Abstract
One of the probable hypotheses for the onset of autoimmunity is molecular mimicry. This study aimed to determine the HLA-II risk alleles for developing Hashimoto’s thyroiditis (HT) in order to analyze the molecular homology between candidate pathogen-derived epitopes and potentially self-antigens (thyroid peroxidase, TPO) based on the presence of HLA risk alleles. HLA-DRB1/-DQB1 genotyping was performed in 100 HT patients and 330 ethnically matched healthy controls to determine the predisposing/protective alleles for HT disease. Then, in silico analysis was conducted to examine the sequence homology between epitopes derived from autoantigens and four potentially relevant pathogens and their binding capacities to HLA risk alleles based on peptide docking analysis. We identified HLA-DRB1*03:01, *04:02, *04:05, and *11:04 as predisposing alleles and DRB1*13:01 as a potentially predictive allele for HT disease. Also, DRB1*11:04 ~ DQB1*03:01 (Pc = 0.002; OR, 3.97) and DRB1*03:01 ~ DQB1*02:01 (Pc = 0.004; OR, 2.24) haplotypes conferred a predisposing role for HT. Based on logistic regression analysis, carrying risk alleles increased the risk of HT development 4.5 times in our population (P = 7.09E-10). Also, ROC curve analysis revealed a high predictive power of those risk alleles for discrimination of the susceptible from healthy individuals (AUC, 0.70; P = 6.6E-10). Analysis of peptide sequence homology between epitopes of TPO and epitopes derived from four candidate microorganisms revealed a homology between envelop glycoprotein D of herpes virus and sequence 151–199 of TPO with remarkable binding capacity to HLA-DRB1*03:01 allele. Our findings indicate the increased risk of developing HT in those individuals carrying HLA risk alleles which can also be related to herpes virus infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hashimoto’s thyroiditis (HT) and Graves’ disease (GD) are two common thyroid-specific autoimmune diseases (AITD), with a prevalence rate ranging from 1 to 5% (Cho et al. 2011). While the clinical manifestations of GD and HT differ, the underlying mechanism involving the breakdown of tolerance to thyroid autoantigens and the subsequent autoimmune responses is similar in both diseases (Lee et al. 2023; Wiersinga 2014). HT is characterized by the infiltration of autoreactive T lymphocytes into the thyroid, resulting in the destruction of thyroid follicle cells through the induction of apoptosis. This process also leads to the production of autoantibodies against the thyroid peroxidase (TPO) autoantigen and ultimately, leads to hypothyroidism (Ralli et al. 2020).
The development of autoimmune thyroid diseases (AITD) is influenced by a combination of genetic, epigenetic, and environmental factors (Lee et al. 2023; Liontiris and Mazokopakis 2017). Genome-wide association studies (GWAS) have identified several gene loci associated with the development of AITD (Zhang et al. 2020; Hasham and Tomer 2012). These genes can be classified into two main categories: immune-related genes, such as human leukocyte antigen (HLA), CTLA4, PTPN22, CD40, and FOXP3, and genes responsible for encoding the primary thyroid autoantigens, such as thyroid peroxidase (TPO) and thyroid-stimulating hormone receptor (TSHR). These genes play pivotal roles in the onset of AITD (Lee et al. 2023; Hasham and Tomer 2012). Furthermore, investigations have indicated that the induction of interferon-alpha (IFN-α) during viral infections may lead to the downregulation of thyroglobulin (Tg) expression. This observation suggests a potential link between infectious triggers and the development of AITD (Lee et al. 2023; Hasham and Tomer 2012).
One of the most relevant genetic loci predisposing individuals to various autoimmune diseases, such as multiple sclerosis (MS), systemic lupus erythematosus (SLE), type 1 diabetes (T1D), and autoimmune thyroid diseases (AITD), is the HLA gene complex. The HLA genes are also among the most polymorphic genes in humans (Al Yafei et al. 2022; Rasouli-Saravani et al. 2021; Enz et al. 2020). These genes encode two sets of classical HLA molecules, HLA-I and HLA-II, which are responsible for presenting peptide antigens to T lymphocytes. T-CD8 + and T-CD4 + lymphocytes recognize antigenic peptides derived from either foreign or self-antigens through HLA-I and HLA-II molecules, respectively (Bodis et al. 2018). The polymorphism in the HLA-II genes, specifically the DRB and DQB genes, is closely associated with the development of autoimmune diseases as it influences the activation of autoreactive T-CD4+ lymphocytes (Arango et al. 2017; Naito and Okada 2022). These autoreactive T-CD4+ lymphocytes play a pivotal role in triggering the activation of autoreactive T-CD8+ lymphocytes and the differentiation of autoreactive B lymphocytes into plasma cells, leading to the promotion and enhancement of autoinflammatory responses and autoantibody production.
Studies conducted on Caucasians in a case–control genotyping setting have revealed that HLA-DRB1*03 ~ DQB1*02, HLA-DRB1*04 ~ DQB1*03, and HLA-DRB1*08 ~ DQB1*04 haplotypes are positively associated with HT. In contrast, HLA-DRB1*07 ~ DQB1*02, HLA-DRB1*13 ~ DQB1*06, and HLA-DRB1*15 ~ DQB1*06 haplotypes appear to confer a protective role against HT (Zeitlin et al. 2008). Studies among East Asians have shown that HLA-DRB1*08:03 ~ DQB1*06:01 and HLA-DRB1*09:01 ~ DQB1*03:03 haplotypes may predispose individuals to HT, while HLA-DRB1*13:02 ~ DQB1*06:04 and HLA-DRB1*15:01 ~ DQB1*06:02 haplotypes offer strong protection against HT (Katahira et al. 2013). In this context, assessing the probability of developing HT in individuals with genetic susceptibility requires the calculation of a genetic risk score (GRS). While clinical, demographic, and other contributory factors can be used for GRS calculation, it is typically evaluated based on genetic variants (Schultheiss et al. 2015; Qu et al. 2021). Additionally, environmental factors, including smoking, viral and bacterial infections, and excessive iodine intake, contribute to the development of HT in genetically susceptible individuals (Wiersinga 2014; Liontiris and Mazokopakis 2017).
