Abstract
Bacillus subtilis and its closely related species are important strains for industry, agriculture, and medicine. However, it is difficult to perform genetic manipulations using the endogenous recombination machinery. In many bacteria, phage recombineering systems have been employed to improve recombineering frequencies. To date, an efficient phage recombineering system for B. subtilis has not been reported. Here, we, for the first time, identified that GP35 from the native phage SPP1 exhibited a high recombination activity in B. subtilis. On this basis, we developed a high-efficiency GP35-meditated recombineering system. Taking single-stranded DNA (ssDNA) as a recombineering substrate, ten recombinases from diverse sources were investigated in B. subtilis W168. GP35 showed the highest recombineering frequency (1.71 ± 0.15 × 10−1). Besides targeting the purine nucleoside phosphorylase gene (deoD), we also demonstrated the utility of GP35 and Beta from Escherichia coli lambda phage by deleting the alpha-amylase gene (amyE) and uracil phosphoribosyltransferase gene (upp). In all three genetic loci, GP35 exhibited a higher frequency than Beta. Moreover, a phylogenetic tree comparing the kinship of different recombinase hosts with B. subtilis was constructed, and the relationship between the recombineering frequency and the kinship of the host was further analyzed. The results suggested that closer kinship to B. subtilis resulted in higher frequency in B. subtilis. In conclusion, the recombinase from native phage or prophage can significantly promote the genetic recombineering frequency in its host, providing an effective genetic tool for constructing genetically engineered strains and investigating bacterial physiology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Recombineering is an efficient homologous recombination (HR)-based method for in vivo genetic engineering, which allows the precise insertion, deletion, or alteration of any DNA sequence (Sharan et al. 2009). However, HR is a rare event in most organisms and the desired recombinant is often difficult to obtain using the endogenous recombination machinery (e.g., mediated by RecA). To solve this problem, phage recombination proteins have been used to improve recombineering frequency (Zhang et al. 1998; Muyrers et al. 1999; Ellis et al. 2001; Jiang et al. 2013), though researchers have found that the efficiency of the same recombinase varies greatly in different bacteria (van Kessel and Hatfull 2008). Therefore, seeking the most suitable recombinase for each bacterium is very important.
Recombineering using recombination proteins is a powerful system for gene inactivation in many bacteria. In Escherichia coli and some species of Salmonella and Shigella (Court et al. 2002; Datta et al. 2006; Maresca et al. 2010; Mosberg et al. 2010; Ranallo et al. 2006), recombineering systems encoded by the λ Red system or the recE and recT genes of the Rac prophage greatly enhance the HR frequency. The mycobacterial phage-encoded protein GP60 was developed for mycobacterial recombineering (van Kessel et al. 2008). In addition, a recent study showed that HR also occurs in lactic acid bacteria when mediated by the native recombinase RecT1 from Lactobacillus reuteri (van Pijkeren and Britton 2012).
Bacillus subtilis, as a model organism, is an important cell factory for the high-level production of enzymes for many industries, antibiotics, insecticides, etc. (Deng et al. 2010). Although the genetic manipulation could be accomplished using natural transformation, the preparation of efficient competent cells is difficult for some Bacillus strains (Nijland et al. 2010). Electroporation is a convenient and universal method, and the transformation efficiency of Bacillus is generally about 105 CFU/μg DNA (Brigidi et al. 1990; Stephenson and Jarrett 1991; Xue et al. 1999) which is similar to other Gram-positive bacteria, such as Mycobacterium tuberculosis (van Kessel and Hatfull 2007) and Staphylococcus aureus (Schenk and Laddaga 1992), but it is not efficient enough for gene manipulation based on the endogenous recombination machinery (Bashkirov et al. 1987; Hofemeister et al. 1983). In order to solve this problem, some recombinases have been investigated to improve the frequency of HR in B. subtilis. A few studies have accomplished genome editing in B. subtilis by taking advantage of an exogenous recombineering system, e.g., the Cre system and λ Red proteins, coupled with electroporation (Wang et al. 2012; Yan et al. 2008). However, the E. coli-based systems appear to be less efficient in distantly related bacteria, especially B. subtilis, which belongs to the class of Gram-positive bacteria (van Kessel and Hatfull 2007). It has been reported that the protein GP35 from the B. subtilis bacteriophage SPP1 shows homologous-pairing activity (Ayora et al. 2002) but exhibits a low recombination activity in E. coli (Datta et al. 2008). However, whether it exhibits recombination activity in B. subtilis has not been reported.
Aside from the recombinase, the construct of transformation substrate is another key factor affecting recombineering frequency. Wen et al. compared the recombineering frequencies of six different DNA constructs in B. subtilis (Wen et al. 2013), and ssDNA exhibited the highest recombineering frequency. Wang et al. successfully applied lambda Beta protein to improve the short homology-directed HR in B. subtilis using ssDNA (Wang et al. 2012). However, the length of the homologous arm was only 70 bp, leading to a low recombineering frequency about 10−3 (104 recombinants/107 transformants). Although allelic exchange events could be recovered with short homologous segments (50 bp) in Mycobacterium smegmatis, the greater recombineering frequency obtained with larger target homologous segments (500 bp) suggests that substrates with longer homologous regions are preferred (van Kessel and Hatfull 2007). Although the ssDNA synthesis technology has been refined, the synthesis of ssDNA longer than 59 nt is very expensive. Furthermore, higher recombineering frequency can be obtained with longer homologous arms. Therefore, the low-cost preparation of long homologous regions of ssDNA via PCR would be the preferred method.
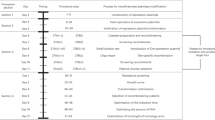
In this study, ten exogenous recombinase genes from diverse phages were expressed under the control of a xylose-inducible promoter, and their recombineering frequencies were assayed. For the first time, we identified that GP35 from the native phage SPP1 exhibited a high recombination activity in B. subtilis and showed the highest recombineering frequency in B. subtilis when compared with the other nine recombinases. Phylogenetic analysis based on the 16S ribosomal RNA (rRNA) gene sequence of ten recombinase hosts showed that GP35 functions significantly better than other distantly related recombinases in B. subtilis (the flowchart of this work is shown in Fig. 1).
