster behaves like a typical pathogenicity island.
Yersiniabactin—not just for iron
Our recent sequencing of E. coli strains isolated from copper-fed pigs allowed us to identify another determinant conferring increased copper resistance—a ten-gene yersiniabactin synthesis cluster (Lüthje et al. 2014). The yersiniabactin determinant is a well-known virulence factor responsible for copper binding (Chaturvedi et al. 2014) that can be present in pathogens such as K. pneumoniae (Fodah et al. 2014); Salmonella (Aviv et al. 2014); E. coli (Schubert et al. 2004), including the EHEC O104:H4 outbreak strain; and the highly virulent Yersinia pestis (Rakin et al. 2012). Interestingly, several strains of Klebsiella and E. coli appear to have the sil/pco determinant in addition to the yersiniabactin synthesis cluster (Table 1).
Copper and zinc resistance in Gram-positive bacteria
Pathogenicity of Gram-positive bacteria such as Enterococcus faecium, Enterococcus faecalis, Staphylococcus aureus, and Staphylococcus haemolyticus is also linked to transition metal resistance. Currently, we have sequenced the genomes of three highly copper-resistant E. faecium and three E. faecalis strains isolated from copper-fed pigs in Denmark (Zhang et al. 2015). As a result, we have identified additional copper resistance determinants characteristic for many pathogenic enterococci, tcrYAZB, encoding a negative transcriptional regulator, a copper chaperone, and two P1B-type ATPases flanked by mobile elements (Hasman 2005). In E. faecalis, this determinant has often been found in the vicinity of a gene encoding a multicopper oxidase resembling CueO, an adjacently encoded two-component system, and possibly CopY. Whether CueO is regulated by the adjacent two-component system or CopY is not known. In this genome region, there are also several putative copper chaperones and a prolipoprotein diacylglyceryl transferase, which has been associated with virulence (Cho et al. 2013; Reffuveille et al. 2012) (Fig. 3).
Copper fitness island in Enterococcus faecalis. Proposed genes and protein products forming the molecular mechanisms of copper detoxification in E. faecalis. The bottom line indicates the genes and their transcriptional and translational directions. TcrY regulates expression of tcrYAZB, encoding for the repressor, a cytoplasmic chaperone (TcrZ), and two P-type ATPases (TcrA and TcrB) responsible for Cu+ export. In close proximity to tcr genes encoding a two-component regulatory system (CueRS), a multicopper oxidase (CueO), a predicted metal chaperone (no homology to TcrZ and labeled “C”), and transcriptional repressor (CopY) have been identified. CueO is predicted to oxidize Cu+ to Cu2+. It is not clear to what extent the predicted chaperon (C) might be involved in copper detoxification. It has not yet been established if transcription of these genes is controlled by the two-component regulatory system (CusRS) responding to external copper concentration, CopY, as a response to internal copper concentrations, or both. Adjacent to and separating the two copper resistance determinants, genes encoding prolipoprotein diacylglyceryl transferase (A), integral membrane protein (B), hypothetical proteins (D), transposase (E), disrupted P-type ATPase (F), integrase (G), adenylate kinase (I), and resolvase (I) have been identified. The extent to which some of these proteins might be involved in copper detoxification has not been analyzed
In addition to copper, zinc resistance has also been linked to virulence and increased survival rates of pathogens. For example, in group A, Streptococcus czcD and gczA deletion mutants characterized by higher zinc sensitivity had shown much lower survival rates in the presence of neutrophils compared to wild-type strains (Ong et al. 2014). Certain correlation between the presence of the zinc resistance gene czrC, methicillin resistance, and virulence has been found in many S. aureus strains (Slifierz et al. 2014; Aarestrup et al. 2010). The gene crzC encodes a Zn2+-translocating P-type ATPase and is located next to a gene encoding a possible transcriptional regulator of the ArsR/SmtB family and a gene encoding a putative iron/zinc permease. According to genomic similarities, the latter might be a distant homolog of the zinc/iron importer ZupT. Sequencing of the