Abstract
Intrabreed and interbreed variation of BOLA-DRB3 exon 2 (BOLA-DRB3.2) was for the first time studied in the Kostroma and Yaroslavl cattle breeds by PCR-RFLP. These breeds are among the best Russian breeds and were developed as dairy–beef and dairy cattle, respectively. Twenty-nine alleles were observed in five Kostroma samples, and 14 of them proved unique in comparison with two Yaroslavl samples, in which 25 alleles were detected, and 10 of them were unique. The total frequency of bovine leukemia virus (BLV) resistance alleles (*11, *23, and *28) was 23.2% in the Kostroma, while the total frequency of BLV susceptibility alleles (*8, *16, *22, *24) was low, 8.4%. The frequencies were 25.8 and 30.1%, respectively, in Yaroslavl cattle. Testing Hardy–Weinberg equilibrium revealed a significant deficit of heterozygotes: the observed (Ho) and expected (He) heterozygosities were, respectively, 0.734 and 0.859 in Kostroma cattle and 0.613 and 0.886 in Yaroslavl cattle. The intrabreed differentiation (FST) in the Kostroma (4.5%, P = 0.001) was substantially higher than in the Yaroslavl (0.5%, P = 0.158), between the two breeds was 8.2% (P = 0.001). The Bayesian clustering approach showed an intrabreed structure for each of the breeds, with the most probable number of clusters being 2 in the Kostroma and 3 in the Yaroslavl. The structure observed in the Kostroma remained the same when the breed was analyzed together with six additional breeds. Our data provide important clues toward the understanding of the genetic structure of indigenous breeds.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Indigenous cattle breeds are of considerable interest to study because they have distinct adaptations to their environmental conditions (Agyemang 2005; Lazebnaya et al. 2010; Mapiye et al. 2019). The set includes climatic conditions, the epidemiological situation, specifics of the forage base, the rearing and breeding conditions, and veterinary support and depends, in particular, on economic performance. To balance the breeding for higher productivity, which is often accompanied by a decrease in health parameters, with preservation of adaptive traits, the levels of intrabreed and interbreed genetic variation are necessary to know for indigenous breeds (FAO 2010; Lazebnaya et al. 2013; Lazebnaya et al. 2018).
In addition, global climatic changes and consequent shifts in the conventional regions of known disease agents, as well as the emergence of new ones, may require the maximal readiness of the immune system. A broad allelic variation of relevant genes is a factor that ensures such adaptation (Giovambattista et al. 2013). The set includes primarily major histocompatibility complex (MHC) genes and, in particular, BOLA-DRB3 (NCBI, Gene ID: 282530, 23q21) as one of the most variable MHC class II genes (Oprzadek et al. 2018). BOLA-DRB3 is expressed in antigen-presenting cells, including B cells, dendritic cells, and macrophages. The gene codes for the DR beta chain, which interacts with the alpha chain to form a transmembrane heterodimer. An extracellular domain of the beta chain is encoded by BOLA-DRB3 exon 2 (BOLA-DRB3.2) and determines the antigen-binding specificity. Polymorphism of exon 2 is, therefore, involved in the formation of immunity (Behl et al. 2012).
A total of 136 BOLA-DRB3.2 alleles are known in cattle (https://www.ebi.ac.uk/ipd/mhc/group/BoLA/) and are detected mostly by the PCR-RFLP and PCR-SBT techniques (Takeshima et al. 2015). Particular alleles of the gene have been associated with morbidity, such as leukemia, foot-and-mouth disease, mastitis, dermatophilosis, reproductive disorders, paratuberculosis, and joint pathologies, and with resistance to ixodic tick-borne pathogens (Behl et al. 2012).
According to FAO data, breed diversity grows lower, and population sizes decrease in certain breeds and primarily the best-fit indigenous breeds (Shabtay 2015) because of various factors, including economic ones. Many such breeds are low productive and seem unprofitable to rear. The trend is observed in Russia as well. The number of Russian cattle breeds was halved over the last 20 years of the past century and was 33 by the start of the twenty-first century (Altukhov et al. 2004). The Yaroslavl and Kostroma cattle breeds are among the best Russian dairy and dairy–beef breeds, respectively. The regions of their origin, which gave names to the breeds, border each other in the central part of European Russia. Yaroslavl cows produce milk with high protein (3.4–3.6%) and high fat (4.37–5.0%) contents (FAO 1989), which are of importance for manufacturing popular dairy products, such as cottage cheese and sour cream. The Kostroma dairy–beef breed is characterized by a strong constitution, hardiness, gaining weight rapidly, and high-quality milk for cheese making (Ruzina et al. 2010). The protein and fat contents of milk are 3.6 and 3.9%, respectively (FAO 1989).