A schematic illustration of the work performed in this paper. a A screen for the optimal field strength for the electroporation transformation of B. subtilis. b A screen for an appropriate promoter to express and identify the recombinase in B. subtilis. c Analysis of the effects of different DNA constructs on the recombineering frequency. d Recombineering with ssDNA mediated by different recombinases
Materials and methods
Bacterial strains, plasmids, and primers
The bacterial strains and plasmids used in this study are listed in Table S1. The primer synthesis (Table S2) and DNA sequencing were performed by Invitrogen (Shanghai, China) and BGI (Beijing, China). DNA manipulation and E. coli transformation were performed using standard techniques (Sambrook et al. 1989). The restriction enzymes, T4 ligases, and DNA markers were purchased from Takara (Dalian, China). The PCR amplicons used for cloning purposes were generated with the KOD DNA hot start polymerase (Toyobo Co., Ltd, Japan), whereas the PCR reactions used for screening purposes were performed using the Taq DNA polymerase. All of the recombinase genes that are seen in phages or prophages of B. subtilis (gp35, accession number X97918, nucleotide 32175–33038), M. smegmatis (gp61, accession number AY129333, nucleotide 43643–44704), Listeria monocytogenes (orf48, accession number AJ242593, nucleotide 32773–33588), Lactococcus lactis (orf245, accession number AF212847, nucleotide 1678–2415), S. aureus (gp20, accession number NC_008617, nucleotide 10317–11237), Vibrio cholerae (s065, accession number AY055428, nucleotide 72817–73635), Photorhabdus luminescens (plu2935, accession number BX571868, nucleotide 324693–325613), Legionella pneumophila (orfC, accession number AJ277755, nucleotide 1415–2299), and E. coli (beta and recT, accession number NC_001416 and NC_000913, nucleotide 32025–32810 and 1412008–1412817) were retrieved from the GenBank database. Taken the chromosomes of SIMD40–43 and SIMD44–50 as templates (Datta et al. 2008), all of the recombinase and recombinase-gfpmut3a fusion genes were PCR amplified using primers as listed in the Table S2 and then digested with BamHI, with the exception of gp61 (digested with SacII). The purified DNA fragment was finally inserted into the pAX01 plasmid digested with the same restriction enzymes to generate pAX01-recombinase (-gfp) vector (Fig. 2d). The restriction sites are underlined in the primer sequences as shown in Table S2. All of the exogenous genes were integrated into the lacA locus of the B. subtilis chromosome by natural competence transformation (Anagnostopoulos and Spizizen 1961), and the protein expression was induced using 1 % xylose. In addition, the gp35 gene was PCR amplified using primers WB789 and WB790 as listed in the Table S2 and then digested with BamHI and SmaI. The purified DNA fragment was finally inserted into the pHCMC04 plasmid digested with the same restriction enzymes to generate pWYE836. Then, the plasmid pWYE836 was electro-transformed into the B. subtilis ATCC 6633 (Duitman et al. 1999) to generate WYB263.
Expression and identification of the recombinase. a A real-time (RT) qPCR assay of the transcription levels. The relative fold changes of the transcriptional levels of gp35 were analyzed by quantitative real-time PCR with the P spac promoter. The relative mRNA level represents the transcription level of gp35 in cells controlled by different promoters. The data shown are mean values from three biological replicates and three technical replicates with their standard deviations. Different letters indicate significant differences among different recombinases (Duncan’s multiple range test, p < 0.05). b Western blot assay for the expression of GP35 controlled by three different promoters: P 43 , a strong constitutive promoter from B. subtilis W168; P xylA , a xylose-inducible promoter (+xylose, with 1 % xylose; -xylose, without xylose); P spac , an IPTG-inducible promoter (+IPTG, with 1 % IPTG; −IPTG, without IPTG). ‘−’, negative control; ‘+’, positive control. c GP35-mediated recombination using ssDNA. BS060 was a GP35-mediated recombineering strain, and W168 was used as a control. All of the experiments were carried out in triplicate. Different letters indicate significant differences among different recombinases (Duncan’s multiple range test, p < 0.05). Recombineering frequency is expressed as the number of recombinants per microgram/cell competency using deoD ssDNA as a recombineering substrate. d The plasmids used for the construction of the genetic engineering strains. Various recombinase genes (gp35, beta, s65, recT, plu2935, Orf48, Orf245, OrfC, gp61, and gp20) were cloned into pAX01, which has a xylose-inducible expression promoter (P xylA ), and were integrated into the B. subtilis chromosome at the lacA locus. T and T1T2 mean transcriptional terminators
Culture and growth conditions
E. coli was routinely cultured in Luria-Bertani (LB) liquid medium at 200 rpm or on LB agar plates at 37 °C. The B. subtilis strains were cultured in LB liquid medium at 220 rpm or on LB agar plates at 30 °C. When appropriate, ampicillin (100 μg/ml for E. coli), kanamycin (10 μg/ml for B. subtilis), chloramphenicol (5 μg/ml for B. subtilis), or MLS (0.5 μg/ml erythromycin plus 12.5 μg/ml lincomycin for B. subtilis) was added to the medium.
Construction of the integration plasmids for the expression of GP35 with different promoters
Multiple genes were rapidly assembled by taking advantage of the high DNA recombination activity in Saccharomyces cerevisiae (Shao et al. 2009). The genes were amplified using PCR primers that contained 50 bp of overlapping sequence to the adjacent gene from their corresponding pWYE724 plasmids (Zhang et al. 2012). S. cerevisiae DAY414 was transformed with the DNA fragments, including mutS upstream and downstream of B. subtilis W168, different promoters (P 43 , a strong constitutive promoter from B. subtilis (Wang et al. 2004; Wang and Doi 1984), P xylA and P spac ), the gp34.1-35-36 genes, the chloromycetin gene, and the pWYE724 plasmid linearized at the EcoRI and SalI loci. The S. cerevisiae DAY414 was then selected for tryptophan autotrophy on synthetic complete (SC) medium lacking tryptophan. The S. cerevisiae transformation was performed using the lithium acetate method (Gietz and Woods 2002). The plasmids were rescued into E. coli TOP10 cells as described by Robzyk and Kassir (1992). All of the recombinant plasmids were verified by restriction digestion and DNA sequencing before any subsequent use. The plasmids carrying the different promoters, P 43 , P xylA , and P spac , were named pWYE597, pWYE598, and pWYE599, respectively. Then, the three plasmids were transformed into W168 using the natural competence transformation method (Anagnostopoulos and Spizizen 1961) to generate BS045, BS047, and BS046, respectively.