One of the strong hypotheses regarding the onset of autoimmune diseases is known as molecular mimicry (Ramasamy et al. 2020; Bogers et al. 2023; Ghobadi et al. 2021; Laron et al. 2023; Martins et al. 2023). According to this theory, autoreactive T cells become activated by antigenic peptides originating from microorganisms. This activation occurs because of the similarity between antigens derived from pathogens and self-antigens (Ramasamy et al. 2020; Martins et al. 2023). Recent studies have demonstrated the presence of viral infections caused by Herpesviruses and Enteroviruses in the thyroid biopsies of patients with autoimmune thyroid diseases (AITD) (Weider et al. 2022; Seyyedi et al. 2019; Hammerstad et al. 2013; Cuan-Baltazar and Soto-Vega 2020). Investigations have also suggested that bacterial pathogens such as Yersinia enterocolitica, Helicobacter pylori, and Mycobacterium avium ssp. paratuberculosis might be associated with the onset of HT (Figura et al. 2019; Choi et al. 2017; Moghadam et al. 2022; Zangiabadian et al. 2021). The current case–control and in silico studies were designed to explore the genetic susceptibility and protective patterns in our HT patients. This exploration was based on HLA class II genotyping, and it aimed to examine molecular homology through the analysis of peptide sequence similarities. Specifically, the study assessed the similarity between candidate pathogen-derived epitopes and potential self-antigens in the patients with HT.
Patients and methods
Subjects
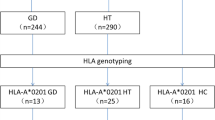
This study enrolled a total of 100 participants, comprising 83 females and 17 males, who were unrelated and diagnosed with Hashimoto’s thyroiditis (HT). The mean age of the participants was 39.8 ± 11.8 years, and they were referred to our outpatient clinic between February 2022 and April 2023. Diagnosis of HT was made in accordance with international guidelines and was confirmed by a specialist. The diagnosis was based on the manifestation of hypothyroidism, which included symptoms such as dry skin, hair loss (calvities), and coldness in peripheral organs at the time of diagnosis. Paraclinical confirmation of the disease was achieved through the measurement of thyroid hormones and the presence of anti-thyroid peroxidase antibody (Ralli et al. 2020; Caturegli et al. 2014). All HT patients were receiving treatment with levothyroxine replacement therapy. The study also collected laboratory data, including TSH (thyroid-stimulating hormone), FT4 (free thyroxine), and anti-thyroid peroxidase antibody (anti-TPO) levels, from the medical records of each patient. Additionally, 330 unrelated and ethnically matched healthy individuals who exhibited normal thyroid function tests and without medical complaints were recruited as a control group. A flowchart of the study design is shown in Fig. 1.
All participants willingly consented to take part in the study, and the research was conducted following the approval of our institutional ethics committee (IR.UMSHA.REC.1401.1028) and in accordance with the principles of the Helsinki Declaration.
DNA extraction and HLA typing
Genomic DNA was extracted from whole blood samples collected using the chloroform-based salting-out method (Moradi et al. 2014). To achieve 2-field resolution HLA-typing, we employed the PCR with sequence-specific oligonucleotide probe (PCR-SSOP) technique. Commercial kits (HISTO SPOT DR and DQ BAG Diagnostics, Germany) were utilized following the manufacturer’s instructions. The process began with the amplification step, where locus-specific biotinylated primers were used in PCR. Subsequently, the PCR products were denatured to produce single-stranded amplicons and then subjected to hybridization with oligonucleotide probes using the MR.SPOT®/MR.SPOT® 2.0 processor. Finally, the patterns of spots were interpreted using specific software provided by the same company (HISTO MATCH interpretation software) to determine the specific DRB1/B3/B4/B5 and DQB1 alleles for each subject. In addition, HLA-DRB1 ~ DQB1 haplotypes were statistically determined using an Expectation–Maximization (EM) algorithm. This algorithm was implemented within the R statistical computing environment (http://www.R-project.org) software.
Estimation of genetic risk score (GRS) for HT
To predict the likelihood of disease development in those individuals carrying HLA risk alleles, we computed the cumulative effect of several risk alleles based on a logistic regression model to generate the genetic risk score (GRS). Subsequently, odds ratios were calculated for both the risk and protective groups compared to the neutral subjects. This analysis allowed us to quantify the risk and protective effects of different alleles. Furthermore, to assess the prediction accuracy of GRS and to determine the predictive power of risk alleles in discrimination of patients from healthy individuals in our population, we measured the area under receiver operating characteristic (ROC) curve (AUC).
Epitope prediction
Based on previous studies (Weider et al. 2022; Figura et al. 2019; Choi et al. 2017; Moghadam et al. 2022; Zangiabadian et al. 2021), we selected four microorganisms, including enterovirus, herpesvirus, Helicobacter pylori, and Yersinia enterocolitica, for the analysis of molecular mimicry between antigens derived from these potentially relevant pathogens and thyroid autoantigens (TPO and Tg). The analysis was conducted in several steps. First, we utilized the Immune Epitope Database (IEDB) (https://www.iedb.org) to extract confirmed epitopes of the selected microorganisms that could bind to HLA-II molecules. Subsequently, we obtained the complete protein sequences related to these confirmed epitopes from the UniProt database (https://www.uniprot.org). We also extracted the complete protein sequences of TPO and Tg from the UniProt database. To predict the binding of peptides derived from the microorganisms, TPO, and Tg to HLA-DRB1*03:01, which is one of the predisposing HLA alleles for developing HT, we used the NetMHCIIpan-4.0 server (https://services.healthtech.dtu.dk/services/NetMHCIIpan-4.0). We applied a filtering criterion based on the percentile rank (% Rank), with a threshold for strong and low binders set at < 1.0 and > 1.0, respectively. A high percentile rank indicated a poor binding capacity, while a low percentile rank indicated a strong binding capacity. Epitopes derived from the microorganisms and Tg proteins that exhibited strong binding (SB) to HLA-DRB1*03:01 were selected from the NetMHCIIpan-4.0 results. In the case of TPO-derived epitopes, none showed strong binding (SB) with HLA-DRB1*03:01, so all epitopes with weak binding (WB) were included in the subsequent analysis.