S. haemolyticus SH32 clinical strain has identified two incomplete staphylococcal cassette chromosome (SCC) elements, with one of them, ψSCCmec(SH32), encoding a Cu(I)-translocating P-type ATPase (Yu et al. 2014). This strain was also shown to contain a putative cadmium resistance determinant cadXD, encoding for a Cd(II) transporter as well as a transcription regulator of the ArsR family (Yu et al. 2014). A recent study has reported that plasmid SAP078A in methicillin-resistant S. aureus CC22 SCCmecIV (EMRSA-15) contains cadCA, mco, and copB in addition to an ars operon conferring resistance to cadmium/zinc, copper, and arsenic, respectively (Loeffler et al. 2013). It was also shown that plasmid SAP078A is widespread among both human and animal isolates of S. aureus (Loeffler et al. 2013). Moreover, given the role of transition metals in the mammalian immune response, the presence of cadCA and copB/mco provides corresponding strains with an advantage (Hood and Skaar 2012). Interestingly, the epidemic ST22-IV has been replacing other MRSA clones from hospitals possibly due to enhanced virulence. Detailed studies revealed that ST22-IV has a significantly higher capacity to invade the A549 cells and a higher virulence in a murine model of acute lung infection causing severe inflammation and determining death in all the mice within 60 h (Baldan et al. 2012, 2015). We suggest that severe pathogenicity of ST22-IV might be partially attributed to increased transition metal resistance of this strain.
Cadmium and silver resistance—mutation plus selection equals evolution to new resistance?
Due to much lower environmental distribution of cadmium and silver, they are not essential micronutrients for bacteria and no data has been published on their involvement in bacteria/host interactions. Therefore, it is unlikely for bacteria to develop specific resistance mechanisms to these metals to a large extent. At the same time, most of the metal resistance determinants described to date are involved in detoxification of multiple substrates, e.g., conferring resistance to both copper and silver as well as to both zinc and cadmium (Rensing et al. 1999; Rosenzweig and Arguello 2012). Moreover, both E. coli and Salmonella contain detoxification systems for zinc and copper in addition to the possible plasmid-encoded resistance determinants on their chromosomes. In other words, it is quite unlikely that the evolutionary pressure comes exclusively from metal-contaminated environments. Rather, resistance to silver is a by-product of copper resistance and cadmium is a by-product of zinc resistance. In fact, resistance studies with S. aureus were unable to produce silver-resistant strains even after 42 days of continuous passage in the presence of AgNO3 (Randall et al. 2013) Similar results were observed for some Gram-negative organisms, whereas in E. coli strains, silver resistance arises as a result of mutations in both ompR and cusS or mediated through the sil system (Randall et al. 2015).
Metal homeostasis in bacteria is a delicate balance, especially since metals like zinc and iron have been found to be essential for the pathogenicity of these organisms (Cerasi et al. 2014; Gradassi et al. 2013; Pesciaroli et al. 2011). The fact that protozoan grazing might be a strong force on keeping or gaining resistances against copper or zinc in nature together with the data suggesting that the concentrations necessary to maintain resistance plasmids within a population are well below the minimal inhibitory concentration (MIC) of the non-plasmid containing susceptible stain (Gullberg et al. 2014) might explain the prevalence of resistance mechanisms. Increasing metal contamination caused by anthropogenic metal use in agriculture and other sectors might also result in increased ability of protozoans to utilize copper and zinc to kill phagocytized bacteria. This, in turn, can cause a higher level of resistance in bacteria—a mechanism that has been primarily selected to avoid protozoan killing. In addition, copper is known to induce viable but non-culturable (VBNC) state in some bacteria, causing an increase in their survival rates (Ordax et al. 2006). It has been reported that cells of the VBNC state might have been responsible for the recent E. coli O104:H4 outbreak (Aurass et al. 2011).