The variation of BOLA-DRB3.2 has been studied in many breeds representing both Bos indicus and B. taurus (Behl et al. 2012). The Yakut, also known as Yakutian; Kalmyk; Mongol, also known as Mongolian (Ruzina et al. 2010); Russian Black-and-White (Udina et al. 2003); and several others have been examined among Russian breeds. Some of the breeds are resistant to the bovine leukemia virus (BLV) and are included as resistant in the FAO list because anti-BLV antibodies have been detected in blood samples from animals tested by enzyme-linked immunosorbent assay (ELISA) (FAO 2007). The Kostroma cattle breed has not been included in the list, in contrast to the Yaroslavl breed, possibly because the relevant data are insufficient. However, economic losses due to BLV-induced leukemia may be substantial even when the disease incidence is low because the productivity decreases; additional veterinary care is necessary; and other animals, including the youth, may be infected. It is therefore of importance to study the intrabreed distribution of anti-BLV immunity.
The BOLA-DRB3.2 variation has already been studied in a pooled sample from two Yaroslavl herds (Mohammadabadi et al. 2004) and, with our participation, in several Kostroma samples (Sulimova et al. 2011; Sulimova et al. 2014), but the studies were not aimed at a detailed population genetic analysis of the intra- and interbreed variations of BOLA-DRB3.2 alleles or BLV resistance and susceptibility alleles in the respective breeds.
The objectives of this work were to perform a molecular genetic analysis of BOLA-DRB3.2 in a new Kostroma breed sample; to evaluate the intrabreed variation using new and our previous data; and to carry out a joint analysis of the intrabreed variation for the Kostroma and the Yaroslavl breeds, which was not examined in this respect earlier. A separate objective was to compare the two breeds with six other breeds whose genotype data were available.
Materials and methods
Sample populations and PCR-RFLP genotyping
Blood samples were collected during scheduled veterinary examinations of Kostroma cattle (cows, N = 112) at the Karavaevo breeding farm of Kostroma Oblast. Polymorphism of a 284-bp BOLA-DRB3.2 fragment, including exon 2, was analyzed by PCR-RFLP as described by Ruzina et al. (2010). To evaluate the intrabreed variation, we additionally used the genotyping data obtained previously with our participation by PCR-RFLP in Kostroma samples from other breeding farms of Kostroma Oblast: Minskoe (cows, N = 20), Gridino (cows, N = 42), Louzhky (cows, N = 56) (Sulimova et al. 2011), and Kostromskoe (bulls, N = 78) (Sulimova et al. 2014). The samples are hereafter designated by the respective farm names. The results obtained in the current study were compared with intrabreed variation estimates that we calculated for two Yaroslavl cattle samples using available genotypic data (Mohammadabadi et al. 2004). The samples were from the breeding farms Mikhailovskoe (cows, N = 44) and Gorshikha (cows, N = 49) of Yaroslavl Oblast and were examined by the PCR-RFLP method as a pooled sample by Mohammadabadi et al. (2004). After considering the intra- and interbreed variations of these breeds, the test group was expanded to include six cattle breeds: the Yakut, Mongol, Kalmyk, the Black-and-White, and Red-and-White Russian breeds, and the Golpayegani from Iran, data for which, except for the Golpayegani breed (Mosafer and Nassiry 2005), were provided by Prof. G.E. Sulimova. The sample size of the considered breeds amounted to 80, 32, 62, 61, 35, and 50, respectively. In total, genetic variation parameters were analyzed for 13 samples of eight breeds, of which only the Kostroma and Yaroslavl were each represented by more than one sample.
Statistical analyses
Results were statistically processed using the software packages GenAlEx 6.503 (http://biology-assets.anu.edu.au/GenAlEx/Download_files/GenAlEx%206.503%20Download.zip), STRUCTURE 2.3.4. (https://web.stanford.edu/group/pritchardlab/structure_software/release_versions/v2.3.4/html/structure.html), and Statistica 10.0.
Nei’s unbiased genetic distances D (Nei 1978) were calculated from the allele frequencies with GenAlEx to perform a multidimensional scaling analysis in Statistica 10.0 (StatSoft 2011). Genotype frequencies were used to perform analysis of molecular variance (AMOVA) using GenAlEx V6.503 (Peakall and Smouse 2012).
The observed (Ho) and expected (He) heterozygosities at the BOLA-DRB3.2 locus under study were estimated using the GenAlEx software for population genetic analyses. Potential deviations from the Hardy–Weinberg equilibrium (HWE) were estimated using GenAlEx for each sample and breed. The genetic structure and genetic differentiation of the breeds were assessed using standard Wright’s FST statistics and the exact G test for population differentiation. The parameters were estimated using GenAlEx. Levels of genetic differentiation between populations were described using the population pairwise FST indices and corresponding probability values for the BOLA-DRB3 gene, and represented graphically using the Heatmapper software (http://heatmapper.ca/). Note that the chart was constructed only on the basis of significant FST values.
The probability values (PFst, PG test, and PHWE) were adjusted using the Benjamini–Hochberg correction (q*) for multiple testing (Benjamini and Hochberg 1995).
The genotypes observed and the STRUCTURE 2.3.4. software (Pritchard et al. 2000) were used to carry out a model-based clustering analysis and to assign individuals to populations as described by Martínez et al. (2012). For each ancestral K value, we performed ten to 20 independent simulations, from K = 2 to K = 15, using a burn-in of 100,000 iterations and a run length of 1,000,000 iterations. The parameter alpha (degree of admixture) was inferred from the data by using the default settings and an admixture model. The method of Evanno et al. (2005) was used to determine the modal distribution of ΔK for the Kostroma and Yaroslavl breeds; and the method of Pritchard et al. (2000) was used in the cases of two and eight breeds.