Electroporation and recombineering in B. subtilis
The electroporation of B. subtilis was carried out according to the method described by Xue et al. (1999) and Zhang et al. (2011) with minor modifications. An overnight LB culture of the B. subtilis cells was diluted 16-fold into fresh growth medium (GM, LB plus 0.5 M sorbitol) and shaken at 30 °C. The cultures were supplemented with 1 % xylose to express the recombinases genes when the optical density (OD)600 reached 0.25–0.35. Upon reaching an OD600 of 0.85–0.95, the culture was supplemented with dl-threonine, glycine, and Tween 80 at final concentrations of 1.0, 2.0, and 0.03 %, respectively, and was shaken for 1 h. The cells were cooled on ice at 4 °C and then centrifuged at 5000×g for 10 min. Following four washes in ice-cold electroporation medium (0.5 M sorbitol, 0.5 M mannitol, and 10 % v/v glycerol), the cells were suspended in 1/40 (v/v) electroporation medium. If necessary, the competent cells were stored at −80 °C for future use. For the electroporation, 100 μl of the competent cells was mixed with 50 ng ssDNA or 100 ng pUB110 and then transferred to an ice-cold electroporation cuvette (1-mm electrode gap). After a brief incubation on ice, the cell-DNA mixture was shocked by a single 18 kV/cm pulse generated by an BTX ECM399 electroporator (BTX, Harvard Apparatus, Holliston, MA, USA) with a time constant of 5–5.8 ms. The cells were immediately diluted into 1 ml of recovery medium (GM plus 0.38 M mannitol), shaken gently at 30 °C for 4 h to allow expression of the antibiotic-resistant genes, and aliquots of the dilutions were then spread onto LB agar plates supplemented with the appropriate antibiotics. The transformation efficiencies were calculated by counting the colonies on the plates after incubation at 30 °C for 16 h. The competence of cells was determined by transformation with 100 ng of pUB110 and expressed as colony-forming units per microgram DNA (CFU/μg DNA). Recombineering frequency was expressed as the number of recombinants per microgram/cell competency (van Kessel and Hatfull 2007). Some of the transformants were verified by plasmid extraction and restriction enzyme digestion.
Recombineering substrates
The recombineering substrates were generated by PCR amplification using the KOD plus according to the method described by Wen et al. (Wen et al. 2013). The sequences of all of the primers used in this study are shown in Table S2. The fragment size and integrity of all of the PCR-generated substrates were confirmed by agarose gel electrophoresis. We designed three cassettes (deoD::kan R, upp::kan R, and amyE::Cm R), consisting of a kanamycin resistance gene (kan R) flanked by 500-bp regions homologous to the deoD gene coding purine nucleoside phosphorylase and upp gene coding uracil phosphoribosyltransferase and a chloramphenicol resistance gene (Cm R) flanked by 500-bp regions homologous to the amyE gene coding alpha-amylase on the B. subtilis chromosome. The recombineering substrate of deoD was generated using pWYE753 as a PCR template, the upp and amyE gene using upp::kan R and amyE::Cm R PCR products as the templates for ssDNA production (defined as deoD ssDNA, upp ssDNA, and amyE ssDNA, respectively). The ssDNA of the PCR products was generated according to the method established by Tang et al. (Tang et al. 2006) except for the following modification: 100 pg of the PCR product amplified in 50 μl of the PCR solution was used as the template for the generation of the ssDNA in the same PCR system containing one primer. The cycling program was 94 °C for 5 min for the DNA denaturation, followed by 30 cycles of 94 °C for 30 s, 54 °C for 30 s, and 72 °C for 1 min (deoD ssDNA gel electrophoresis was shown in Fig. S2). The ssDNA was purified using the TIANquick Midi Purification Kit (Tiangen, China). All of the constructs that were used as templates to produce the recombineering substrates were confirmed by restriction digestion and/or sequencing.
RNA preparation and quantitative real-time RT-PCR
B. subtilis BS045, BS046, and BS047 were cultivated in GM, and the BS046 and BS047 were supplemented with 1 % xylose and 1 mM IPTG, respectively, when the OD600 reached 0.25–0.35. The cells were harvested when the OD600 reached 0.85–0.95, and the total RNA was isolated using the RNAprep Pure Cell/Bacteria Kit (Tiangen, China). The reverse transcription (RT) of approximately 200 ng of RNA was performed with the specific primers listed in Table S2 in the supplemental material using the FastQuant RT Kit (Tiangen, China). The quantitative PCR was performed with Bryt Green from the GoTaq qPCR master mix (Promega, USA) and the Rotor-Gene Q Real-Time PCR Detection System (Qiagen, Germany). The B. subtilis 16S rRNA gene was used as the reference to normalize the gp35 mRNA levels. Negative controls were used in each PCR run to exclude DNA and other contamination. The quantitative PCR (qPCR) products were verified with a melting curve analysis. The data collection and analysis were facilitated by the Rotor-Gene Q Series software, version 2.0.3, according to the 2−ΔΔCT method (Schmittgen and Livak 2008).
Western blot analysis
The cells were harvested as described in the “RNA preparation and quantitative real-time RT-PCR” section, washed with phosphate-buffered saline (PBS, pH 7.2), and resuspended in the same buffer. The W168 and purified GP35 proteins were used as negative and positive controls, respectively. The protein concentration was determined with NanoDrop 2000C spectrophotometer (Thermo, USA). Samples from different culture conditions were analyzed by SDS-polyacrylamide gel electrophoresis. The proteins were electro-transferred to a PVDF membrane and probed with rabbit polyclonal antibodies raised against a synthetic peptide analogue of the GP35 protein. The blots were visualized with a peroxidase-coupled goat anti-rabbit secondary antibody and the ECL color development reagent (GE, USA).