It is worth noting here that copper-induced resistance combined with high toxicity and non-selectivity of redox processes induced by copper presence in cells present major challenges for developing copper-based antimicrobial therapy. Recently, Festa et al. have published a novel approach that allows accumulation of copper in pathogen cells without activating its Cu-resistant mechanisms and significantly increases selectivity of the treatment (Festa et al. 2014). In other strategies, Cu(II) ions have been either utilized as carriers for known antibiotics, allowing them to bypass existing efflux-mediated resistance to drugs (Manning et al. 2014; Lopes et al. 2013; Shams et al. 2014), or as chelators that upon binding to a ligand, change its conformation to the “active” mode (Haeili et al. 2014). Several potent copper chelators with activity against MRSA and Mycobacterium tuberculosis strains have been identified through drug screening assays specifically designed for identification of copper-dependent antimicrobial compounds (Speer et al. 2013) with, potentially, more discoveries on the way.
References
Aarestrup FM, Cavaco L, Hasman H (2010) Decreased susceptibility to zinc chloride is associated with methicillin resistant Staphylococcus aureus CC398 in Danish swine. Vet Microbiol 142:455–457. doi:10.1016/j.vetmic.2009.10.021
Aurass P, Prager R, Flieger A (2011) EHEC/EAEC O104:H4 strain linked with the 2011 German outbreak of haemolytic uremic syndrome enters into the viable but non-culturable state in response to various stresses and resuscitates upon stress relief. Environ Microbiol 13:3139–3148. doi:10.1111/j.1462-2920.2011.02604.x
Aviv G, Tsyba K, Steck N, Salmon-Divon M, Cornelius A, Rahav G, Grassl GA, Gal-Mor O (2014) A unique megaplasmid contributes to stress tolerance and pathogenicity of an emergent Salmonella enterica serovar Infantis strain. Environ Microbiol 16:977–994. doi:10.1111/1462-2920.12351
Baldan R, Testa F, Lore NI, Bragonzi A, Cichero P, Ossi C, Biancardi A, Nizzero P, Moro M, and Cirillo DM (2012) Factors contributing to epidemic MRSA clones replacement in a hospital setting. PLoS One, e43153. doi:10.1371/journal.pone.0043153
Baldan R, Rancoita PM, Di Serio C, Mazzotti M, Cichero P, Ossi C, Biancardi A, Nizzero P, Saracco A, Scarpellini P, Cirillo DM (2015) Epidemic MRSA clone ST22-IV is more resistant to multiple host- and environment-related stresses compared with ST228-I. J Antimicrob Chemother 70:757–765. doi:10.1093/jac/dku467
Bonis M, Ecobichon C, Guadagnini S, Prevost MC, Boneca IG (2010) A M23B family metallopeptidase of Helicobacter pylori required for cell shape, pole formation and virulence. Mol Microbiol 78:809–819. doi:10.1111/j.1365-2958.2010.07383.x
Braymer JJ, Giedroc DP (2014) Recent developments in copper and zinc homeostasis in bacterial pathogens. Curr Opin Chem Biol 19:59–66. doi:10.1016/j.cbpa.2013.12.021
Brown NL, Barrett SR, Camakaris J, Lee BT, Rouch DA (1995) Molecular genetics and transport analysis of the copper-resistance determinant (pco) from Escherichia coli plasmid pRJ1004. Mol Microbiol 17:1153–1166
Burlando B, Evangelisti V, Dondero F, Pons G, Camakaris J, Viarengo A (2002) Occurrence of Cu-ATPase in Dictyostelium: possible role in resistance to copper. Biochem Biophys Res Commun 291:476–483. doi:10.1006/bbrc.2002.6463