Results
Analysis of the Kostroma and Yaroslavl cattle samples
Distributions of BOLA-DRB3.2 alleles
The BOLA-DRB3.2 allele frequencies in samples of the Kostroma and Yaroslavl breeds are summarized in Table 1. The number of alleles per sample varied from 13 (Minskoe) to 19 (Karavaevo) in the Kostroma samples. Alleles *10, *11, *12, and *28 were found in all samples of the breed; their total frequency ranged from 47% (Louzhky) to 61% (Karavaevo). Sample-specific alleles were observed in three Kostroma cattle samples: five in the Karavaevo sample, three alleles in the Louzhky sample, and one allele in the Gridino sample.
In the Yaroslavl breed, 21 and 17 alleles were observed in the Mikhailovskoe and Gorshikha samples, respectively (Table 1). Eight and four alleles were specific for the respective sample, while 13 alleles were common for them. The total frequencies of the common alleles were 81.8% in the Mikhailovskoe sample and 93.9% in the Gorshikha sample. Note that substantial frequencies were observed for alleles *24 and *28: 19.3 and 26.1%, respectively, in the Mikhailovskoe sample and 23.5 and 16.3%, respectively, in the Gorshikha sample.
The number of BOLA-DRB3.2 alleles with frequencies higher than 5% varied in the Kostroma samples from five (Karavaevo) to eight (Gridino and Kostromskoe) (see Supplementary Material; Table S1). Alleles *10, *11, and *28 were common for all samples in this allele group. In the Yaroslavl cattle, six and eight alleles occurred at a frequency higher than 5% in the Mikhailovskoe and Gorshikha samples, respectively. Five alleles were common for the two samples. Substantial interbreed differences were observed in this parameter. Thus, the set of alleles with frequencies higher than 5% included seven alleles in each of the breeds, but only one allele, *28, was common for the two sets. The frequency of allele *28 in the Kostroma cattle was half as high (9%) as in the Yaroslavl cattle (21%) (Table 1).
The genotype frequency distribution did not obey the HWE in the majority of the Kostroma samples. A deviation from the HWE was significant (q* = 0.04) and due to a lower heterozygote frequency (Table 1). The same pattern was observed in the Yaroslavl breed samples.
A total of 29 alleles were observed in the Kostroma cattle and 25 alleles in the Yaroslavl cattle (Table 1). Fifteen alleles were detected in both of the breeds. Of these alleles, *10 (31.8%) and *11 (10.6%) were the most frequent in the Kostroma and *28 (21%) was the most frequent in the Yaroslavl. Among the 14 alleles that were unique for the Kostroma breed, alleles *1, *7, *8, and *18 had frequencies higher than 5% (Supplementary Table S1). Among the 10 alleles specific for the Yaroslavl breed, alleles *16, *24, *40, and *44 had frequencies higher than 5%.
Frequency distributions of resistance and susceptibility alleles
We studied the distribution of BOLA-DRB3.2 alleles associated with BLV resistance or susceptibility in the two breeds (Fig. 1a, b). Alleles *11, *23, and *28 are the best-known alleles associated with BLV resistance. Of these, alleles *11 and *28 were observed in all of the Kostroma samples (Fig. 1a). Allele *23 was detected only in the Kostromskoe and Karavaevo samples. In the Yaroslavl, alleles *23 and *28 were common in the two samples. The third BLV resistance allele, *11, was additionally detected in the Mikhailovskoe sample. Note that the total frequency of the resistance alleles was relatively high in both of the breeds, 23.2% in the Kostroma and 25.8% in the Yaroslavl; in addition, a significant contribution to BLV resistance was due to alleles *11 (10.6%) and *28 (9.1%) in the Kostroma cattle and allele *28 (21%) in the Yaroslavl cattle.
Alleles *8, *16, *22, and *24 are major BLV susceptibility alleles. Two of them, *8 and *22, were observed in most of the Kostroma breed samples (Fig. 1b), besides the Minskoe and Louzhky samples, respectively. In the Yaroslavl, alleles *16, *22, and *24 out of four BLV susceptibility alleles were observed in both of the samples. The total frequencies of the susceptibility alleles were 8.4% in the Kostroma and 30.1% in the Yaroslavl.
Comparison of the BOLA-DRB3.2 genotype frequency distribution
Intrabreed and interbreed pairwise comparisons of the BOLA-DRB3.2 genotype frequency distribution were performed by the G test. The results and respective probability estimates are summarized in Supplementary Table S2. Significant G test values were obtained in the majority of cases (q* = 0.05), except for Minskoe–Gridino and Minskoe–Kostromskoe comparisons.
Genetic diversity
The expected heterozygosity He varied in the Kostroma from 0.764 (Karavaevo) to 0.875 (Gridino) (Table 2). A lack of heterozygotes was observed in four samples, except for the Minskoe sample. The He estimates obtained for the two Yaroslavl cattle samples were higher than the maximal values observed in the Kostroma samples. A lack of heterozygotes was substantial in the Gorshikha sample, where the Ho was 0.531, while Ho was 0.705 in the Mikhailovskoe sample. In general, the He values were 0.859 and 0.886 in the Kostroma and Yaroslavl cattle, respectively, while the FIS values differed threefold between the breeds (0.095 and 0.296, respectively).