Flow cytometry analysis
The gfpmut3a gene, encoding green fluorescent protein (GFP), which has a more intense fluorescence, was used as a reporter gene to investigate the expression level of different recombinases. The gfpmut3a gene from the pAD43-25 vector was fused to the recombinase genes using SOE-PCR and then inserted into the pAX01 plasmid. The flow cytometry was performed on a BD FACSCalibur flow cytometer equipped with an argon laser. The cells were diluted to an OD600 of 0.4 using PBS buffer and placed on ice prior to analysis. For each sample, 20,000 events were collected. The cells from W168 were used as a control to determine the background fluorescence.
Phylogenetic analysis
The 16S rRNA gene sequences from different recombinase phage hosts were obtained from the National Center for Biotechnology Information database. Gene sequence alignments were performed with the ClustalX program, and a phylogenetic tree was constructed using MEGA 5.1 software according to the neighbor-joining method (Tamura et al. 2011).
Results
Determination of the suitable transformation level by optimizing field strength
In previous studies, different field strengths have been used for the transformation of B. subtilis (Stephenson and Jarrett 1991; Wang et al. 2012; Xue et al. 1999). To optimize the transformation conditions, the field strengths ranging from 12 to 21 kV/cm were applied when transforming the B. subtilis W168 with the pUB110 replicative plasmid. As shown in Figs. 1a and S1, the transformation efficiency of 2.36 ± 0.35 × 105 CFU/μg DNA was obtained at the field strength of 18 kV/cm which was used for the following study.
The expression and recombination activity of SPP1 GP35 in B. subtilis
In order to select an appropriate expression system for the recombinase GP35, three promoters (P 43 , P xylA , and P spac ) with different strengths were used for controlling the gp35 gene expression in B. subtilis. The quantitative real-time RT-PCR and Western blotting showed that the highest expression level of GP35 was obtained under the control of the P 43 promoter, but it had a negative effect on cell growth (data not shown). The expression of gp35 was maintained at a moderate level when under the control of P xylA , while the expression was maintained at a low level under the control of P spac promoter (Figs. 1b and 2a, b). Therefore, P xylA was chosen to control the GP35 expression.
To test whether GP35 could work in B. subtilis, the ssDNA was transformed into BS060 (the gp35 gene was integrated into the lacA locus of W168 under the control of P xylA ) and W168. As shown in Fig. 2c, when compared to W168, the recombineering frequency (1.71 ± 0.15 × 10−1) mediated by GP35 was significantly improved. This suggests that GP35 can undergo high recombination activity in the phage SPP1 host.
The effect of different constructs of exogenous DNA on the recombineering frequency
To investigate the effects of different DNA constructs, strand bias, or phosphorothioation on recombineering frequency, linear double-stranded DNA (dsDNA) constructs and leading- and lagging-strand annealing ssDNA substrates with or without phosphorothioation were transformed into B. subtilis W168. The lagging-strand targeting ssDNA construct was found to show significantly higher recombineering frequency than the other forms of DNA (Figs. 1c and 3).
The effect of various constructs on the recombineering substrates. Eight different gene replacement DNA cassettes (kan R) were tested to determine their recombineering frequency. 1, unmodified dsDNA; 2, leading-targeting phosphorothioated dsDNA; 3, lagging-targeting phosphorothioated dsDNA; 4, dsDNA with both strands phosphorothioated; 5, leading-targeting ssDNA; 6, lagging-targeting ssDNA; 7, phosphorothioated leading-targeting ssDNA; 8, phosphorothioated lagging-targeting ssDNA. All of the experiments were carried out in triplicate. Different letters indicate significant differences among different recombinases (Duncan’s multiple range test, p < 0.05). Recombineering frequency is expressed as the number of recombinants per microgram/cell competency using deoD ssDNA as a recombineering substrate
The data on the leading- and lagging-strand ssDNA recombineering demonstrated that the recombineering frequency of the lagging-strand ssDNA was higher than the leading-strand ssDNA (Fig. 3), which agreed with previous observations on the strand bias of ssDNA-mediated λ Red HR (Wang et al. 2012) due to the greater accessibility of the lagging strand for annealing (Li et al. 2003). The leading- and lagging-strand ssDNA constructs were subjected to phosphorothioation with four consecutive phosphorothioate linkages at the 5′ ends, to test the effect of strand protection on the recombineering frequency, because the phosphorothioate bond has been shown to protect DNA from exonuclease degradation while being permissive of the subsequent recombination steps (Mosberg et al. 2010; Maresca et al 2010; van Kessel et al. 2008). Therefore, the recombineering frequency of the lagging-strand targeting ssDNA construct was significantly improved with the phosphorothioate modifications, with improvements up to 8.41 ± 0.95 × 10−3. In addition, no obvious effect on the recombineering frequency was observed with the 5′-phosphorothioation modification of the leading-strand targeting ssDNA or the dsDNA construct (Fig. 3). Based on these observations, we could conclude that the phosphorothioated lagging-strand targeting ssDNA construct was readily transformable and highly recombinogenic in B. subtilis and was used for the following study.
Expression of the different recombinases in B. subtilis
To assess the expression levels of different recombinase genes, gfpmut3a was fused to the recombinase gene and used as a reporter gene. When compared to the control, all of the recombinases were capable of being expressed in B. subtilis (Fig. 4a). The expression levels of all of the recombinases were divided into two groups. The first group had low expression levels, including GP35, Beta, S65, RecT, and Plu2935, and their fluorescence intensity ranged from 13.65 to 16.79 (Fig. S3b). The second group, which included Orf48, Orf245, OrfC, GP61, and GP20, showed high expression levels. The expression level of OrfC was the highest, with a fluorescence intensity of 64.21 (Fig. S3b).