Homology and alignment search
We performed a simultaneous alignment and clustering of multiple peptide sequences using the GibbsCluster—2.0 (https://services.healthtech.dtu.dk/services/GibbsCluster-2.0) for the selected epitopes derived from microorganisms, TPO and Tg as obtained from the NetMHCIIpan-4.0 results. Also, to investigate the homology among TPO and Tg proteins and confirmed protein sequences of microorganisms, we utilized the BLASTp program (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
HLA-peptide docking
We conducted HLA-peptide docking to assess the HLA-II binding promiscuity of human and microorganism-mimicking peptides using the HPEPDOCK 2.0 server (http://huanglab.phys.hust.edu.cn/hpepdock). This process is based on flexible peptide-protein docking, involving the fast modeling of peptide conformations and global/local sampling of binding orientations. Notably, HPEPDOCK 2.0 does not require the 3D structure of the peptide, making the docking independent of the prediction error of homologous peptide structures. Docking scores are represented based on Gibbs free energy (kcal/mol). In the initial step, the HLA-DRB1*03-peptide binding complex (1A6A) was extracted from the Protein Data Bank (https://www.rcsb.org). To prepare for peptide docking, water and other atoms were removed from the complex. Molecular docking was carried out to confirm and identify the critical residues of epitopes from self and non-self proteins in conjunction with HLA-DRB1*03:01 molecule. To visualize the docked models, we utilized Discovery Studio 2016 and PyMol 1.7.4 to generate molecular graphic images.
Statistical analysis
We employed a range of descriptive statistics, including measures such as the mean and standard deviation for quantitative variables, and empirical distribution for qualitative variables. To compare proportions between two independent groups, we utilized the chi-squared test or, when appropriate, the Fisher-exact test. To assess the risk associated with haplotypes and genotypes, we calculated odds ratios (OR) and their corresponding 95% confidence intervals. In order to address the issue of multiple testing and control the false discovery rate (FDR), we applied the Benjamini–Hochberg method for multiple comparisons (Benjamini and Hochberg, revised version 2010). All computations and analyses were performed using the R program, version 4.3.0. For data management and some visualizations, we utilized the tidyverse package, while the pROC package was employed for the calculation of the area under the ROC curve.
Results
We conducted an analysis on a total of 100 HT patients, consisting of 17 males and 83 females, with a mean age of 39.81 ± 11.84 years. The mean age of disease onset was 30.30 ± 10.74 years and the mean time for disease duration was 9.50 ± 8.38 years. At the time of diagnosis, the majority of patients exhibited various symptoms, including fatigue and weakness (97.0%), calvities (92.0%), coldness of peripheral organs (90.0%), and dry skin and diffuse alopecia (88.0%). Additionally, 80.0% of patients experienced a loss of appetite and weight gain, 68.0% felt cold, 51.0% had constipation, and 46.0% exhibited puffiness in their face, hands, and feet. Carpal tunnel syndrome and paresthesia were present in 27.0% and 23.0% of patients, respectively. Among female patients, 67 (80.7%) experienced oligomenorrhea. In the patient group, the mean serum levels of TSH, T4, and anti-TPO antibody were 85.35 ± 24.68 mlU/L, 3.94 ± 0.73 µg/dl, and 605.80 ± 527.09 IU/ml, respectively. We also analyzed a control group consisting of 330 unrelated, ethnically matched healthy individuals including 177 males and 153 females, aged 36.4 ± 12.3 years.
Distributions of DRB1 and DQB1 alleles, genotypes, and haplotypes among HT patients and healthy controls
The highest risk for developing HT was associated with DRB1*04:05 allele (Pc = 0.01; OR, 6.15), followed by DRB1*11:04 (Pc = 0.001; OR, 4.19), DRB1*04:02 (Pc = 0.05; OR, 3.17), and DRB1*03:01 (Pc = 0.003; OR, 2.24) alleles. Additionally, DQB1*02:01 (Pc = 0.006; OR, 2.17) and DQB1*03:02 (Pc = 0.02; OR, 2.10) alleles were found to be associated with an increased risk for HT. Conversely, DRB1*13:01 (Pc = 0.05; OR, 0.28) and DQB1*06:03 (Pc = 0.05; OR, 0.29) alleles exhibited a potential protective effect against the development of HT (Tables 1 and 2 and Supplementary Figs. 1 & 2).
As expected, the haplotypes associated with increased risk for HT were DRB1*11:04 ~ DQB1*03:01 (Pc = 0.002; OR, 3.97) and DRB1*03:01 ~ DQB1*02:01 (Pc = 0.004; OR, 2.24). The only potentially protective haplotype was DRB1*13:01 ~ DQB1*06:03 (Pc = 0.07; OR, 0.30) (Suppl Tables 1 & 2) in out HT patients. Furthermore, when comparing genotype distributions between the patient and control groups, it was found that the major predisposing diplotypes among patients, as compared to controls, were DR3/DR4 (Pc = 0.005; OR, 9.41), followed by DR4/DR11 (Pc = 0.02; OR, 2.12), and DR3/DR11 (Pc = 0.04; OR, 2.11). Conversely, the major protective diplotypes among patients, in comparison to controls, were DRX/DR13 (Pc = 0.02; OR, 0.25) and DRX/DRX (Pc = 0.02; OR, 0.51). Here, DRX denotes the presence of DRB1 alleles other than predisposing (DRB1*03 and *04) or protective (DRB1*13) alleles (Table 3).
Estimation of the cumulative risk effects: GRS estimation and ROC curve analysis
According to the identification of potentially HLA risk alleles (DRB1*03:01, *04:02, *04:05, and *11:04) and protective allele (DRB1*13 allele group) in our HT patients compared to healthy controls, we performed logistic regression analysis to estimate the GRS. In this model, our study subject was classified into four groups: (1) reference group (absence of either risk or protective alleles), (2) risk group (carrying two risk alleles), (3) protective group (carrying of two protective alleles), and (4) risk/protective group. We observed that subjects carrying risk alleles have 4.5 fold of risk to develop HT disease in our population (P = 7.09E-10, Table 4). Also, ROC curve analysis revealed a high predictive power of those risk alleles in discrimination of susceptible from healthy individuals (AUC, 0.70; P = 6.6E-10). Although, the AUC indicated the sensitivity of 60.0% and specificity of 77.0% for this model (Fig. 2).