Wright’s pairwise fixation index FST and AMOVA
Wright’s pairwise fixation index FST was used to study sample differentiation for the Kostroma and Yaroslavl cattle (Table 3). Significant FST values (q* < 0.04) were obtained for the majority of sample pairs, with the exception of Minskoe–Gridino (FST = 0.018, P = 0.058) and Minskoe–Kostromskoe (FST = 0.014, P = 0.108). The Minskoe–Karavaevo sample pair was the most differentiated FST = 0.059 (P = 0.001). In total, the Kostroma breed showed FST = 0.045 (P = 0.001). The Yaroslavl cattle samples showed no differentiation by FST = 0.005 (P = 0.158). Differentiation of the two breeds was observed (FST = 0.082, P = 0.001). Similar results were obtained by means of AMOVA (Fig. 2a–c).
Bayesian clustering
The population structure was modeled for the Kostroma and Yaroslavl cattle, and the two breeds were analyzed together with the use of the STRUCTURE 2.3.4 software and a model with admixture. The results are shown in Fig. 3a–c. Based on ΔK, the most reliable clustering is achieved at K = 2 (Fig. 3a). A structure shown green was predominant in four Kostroma samples (87.2–99.8%). The Louzhky sample differed from the four other samples by having a different proportion of the two structures detected. The portion of the structure shown red was 97.5% in the Louzhky sample.
Bayesian genotypic cluster analysis based on BOLA-DRB3 polymorphism for a the Kostromа, b the Yaroslavl, and c the two breeds together at various K values. Each cluster is designated with a particular color (see the description in the text). X-axis, samples. a K = 2. Kostroma samples: 1, Minskoe; 2, Gridino; 3, Kostromskoe; 4, Karavaevo; 5, Louzhky. b K = 3. Yaroslavl samples: 1, Mikhailovskoe; 2, Gorshikha. c K = 2–5. Samples: 1–5, the Kostroma samples in the same order as in a; 6, 7, the Yaroslavl samples in the same order as in b
In the Yaroslavl cattle, the most likely number of genetic structures was found to be 3 (Fig. 3b). The structures were irregularly distributed in the Mikhailovskoe and Gorshikha samples. The cluster shown red was predominant in both of the samples (62.1% in Mikhailovskoe and 48.6% in Gorshikha). The second most frequent cluster was the cluster shown green in the Mikhailovskoe sample (36.7%), and the cluster shown blue in the Gorshikha sample (35.3%). It should be noted that the within-sample cluster distribution in the Gorshikha sample was more regular than in the Mikhailovskoe sample.
The distribution of the established clusters (K = 2 ÷ 5) was studied using the pooled data on the two breeds while preserving the partitioning of individual samples (Fig. 3c). When the number of clusters was set to be K = 2, interbreed differentiation was the most distinct. The most likely number of clusters was K = 3 for the pooled sample of the two breeds. The Yaroslavl breed remained monomorphic. As for the Kostroma cattle, the Louzhky sample was separated from the other samples, as was seen in the diagram constructed for the Kostroma alone (Fig. 3a).
Analysis of 13 samples from eight cattle
To compare the variation levels observed in the Kostroma and Yaroslavl with the levels characteristic of other cattle breeds, six other breeds were added to the test group: the Yakut, Mongol, Kalmyk, Black-and-White, and Red-and-White Russian breeds, and the Golpayegani from Iran, data for which, except for the Golpayegani breed (Mosafer and Nassiry 2005), were provided by Prof. G.E. Sulimova.
Multidimensional scaling
An ordination plot (Fig. 4) was constructed for the Kostroma and Yaroslavl samples, and the six breeds additionally included in the analysis by multidimensional scaling based on Nei’s pairwise unbiased distances. The Yaroslavl samples are in the area enclosed by a red line and show almost no differentiation from each other by dimension 1, in contrast to the Kostroma samples, which are in the area enclosed by a blue line. Dimension 2 illustrates that the intrabreed variation in the Kostroma cattle (− 0.997 to − 0.288) is far greater than in the Yaroslavl cattle (0.786 to 1.095) and exceeds the interbreed differentiation level observed in the other breeds. It should be noted that the Russian Black-and-White and Kalmyk breeds are in the immediate vicinity of the Yaroslavl and Kostroma breeds on dimension 1. On the dimension 1 scale, the Kostroma and Yaroslavl samples cluster together within a group that includes the majority of the breeds, but not the Mongol and Yakut cattle. In contrast to the Mongol and Yakut breeds, the Kalmyk, which is also of a Turano-Mongolian origin, is closer to the European breeds.