ssDNA recombineering using different recombinases. a Flow cytometry analysis of the expression of the recombinase-GFP fusion protein. One group, which includes GP35, Beta, S65, RecT, and Plu2935, showed low expression levels, while the other group, which includes Orf48, Orf245, OrfC, GP61, and GP20, showed high expression levels. W168 was used as a negative control. b Recombinases were assayed for ssDNA recombineering ability in B. subtilis by their ability to introduce kanamycin resistance. The recombinase gene expression was controlled by a xylose-inducible promoter and integrated into the lacA locus. The recombineering frequencies obtained for each recombinase are shown. All of the experiments were carried out in triplicate. Different letters indicate significant differences among different recombinases (Duncan’s multiple range test, p < 0.05). Recombineering frequency is expressed as the number of recombinants per microgram/cell competency using deoD ssDNA as a recombineering substrate
The ssDNA recombineering mediated by different recombinases
All of the recombinase genes were separately introduced into the B. subtilis W168 chromosome to create BS056-064 and BS068 (Table S1). As shown in Figs. 1d and 4b, the recombination mediated by GP35 from the phage SPP1 (in BS060) generated 1.71 ± 0.15 × 10−1. Orf48 from the A118 phage of L. monocytogenes also showed similar frequency (1.36 ± 0.16 × 10−1) to GP35. GP20 from S. aureus, Orf245 from the L. lactis phage, and GP61 from the M. smegmatis phage had moderate recombineering frequency (8.81 ± 1.57 × 10−2, 6.70 ± 0.92 × 10−2, and 6.05 ± 0.83 × 10−2, respectively), which was lower than GP35 and Orf48. When compared to the above enzymes from Gram-positive bacterial phages, the recombinases from the Gram-negative bacterial phages, including Beta, S065, RecT, Plu2935, and OrfC, showed low recombineering frequency (3.07 ± 0.18 × 10−2, 2.50 ± 0.10 × 10−2, 2.11 ± 0.14 × 10−2, 1.41 ± 0.24 × 10−2 and 1.18 ± 0.27 × 10−2, respectively). GP35 showed a 4.6-fold higher frequency than Beta. Under identical experimental conditions, OrfC from L. pneumophila and Plu2935 from P. luminescens were relatively inefficient, as they could only generate similar frequency to the wild-type W168 (0.92 ± 0.08 × 10−2).
Deletion of the amyE and upp genes mediated by GP35 and Beta
To evaluate differences of recombineering frequency from the same or different recombinases in different genetic loci, we performed the recombineering mediated by GP35 and Beta via inactivating the amyE and upp genes in B. subtilis. The left and right flanking homologous arms (500 bp) were amplified from the upstream and downstream regions of the target gene and then assembled with the resistance gene via fusion PCR.
The PCR and sequencing analyses revealed the successful replacement of the upp gene with the kanamycin resistance gene with a recombineering frequency of 4.17 ± 0.26 × 10−2 (GP35) vs. 2.98 ± 0.62 × 10−3 (Beta), and GP35 showed a 14.0-fold higher frequency than Beta (Fig. 5a, b). Uracil phosphoribosyltransferase converts 5-fluorouracil (5-FU) to 5-fluoro-UMP, which is ultimately metabolized to the toxic compound 5-fluoro-dUMP and is capable of inhibiting the activity of thymidylate synthetase. The upp/5-FU module has been widely used in many bacterial species for the deletion of genes without introducing antibiotic resistance markers (Fabret et al. 2002). In contrast with the BS060 strain, the BS069 strain was able to grow on minimal medium (MM) supplemented with 5-FU (Fig. 5d).
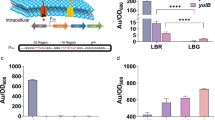
Application of the ssDNA recombineering system mediated by GP35. a Recombineering mediated by GP35 and Beta in different loci. Asterisks indicate significant difference (pairwise Mann-Whitney U test, p < 0.05). b PCR confirmation of the knockout of the upp gene using the WY031 and WY034 primers. BS069 was acquired by transforming ssDNA into BS060. c PCR confirmation of the knockout of the amyE gene using the WB1177 and WB1224 primers. BS070 was acquired by transforming ssDNA into BS060. d Verification of the loss of uracil phosphoribosyltransferase activity in B. subtilis strain BS069. Strain BS069 was capable of growing on MM plates containing 5 mM 5-FU, but the BS060 strain was unable to grow. MM was used as a control. e Verification of the amyE knockout using a starch plate assay. The left image (before staining with iodine) shows a comparison of the colony size between BS060 and the mutant BS070, with an integrated kan-cassette at the amyE locus. The right image shows the same plate after iodine fumigation. The loss of the transparent halo indicates the loss of alpha-amylase activity
Similarly, the BS070 with the disrupted amyE gene lost the ability to degrade amylose and had a high recombineering frequency of 9.86 ± 0.46 × 10−2 (GP35) vs. 6.12 ± 0.84 × 10−3 (Beta), and GP35 showed a 16.1-fold higher frequency than Beta (Fig. 5a). The PCR and sequencing analyses revealed the successful replacement of the amyE gene with the chloramphenicol resistance gene (Fig. 5c). The occurrence of the amyE gene replacement event was examined using a starch plate assay showing the absence of a clear area around the mutant colony (Fig. 5e).
Recombineering mediated by GP35 in B. subtilis with poor natural competence
Compared to other strains of the genus Bacillus, 168 derivative strains possess high natural competence. To enlarge the application of the recombineering system mediated by GP35 in other Bacillus strains, the upp ssDNA was electro-transformed into an originally poor competence strain B. subtilis ATCC 6633 (Fig. S4a) and a high recombineering frequency (0.92 ± 0.15 × 10−1) was also acquired (Fig. S4b). The recombineering frequency of ATCC 6633/pHCMC04-gp35 was increased more than 40-fold compared to that of wild type (2.30 ± 0.80 × 10−3). The result suggested that the GP35 could also perform high-efficiency recombination in Bacillus strains with poor natural competence.
Recombinases perform better in their native hosts
An amino acid sequence alignment of all of the recombinases showed the greatest similarity between RecT and GP35, but its recombineering frequency was noticeably lower than that of GP35 and some of the other recombinases (Fig. S3a). Therefore, the recombineering frequency was not related to the homology levels of the various recombinases. Moreover, there was no direct relationship between the recombineering frequency and the expression level: high expression level, such as OrfC, resulted in low frequency and low expression level resulted in high frequency, as was observed in GP35 (Fig. S3b). There may be two possible scenarios if a threshold exists (Deans et al. 2007). In one scenario, the excess expression is not involved in the recombination processes when the expression reaches the threshold. In the second scenario, if the expression level does not reach the threshold, the activities of the recombinases can be denoted by the recombineering frequency/amount of recombinase.
