Abstract
A decrease in the incidence of bovine mastitis, the costliest disease in the dairy industry, can be facilitated through genetic marker-assisted selective breeding programs. Identification of genomic variants associated with mastitis resistance is an ongoing endeavor for which genome-wide association studies (GWAS) using high-density arrays provide a valuable tool. We identified single nucleotide polymorphisms (SNPs) in Holstein dairy cattle associated with mastitis resistance in a GWAS by using a high-density SNP array. Mastitis-resistant (15) and mastitis-susceptible (28) phenotypic extremes were identified from 224 lactating dairy cows on commercial dairy farm located in Utah based on multiple criteria of mastitis resistance over an 8-month period. Twenty-seven quantitative trait loci (QTLs) for mastitis resistance were identified based on 117 SNPs suggestive of genome-wide significance for mastitis resistance (p ≤ 1 × 10−4), including 10 novel QTLs. Seventeen QTLs overlapped previously reported QTLs of traits relevant to mastitis, including four QTLs for teat length. One QTL includes the RAS guanyl-releasing protein 1 gene (RASGRP1), a candidate gene for mastitis resistance. This GWAS identifies 117 candidate SNPs and 27 QTLs for mastitis resistance using a selective genotyping approach, including 10 novel QTLs. Based on overlap with previously identified QTLs, teat length appears to be an important trait in mastitis resistance. RASGRP1, overlapped by one QTL, is a candidate gene for mastitis resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mastitis, defined as inflammation of the mammary gland, is the costliest disease in the dairy industry (Kaneene and Scott Hurd 1990; Seegers et al. 2003). In the USA, the cost of bovine mastitis is estimated at a value of approximately 10% of total milk sales (Nash et al. 2000). Associated costs include loss of milk production, decreased milk quality, discarded milk, labor, veterinary treatments, mastitis-related culls, diagnostics, and preventative measures (Halasa et al. 2007).
Conventional methods to reduce the incidence of mastitis within a herd encompass both management practices and selection for mastitis-resistant phenotypes. Recent technical advancement in cattle genomics, such as genome-wide association studies (GWAS), has led to the identification of quantitative trait loci (QTLs) associated with mastitis traits (Holmbeg and Andersson-Eklund 2004; Meredith et al. 2013; Tiezzil et al. 2015). Genetic marker-assisted selection for mastitis traits provides a valuable tool for decreasing mastitis incidence, as it leads to a higher level of discrimination between phenotypes and a greater uniformity than does conventional selection (Kühn et al. 2008). Genome-wide association studies are well-suited to identifying genetic markers of complex traits such as mastitis, enabling genotyping of large numbers of potential genetic markers, such single nucleotide polymorphisms (SNPs), across the genome (Ziegler et al. 2008; Bush and Moore 2012). Indeed, GWAS carried out over the past several years have identified genetic markers, candidate genes, and QTLs for individual mastitis traits such as somatic cell count (SCC), somatic cells score (SCS) and clinical mastitis (Sodeland et al. 2011; Meredith et al. 2013; Wu et al. 2015). Many of these studies use low- or medium-marker density arrays to detect genetic markers (Sodeland et al. 2011; Sahana et al. 2013; Tiezzil et al. 2015). High-density bovine arrays capable of genotyping close to one million SNPs are available for cattle and offer the advantage of increased genomic coverage and statistical power (Wu et al. 2015). Studies using such high-density arrays have the potential to identify novel genetic markers as well as verify the significance of previously-identified markers.
In this study, we performed a GWAS using a high-density array to identify SNP genetic markers and define QTLs of mastitis resistance in Holstein dairy cows. We used a selective genotyping approach, identifying the most mastitis-resistant and mastitis-susceptible animals within the sample population. This approach facilitated detection of causative alleles due to an enrichment effect of these alleles among phenotypically extreme individuals (Guey et al. 2011). Phenotypic characterization was based on multiple criteria of intramammary infection status in order to achieve more accurate characterization of phenotypic extremes of mastitis resistance and mastitis susceptibility than could be achieved with use of a single measure of mastitis alone.
Methods
Selection of phenotypically extreme cattle
Cattle used in the study were adult lactating Holstein cattle from a single farm, and phenotypically extreme individuals of mastitis resistance and mastitis susceptibility were identified and selected for genotyping. Phenotypic characterization was based on a combination of milk bacterial culture, observation for clinical mastitis, and SCC evaluation over an eight-month period. Subclinical mastitis was defined as cases in which intramammary infection was detected by bacterial culture of milk but no changes were detected in the appearance of the mammary gland or milk. Clinical mastitis was defined as intramammary infection accompanied by clinically detectable inflammatory changes in the mammary gland and/or changes in the consistency or color of the milk.