In silico results
Epitope prediction
We extracted confirmed epitopes from microorganisms that demonstrated binding capacity to HLA-II molecule (DRB1*03:01) from IEDB. Subsequently, we subjected the global proteins of TPO and Tg, along with the proteins containing the confirmed epitopes from microorganisms, to identify 15-amino acid peptides potentially presented by HLA-DRB1*03:01 molecule. We employed NetMHCIIpan as a predictive tool for this analysis. The selection of predicted peptides was based on the percentile rank (%Rank). A peptide was categorized as a strong binder if the percentile rank was below 1.0, and as a weak binder if it exceeded the 1.0 specified thresholds. Our predictions indicated that 42, 30, and 6 epitopes from Herpesviruses, Enteroviruses, and Yersinia enterocolitica, respectively, were strong binders for HLA-DRB1*03:01, as shown in Suppl Tables 3. Additionally, 7 epitopes from Helicobacter pylori were strongly bound to HLA-DRB1*03:01, while other predicted epitopes for this pathogen exhibited weak binding. All 50 predicted TPO-derived epitopes were classified as weak binders, and those epitopes containing V-D, I-D, and L-D anchor residues are illustrated in Suppl Table 4. Among the 63 predicted Tg-derived epitopes, 6 were strongly bound to HLA-DRB1*03:01, while the remaining epitopes were considered weak binders.
Sequence homology analysis among predicted epitopes derived from TPO, Tg, and four microorganisms
Following the Gibbs simultaneous alignment and clustering algorithm, 247 predicted self and non-self-epitopes, were grouped into three initial meaningful clusters for motif identification in the peptide dataset. The applied algorithm efforts at maximizing the information content of individual matrixes while minimizing the overlap between distinct clusters, so that each cluster is represented by a position-specific scoring matrix (PSSM). Also, to identify the optimal local sequence alignment in each cluster Kullback–Leibler distance (KLD) sum of the alignments was measured by Gibbs Cluster—2.0. The KLD measures the information gain of an observed amino acid distribution compared with a background distribution (the frequency of each amino acid in random protein sequences) (Suppl Fig. 3).
The anchor residues of peptides for the resulting clusters are depicted in Fig. 3. In the first cluster, consisting of 247 sequences, amino acid preferences at the 2nd, 5th, 7th, and 9th positions were represented by [LIM], [DEN], [KR], and [RP], respectively. Within group 1 of the second cluster, which included 145 sequences from the total of 247 peptides, predominant amino acids at the 3rd and 6th positions were [LVI] and [DEN], respectively. Also, in group 2 of the second cluster, comprising 102 peptides, the 2nd and 5th positions were mainly occupied by [IVL] and [D] amino acids. Additionally, group 1 of the third cluster, consisting of 123 sequences from all 247 peptides displayed preferences for [LVI] and [DNE] amino acids at the 1st and 4th positions, respectively. Finally, group 2 of the third cluster exhibited more promiscuity in anchor positions, although [D] and [R] amino acids were predominantly favored at the 6th and 9th positions, respectively (Fig. 3).
Sequence Logos of the selected epitopes from thyroid peroxidase (TPO), thyroglobulin (Tg), and four microorganisms by Gibbs Cluster 2.0. The height of the stack indicates the conserved residues at specific position, while the height of symbols within the stack indicates the relative frequency of each residue at that position
Global protein homology
We used the BLASTp program to analyze the homology between the global proteins of TPO, Tg, and the proteins of microorganisms containing confirmed epitopes. The results revealed that none of the confirmed proteins from microorganisms exhibited homology with Tg and TPO, except for the presence of homology between the envelope glycoprotein D of herpes virus and TPO. Specifically, as shown in Suppl Table 5, a homologous sequence within the envelope glycoprotein D from the herpes virus (sequence 145–180) was identified with an E value of 0.029, sequence coverage of 9%, and sequence percentage identity of 32.65%. This sequence corresponded to sequence 151–199 of TPO.
Molecular docking
TPO-derived epitopes and herpes virus-derived epitopes were randomly chosen for docking. The results of docking simulations revealed that those epitopes feasibly interacted with the HLA-DR3 peptide binding cleft, forming multiple molecular interactions, particularly hydrogen and electrostatic bonds. The scores obtained from the peptide docking simulations indicated the binding affinities, where higher scores indicated lower binding affinities and vice versa. Based on molecular docking analysis, we found that Asnβ82, Arg β74, Glnβ70, and Serα53 and Asnα62 amino acid residues of HLA-DR3 molecule are critical for binding to the anchor residues of V and D in the epitopes. Additionally, anchor residues of L and D in the epitopes bind to Lysβ71 and Argβ74, along with Asnα62 and Asnα69 of HLA-DR3 molecule. Moreover, Lysβ71, Argβ74, and Trpβ61, as well as Asnα62 and Asnα69 are important residues for binding to I and D residues in the epitopes (Suppl Fig. 4).