Cerasi M, Liu JZ, Ammendola S, Poe AJ, Petrarca P, Pesciaroli M, Pasquali P, Raffatellu M, Battistoni A (2014) The ZupT transporter plays an important role in zinc homeostasis and contributes to Salmonella enterica virulence. Metallomics 6:845–853. doi:10.1039/c3mt00352c
Chaturvedi KS, Hung CS, Giblin DE, Urushidani S, Austin AM, Dinauer MC, Henderson JP (2014) Cupric yersiniabactin is a virulence-associated superoxide dismutase mimic. ACS Chem Biol 9:551–561. doi:10.1021/cb400658k
Chen YT, Chang HY, Lai YC, Pan CC, Tsai SF, Peng HL (2004) Sequencing and analysis of the large virulence plasmid pLVPK of Klebsiella pneumoniae CG43. Gene 337:189–198. doi:10.1016/j.gene.2004.05.008
Cho K, Arimoto T, Igarashi T, Yamamoto M (2013) Involvement of lipoprotein PpiA of Streptococcus gordonii in evasion of phagocytosis by macrophages. Mol Oral Microbiol 28:379–391. doi:10.1111/omi.12031
Crossman LC, Chaudhuri RR, Beatson SA, Wells TJ, Desvaux M, Cunningham AF, Petty NK, Mahon V, Brinkley C, Hobman CL, Savarino SJ, Turner SM, Pallen MJ, Penn CW, Parkhill J, Turner AK, Johnson TJ, Thomson NR, Smith SG, Henderson IR (2010) A commensal gone bad: complete genome sequence of the prototypical enterotoxigenic Escherichia coli strain H10407. J Bacteriol 192:5822–5831. doi:10.1128/JB.00710-10
Dupont CL, Grass G, Rensing C (2011) Copper toxicity and the origin of bacterial resistance—new insights and applications. Metallomics 3:1109–1118. doi:10.1039/c1mt00107h
Festa RA, Thiele DJ (2012) Copper at the front line of the host-pathogen battle. PLoS Pathog 8:e1002887. doi:10.1371/journal.ppat.1002887
Festa RA, Helsel ME, Franz KJ, Thiele DJ (2014) Exploiting innate immune cell activation of a copper-dependent antimicrobial agent during infection. Chem Biol 21:977–987. doi:10.1016/j.chembiol.2014.06.009
Fodah RA, Scott JB, Tam HH, Yan P, Pfeffer TL, Bundschuh R, Warawa JM (2014) Correlation of Klebsiella pneumoniae comparative genetic analyses with virulence profiles in a murine respiratory disease model. PLoS One 9:e107394. doi:10.1371/journal.pone.0107394
Gehring AM, DeMoll E, Fetherston JD, Mori I, Mayhew GF, Blattner FR, Walsh CT, Perry RD (1998) Iron acquisition in plague: modular logic in enzymatic biogenesis of yersiniabactin by Yersinia pestis. Chem Biol 5:573–586
German N, Doyscher D, Rensing C (2013) Bacterial killing in macrophages and amoeba: do they all use a brass dagger? Future Microbiol 8:1257–1264. doi:10.2217/fmb.13.100
Gilmour MW, Thomson NR, Sanders M, Parkhill J, Taylor DE (2004) The complete nucleotide sequence of the resistance plasmid R478: defining the backbone components of incompatibility group H conjugative plasmids through comparative genomics. Plasmid 52:182–292
Gradassi M, Pesciaroli M, Martinelli N, Ruggeri J, Petrucci P, Hassan WH, Raffatellu M, Scaglione FE, Ammendola S, Battistoni A, Alborali GL, Pasquali P (2013) Attenuated Salmonella enterica serovar Typhimurium lacking the ZnuABC transporter: an efficacious orally-administered mucosal vaccine against salmonellosis in pigs. Vaccine 31:3695–3701. doi:10.1016/j.vaccine.2013.05.105
Gullberg E, Albrecht LM, Karlsson C, Sandegren L, Andersson DI (2014) Selection of a multidrug resistance plasmid by sublethal levels of antibiotics and heavy metals. MBio 5:e01918