To detect clinical and subclinical mastitis, bacterial cultures were performed by using aseptically collected milk samples. During one milking per month, composite milk samples were collected from all lactating cows that had no evidence of clinical mastitis. Clinical mastitis was monitored during that milking and, additionally, at bi-monthly evaluations specifically for clinical mastitis during another milking each month. Clinical mastitis examination was carried out by veterinarians and assistants trained by veterinarians along with continuous monitoring by farm personnel. Clinical veterinary examinations consisted of careful visual and tactile inspection of all mammary gland quarters for alterations in color, consistency, and temperature, and visual inspection of milk from all quarters for alterations in color or consistency. When mammary gland or milk abnormality (clinical mastitis) was detected during the monthly sampling of all cows, at the bi-monthly clinical mastitis examinations, or by farm personnel at any other milking time, milk samples were collected from affected quarters for bacterial culture. A composite sample was also collected from the remaining unaffected quarters. Milk microbial culture was carried out according to the guidelines outlined by the National Mastitis Council (1999). Isolation of at least one bacterial colony from a 0.01-ml inoculum of a single quarter or composite milk culture sample was considered sufficient to diagnose intramammary infection, as proposed for individual quarter samples by the Mastitis Research Workers (Dohoo et al. 2011). Composite milk sample cultures have sensitivity of 72%, specificity of 81% (Souza et al. 2016), and positive and negative predictive values of 88.2 to 100% (Reyher and Dohoo 2011) for most mastitis pathogens, when individual quarter samples are considered a “gold standard.” Monthly SCCs (< 250,000 cells/ml) were used as supplementary evidence for the absence of intramammary infection in animals from which no bacteria were isolated from milk samples and no clinical mastitis was detected. Monthly SCC measures were obtained from the Dairy Herd Improvement Association.
Criteria for classification as mastitis-resistant included an absence of clinical mastitis, an absence of bacteria cultured from composite milk samples throughout the 8-month period, and consistently low SCCs (< 250,000 cells/ml). The criterion for classification as mastitis-susceptible was the detection of at least four cases of mastitis. Cases of mastitis were defined by isolation of one or more mastitis pathogens from a composite or individual quarter milk sample and/or detection of clinical mastitis. Isolates from more than one quarter on one date could contribute as many mastitis cases as there were culture-positive quarters. Clinical mastitis detected in more than one quarter on one date contributed as many mastitis cases as there were clinically mastitic quarters. All three methods of mastitis detection were applied to all cows in the study. Animals were classified as mastitis-resistant only if no indications of mastitis by any of the three methods were observed, while animals were classified as mastitis-susceptible if four cases of mastitis were detected clinically or by milk bacterial culture alone or in combination.
DNA isolation
Genomic DNA of cows characterized as mastitis-resistant or mastitis-susceptible was isolated from ear notches or hair follicles. Isolation and purification of DNA was carried out using the Gentra Puregene Tissue Kit (Qiagen, Valencia, CA) according to the manufacturer’s instructions.
SNP genotyping
Genotype calling was carried out by the Core Facility at the University of Utah for SNP genotyping using the Illumina BovineHD BeadChip (part no. WG-450-1002; Illumina Inc., San Diego, CA), an array with 777,962 SNPs that uniformly span the entire bovine genome. Bead chips were processed according to the Infinium protocol from Illumina, and scanning was carried out by the iScan scanner (Illumina Inc., San Diego, CA). Quality control measures included removal of animals with low call rates (< 96%), SNPs with low call rates (< 0.95), and SNPs with low minor allele frequencies (< 5%). After quality control and allele frequency filtering, a total of 585,949 SNPs were used for association testing.
Statistical analysis
Significant associations between SNPs and mastitis resistance were detected using a single locus mixed model approach as implemented by the SNP and Variation Suite software (SVS version 8.4, Golden Helix, Bozeman, MT). Familial relatedness was corrected for as a random effect by incorporation of a genomic best linear unbiased prediction (gBLUP) kinship matrix (Clark and van der Werf 2013) into the model, constructed from genome-wide SNPs after pruning for linkage disequilibrium (LD). Genome-wide association mapping used a mixed linear model analysis (Segura et al. 2012) based on the gBLUP matrix to correct for cryptic relatedness, with mastitis resistance/susceptibility coded as a binary phenotype. A genome-wide suggestive threshold was set at an uncorrected p value of p ≤ 1 × 10−4, with p ≤ 1 × 10−3 considered nominal. A genome-wide significance threshold was set at an uncorrected p value of p ≤ 7.65 × 10−7 (−log10[p value] ≥ 6.12), determined empirically using the simpleM method (Gao et al. 2010) to calculate the effective number of independent tests (= 65,386) after adjusting for linkage disequilibrium.