Discussion
Allelic distributions of DRB1/DQB1 genes among HT patients and healthy controls
We analyzed the frequencies of HLA-DRB1 and –DQB1 alleles, genotypes, and haplotypes in 100 patients with Hashimoto’s thyroiditis (HT) and 330 ethnically matched healthy individuals. Our findings are in line with similar studies conducted in Caucasian (Farid and Thompson 1986), Hungarian (Stenszky et al. 1987), and Italian (Petrone et al. 2001) populations, which demonstrated the predisposing role of HLA-DRB1*03 and HLA-DRB1*04 alleles in the development of HT. Consistently, we observed that HLA-DRB1*03:01, DRB1*04:02 and DRB1*04:05 as well as DRB1*11:04 alleles may be associated with an increased susceptibility to HT. Furthermore, our study revealed a potentially predisposing role for HLA-DQB1*02:01 and DQB1*03:02 alleles, along with a potential protective effect of DRB1*13:01 and DQB1*06:03 alleles in our HT patients. In contrast to Zeitlin et al. study (Zeitlin et al. 2008), we observed a susceptibility role for DRB1*11:04 alleles and similarly a protective role for DRB1*13:01 allele in our HT patients. Also, in agreement with a study on the Greek population (Kokaraki et al. 2009), our study demonstrated that DRB1*04:05, DQB1*02:01, and DQB1*03:02 alleles are associated with an increased risk for developing HT in Iranians. Additionally, our study identified more predisposing alleles, including DRB1*03:01 and DRB1*04:02, and suggested a possibly neutral effect of DRB1*07 alleles in our patients which is inconsistent with the results of a study conducted in the Greek population (Kokaraki et al. 2009).
Furthermore, our findings corroborate the results of the Zeitlin et al. study (Zeitlin et al. 2008) in terms of highlighting the predisposing effect of DRB1*03:01 ~ DQB1*02:01 haplotype as well as the marginally protective effect of the DRB1*13:01 ~ DQB1*06:03 haplotype against the development of HT. Among the major predisposing diplotypes in our patients, we observed that DR3/DR4, DR3/DRX, and DR4/DRX diplotypes, where “X” represents any alleles except DRB1*03, *04, *11, and *13, conferred higher risk for disease development. Notably, within the DR3/DRX and DR4/DRX diplotypes, DR3/DR11 and DR4/DR11 were identified as two significantly predisposing diplotypes in our patients. Additionally, the estimation of GRS and ROC curve analysis indicated that the HLA risk alleles for HT in our population can be used for screening programs and to improve the preventive strategies among high-risk individuals.
In silico analysis
One of the probable hypotheses for the onset of autoimmunity is molecular mimicry (Ramasamy et al. 2020). Building upon this hypothesis, a potential connection has been demonstrated between coxsackie B virus infection and the initiation of type 1 diabetes (Nekoua et al. 2022). Recent studies have also suggested that certain bacteria and some of common human viruses might be associated with the onset of Hashimoto’s thyroiditis (Weider et al. 2022; Figura et al. 2019; Choi et al. 2017; Moghadam et al. 2022; Zangiabadian et al. 2021). With this in mind, epitopes derived from four microorganisms, which have the capability to bind to the DRB1*03:01 molecule, as risk allele for HT, were analyzed for their similarity to epitopes derived from TPO and Tg, potential autoantigens in HT. Amino acid sequence alignment of the selected epitopes, based on their percentile rank from NetMHCIIpan-4.0, revealed a sequence homology between epitopes derived from TPO and Tg and non-self-epitopes. Comparisons of the sequence logo diagrams created by Gibbs sampling for TPO- and Tg-derived epitopes and the non-self epitopes indicated that aspartic acid is a predominant residue at the 4th, 5th, and 6th positions in both self and non-self epitopes. Meanwhile, the 1st, 2nd, and 3rd positions were primarily occupied by three amino acids: valine, leucine, and isoleucine.
The main focus of the current study was to investigate the interaction between self and non-self peptides with HLA molecules, regardless of their binding capacities for the interaction with TCR and subsequently T cell activation. The peptide binding to HLA molecules and the interaction of HLA-peptide complex with TCR is a crucial step for T cell activation. A recent study (Koukoulitsa et al. 2020) centered on the evaluation of antigenic peptides associated with MS disease has shown the conformational properties of peptides for binding to both HLA-II and TCR molecules. This interaction is orchestrated through anchor residues responsible for binding to HLA, coupled with specific residues from the peptide that play a pivotal role in facilitating TCR binding. Therefore, it can be speculated that the different peptides with sequence similarity in the critical residues may exhibit a capacity for binding to both HLA-II and TCR molecules which, in fact, reflects the cross-reactivity of different epitopes for inducing T cell responses. On the other hand, different peptides with no sequence similarity but with structural similarity in the form of peptide-MHC complex can be a cross-reactive target for specific T cells. This, in turn, indicates that both structural and sequential features of peptide-MHC complex can affect the intensity and directionality of specific or cross-reactive T cell responses (Antunes et al. 2017). These findings may shed some light on the intricate mechanisms governing the recognition of antigens and T cell activation processes during autoimmune responses induced by molecular mimicry.
Additionally, based on data extracted from UniProt, it was observed that TPO-derived epitopes, containing anchor residues L, I, and D, may be involved in binding functions, such as binding to Ca, Heme b, and proton, within this protein. Whereas, epitopes consisting of V and D are derived from the structural portion of TPO. The presence of three epitopes carrying I and D residues at critical positions in TPO protein (associated with binding sites) suggests that those positions might be more dominant to present molecular mimics compared to other residues in the epitopes of TPO. Furthermore, it is worth noting that Enteroviruses and Yersinia enterocolitica have epitopes derived from the genomic polyprotein and chaperonin GroEL, respectively. Similar to both Herpesviruses and Yersinia enterocolitica, Enteroviruses exhibit a higher number of I and D residues in their epitopes. Additionally, two types of glycoprotein epitopes consisting V, I, and D residues, which are involved in mediating herpesvirus entry into host cells and play an immunogenic role, appear to be of significant importance.
When comparing the sequence homology between the TPO protein and the proteins from the four microorganisms, it was found that only the sequence 145–180 of the envelope glycoprotein D in Herpesviruses exhibited homology with the TPO sequence 151–199. Additionally, data extracted from UniProt revealed that one of the essential receptors for the entry of Herpesviruses into human cells is the CD160 (HVEM) molecule, which binds to the envelope glycoprotein D of Herpesviruses. A study conducted by He et al. (2021) has demonstrated that polymorphisms in the CD160 receptor can influence the entry of Herpesviruses into thyroid follicles. Furthermore, an increased load of Herpesviruses within thyroid cells can enhance the presentation of epitopes from the envelope glycoprotein D via the predisposing HLA-DRB1*03:01 molecule, leading to the induction of immune responses. Consequently, due to the presence of homology between glycoprotein D and TPO, T cells activated by the envelop glycoprotein D may also recognize the TPO autoantigen as a foreign antigen, thus leading to the development of anti-TPO antibodies through a process known as molecular mimicry.