The phylogenetic tree analysis showed that the homologous proteins from different bacterial phages could be classified into two groups (Fig. 6a). One group of recombinases derived from Gram-positive bacterial phages showed high recombineering frequency, while another group of recombinases derived from Gram-negative bacterial phages showed low frequency. Alignment of the 16S rRNA gene sequences of the ten recombinase phage hosts demonstrated that the recombinase with the greatest similarity to GP35 acquired a higher recombineering frequency (Fig. 6b). Among all of the recombinases, the highest recombineering frequency in B. subtilis was produced by GP35, which comes from the native Bacillus phage SPP1. Therefore, the recombineering frequency of various recombinases in B. subtilis may be closely related to the phylogenetic relationship of the phage hosts.
Recombinases from close relatives show greater recombineering frequency in the target bacteria. a A phylogenetic tree of the different recombinase phage hosts. The 16S rRNA gene sequences from the ten recombinase phage hosts were obtained from GenBank. The bootstrap values above 50 % are indicated at the nodes and represent the percentages of 100 replicates. The scale bar indicates a 1 % fixed point mutation per nucleotide base. b The relationship between the recombineering frequency of the recombinases and the kinship of the phage hosts. The 16S rRNA gene sequence from B. subtilis W168 was used as a standard, and the 16S rRNA gene sequences from the nine other recombinase phage hosts were aligned with W168. Recombineering frequency is expressed as the number of recombinants per microgram/cell competency using deoD ssDNA as a recombineering substrate
Discussion
Recently, phage recombineering systems have been employed to improve the recombineering frequency in many bacteria (Zhang et al. 1998; Muyrers et al. 1999; Ellis et al. 2001; Jiang et al. 2013), but an efficient B. subtilis phage recombineering system has not yet been reported. In this study, we firstly identified the high recombination activity of GP35 in vivo and found that it is an efficient recombinase for recombineering using ssDNA in B. subtilis. By comparing the recombineering frequencies of ten recombinases from both Gram-negative and Gram-positive bacteria and their phages, we found that GP35 from the native phage SPP1 showed the highest frequency in B. subtilis. On this basis, we developed a high-efficiency GP35-mediated recombineering system using PCR-based ssDNA.
Although the transformation efficiency of some Bacillus strains could achieve 107 CFU/μg DNA (Wang et al. 2012; Zhang et al. 2011), electroporation of B. subtilis W168 and most other Bacillus strains usually produces an efficiency of 105 CFU/μg DNA (Brigidi et al. 1990; Stephenson and Jarrett 1991; Xue et al. 1999). So, the system was developed based on the transformation efficiency of 105 CFU/μg DNA, and the transformation efficiency was achieved only by optimizing the field strength (Figs. 1a and S1). The efficiency could also be achieved easily and reproducibly for many Gram-positive bacteria (Schenk and Laddaga 1992; van Kessel and Hatfull 2007) including other Bacillus strains through simple optimization in most laboratories. Therefore, the recombineering system in this study suggests a universal application.
Using deoD ssDNA as a substrate, GP35 showed the highest recombineering frequency in B. subtilis when compared with nine other recombinases from various phages. To investigate the effect of different genetic loci on recombineering frequency, we also compared the recombineering frequency in amyE and upp loci mediated by GP35 and Beta, which has been widely employed to promote recombineering frequency in many bacteria including B. subtilis (Wang et al. 2012). The result in Fig. 5a showed that different recombineering frequencies existed in different loci mediated by the same recombinase. We speculated that this may be caused by different spatial structures of different genetic loci. Furthermore, in all three loci, Beta exhibited a lower frequency than GP35. This may be attributed to its interactions with the host factors (Datta et al. 2008). We also performed the recombineering mediated by GP35 and Beta at 37 °C, but there were no differences of recombineering frequencies between 37 and 30 °C (data not shown). Therefore, the recombineering system mediated by GP35 performs better than that mediated by Beta in B. subtilis at both 37 and 30 °C.
In addition, the same genotype strain to BS060 (W168△lacA::P xylA -gp35-MLS R) was also obtained by integrating the gp35 gene into the genome of B. subtilis W168 using electro-transformation and showed a similar recombineering frequency to that by natural competence transformation (data not shown). In Bacillus genus, many strains, such as B. subtilis ATCC 6633 (Duitman et al. 1999), possess so poor natural competence that the exogenous DNA cannot be transformed into the cells by natural competence transformation. Whether could the GP35-mediated recombineering system be applied in originally poor natural competence strains completely by electro-transformation? For this purpose, the system was applied by introducing a low-copy plasmid pHCMC04-gp35 into B. subtilis ATCC 6633 and the recombineering frequency was increased more than 40-fold compared to that of wild type (Fig. S4). In a word, the recombineering system mediated by GP35 could perform well in Bacillus strains not only with natural competence, but also with originally poor natural competence.
Furthermore, we attempted to express the gp34.1-35-36 genes (gp34.1, 5′-3′exonuclease gene; gp36, single-strand DNA-binding protein gene) to recombine with dsDNA. However, the recombination mediated by GP34.1-35-36 with the dsDNA was inefficient (data not shown), most likely due to degradation of the dsDNA molecules introduced into the B. subtilis by the rapid and processive AddAB helicase-nuclease (Haijema et al. 1996; Kooistra et al. 1993).
There was an interesting phenomenon that the recombinases GP35 and Orf48, which have been reported to exhibit low recombineering frequency in E. coli (Datta et al. 2008), displayed high frequency in B. subtilis (Fig. 4b). The different behavior of the recombinase GP61 in M. smegmatis and E. coli was also reported previously (Datta et al. 2008; van Kessel and Hatfull 2007, 2008). The fact that the performance of the same recombinase differs in different hosts may be due to specific molecular interactions with the endogenous host factors. We further investigated the relationship between the recombineering frequency of the recombinases and the phage host kinship, expression levels, and amino acid sequence identities. With the exception of host kinship, the expression levels and amino acid sequence identities of the recombinases were not related to the recombineering frequency (Fig. S3a). The analysis of the relatedness of the hosts and the recombineering frequency showed that the recombinases most likely perform better in their native hosts, where specific molecular interactions with host factors may be required for tighter binding in the native hosts. In the native host, these recombinases are encoded by bacterial phages, prophages, or their remnants, and the protein structure is most suitable to interact with other host factors. GP35 from native phage SPP1 is an ATP-independent single-strand annealing (SSA) protein and specifically interacts with SPP1-encoded replicative DNA helicase GP40 and single-strand binding (SSB) protein GP36 (Ayora et al. 2002). Because the DNA helicase DnaC for promoting replication and SSB protein SsbA for protecting ssDNA from degrading, two crucial proteins in DNA strand invasion and exchange process of B. subtilis, share high identities with GP40 and GP36, respectively, we speculate that GP35 might interact with DnaC and SsbA effectively and promote DNA strand invasion and exchange, which results in higher recombineering frequency than other recombinases in B. subtilis. Therefore, endogenous recombinase or its nearest relatives are the best candidates for an efficient genetic manipulation system.