Defining QTLs
Quantitative trait loci were defined as described previously (Meredith et al. 2013). A QTL surrounding each SNP detected as significant (p ≤ 1 × 10−4) was defined based on local LD structure. Pairwise LD between the target SNP and all individual genotyped SNPs within 1 Mb upstream and downstream was calculated using PLINK (Purcell et al. 2007). Within this region, visualized using the ggplot function of the R Studio statistical package (R Core Team 2016), the furthest upstream and downstream SNPs in strong LD with the target SNP (r2 ≥ 0.8) were used to define QTLs. Quantitative trait loci comprised of a single SNP only were excluded. Overlapping QTLs were combined into a single QTL, defined by the furthest upstream and downstream SNPs for the combined region. Once defined, QTLs were used to query the bovine genome (Bos_taurus_3.1.1/bosTau8 assembly (Zimin et al. 2009)) using the University of California Santa Cruz Genome Browser tool (https://genome.ucsc.edu/) to identify genes overlapping these regions. These QTLs were checked for overlap with known bovine QTLs using the cattle QTL database (http://animalgenome.org/cgi-bin/QTLdb/BT/index) as of April 2017 (Hu et al. 2013).
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Results
Sample population
Among cows in a commercial dairy herd of 224 lactating Holstein cows, 15 animals were characterized as mastitis-resistant (Table 1) and 28 animals as mastitis-susceptible (Table 2). All mastitis-susceptible cows with the exception of one had four confirmed cases of mastitis. One cow had three cases of mastitis confirmed by isolation of three separate pathogens, and one tentative case where sample contamination precluded definitive pathogen isolation. Cattle within the mastitis-resistant group ranged from second to sixth lactation, and cattle within the mastitis-susceptible group ranged from first to sixth lactation. Among the mastitis-resistant group, individual composite milk SCCs over the 8-month period ranged from 5000 to 220,000 cells/ml, with an average of 56,300 cells/ml. Among the mastitis-susceptible group, the number of clinical mastitis cases ranged from none to three. Individual quarter and composite milk SCCs ranged from 6000 to 2,676,000 cells/ml, with an average of 303,000 cells/ml. Commonly isolated bacterial species from composite and individual quarter milk samples from mastitis-susceptible cattle included coagulase-negative staphylococci, Streptococcus sp., Corynebacterium sp., and Escherichia coli (Table 2). Among cows classified as mastitis-susceptible, all individual clinical mastitis cases occurred within a single quarter on a given date; there were no instances of two different quarters with clinical mastitis at the same time.
Genome-wide associations
In order to identify SNP genetic markers and QTLs of mastitis resistance, we carried out a GWAS using a selective genotyping approach on the 15 mastitis-resistant cows and the 28 mastitis-susceptible cows.
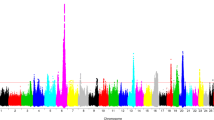
Following data quality control measures, 585,949 SNPs remained for association testing using data from 43 animals (28 mastitis-susceptible, 15 mastitis-resistant), represented in Fig. 1a as a Manhattan plot. Based on deviation from a linear relationship between observed and expected p values, as illustrated in Fig. 1b in a quantile-quantile plot, a p value threshold of p ≤ 1 × 10−4 was set as suggestive of genome-wide significance and a p value of p > 1 × 10−4, ≤ 1 × 10−3 was considered nominal for association. One SNP on chromosome 7, rs43503386 (−log10[p value] = 6.33), exceeded the genome-wide significant threshold of p ≤ 7.65 × 10−7 (−log10[p value] ≥ 6.12), and 116 SNPs were suggestive of genome-wide significance (Table 3).