Gupta A, Matsui K, Lo JF, Silver S (1999) Molecular basis for resistance to silver cations in Salmonella. Nat Med 5:183–188. doi:10.1038/5545
Haeili M, Moore C, Davis CJ, Cochran JB, Shah S, Shrestha TB, Zhang Y, Bossmann SH, Benjamin WH, Kutsch O, Wolschendorf F (2014) Copper complexation screen reveals compounds with potent antibiotic properties against methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother 58:3727–3736. doi:10.1128/AAC.02316-13
Hasman H (2005) The tcrB gene is part of the tcrYAZB operon conferring copper resistance in Enterococcus faecium and Enterococcus faecalis. Microbiology 151:3019–3025
Hobman JL, Crossman LC (2015) Bacterial antimicrobial metal ion resistance. J Med Microbiol. 64:471–497. doi:10.1099/jmm.0.023036-0
Hood MI, Skaar EP (2012) Nutritional immunity: transition metals at the pathogen-host interface. Nat Rev Microbiol 10:525–537. doi:10.1038/nrmicro2836
Johnson TJ, Siek KE, Johnson SJ, Nolan LK (2005) DNA sequence and comparative genomics of pAPEC-O2-R, an avian pathogenic Escherichia coli transmissible R plasmid. Antimicrob Agents Chemother 49:4681–4688
Kashyap DR, Rompca A, Gaballa A, Helmann JD, Chan J, Chang CJ, Hozo I, Gupta D, Dziarski R (2014) Peptidoglycan recognition proteins kill bacteria by inducing oxidative, thiol, and metal stress. PLoS Pathog 10:e1004280. doi:10.1371/journal.ppat.1004280
Kremer AN, Hoffmann H (2012) Subtractive hybridization yields a silver resistance determinant unique to nosocomial pathogens in the Enterobacter cloacae complex. J Clin Microbiol 50:3249–3257. doi:10.1128/JCM.00885-12
Loeffler A, McCarthy A, Lloyd DH, Musilova E, Pfeiffer DU, Lindsay JA (2013) Whole-genome comparison of meticillin-resistant Staphylococcus aureus CC22 SCCmecIV from people and their in-contact pets. Vet Dermatol 24:538–e128. doi:10.1111/vde.12062
Lopes SC, Ribeiro C, Gameiro P (2013) A new approach to counteract bacteria resistance: a comparative study between moxifloxacin and a new moxifloxacin derivative in different model systems of bacterial membrane. Chem Biol Drug Des 81:265–274. doi:10.1111/cbdd.12071
Lüthje FL, Hasman H, Aarestrup FM, Alwathnani HA, Rensing C (2014) Genome sequences of two copper-resistant Escherichia coli strains isolated from copper-fed pigs. Genome Announc 2:e01341. doi:10.1128/genomeA.01341-14
Macomber L, Imlay JA (2009) The iron-sulfur clusters of dehydratases are primary intracellular targets of copper toxicity. Proc Natl Acad Sci U S A 106:8344–8349. doi:10.1073/pnas.0812808106
Manning T, Mikula R, Lee H, Calvin A, Darrah J, Wylie G, Phillips D, Bythell BJ (2014) The copper (II) ion as a carrier for the antibiotic capreomycin against Mycobacterium tuberculosis. Bioorg Med Chem Lett 24:976–982. doi:10.1016/j.bmcl.2013.12.053
McHugh GL, Moellering RC, Hopkins CC, Swartz MN (1975) Salmonella typhimurium resistant to silver nitrate, chloramphenicol, and ampicillin. Lancet 1:235–240
Moreno Switt AI, den Bakker HC, Cummings CA, Rodriguez-Rivera LD, Govoni G, Raneiri ML, Degoricija L, Brown S, Hoelzer K, Peters JE, Bolchacova E, Furtado MR, Wiedmann M (2012) Identification and characterization of novel Salmonella mobile elements involved in the dissemination of genes linked to virulence and transmission. PLoS One 7:e41247. doi:10.1371/journal.pone.0041247