According to a study by Baker et al. (2023), specific residues within the TPO sequence 151–199, including Leu 177, Gly 194, Leu 196, Asn 198, and Gly 199, bind to the Fab region of the anti-TPO antibody’s L, H, H, L, and H chains, respectively. Based on these findings, it can be postulated that the homologous sequence 145–180 of the envelope glycoprotein D in Herpesviruses serves as a critical molecular mimic. This suggests that linear epitopes, such as IREDDQPSS and VTVDSIGML, from this protein may be presented to autoreactive T cells, leading to their activation. It is important to note that antibodies typically recognize conformational epitopes, whereas the studied epitopes in this research are linear. This brings to mind the concept of linked recognition, wherein B-lymphocytes recognize epitopes in their folded, conformational forms, and then present linear epitopes derived from the same antigen to induce T cell responses (Rastogi et al. 2022). In this context, the other linear epitopes studied may also be presented to autoreactive T cells, potentially triggering autoimmune cellular responses.
HLA-II molecules possess nine binding pockets inside the binding cleft termed pockets P1–P9 (Painter and Stern 2012). P1, P4, and P9 binding pockets have key residues for binding to anchor residues of the antigenic peptides. Key residues of HLA-DR3 molecule are Asnβ82 and Serα53 in P1, Lysβ71, Arg β74, and Asnα62 in P4 and Trpβ61 and Asnα69 in P9. Peptides bind to HLA molecules via establishing non-covalent bonds such as hydrogen and electrostatic bonds. In the HLA-DR3 molecule, the key residues inside the binding pockets are polar and positively charged amino acids, while the peptide anchor residues are polar and negatively charged that leads to a strong non-covalent binding between peptide residues and HLA pockets to increase the binding affinity of HLA molecule to the antigenic epitope. The P4 binding pocket was found to be more important than the other pockets, as the key amino acids (Lysβ71, Arg β74, and Asnα62) are located in this pocket and primarily interact with aspartic acid which is commonly found in the various studied peptides. Conversely, the key amino acids in the P1 and P9 pockets bind to different anchor residues of peptides. Any substitution with aspartic acid at this position affects the binding affinity of HLA molecule to the antigenic epitope. The Lysβ71 and Argβ74 primarily form hydrogen and electrostatic bonds with the aspartate at positions 6, 7, and 8 of the peptides which indicated the critical role of aspartate at those positions.
Based on the multiple epitope alignment, it was determined that the four microorganisms-derived epitopes (herpesvirus, enterovirus, Helicobacter pylori, and Yersinia enterocolitica), showed homology with TPO and Tg-derived epitopes. Also, based on docking, alignment epitopes indicated the capacity for binding to the predisposing HLA-DRB1*03 allele. Therefore, the observed similarities between proteins from those pathogens especially envelop glycoprotein D from Herpesviruses and TPO autoantigen might somewhat confirm the molecular mimicry hypotheses for development of HT.
Conclusion
Determining the potentially HLA class II risk alleles for developing Hashimoto’s thyroiditis as well as their predictive power for this thyroid autoimmune disease in our population were carried out. Subsequently, one of the most probable risk allele (DRB1*03:01) was considered for in silico analysis based on a hypothesis raised in connection with the role of microorganisms in the development of HT in the genetically susceptible individuals. The analysis of peptide sequence homology between epitopes of TPO and epitopes derived from four candidate microorganisms revealed a homology between envelop glycoprotein D of herpes virus and sequence 151–199 of TPO with remarkable binding capacity to HLA risk allele. These findings reinforce the hypothesis of molecular mimicry between epitopes derived from self-antigens and pathogens to induce activation of autoreactive T cells specific for thyroid antigens and consequently development of HT disease. Alternatively, it can be indicative for the increased risk of developing Hashimoto’s thyroiditis in the genetically susceptible individuals with a history of herpes virus infection as an environmental risk factor. Altogether, our findings in relation to the potential impact of molecular mimicry for developing HT disease require further investigations by recruiting a larger sample size and evaluating T cell cross-reactivity prediction to clarify the underpinning mechanisms more precisely.
Data availability
All necessary data have been included in the paper either as the main or supplementary files.