a The genome-wide significance threshold is indicated by the solid line (p < 0.0001). Bovine chromosome position is shown on the x-axis. Strength of association for a single-locus mixed model GWAS is shown on the y-axis. Manhattan plot of genome-wide associations for mastitis resistance in 43 Holstein cows. b Quantile-quantile plot of observed and expected p values
Based on the 116 SNPs suggestive of genome-wide significance, we identified 27 QTLs of mastitis resistance, distributed across 14 chromosomes (2, 3, 7, 8, 10, 11, 12, 15, 16, 17, 18, 26, 27, and 28) and overlapping a total of 29 genes (Table 4). Of these QTLs, 10 have not been reported previously. Although SNP rs43503386 exceeded the genome-wide significance threshold, it was not in strong LD (r2 was < 0.8) with any other genotyped SNPs in this study and was therefore not considered in the QTL analysis. This SNP is located 72 kb upstream of the casein kinase 1 gamma 3 (CSNK1G3) gene, involved in post-translational processing of milk casein and other acidic proteins (Buitenhuis et al. 2016).
The three QTLs most highly suggestive of genome-wide significance (−log10(p value) ≥ 5.41) are located on Bos taurus autosome (BTA) 26 and overlap the sortilin-related VSP10 domain containing receptor 3 (SORCS3) gene as well as a previously identified QTL for teat length. The SORCS3 gene has no known function in bovine mastitis. Another QTL suggestive of genome-wide significance overlaps the RAS guanyl-releasing protein 1 (RASGRP1) gene, a candidate gene for mastitis resistance. Seven hundred sixty-three SNPs were nominal for genome-wide significance (Online Resource 1), distributed across all autosomal and the X chromosome.
Seventeen of the QTLs we identified overlap with previously identified QTLs of mastitis traits (somatic cell score, SCC, and clinical mastitis) and/or udder conformation traits (teat length, teat number, udder attachment, and udder depth; Table 5). Our findings reinforce the discovery of these 17 QTLs and provide supporting evidence that these QTLs may influence mastitis resistance. The top three QTLs overlap with a known QTL for teat length, which may provide the basis of mastitis resistance at these regions.
Discussion
We carried out a GWAS using a selective genotyping approach and a high-density bovine SNP array and identified 117 SNPs suggestive of genome-wide association for mastitis resistance in Holstein dairy cattle. Based on these 117 SNPs, we identified 27 QTLs of mastitis resistance, including 10 novel QTLs.
The RASGRP1 gene is located within a QTL we identified on BTA10, defined by eight SNPs genotyped in our GWAS. The RASGRP1 gene is involved in the regulation of lymphocyte development, activation, and function and in T cell receptor signaling (Bonnefont et al. 2011). Differential expression of RASGRP1 as a result of pathogen challenge occurs in primary bMECs (Brand et al. 2011) and in ovine milk somatic cells (Bonnefont et al. 2011), indicating a potential role in mastitis in ruminants. Overlap of the RASGRP1 gene by one of the QTLs indicates this gene as a strong candidate for mastitis resistance, warranting further investigation.
In dairy cattle, both immune functions and udder conformation traits are recognized factors affecting mastitis resistance (Ashwell et al. 2005). Udder attachment and udder depth have been associated previously with SCC and clinical mastitis (Seykora and McDaniel 1986; Rupp and Boichard 2003). Teat placement has been associated with SCC (Seykora and McDaniel 1986), and various studies show conflicting results of the association between teat length and SCC and clinical mastitis (Detilleux 2002). The presence of supernumerary teats is considered a risk factor in bovine mastitis, and their surgical removal at an early age may have a protective effect against subclinical mastitis in heifers (Santman-Berends et al. 2012). Seventeen QTLs that overlap with previously identified QTLs of udder conformation traits (teat length, teat number, udder attachment, and udder depth) as well as mastitis traits (somatic cell score, SCC, and clinical mastitis) were identified in this study. Overall, 11 of these 17 QTLs overlap with QTLs for mastitis traits, and 13 overlap with QTLs for udder conformation traits.
Ten of these 17 QTLs overlap with previously identified QTLs for teat length. Six of these, including the top three where the strongest association signals were detected overall, are located on BTA26 and overlap with a single previously identified QTL for teat length (Ashwell et al. 2005). The remaining overlap with QTLs for teat length on BTA16 (Ashwell et al. 2005), BTA18 (Schnabel et al. 2005), and BTA10 (Schnabel et al. 2005). This finding provides strong supportive evidence for an effect of teat length on bovine mastitis resistance, highlighting the importance of udder conformation traits as factors in the pathogenesis of this disease.