Multidimensional scaling of 13 samples from eight cattle breeds based on Nei’s pairwise unbiased genetic distances. Kostroma samples: K-Minsk, Minskoe; K-Grid, Gridino; K-Kostr, Kostromskoe; K-Karav, Karavaevo; K-Louzh, Louzhky; Yaroslavl samples: Y-Mikh, Mikhailovskoe; Y-Gorsh, Gorshikha. Breeds: Golpay, Golpayegani; B-W, Black-and-White; R-W, Red-and-White. Breeds without abbreviation: Kalmyk, Mongol, Yakut
Analysis of pairwise FST values and analysis of molecular variance
Pairwise FST values based on the extended data set demonstrate high intrabreed differentiation for the Kostroma concerning the BOLA-DRB3.2 region under study. This is evident from the variation range of FST values from 0.012 in the Minskoe–Kostromskoe sample pair to 0.103 in the Minskoe–Karavaevo one (Table 4). In the Yaroslavl, there is almost no difference between the samples by FST (Table 4). The same conclusions can be made from a heat map (Fig. 5), which was based on significant (q* = 0.049) pairwise between-sample FST values. Intervals of the color scale bar indicate that intense blue corresponds to zero FST and intense yellow, to maximal FST in the heat map. A green outline isolates the area that is intense blue, reflecting the low FST values of the Yaroslavl sample pairs (Fig. 5). The Yaroslavl samples are similar in their relationships with the Kostroma samples and the other breeds, while such similarity is not observed for the Kostroma samples. Note that the Yakut breed is most strongly differentiated from the other samples and breeds except for the Mongol and, to a lesser extent, Kalmyk cattle. The most considerable difference is observed between the Yakut breed and the Karavaevo sample of the Kostroma (FST = 0.238) (Table 4).
Heat map of 13 samples from eight cattle breeds based on the significant pairwise FST values. Kostroma samples: K-Minsk or K.Minsk, Minskoe; K-Grid or K.Grid, Gridino; K-Kost or K.Kost, Kostromskoe; K-Karav or K.Karav, Karavaevo; K-Louzh or K.Louzh, Louzhky; Yaroslavl samples: Y-Mikh or Y.Mikh, Mikhailovskoe; Y-Gorsh or Y.Gorsh, Gorshikha. Breeds: Golpay, Golpayegani; B-W, Black-and-White; R-W, Red-and-White. Breeds without abbreviation: Kalmyk, Mongol, Yakut
AMOVA of the eight breeds with the Kostroma and Yaroslavl not separated into individual samples showed a high level of differentiation FST = 9% (P = 0.001).
Bayesian clustering
Figure 6 shows the results of a Bayesian clustering of the total data set at several K values (from 2 to 8). At K = 2, the Kostroma breed (a red cluster) separates from the other sample and breeds. At K = 3, the Yakut breed (a green cluster) is differentiated in addition to the Kostroma. An increase in the number of clusters to K = 4 leads the Yaroslavl forms to separate as a yellow cluster. A further increase to K = 5 leads to differentiation of the Louzhky sample in the Kostroma breed, and a second cluster (green) is observed in addition to the red one, which is major for the breed. The same structure is observed for the Kostroma at K = 6 ÷ 8.
Bayesian genotypic cluster analysis based on BOLA-DRB3 polymorphism for 13 samples of eight cattle breeds at K values varying from 2 to 8. Each cluster is designated with a particular color (see the description in the text). X-axis, samples. Kostroma samples: 1, Minskoe; 2, Gridino; 3, Kostromskoe; 4, Karavaevo; 5, Louzhky; Yaroslavl samples: 6, Mikhailovskoe; 7, Gorshikha. 8, Black-and-White; 9, Red-and-White; 10, Kalmyk; 11, Mongol; 12, Yakut; 13, Golpayegani
The most probable number of clusters was taken to be K = 7 for the set of 13 samples from the eight breeds according to the clustering algorithm used. The results obtained for the Kostroma breed agree with the results described above for the Kostroma analyzed separately (Fig. 3a), while the structure observed for the Yaroslavl matches the structure observed for the Yaroslavl when the two breeds were analyzed together (Fig. 3c).
The diagram constructed with K = 7 (Fig. 6) corresponds to the cluster distribution characterized in Supplementary Table S3. It is of interest to note that only one cluster per breed accounts for a considerable fraction in the majority of other breeds: the fractions of the red cluster are 0.667 in the Russian Black-and-White breed and 0.846 in the Russian Red-and-White breed; the sandy brown cluster is the most prevalent (0.747) in the Iranian Golpaygani, and the Yakut breed, where the magenta cluster dominates (0.997). Note that the Yakut is the most monomorphic among all breeds under study concerning the BOLA-DRB3.2 structure. The Kalmyk is a breed with a major cluster (0.418) and substantial fractions of several clusters.
Discussion
In this study, we investigated the variation of the BOLA-DRB3.2 region in 112 cows of the Kostroma breed (Karavaevo farm) and, to evaluate the intra- and interbreed variations, compared the sample with four other Kostroma samples, which have been genotyped previously with our participation, and two Yaroslavl samples, for which genotyping data were available. To compare the interbreed variation between the two breeds with levels characteristic of other breeds, six other breeds (Mongol, Yakut, Kalmyk, Iranian Golpaygani, Russian Black-and-White, and Russian Red-and-White) with available molecular genetic data were included in the analysis.