Previous studies have shown that different recombinase homologs have the ability to function in a wide range of hosts (Datta et al. 2008; van Kessel and Hatfull 2008; Wang et al. 2012); therefore, choosing an effective recombinase would be a useful approach for improving ssDNA recombineering in all bacteria. We focused on B. subtilis and the identification of GP35, which may serve as a useful tool for improving recombineering. In conclusion, the recombinase GP35 from the native phage SPP1 was identified to promote the ssDNA recombineering frequency in B. subtilis. Further discovery of high-performance recombinases from native hosts in target bacteria may provide an effective genetic tool for constructing genetically engineered strains and investigating bacterial physiology.
References
Anagnostopoulos C, Spizizen J (1961) Requirements for transformation in Bacillus subtilis. J Bacteriol 81(5):741–746
Ayora S, Missich R, Mesa P, Lurz R, Yang SX, Egelman EH, Alonso JC (2002) Homologous-pairing activity of the Bacillus subtilis bacteriophage SPP1 replication protein G35P. J Biol Chem 277(39):35969–35979. doi:10.1074/jbc.M204467200
Bashkirov VI, Khasanov FK, Prozorov AA (1987) Illegitimate recombination in Bacillus subtilis nucleotide sequences at recombinant DNA junctions. Mol Gen Genet 210(3):578–580. doi:10.1007/Bf00327215
Brigidi P, De Rossi E, Bertarini ML, Riccardi G, Matteuzzi D (1990) Genetic transformation of intact cells of Bacillus subtilis by electroporation. FEMS Microbiol Lett 55(1–2):135–138
Court DL, Sawitzke JA, Thomason LC (2002) Genetic engineering using homologous recombination. Annu Rev Genet 36:361–388. doi:10.1146/annurev.genet.36.061102.093104
Datta S, Costantino N, Court DL (2006) A set of recombineering plasmids for gram-negative bacteria. Gene 379:109–115. doi:10.1016/j.gene.2006.04.018
Datta S, Costantino N, Zhou XM, Court DL (2008) Identification and analysis of recombineering functions from Gram-negative and Gram-positive bacteria and their phages. Proc Natl Acad Sci U S A 105(5):1626–1631. doi:10.1073/pnas.0709089105
Deans TL, Cantor CR, Collins JJ (2007) A tunable genetic switch based on RNAi and repressor proteins for regulating gene expression in mammalian cells. Cell 130(2):363–372. doi:10.1016/j.cell.2007.05.045
Deng AH, Wu J, Zhang Y, Zhang GQ, Wen TY (2010) Purification and characterization of a surfactant-stable high-alkaline protease from Bacillus sp B001. Bioresour Technol 101(18):7100–7106. doi:10.1016/j.biortech.2010.03.130
Duitman EH, Hamoen LW, Rembold M, Venema G, Seitz H, Saenger W, Bernhard F, Reinhardt R, Schmidt M, Ullrich C, Stein T, Leenders F, Vater J (1999) The mycosubtilin synthetase of Bacillus subtilis ATCC6633: a multifunctional hybrid between a peptide synthetase, an amino transferase, and a fatty acid synthase. Proc Natl Acad Sci U S A 96(23):13294–13299. doi:10.1073/pnas.96.23.13294
Ellis HM, Yu DG, DiTizio T, Court DL (2001) High efficiency mutagenesis, repair, and engineering of chromosomal DNA using single-stranded oligonucleotides. Proc Natl Acad Sci U S A 98(12):6742–6746. doi:10.1073/pnas.121164898
Fabret C, Ehrlich SD, Noirot P (2002) A new mutation delivery system for genome-scale approaches in Bacillus subtilis. Mol Microbiol 46(1):25–36. doi:10.1046/j.1365-2958.2002.03140.x
Gietz RD, Woods RA (2002) Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol 350:87–96
Haijema BJ, Venema G, Kooistra J (1996) The C terminus of the AddA subunit of the Bacillus subtilis ATP-dependent DNase is required for the ATP-dependent exonuclease activity but not for the helicase activity. J Bacteriol 178(17):5086–5091
Hofemeister J, Israelireches M, Dubnau D (1983) Integration of plasmid-pE194 at multiple sites on the Bacillus subtilis chromosome. Mol Gen Genet 189(1):58–68. doi:10.1007/Bf00326055
Jiang WY, Bikard D, Cox D, Zhang F, Marraffini LA (2013) RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat Biotechnol 31(3):233–239. doi:10.1038/Nbt.2508
Kooistra J, Haijema BJ, Venema G (1993) The Bacillus subtilis addAB genes are fully functional in Escherichia coli. Mol Microbiol 7(6):915–923. doi:10.1111/j.1365-2958.1993.tb01182.x
Li XT, Costantino N, Lu LY, Liu DP, Watt RM, Cheah KSE, Court DL, Huang JD (2003) Identification of factors influencing strand bias in oligonucleotide-mediated recombination in Escherichia coli. Nucleic Acids Res 31(22):6674–6687. doi:10.1093/nar/gkg844
Maresca M, Erler A, Fu J, Friedrich A, Zhang YM, Stewart AF (2010) Single-stranded heteroduplex intermediates in lambda Red homologous recombination. BMC Mol Biol 11(54):54. doi:10.1186/1471-2199-11-54
Mosberg JA, Lajoie MJ, Church GM (2010) Lambda red recombineering in Escherichia coli occurs through a fully single-stranded intermediate. Genetics 186(3):791–799. doi:10.1534/genetics.110.120782
Muyers JPP, Zhang YM, Testa G, Stewart AF (1999) Rapid modification of bacterial artificial chromosome by ET-recombination. Nucleic Acids Res 27(6):1555–1557. doi:10.1093/nar/27.6.1555