In this study, multiple measures were used to determine intramammary infection status over time and identify mastitis-resistant and mastitis-susceptible phenotypic extremes. The effectiveness of selection for mastitis resistance increases when more than a single trait is measured for determination of intramammary infection status. For example, the use of SCC and clinical mastitis together is approximately 20% more effective than the use of either of these traits alone in selecting for mastitis resistance (Philipsson et al. 1995; Odegård et al. 2003). The use of multiple measures to detect mastitis helps to overcome limitations of any one method. For example, patterns of bacterial shedding in milk during the course of infection may affect the sensitivity of milk bacterial culture to detect intramammary infection (Sears et al. 1990). Examination for clinical mastitis alone by definition excludes cases of subclinical mastitis, potentially excluding a substantial number of intramammary infections from being detected. Indirect measures such as SCC or its derivatives (linear score and estimated breeding values for these traits) can be influenced by a number of management and cow-dependent factors such as immune status, parity, lactation stage, diurnal variation, and sudden changes in feed or water management (Schultz 1977; Reneau et al. 1986; Harmon et al. 1994). Additionally, although low SCC is commonly accepted as indicative of an absence of intramammary infection, some studies have demonstrated low SCC as a risk factor in the subsequent development of clinical mastitis (Waage et al. 1998; Suriyasathaporn et al. 2000). The use of SCC alone to detect mastitis therefore has the potential to result in false negatives if not supplemented by additional measures.
In consideration of the above limitations, phenotypic characterization in this study was based on multiple criteria in order to accurately identify phenotypic extremes of mastitis resistance and susceptibility. Reliable determination of intramammary infection status is best achieved through a combination of SCC measurement, bacterial culture, and clinical detection (Dohoo et al. 2011), as used in this study. Regular monitoring using these three parameters facilitates detection of clinical and subclinical mastitis, including infections resulting in minor increases in SCC. Additionally, identification of the causative bacteria allows distinction between continuing and new intramammary infections, yielding a more accurate picture of the frequency of intramammary infection in individual cows (i.e., whether increased SCC or clinical mastitis over time represents an ongoing infection or multiple separate infections). As discussed above, the use of SCC alone to detect mastitis may result in false negatives and, thus, potentially misclassification of individual animals as mastitis-resistant or mastitis-susceptible. Therefore, in this study, SCC, milk bacterial culture, and screening for clinical mastitis were used in concert among all cows to minimize false classification of cattle as mastitis-resistant or mastitis-susceptible. Consistently low SCC in the face of multiple cases of mastitis detected by bacterial culture and/or clinical mastitis screening did not exclude an individual cow from being classified as mastitis susceptible, as low SCC may be a predisposing factor to the development of mastitis (Suriyasathaporn et al. 2000). All cows within the current study were within the same herd and were subjected to the same management conditions. Thus, effects of environmental variables on mastitis susceptibility are expected to be low relative to studies in which cattle from different farms and thereby under different environmental and management conditions are included. Direct (milk bacterial culture and evaluation for clinical mastitis) and indirect (SCC) measures for mastitis detection were used in the identification of mastitis-resistant and mastitis-susceptible cows and may have facilitated the identification of 10 novel QTLs of mastitis resistance in this study.
A potential limitation to the current study is the relatively small sample size. Out of 224 lactating cows, 43 were characterized as phenotypic extremes for mastitis resistance or susceptibility. In GWAS, sample size is one of the factors influencing statistical power, and sample sizes in the thousands are often used (Pearson and Manolio 2008). In this study, meticulous phenotypic characterization was chosen at the expense of large sample size in order to identify individual cattle representative of phenotypic extremes. Genotyping only the individuals that represent phenotypic extremes for a trait (no more than 20–25% of the sample population) can be used to detect QTLs for single traits among a small sample size while preserving statistical power in a selective genotyping approach (Lander and Botstein 1989; Darvasi 1997). Out of 224 cows, only the highest and lowest extremes for mastitis resistance of the population at 6.7 and 12.5%, respectively, were genotyped. The use of selective genotyping provides an enrichment effect, as causal and protective variants are more likely to be concentrated in these individuals as compared with individuals sampled randomly from the population. Thus, the power to detect causal and protective variants, particularly rare variants, is increased, although the effect size will be overestimated (Guey et al. 2011). Follow-on studies to replicate results are therefore important (Guey et al. 2011). We believe that, in addition to phenotypic characterization methods, the use of selective genotyping along with a high-density SNP array facilitated identification of the 10 novel QTLs.