Mourão J, Novais C, Machado J, Peixe L, Antunes P (2015) Metal tolerance in emerging clinical relevant multidrug-resistant Salmonella enterica serotype 4,[5], 12:i:- clones circulating in Europe. Int J Antimicrob Agents 45:610–616. doi:10.1016/j.ijantimicag.2015.01.013
Neyrolles O, Wolschendorf F, Mitra A, Niederweis M (2015) Mycobacteria, metals, and the macrophage. Immunol Rev 264:249–263. doi:10.1111/imr.12265
Ong C-IY, Gillen CM, Walkera MJ, McEwan AG (2014) An antimicrobial role for zinc in innate immune defense against Group A Streptococcus. J Infect Dis 209:1500–1508. doi:10.1093/infdis/jiu053
Ordax M, Marco-Noales E, Lopez MM, Biosca EG (2006) Survival strategy of Erwinia amylovora against copper: induction of the viable-but-nonculturable state. Appl Environ Microbiol 72:3482–3488. doi:10.1128/AEM.72.5.3482-3488.2006
Pesciaroli M, Aloisio F, Ammendola S, Pistoia C, Petrucci P, Tarantino M, Francia M, Battistoni A, Pasquali P (2011) An attenuated Salmonella enterica serovar Typhimurium strain lacking the ZnuABC transporter induces protection in a mouse intestinal model of Salmonella infection. Vaccine 29:1783–1790. doi:10.1016/j.vaccine.2010.12.111
Peters JE, Fricker AD, Kapili BJ, Petassi MT (2014) Heteromeric transposase elements: generators of genomic islands across diverse bacteria. Mol Microbiol 93:1084–1092. doi:10.1111/mmi.12740
Qin Y, Hasman H, Aarestrup FM, Alwathnani HA, and Rensing C (2014) Genome sequences of three highly copper-resistant Salmonella enterica subsp. I serovar Typhimurium strains isolated from pigs in Denmark. Genome Announc 2. doi:10.1128/genomeA.01334-14
Rakin A, Schneider L, Podladchikova O (2012) Hunger for iron: the alternative siderophore iron scavenging systems in highly virulent Yersinia. Front Cell Infect Microbiol 2:151/1. doi:10.3389/fcimb.2012.00151
Randall CP, Oyama LB, Bostock JM, Chopra I, O’Neill AJ (2013) The silver cation (Ag+): antistaphylococcal activity, mode of action and resistance studies. J Antimicrob Chemother 68:131–138. doi:10.1093/jac/dks372
Randall CP, Gupta A, Jackson N, Busse D, O’Neill AJ (2015) Silver resistance in Gram-negative bacteria: a dissection of endogenous and exogenous mechanisms. J Antimicrob Chemother 70:1037–1046. doi:10.1093/jac/dku523
Reffuveille F, Serror P, Chevalier S, Budin-Verneuil A, Ladjouzi R, Bernay B, Auffray Y, Rince A (2012) The prolipoprotein diacylglyceryl transferase (Lgt) of Enterococcus faecalis contributes to virulence. Microbiology 158:816–825. doi:10.1099/mic.0.055319-0
Ren Y, Ren Y, Zhou Z, Guo X, Li Y, Feng L, Wang L (2010) Complete genome sequence of Enterobacter cloacae subsp. cloacae type strain ATCC 13047. J Bacteriol 192:2463–2464. doi:10.1128/JB.00067-10
Rensing C, Ghosh M, Rosen BP (1999) Families of soft-metal-ion-transporting ATPases. J Bacteriol 181:5891–5897
Rosenzweig AC, Arguello JM (2012) Toward a molecular understanding of metal transport by P(1B)-type ATPases. Curr Top Membr 69:113–136. doi:10.1016/B978-0-12-394390-3.00005-7
Sandegren L, Linkevicius M, Lytsy B, Melhus A, Andersson DI (2012) Transfer of an Escherichia coli ST131 multiresistance cassette has created a Klebsiella pneumoniae-specific plasmid associated with a major nosocomial outbreak. J Antimicrob Chemother 67:74–83. doi:10.1093/jac/dkr405