References
Al Yafei Z, Mack SJ, Alvares M, Ali BR, Afandi B, Beshyah SA, Sharma C, Osman W, Mirghani R, Nasr A, Al Remithi S, Al Jubeh J, Almawi WY, Alkaabi J, Elghazali G (2022) HLA-DRB1 and -DQB1 alleles, haplotypes and genotypes in Emirati patients with type 1 diabetes underscores the benefits of evaluating understudied populations. Front Genet 13:841879. https://doi.org/10.3389/fgene.2022.841879
Antunes DA, Rigo MM, Freitas MV, Mendes MFA, Sinigaglia M, Lizée G, Kavraki LE, Selin LK, Cornberg M, Vieira GF (2017) Interpreting T-cell cross-reactivity through structure: implications for TCR-based cancer immunotherapy. Front Immunol 8:1210. https://doi.org/10.3389/fimmu.2017.01210
Arango MT, Perricone C, Kivity S, Cipriano E, Ceccarelli F, Valesini G, Shoenfeld Y (2017) HLA-DRB1 the notorious gene in the mosaic of autoimmunity. Immunol Res 65:82–98. https://doi.org/10.1007/s12026-016-8817-7
Baker S, Miguel RN, Thomas D, Powell M, Furmaniak J, Smith BR (2023) Cryo-electron microscopy structures of human thyroid peroxidase (TPO) in complex with TPO antibodies. J Mol Endocrinol 70(3). https://doi.org/10.1530/jme-22-0149
Bodis G, Toth V, Schwarting A (2018) Role of human leukocyte antigens (HLA) in autoimmune diseases. Rheumatol Ther 5:5–20. https://doi.org/10.1007/s40744-018-0100-z
Bogers L, Kuiper KL, Smolders J, Rip J, Van Luijn MM (2023) Epstein-Barr virus and genetic risk variants as determinants of T-bet(+) b cell-driven autoimmune diseases. Immunol Lett 261:66–74. https://doi.org/10.1016/j.imlet.2023.07.006
Caturegli P, De Remigis A, Rose NR (2014) Hashimoto thyroiditis: clinical and diagnostic criteria. Autoimmun Rev 13:391–397. https://doi.org/10.1016/j.autrev.2014.01.007
Cho WK, Jung MH, Choi EJ, Choi HB, Kim TG, Suh BK (2011) Association of HLA alleles with autoimmune thyroid disease in Korean children. Horm Res Paediatr 76:328–334. https://doi.org/10.1159/000331134
Choi YM, Kim TY, Kim EY, Jang EK, Jeon MJ, Kim WG, Shong YK, Kim WB (2017) Association between thyroid autoimmunity and helicobacter pylori infection. Korean j Intern Med 32:309–313. https://doi.org/10.3904/kjim.2014.369
Cuan-Baltazar Y, Soto-Vega E (2020) Microorganisms associated to thyroid autoimmunity. Autoimmun Rev 19:102614. https://doi.org/10.1016/j.autrev.2020.102614
Enz LS, Zeis T, Schmid D, Geier F, Van Der Meer F, Steiner G, Certa U, Binder TMC, Stadelmann C, Martin R, Schaeren-Wiemers N (2020) Increased HLA-DR expression and cortical demyelination in MS links with HLA-DR15. Neurol Neuroimmunol Neuroinflamm 7(2). https://doi.org/10.1212/nxi.0000000000000656
Farid NR, Thompson C (1986) HLA and autoimmune endocrine disease 1985. Mol Biol Med 3:85–97
Figura N, Di Cairano G, Moretti E, Iacoponi F, Santucci A, Bernardini G, Gonnelli S, Giordano N, Ponzetto A (2019) Helicobacter pylori infection and autoimmune thyroid diseases: the role of virulent strains. Antibiotics (Basel) 9(1). https://doi.org/10.3390/antibiotics9010012
Ghobadi MZ, Izadi S, Teymoori-Rad M, Farahmand M, Mozhgani SH, Labbaf N, Shokri F, Marashi SM (2021) Potential role of viral infection and B cells as a linker between innate and adaptive immune response in systemic lupus erythematosus. Immunol Res 69:196–204. https://doi.org/10.1007/s12026-021-09186-4
Hammerstad SS, Jahnsen FL, Tauriainen S, Hyöty H, Paulsen T, Norheim I, Dahl-Jørgensen K (2013) Inflammation and increased myxovirus resistance protein A expression in thyroid tissue in the early stages of Hashimotoʼs thyroiditis. Thyroid 23:334–341. https://doi.org/10.1089/thy.2012.0264
Hasham A, Tomer Y (2012) Genetic and epigenetic mechanisms in thyroid autoimmunity. Immunol Res 54:204–213. https://doi.org/10.1007/s12026-012-8302-x
He W, Zhao J, Liu X, Li S, Mu K, Zhang J, Zhang JA (2021) Associations between CD160 polymorphisms and autoimmune thyroid disease: a case-control study. Bmc Endocr Disord 21:148. https://doi.org/10.1186/s12902-021-00810-w
Katahira M, Hanakita M, Ito T, Suzuki M (2013) Effect of human leukocyte antigen class II genes on Hashimotoʼs thyroiditis requiring replacement therapy with levothyroxine in the Japanese population. Hum Immunol 74:607–609. https://doi.org/10.1016/j.humimm.2013.01.026
Kokaraki G, Daniilidis M, Yiangou M, Arsenakis M, Karyotis N, Tsilipakou M, Fleva A, Gerofotis A, Karadani N, Yovos JG (2009) Major histocompatibility complex class II (DRB1*, DQA1*, and DQB1*) and DRB1*04 subtypesʼ associations of Hashimotoʼs thyroiditis in a Greek population. Tissue Antigens 73:199–205. https://doi.org/10.1111/j.1399-0039.2008.01182.x
Koukoulitsa C, Chontzopoulou E, Kiriakidi S, Tzakos AG, Mavromoustakos T (2020) A journey to the conformational analysis of T-cell epitope peptides involved in multiple sclerosis. Brain Sci 10(6). https://doi.org/10.3390/brainsci10060356
Laron Z, Shulman L, Hampe C, Blumenfeld O (2023) Hypothesis: viral infections of pregnant women may be early triggers of childhood type 1 diabetes and other autoimmune disease. J Autoimmun 135:102977. https://doi.org/10.1016/j.jaut.2022.102977
Lee HJ, Stefan-Lifshitz M, Li CW, Tomer Y (2023) Genetics and epigenetics of autoimmune thyroid diseases: translational implications. Best Pract Res Clin Endocrinol Metab 37:101661. https://doi.org/10.1016/j.beem.2022.101661
Liontiris MI, Mazokopakis EE (2017) A concise review of Hashimoto thyroiditis (HT) and the importance of iodine, selenium, vitamin D and gluten on the autoimmunity and dietary management of HT patients. Points that need more investigation. Hell j Nucl Med 20:51–56. https://doi.org/10.1967/s002449910507
Martins YC, Jurberg AD, Daniel-Ribeiro CT (2023) Visiting molecular mimicry once more: pathogenicity, virulence, and autoimmunity. Microorganisms 11(6). https://doi.org/10.3390/microorganisms11061472