Conclusions
One hundred seventeen candidate SNPs and 27 QTLs associated with mastitis resistance within a population of phenotypically well-characterized dairy cattle were identified. The three QTLs most suggestive of genome-wide significance are located on BTA26 and overlap the SORCS3 gene and a previously identified QTL for teat length. Ten of the 27 QTLs have not been reported previously, while 17 overlap previously identified QTLs for mastitis or udder conformation traits relevant to mastitis. One QTL on BTA10 overlaps the RASGRP1 gene, considered a candidate gene of mastitis resistance requiring further study. Validation of these QTLs as genetic markers of mastitis resistance in an expanded population is required.
References
Ashwell MS, Heyen DW, Weller JI, Ron M, Sonstegard TS, van Tassell CP, Lewin HA (2005) Detection of quantitative trait loci influencing conformation traits and calving ease in Holstein-Friesian cattle. J Dairy Sci 88:4111–4119. https://doi.org/10.3168/jds.S0022-0302(05)73095-2
Bonnefont CMD, Toufeer M, Caubet C, Foulon E, Tasca C, Aurel MR, Bergonier D, Boullier S, Robert-Granié C, Foucras G, Rupp R (2011) Transcriptomic analysis of milk somatic cells in mastitis resistant and susceptible sheep upon challenge with Staphylococcus epidermidis and Staphylococcus aureus. BMC Genomics 12:208. https://doi.org/10.1186/1471-2164-12-208
Brand B, Hartmann A, Repsilber D, Griesbeck-Zilch B, Wellnitz O, Kühn C, Ponsuksili S, Meyer HHD, Schwerin M (2011) Comparative expression profiling of E. coli and S. aureus inoculated primary mammary gland cells sampled from cows with different genetic predispositions for somatic cell score. Genet Sel Evol 43:24. https://doi.org/10.1186/1297-9686-43-24
Buitenhuis B, Poulsen NA, Gebreyesus G, Larsen LB (2016) Estimation of genetic parameters and detection of chromosomal regions affecting the major milk proteins and their post translational modifications in Danish Holstein and Danish Jersey cattle. BMC Genet 17:114. https://doi.org/10.1186/s12863-016-0421-2
Bush WS, Moore JH (2012) Chapter 11: genome-wide association studies. https://doi.org/10.1371/journal.pcbi.1002822
Clark SA, van der Werf J (2013) Genomic best linear unbiased prediction (gBLUP) for the estimation of genomic breeding values. pp 321–330
Darvasi A (1997) The effect of selective genotyping on QTL mapping accuracy. Mamm Genome 8:67–68. https://doi.org/10.1007/s003359900353
Detilleux JC (2002) Genetic factors affecting susceptibility of dairy cows to udder pathogens. Vet Immunol Immunopathol 88:103–110. https://doi.org/10.1016/S0165-2427(02)00138-1
Dohoo IR, Smith J, Andersen S, Kelton DF, Godden S, Mastitis Research Workers’ Conference (2011) Diagnosing intramammary infections: evaluation of definitions based on a single milk sample. J Dairy Sci 94:250–261. https://doi.org/10.3168/jds.2010-3559
Gao X, Becker LC, Becker DM, Starmer JD, Province MA (2010) Avoiding the high Bonferroni penalty in genome-wide association studies. Genet Epidemiol 34:100–105. https://doi.org/10.1002/gepi.20430
Guey LT, Kravic J, Melander O, Burtt NP, Laramie JM, Lyssenko V, Jonsson A, Lindholm E, Tuomi T, Isomaa B, Nilsson P, Almgren P, Kathiresan S, Groop L, Seymour AB, Altshuler D, Voight BF (2011) Power in the phenotypic extremes: a simulation study of power in discovery and replication of rare variants. Genet Epidemiol Swedish Res Counc (Scania Diabetes Regist) 35:236–246. https://doi.org/10.1002/gepi.20572
Halasa T, Huijps K, Østerås O, Hogeveen H (2007) Economic effects of bovine mastitis and mastitis management: a review. Vet Q 29:18–31. https://doi.org/10.1080/01652176.2007.9695224
Harmon RJ, Anderson KL, Kindahl H et al (1994) Physiology of mastitis and factors affecting somatic cell counts. J Dairy Sci 77:2103–2112. https://doi.org/10.3168/jds.S0022-0302(94)77153-8
Holmbeg M, Andersson-Eklund L (2004) Quantitative trait loci affecting health traits in Swedish dairy cattle. J Dairy Sci 87:2653–2659. https://doi.org/10.3168/jds.S0022-0302(04)73391-3