Schubert S, Dufke S, Sorsa J, Heesemann J (2004) A novel integrative and conjugative element (ICE) of Escherichia coli: the putative progenitor of the Yersinia high-pathogenicity island. Mol Microbiol 51:837–848. doi:10.1046/j.1365-2958.2003.03870.x
Shams S, Ali B, Afzal M, Kazmi I, Al-Abbasi FA, Anwar F (2014) Antifungal effect of gatifloxacin and copper ions combination. J Antibiot (Tokyo) 67:499–504. doi:10.1038/ja.2014.35
Slifierz MJ, Park J, Friendship RM, Weese JS (2014) Zinc-resistance gene CzrC identified in methicillin-resistant Staphylococcus hyicus isolated from pigs with exudative epidermitis. Can Vet J 55:489–490
Speer A, Shrestha TB, Bossmann SH, Basaraba RJ, Harber GJ, Michalek SM, Niederweis M, Kutsch O, Wolschendorf F (2013) Copper-boosting compounds: a novel concept for antimycobacterial drug discovery. Antimicrob Agents Chemother 57:1089–1091. doi:10.1128/AAC.01781-12
Subashchandrabose S, Hazen TH, Brumbaugh AR, Himpsl SD, Smith SN, Ernst RD, Rasko DA, Mobley HL (2014) Host-specific induction of Escherichia coli fitness genes during human urinary tract infection. Proc Natl Acad Sci U S A 111:18327–18332. doi:10.1073/pnas.1415959112
Wu KM, Li LH, Yan JJ, Tsao N, Liao TL, Tsai HC, Fung CP, Chen HJ, Liu YM, Wang JT, Fang CT, Chang SC, Shu HY, Liu TT, Chen YT, Shiau YR, Lauderdale TL, Su IJ, Kirby R, Tsai SF (2009) Genome sequencing and comparative analysis of Klebsiella pneumoniae NTUH-K2044, a strain causing liver abscess and meningitis. J Bacteriol 191:4492–4501. doi:10.1128/JB.00315-09
Xu FF, Imlay JA (2012) Silver(I), mercury(II), cadmium(II) and zinc(II) target exposed enzymic iron-sulfur clusters when they toxify Escherichia coli. Appl Env Microbiol 78:3614–3621. doi:10.1128/AEM.07368-11
Yu D, Pi B, Chen Y, Wang Y, Ruan Z, Otto M, Yu Y (2014) Characterization of the staphylococcal cassette chromosome composite island of Staphylococcus haemolyticus SH32, a methicillin-resistant clinical isolate from China. PLoS One 9:e87346. doi:10.1371/journal.pone.0087346
Zhang S, Wang D, Wang Y, Hasman H, Aerestrup FM, Alwathnani HA, Zhu YG, Rensing C (2015) Genome sequences of copper resistant and sensitive Enterococcus faecalis strains isolated from copper-fed pigs. Stand. Genomic Sci (in press)
Zimmermann M, Udagedara SR, Sze CM, Ryan TM, Howlett GJ, Xiao Z, Wedd AG (2012) PcoE - A metal sponge expressed to the periplasm of copper resistance Escherichia coli. Implication of its function role in copper resistance. J Inorg Biochem 115:186–197. doi:10.1016/j.jinorgbio.2012.04.009
Acknowledgments
This work was supported in part by the Center for Environmental and Agricultural Microbiology (CREAM) funded by the Villum Kann Rasmussen Foundation.
Conflict of interest
The authors declare that they have no competing interests.
Compliance with ethical standards
This article does not contain any studies with human participants or animals performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hao, X., Lüthje, F.L., Qin, Y. et al. Survival in amoeba—a major selection pressure on the presence of bacterial copper and zinc resistance determinants? Identification of a “copper pathogenicity island”. Appl Microbiol Biotechnol 99, 5817–5824 (2015). https://doi.org/10.1007/s00253-015-6749-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6749-0