Moghadam M, Ghaemi EA, Akbari H, Razavi Nikoo H, Zamani S (2022) Mycobacterium avium subsp. Paratuberculosis and Hashimotoʼs thyroiditis: is map the trigger? Front cell infect microbiol 12:972929. https://doi.org/10.3389/fcimb.2022.972929
Moradi MT, Yari K, Khodarahmi R (2014) A novel, efficient, fast and inexpensive DNA extraction protocol from whole blood applicable for studying drug-DNA interaction. J Rep Pharm Sci 3:80–84
Naito T, Okada Y (2022) HLA imputation and its application to genetic and molecular fine-mapping of the MHC region in autoimmune diseases. Semin Immunopathol 44:15–28. https://doi.org/10.1007/s00281-021-00901-9
Nekoua MP, Alidjinou EK, Hober D (2022) Persistent coxsackievirus B infection and pathogenesis of type 1 diabetes mellitus. Nat Rev Endocrinol 18:503–516. https://doi.org/10.1038/s41574-022-00688-1
Painter CA, Stern LJ (2012) Conformational variation in structures of classical and non-classical MHCII proteins and functional implications. Immunol Rev 250:144–157. https://doi.org/10.1111/imr.12003
Petrone A, Giorgi G, Mesturino CA, Capizzi M, Cascino I, Nistico L, Osborn J, Di mario U, Buzzetti R (2001) Association of DRB1*04-DQB1*0301 haplotype and lack of association of two polymorphic sites at CTLA-4 gene with Hashimotoʼs thyroiditis in an Italian population. Thyroid 11:171–175. https://doi.org/10.1089/105072501300042901
Qu HQ, Qu J, Bradfield J, Glessner J, Chang X, March M, Mentch FD, Roizen JD, Connolly JJ, Sleiman P, Hakonarson H (2021) Combined application of genetic and polygenic risk scores for type 1 diabetes risk prediction. Diabetes Obes Metab 23:2001–2003. https://doi.org/10.1111/dom.14419
Ralli M, Angeletti D, Fiore M, D’aguanno V, Lambiase A, Artico M, De Vincentiis M, Greco A (2020) Hashimotoʼs thyroiditis: an update on pathogenic mechanisms, diagnostic protocols, therapeutic strategies, and potential malignant transformation. Autoimmun Rev 19:102649. https://doi.org/10.1016/j.autrev.2020.102649
Ramasamy R, Mohammed F, Meier UC (2020) HLA DR2B-binding peptides from human endogenous retrovirus envelope, Epstein-Barr virus and brain proteins in the context of molecular mimicry in multiple sclerosis. Immunol Lett 217:15–24. https://doi.org/10.1016/j.imlet.2019.10.017
Rasouli-Saravani A, Tahamoli-Roudsari A, Basiri Z, Babaei M, Fazaeli A, Roshanaei G, Hajilooi M, Solgi G (2021) Relevance of autoantibody profile with Hla-Drb1 and -DQB1 alleles in a group of Iranian systemic lupus erythematosus patients. Immunol Lett 237:11–16. https://doi.org/10.1016/j.imlet.2021.06.004
Rastogi I, Jeon D, Moseman JE, Muralidhar A, Potluri HK, Mcneel DG (2022) Role of B cells as antigen presenting cells. Front Immunol 13:954936. https://doi.org/10.3389/fimmu.2022.954936
Schultheiss UT, Teumer A, Medici M, Li Y, Daya N, Chaker L, Homuth G, Uitterlinden AG, Nauck M, Hofman A, Selvin E, Völzke H, Peeters RP, Köttgen A (2015) A genetic risk score for thyroid peroxidase antibodies associates with clinical thyroid disease in community-based populations. J Clin Endocrinol Metab 100:e799-807. https://doi.org/10.1210/jc.2014-4352
Seyyedi N, Dehbidi GR, Karimi M, Asgari A, Esmaeili B, Zare F, Farhadi A, Dabbaghmanesh MH, Saki F, Behzad-Behbahani A (2019) Human herpesvirus 6A active infection in patients with autoimmune Hashimotoʼs thyroiditis. Braz j Infect Dis 23:435–440. https://doi.org/10.1016/j.bjid.2019.10.004
Stenszky V, Balázs C, Kraszits E, Juhasz F, Kozma L, Balázs G, Farid NR (1987) Association of goitrous autoimmune thyroiditis with HLA-DR3 in eastern hungary. J Immunogenet 14:143–148. https://doi.org/10.1111/j.1744-313x.1987.tb00374.x
Weider T, Genoni A, Broccolo F, Paulsen TH, Dahl-Jørgensen K, Toniolo A, Hammerstad SS (2022) High prevalence of common human viruses in thyroid tissue. Front Endocrinol (lausanne) 13:938633. https://doi.org/10.3389/fendo.2022.938633
Wiersinga WM (2014) Thyroid Autoimmunity Endocr Dev 26:139–157. https://doi.org/10.1159/000363161
Zangiabadian M, Mirsaeidi M, Pooyafar MH, Goudarzi M, Nasiri MJ (2021) Associations of Yersinia enterocolitica infection with autoimmune thyroid diseases: a systematic review and meta-analysis. Endocr Metab Immune Disord Drug Targets 21:682–687. https://doi.org/10.2174/1871530320666200621180515
Zeitlin AA, Heward JM, Newby PR, Carr-Smith JD, Franklyn JA, Gough SC, Simmonds MJ (2008) Analysis of HLA class II genes in Hashimotoʼs thyroiditis reveals differences compared to Gravesʼ disease. Genes Immun 9:358–363. https://doi.org/10.1038/gene.2008.26
Zhang QY, Liu W, Li L, Du WH, Zuo CL, Ye XP, Zhou Z, Yuan FF, Ma YR, Sun F, Yu SS, Xie HJ, Zhang CR, Ying YX, Yuan GY, Gao GQ, Liang J, Zhao SX, Song HD (2020) Genetic study in a large cohort supported different pathogenesis of Gravesʼ disease and Hashimotoʼs hypothyroidism. J Clin Endocrinol Metab 105(7). https://doi.org/10.1210/clinem/dgaa170
Funding
This study has received financial support from Hamadan University of Medical Sciences (Grant No. 1401120910807).
Author information
Authors and Affiliations
Contributions
GS, SB, AS, and ZR: conceptualization and methodology, funding acquisition, resources; GS, AT, and JF: data analysis and software; AS and GS: supervision, sampling, laboratory tests; GS and SB: data validation, reviewing and editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shirizadeh, A., Borzouei, S., Razavi, Z. et al. Determination of HLA class II risk alleles and prediction of self/non-self-epitopes contributing Hashimoto’s thyroiditis in a group of Iranian patients. Immunogenetics 76, 175–187 (2024). https://doi.org/10.1007/s00251-024-01339-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-024-01339-7