Hu ZL, Park CA, Wu XL, Reecy JM (2013) Animal QTLdb: an improved database tool for livestock animal QTL/association data dissemination in the post-genome era. Nucleic Acids Res 41:D871–D879. https://doi.org/10.1093/nar/gks1150
Kaneene JB, Scott Hurd H (1990) The national animal health monitoring system in Michigan. III. Cost estimates of selected dairy cattle diseases. Prev Vet Med 8:127–140. https://doi.org/10.1016/0167-5877(90)90006-4
Kühn C, Reinhardt F, Schwerin M (2008) Marker assisted selection of heifers improved milk somatic cell count compared to selection on conventional pedigree breeding values. Arch Tierz Dummerstorf 51:23–32
National Mastitis Council (1999) Laboratory Handbook on Bovine Mastitis, Revised. National Mastitis Council, Madison, WI
Lander ES, Botstein S (1989) Mapping Mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121:185–674. https://doi.org/10.1038/hdy.2014.4
Meredith BK, Berry DP, Kearney F, Finlay EK, Fahey AG, Bradley DG, Lynn DJ (2013) A genome-wide association study for somatic cell score using the Illumina high-density bovine beadchip identifies several novel QTL potentially related to mastitis susceptibility. Front Genet 4:229. https://doi.org/10.3389/fgene.2013.00229
Nash DL, Rogers GW, Cooper JB, Hargrove GL, Keown JF, Hansen LB (2000) Heritability of clinical mastitis incidence and relationships with sire transmitting abilities for somatic cell score, udder type traits, productive life, and protein yield. J Dairy Sci 83:2350–2360. https://doi.org/10.3168/jds.S0022-0302(00)75123-X
Odegård J, Klemetsdal G, Heringstad B et al (2003) Genetic improvement of mastitis resistance: validation of somatic cell score and clinical mastitis as selection criteria. J Dairy Sci 86:4129–4136. https://doi.org/10.3168/jds.S0022-0302(03)74027-2
Pearson TA, Manolio TA (2008) How to interpret a genome-wide association study. JAMA Stat Res Methods; Genet Genet 29929911:1335–1344. https://doi.org/10.1001/jama.299.11.1335
Philipsson J, Ral G, Berglund B (1995) Somatic cell count as a selection criterion for mastitis resistance in dairy cattle. Livest Prod Sci 41:195–200. https://doi.org/10.1016/0301-6226(94)00067-H
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, de Bakker PIW, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575. https://doi.org/10.1086/519795
R Core Team RF for SC (2016) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/
Reneau JK, Andrews RJ, Kitchen BJ et al (1986) Effective use of dairy herd improvement somatic cell counts in mastitis control. J Dairy Sci 69:1708–1720. https://doi.org/10.3168/jds.S0022-0302(86)80590-2
Reyher KK, Dohoo IR (2011) Diagnosing intramammary infections: evaluation of composite milk samples to detect intramammary infections. J Dairy Sci 94:3387–3396. https://doi.org/10.3168/jds.2010-3907
Rupp R, Boichard D (2003) Genetics of resistance to mastitis in dairy cattle. 671. Vet Res 34:671–688. https://doi.org/10.1051/vetres:2003020
Sahana G, Guldbrandtsen B, Thomsen B, Lund MS (2013) Confirmation and fine-mapping of clinical mastitis and somatic cell score QTL in Nordic Holstein cattle. Anim Genet 44:620–626. https://doi.org/10.1111/age.12053
Santman-Berends IM, Olde Riekerink RG, Sampimon OC et al (2012) Incidence of subclinical mastitis in Dutch dairy heifers in the first 100 days in lactation and associated risk factors. J Dairy Sci 95:2476–2484. https://doi.org/10.3168/jds.2011-4766
Schnabel RD, Sonstegard TS, Taylor JF, Ashwell MS (2005) Whole-genome scan to detect QTL for milk production, conformation, fertility and functional traits in two US Holstein families. Anim Genet 36:408–416. https://doi.org/10.1111/j.1365-2052.2005.01337.x
Schultz LH (1977) Somatic cells in milk-physiological aspects and relationship to amount and composition of Milk. J of’Food Prot 40:125–131
Sears PM, Smith BS, English PB, Herer PS, Gonzalez RN (1990) Shedding pattern of Staphylococcus aureus from bovine intramammary infections. J Dairy Sci 73:2785–2789. https://doi.org/10.3168/jds.S0022-0302(90)78964-3
Seegers H, Fourichon C, Beaudeau F (2003) Production effects related to mastitis and mastitis economics in dairy cattle herds. Vet Res 34:475–491