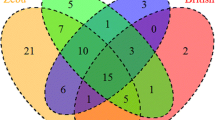
The intra- and interbreed variations with respect to the alleles detected and alleles responsible for BLV resistance or susceptibility were evaluated in the Kostroma and Yaroslavl breeds because such studies had not been performed earlier. The total number of BOLA-DRB3.2 alleles found in the Kostroma breed (29) was slightly higher than in the Yaroslavl (25). The number of common alleles and their total frequency reflected the similarity between the samples and were substantially lower in the Kostroma breed compared with those in the Yaroslavl (4 vs. 13 common alleles and 47–61% vs. 81.8–93.9% total common allele frequency, respectively).
The proviral load is known to be lower in the heterozygous animals that carry a resistance allele regardless of the nature of the other allele (a neutral or susceptibility allele) and higher in homozygotes or heterozygotes for susceptibility alleles (Miyasaka et al. 2013). Although different sets of resistance alleles (*11, *23, and *28) with different allele frequencies are responsible for BLV resistance in the Kostroma and Yaroslavl breeds, the total resistance allele frequency is similar in the two breeds (23.2 and 25.8%, respectively). Given additionally that the total allele number is lower and the total frequency (30.1%) of susceptibility alleles (*8, *16, *22, and *24) is three times higher in the Yaroslavl breed, the Kostroma may generally be somewhat superior to the Yaroslavl in terms of BLV resistance. Note that the total frequency of the above BLV resistance alleles in the additional group of the six breeds ranged from zero in the Yakut to 19.4% in the Kalmyk, thus being lower than in the Yaroslavl and Kostroma breeds. The total frequency of the susceptibility alleles ranged from zero in the Yakut breed to 30% in the Red-and-White breed. The Kostroma breed is intermediate in total susceptibility allele frequency (8.4%), and the estimate obtained for the Yaroslavl breed does not exceed the maximal total susceptibility allele frequency observed in the comparison group.
A number of population genetic parameters point to a higher level of intrabreed differences for the Kostroma cattle and a lower level for the Yaroslavl cattle, as was evident from Nei’s distance and Wright’s pairwise fixation index: D = 0.089 ÷ 1.979 и D = 0.064; FST = 0.045 и FST = 0.005, respectively. The interbreed difference between the two breeds was substantial (D = 1.309, FST = 0.082). This was similarly observed in a joint analysis of the Kostroma and Yaroslavl samples with the other breeds and reflected in the heat map based on the significant pairwise FST values. Thus, the two Yaroslavl samples, Mikhailovskoe and Gorshikha, show similar patterns of differentiation from the other breeds as a result of their high similarity to each other. Louzhki and Karavaevo are distinct among all Kostroma samples in being differentiated to a greater extent from both other Kostroma samples (FST = 0.026 ÷ 0.042 and FST = 0.020 ÷ 0.059, respectively) and the other breeds (FST = 0.032 ÷ 0.119 and FST = 0.052 ÷ 0.139, respectively). Note that FST = 0.045 established for the Kostroma breed coincides with the high level of intrabreed differentiation (0.044) that has been obtained for the Colombian Creole cattle (Hernández-Herrera et al. 2013), which is also a double-purpose breed, by the same method as in our work.
Yurchenko et al. (2018) have performed a Bayesian cluster analysis of a large group of Russian breeds on the basis of data from a genome-wide SNP study, and the differentiation pattern reported in their work is generally similar to the pattern that we established on the basis of BOLA-DRB3.2 for the Kostroma, Yaroslavl, Kalmyk, and Yakut breeds. We additionally observed differentiation of the Golpayegani and Mongol breeds, which have not been examined by Yurchenko et al. (2018). Note that only one sample was tested for each breed, including the Kostroma and Yaroslavl, and that the intrabreed variation was not evaluated in the study by Yurchenko et al. (2018). In our study, a stable intrabreed genetic structure was observed for the Kostroma breed and remained the same in its joint analyses with the Yaroslavl breed or the additional breeds. The genetic structure established for the Yaroslavl breed became monomorphic in the joint analysis with the other breeds.
Our study is the first to detect a high variation of the BOLA-DRB3.2 region in the Kostroma and Yaroslavl Russian indigenous cattle breeds. Intrabreed differentiation was found to be extremely low in the Yaroslavl and substantial in the Kostroma. The difference is possibly explained by the different breeding purposes; i.e., the former is a single-purpose breed, while the latter is a dual-purpose breed. The assumption is supported by the similar observations reported for the Holstein, one of the best international transboundary dairy breeds, in a study where the intrabreed variation of the BOLA-DRB3.2 region has been evaluated by the PCR-SBT method (Takeshima et al. 2015). High intrabreed differentiation of the Kostroma probably results from its breeding for both dairy and beef productivity traits. Our assumption is further supported by the separation of the Louzhky sample, which has been bred favoring beef traits over dairy traits, from the other Kostroma samples on evidence of Bayesian modeling and Wright’s pairwise fixation index FST. The Karavaevo Kostroma sample, which had better dairy productivity parameters as compared with the other samples of the Kostroma breed, showed a more monomorphic structure in the cluster analysis and differed to the greatest extent (FST) from the other samples and even breeds. Breeding for dairy traits and breeding for beef traits may each be to a greater extent associated with a particular set of infectious diseases and, therefore, a particular set of disease resistance alleles. In line with the idea, the spectrum and frequencies of BLV resistance alleles were similar in the Yaroslavl samples and varied among the Kostroma samples. Like in the Kostroma and Yaroslavl, we revealed genetic structure specifics detectable by clustering, in particular, in several other breeds (Yakut, Mongol, Golpayegani) included in our analysis of BOLA-DRB3.2 polymorphism.