Nijland R, Burgess JG, Errington J, Veening JW (2010) Transformation of environmental Bacillus subtilis isolates by transiently inducing genetic competence. PLoS ONE 5(3):e9724. doi:10.1371/journal.pone.0009724
Ranallo RT, Barnoy S, Thakkar S, Urick T, Venkatesan MM (2006) Developing live Shigella vaccines using lambda red recombineering. FEMS Immunol Med Microbiol 47(3):462–469. doi:10.1111/j.1574-695X.2006.00118.x
Robzyk K, Kassir Y (1992) A simple and highly efficient procedure for rescuing autonomous plasmids from yeast. Nucleic Acids Res 20(14):3790
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, pp 53–84
Schenk S, Laddaga RA (1992) Improved method for electroporation of Staphylococcus aureus. FEMS Microbiol Lett 73(1–2):133–138
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C-T method. Nat Protoc 3(6):1101–1108. doi:10.1038/nprot.2008.73
Shao Z, Zhao H, Zhao H (2009) DNA assembler, an in vivo genetic method for rapid construction of biochemical pathways. Nucleic Acids Res 37(2):e16. doi:10.1093/nar/gkn991
Sharan SK, Thomason LC, Kuznetsov SG, Court DL (2009) Recombineering: a homologous recombination-based method of genetic engineering. Nat Protoc 4(2):206–223. doi:10.1038/nprot.2008.227
Stephenson M, Jarrett P (1991) Transformation of Bacillus subtilis by electroporation. Biotechnol Tech 5(1):9–12. doi:10.1007/Bf00152746
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739. doi:10.1093/molbev/msr121
Tang X, Morris SL, Langone JJ, Bockstahler LE (2006) Simple and effective method for generating single-stranded DNA targets and probes. Biotechniques 40(6):759–763. doi:10.2144/000112154
van Kessel JC, Hatfull GF (2007) Recombineering in Mycobacterium tuberculosis. Nat Methods 4(2):147–152. doi:10.1038/nmeth996
van Kessel JC, Hatfull GF (2008) Efficient point mutagenesis in mycobacteria using single-stranded DNA recombineering: characterization of antimycobacterial drug targets. Mol Microbiol 67(5):1094–1107. doi:10.1111/j.1365-2958.2008.06109.x
van Kessel JC, Marinelli LJ, Hatfull GF (2008) Recombineering mycobacteria and their phages. Nat Rev Microbiol 6(11):851–857. doi:10.1038/Nrmicro2014
van Pijkeren JP, Britton RA (2012) High efficiency recombineering in lactic acid bacteria. Nucleic Acids Res 40(10):e76. doi:10.1093/nar/gks147
Wang PZ, Doi RH (1984) Overlapping promoters transcribed by Bacillus subtilis sigma-55 and sigma-37 RNA-polymerase holoenzymes during growth and stationary phases. J Biol Chem 259(13):8619–8625
Wang JJ, Rojanatavorn K, Shih JCH (2004) Increased production of Bacillus keratinase by chromosomal integration of multiple copies of the kerA gene. Biotechnol Bioeng 87(4):459–464. doi:10.1002/Bit.20145
Wang Y, Weng J, Waseem R, Yin XH, Zhang RF, Shen QR (2012) Bacillus subtilis genome editing using ssDNA with short homology regions. Nucleic Acids Res 40(12):e91. doi:10.1093/nar/gks248
Wen S, Yang JG, Tan TW (2013) Full-length single-stranded PCR product mediated chromosomal integration in intact Bacillus subtilis. J Microbiol Methods 92(3):273–277. doi:10.1016/j.mimet.2012.11.012
Xue GP, Johnson JS, Dalrymple BP (1999) High osmolarity improves the electro-transformation efficiency of the gram-positive bacteria Bacillus subtilis and Bacillus licheniformis. J Microbiol Methods 34(3):183–191. doi:10.1016/S0167-7012(98)00087-6
Yan X, Yu HJ, Hong Q, Li SP (2008) Cre/lox system and PCR-based genome engineering in Bacillus subtilis. Appl Environ Microbiol 74(17):5556–5562. doi:10.1128/Aem. 01156-08
Zhang YM, Buchholz F, Muyrers JPP, Stewart AF (1998) A new logic for DNA engineering using recombination in Escherichia coli. Nat Genet 20(2):123–128. doi:10.1038/2417
Zhang GQ, Bao P, Zhang Y, Deng AH, Chen N, Wen TY (2011) Enhancing electro-transformation competency of recalcitrant Bacillus amyloliquefaciens by combining cell-wall weakening and cell-membrane fluidity disturbing. Anal Biochem 409(1):130–137. doi:10.1016/J.Ab.2010.10.013
Zhang GQ, Wang WZ, Deng AH, Sun ZP, Zhang Y, Liang Y, Che YS, Wen TY (2012) A mimicking-of-DNA-methylation-patterns pipeline for overcoming the restriction barrier of bacteria. PLoS Genet 8(9):e1002987. doi:10.1371/journal.pgen.1002987
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (31170103) and the Chinese Academy of Sciences (KSCX2-EW-J-6). We thank Prof. Donald L. Court (National Cancer Institute at Frederick) for kindly providing the chromosomes of strains SIMD40-43 and SIMD44-50 for cloning recombinases and Dr. Mingchun Li for S. cerevisiae DAY414 and BGSC for various shuttle plasmids. We are grateful to Tong Zhao for the flow cytometry analysis.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 2411 kb)
Rights and permissions
About this article
Cite this article
Sun, Z., Deng, A., Hu, T. et al. A high-efficiency recombineering system with PCR-based ssDNA in Bacillus subtilis mediated by the native phage recombinase GP35. Appl Microbiol Biotechnol 99, 5151–5162 (2015). https://doi.org/10.1007/s00253-015-6485-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6485-5