Segura V, Vilhjálmsson BJ, Platt A, Korte A, Seren Ü, Long Q, Nordborg M (2012) An efficient multi-locus mixed-model approach for genome-wide association studies in structured populations. Nat Genet 44:825–830. https://doi.org/10.1038/ng.2314
Seykora AJ, McDaniel BT (1986) Genetics statistics and relationships of teat and udder traits, somatic cell counts, and milk production. J Dairy Sci 69:2395–2407. https://doi.org/10.3168/jds.S0022-0302(86)80679-8
Sodeland M, Kent MP, Olsen HG, Opsal MA, Svendsen M, Sehested E, Hayes BJ, Lien S (2011) Quantitative trait loci for clinical mastitis on chromosomes 2, 6, 14 and 20 in Norwegian red cattle. Anim Genet 42:457–465. https://doi.org/10.1111/j.1365-2052.2010.02165.x
Souza FN, Cunha AF, Rosa DLSO, Brito MAV, Guimarães AS, Mendonça LC, Souza GN, Lage AP, Blagitz MG, Libera AMMPD, Heinemann MB, Cerqueira MMOP (2016) Somatic cell count and mastitis pathogen detection in composite and single or duplicate quarter milk samples. Pesqui Vet Bras 36:811–818. https://doi.org/10.1590/S0100-736X2016000900004
Suriyasathaporn W, Schukken YH, Nielen M, Brand A (2000) Low somatic cell count: a risk factor for subsequent clinical mastitis in a dairy herd. J Dairy Sci 83:1248–1255. https://doi.org/10.3168/jds.S0022-0302(00)74991-5
Tiezzil F, Parker-Gaddis KL, Cole JB et al (2015) A genome-wide association study for clinical mastitis in first parity US Holstein cows using single-step approach and genomic matrix re-weighting procedure. PLoS One 10:e0114919. https://doi.org/10.1371/journal.pone.0114919
Waage S, Sviland S, Ødegaard SA (1998) Identification of risk factors for clinical mastitis in dairy heifers. J Dairy Sci 81:1275–1284. https://doi.org/10.3168/jds.S0022-0302(98)75689-9
Wu X, Lund MS, Sahana G, Guldbrandtsen B, Sun D, Zhang Q, Su G (2015) Association analysis for udder health based on SNP-panel and sequence data in Danish Holsteins. Genet Sel Evol 47:50. https://doi.org/10.1186/s12711-015-0129-1
Ziegler A, König IR, Thompson JR (2008) Biostatistical aspects of genome-wide association studies. Biom J 50:8–28. https://doi.org/10.1002/bimj.200710398
Zimin AV, Delcher AL, Florea L, Kelley DR, Schatz MC, Puiu D, Hanrahan F, Pertea G, van Tassell CP, Sonstegard TS, Marçais G, Roberts M, Subramanian P, Yorke JA, Salzberg SL (2009) A whole-genome assembly of the domestic cow, Bos taurus. Genome Biol 10:R42. https://doi.org/10.1186/gb-2009-10-4-r42
Acknowledgments
The support and resources from the Center for High Performance Computing at the University of Utah are gratefully acknowledged.
Funding
Funding for sample collection and data analysis was provided by the Utah Agriculture Experiment Station (Utah State University Extension Grants Program to Zhongde Wang) and the Utah Department of Agriculture and Food (Cap Ferry Agricultural Grant Fund to Zhongde Wang). The funding body had no role in the design of the study and collection, analysis, and interpretation of data or in writing the manuscript. The computational resources used were partially funded by the NIH Shared Instrumentation Grant 1S10OD021644-01A1.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The use of animals in this study was approved by the Utah State University Institutional Animal Care and Use Committee (protocol IACUC-2282), and permission was obtained from the cattle owner. All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. All procedures performed in studies involving animals were in accordance with the ethical standards of the institution or practices at which the studies were conducted.
Conflict of interest
The authors declare that they have no conflicts of interest.
Electronic supplementary material
Online Resource 1
SNPs nominal for genome-wide significance for bovine mastitis resistance. (XLSX 43 kb)
Rights and permissions
About this article
Cite this article
Kurz, J.P., Yang, Z., Weiss, R.B. et al. A genome-wide association study for mastitis resistance in phenotypically well-characterized Holstein dairy cattle using a selective genotyping approach. Immunogenetics 71, 35–47 (2019). https://doi.org/10.1007/s00251-018-1088-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-018-1088-9