Our finding testifies again that indigenous breeds are necessary to study comprehensively because their genetic features might be of importance for sustainable development of cattle farming when the climate changes to affect the forage base, resistance to known pathogens decreases, or resistance to new infectious agents is lacking. In addition, our data might be useful for developing programs to preserve and improve the Kostroma and Yaroslavl cattle breeds without losing BOLA-DRB3.2 allelic diversity, which contributes to the immune protection against viruses and bacteria.
Data availability
Data are presented in the manuscript.
References
Agyemang K (2005) Trypanotolerant livestock in the context of trypanosomiasis intervention strategies, vol 7. Food & Agriculture Org, Rome
Altukhov YuP, Salmenkova EA, Kurbatova OL, Pobedonostseva EYu, Politov DV, Evsyukov AN, Zhukova OV, Zakharov IA, Moiseeva IG, Stolpovsky YuA, Pukhal’skiy VA, Pomortsev AA, Upel’niek VP, Kalabushkin BA (2004) Dinamika populyatsionnykh genofondov zhivotnykh [Dynamics of population animal gene pools] Pod red. Altukhova YuP [In: Altukhov YuP (ed)] Dinamika populyatsionnykh genofondov pri antropogennykh vozdeystviyakh [The dynamics of population gene pools under human actions], Nauka Publ, Moscow, pp 110-294 (in Russian)
Behl JD, Verma NK, Tyagi N, Mishra P, Behl R, Joshi BK (2012) The major histocompatibility complex in bovines: a review. ISRN Vet Sci 2012:1–12. https://doi.org/10.5402/2012/872710
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Stat Methodol 57:289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
FAO (1989) Animal genetic resources of the USSR. In: Dmitriev NG, Ernst LK (eds) Animal production and health (issue 65). FAO, Rome
FAO (2007) Section E. Animal genetic resources and resistance to disease. In: Rischkowsky B, Pilling D (eds) The State of the World’s Animal Genetic Resources for Food and Agriculture. Food and Agriculture Organization of the United Nations, Rome
FAO (2010) In: Hiemstra SJ, de Haas Y, MaKi-Tanila A, Gandini G (eds) Local cattle breeds in Europe: development of policies and strategies for self-sustaining breeds. Wageningen Academic Pub, Wageningen
Giovambattista G, Takeshima SN, Ripoli MV, Matsumoto Y, Franco LA, Saito H, Onuma M, Aida Y (2013) Characterization of bovine MHC DRB3 diversity in Latin American Creole cattle breeds. Gene 519:150–158. https://doi.org/10.1016/j.gene.2013.01.002
Hernández-Herrera D, Posso-Terranova A, Muñoz-Florez J, Giovambattista G, Álvarez-Franco L (2013) Polymorphism of BOLA-DRB3.2* gene in creole Colombian breeds. Rev MVZ Córdoba 18:3665–3671. https://doi.org/10.21897/rmvz.133
Lazebnaya IV, Lazebny OE, Sulimova GE (2010) Study of genetic variation in Yakutian cattle (Bos taurus L.) using the prolactin bPRL, growth hormone bGH, and transcription factor bPit-1 genes. Russ J Genet 46:377–380
Lazebnaya IV, Lazebny OE, Khatami SR, Sulimova GE (2013) Use of the bovine prolactin gene (bPRL) for estimating genetic variation and milk production in aboriginal Russian breeds of Bos taurus. In: Nagy GM, Toth BE (eds) Prolactin. InTech, Rijeka, Croatia, pp 35–52. https://doi.org/10.5772/54756
Lazebnaya IV, Perchun AV, Lhasaranov BB, Lazebny OE, Stolpovskiy YA (2018) Analysis of GH1, GHR and PRL gene polymorphisms for estimation of the genetic diversity of Buryat and Altai cattle breeds. Vavilov J Genet Breed 22:734–741. https://doi.org/10.18699/VJ18.417
Mapiye C, Chikwanha OC, Chimonyo M, Dzama K (2019) Strategies for sustainable use of indigenous cattle genetic resources in Southern Africa. Diversity 11:214. https://doi.org/10.3390/d11110214
Martínez AM, Gama LT, Cañón J, Ginja C, Delgado JV, Dunner S, Landi V, Martín-Burriel I, Penedo MC, Rodellar C, Vega-Pla JL, Acosta A, Alvarez LA, Camacho E, Cortés O, Marques JR, Martínez R, Martínez RD, Melucci L, Martínez-Velázquez G, Muñoz JE, Postiglioni A, Quiroz J, Sponenberg P, Uffo O, Villalobos A, Zambrano D, Zaragoza P (2012) Genetic footprints of Iberian cattle in America 500 years after the arrival of Columbus. PLoS One 7:e49066. https://doi.org/10.1371/journal.pone.0049066
Miyasaka T, Takeshima SN, Jimba M, Matsumoto Y, Kobayashi N, Matsuhashi T, Sentsui H, Aida Y (2013) Identification of bovine leukocyte antigen class II haplotypes associated with variations in bovine leukemia virus proviral load in Japanese Black cattle. Tissue Antigens 81:72–82. https://doi.org/10.1111/tan.1204
Mohammadabadi MR, Shaikhaev GO, Sulimova GE, Rahman O, Mozafari MR (2004) Detection of bovine leukemia virus proviral DNA in Yaroslavl, Mongolian and Black Pied cattle by PCR. Cell Mol Biol Lett 9:766–768
Mosafer J, Nassiry MR (2005) Identification of bovine lymphocyte antigen DRB3.2 alleles in Iranian Golpayegani cattle by DNA test. Asian-Australas J Anim Sci 18:1691–1695. https://doi.org/10.5713/ajas.2005.1691
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 76:379–390
Oprzadek JM, Brzozowska AM, Urtnowski P, Rutkowska K, Lukaszewicz M (2018) Association of BOLA-DRB3 genotype with somatic cell count in milk of Polish Holstein cattle. R Bras Zootec 47. https://doi.org/10.1590/rbz4720150290
Peakall R, Smouse PE (2012) GenAlEx tutorials-part 2: genetic distance and analysis of molecular variance (AMOVA). Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ruzina MN, Shtyfurko TA, Mohammadabadi MR, Gendzhieva OB, Tsedev T, Sulimova GE (2010) Polymorphism of the BOLA-DRB3 gene in the Mongolian, Kalmyk, and Yakut cattle breeds. Russ J Genet 46:456–463. https://doi.org/10.1134/S1022795410040113
Shabtay A (2015) Adaptive traits of indigenous cattle breeds: the Mediterranean Baladi as a case study. Meat Sci 109:27–39. https://doi.org/10.1016/j.meatsci.2015.05.014
StatSoft, Inc. (2011) STATISTICA for Windows [Computer program manual]. StatSoft, Inc., Tulsa
Sulimova GE, Lazebnaja IV, Perchun AV, Voronkova VN, Ruzina MN, Badin GA (2011) Uniqueness of Kostroma breed of cattle from a position of molecular genetics. Adv Sci Technol Agro-Industr Complex 9:52–54
Sulimova GE, Lazebnaya IV, Ruzina MN, Belokurov SG, Perchun AV (2014) Polymorphism of the BOLA-DRB3 gene in bull sires of the Kostroma breed as genetic factor of the resistance to leukemia. J Vet Dent 6:24–27
Takeshima SN, Giovambattista G, Okimoto N, Matsumoto Y, Rogberg-Muñoz A, Acosta TJ, Onuma M, Aida Y (2015) Characterization of bovine MHC class II DRB3 diversity in South American Holstein cattle populations. Tissue Antigens 86:419–430. https://doi.org/10.1111/tan.12692
Udina IG, Karamysheva EE, Turkova SO, Orlova AR, Sulimova GE (2003) Genetic mechanisms of resistance and susceptibility to leukemia in Ayrshire and black pied cattle breeds determined by allelic distribution of gene BOLA-DRB3. Russ J Genet 39:306–317. https://doi.org/10.1023/A:1023279818867
Yurchenko A, Yudin N, Aitnazarov R, Plyusnina A, Brukhin V, Soloshenko V, Lhasaranov B, Popov R, Paronyan IA, Plemyashov KV, Larkin DM (2018) Genome-wide genotyping uncovers genetic profiles and history of the Russian cattle breeds. Heredity 20:125–137. https://doi.org/10.1038/s41437-017-0024-3
Funding
This work has been funded by government program of basic research in the Vavilov Institute of General Genetics of the Russian Academy of Sciences in 2020, 0112-2016-0003, and partly by the Russian Science Foundation under grant 19-76-20061. The work of OEL was conducted under the IDB RAS Government basic research program in 2020, № 0108-2019-0007.
Author information
Authors and Affiliations
Contributions
Irina V. Lazebnaya contributed to the study conception and design. Sample collection and genotyping were performed by Aleksey V. Perchun and Irina V. Lazebnaya. Data analysis was conducted by Irina V. Lazebnaya and Oleg E. Lazebny. The first draft of the manuscript was written by Irina V. Lazebnaya, and all of the authors commented on previous versions of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
Biological samples were collected during routine veterinary checkups in the framework of official health control programs and with the agreement of breeders.
Consent to participate
Not applicable.
Consent for publication
All authors read and approved the manuscript.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lazebnaya, I.V., Perchun, A.V. & Lazebny, O.E. Intrabreed and interbreed variation of the BOLA-DRB3.2 gene in the Kostroma and Yaroslavl indigenous Russian cattle breeds. Immunogenetics 72, 355–366 (2020). https://doi.org/10.1007/s00251-020-01173-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-020-01173-7