Abstract
High mountain freshwater systems are particularly sensitive to the impacts of global warming and relevant environmental changes. Microorganisms contribute substantially to biogeochemical processes, yet their distribution patterns and driving mechanism in alpine streams remain understudied. Here, we examined the bacterial and fungal community compositions in stream biofilm along the elevational gradient of 745–1874 m on Mt. Kilimanjaro and explored their alpha and beta diversity patterns and the underlying environmental drivers. We found that the species richness and evenness monotonically increased towards higher elevations for bacteria, while were non-significant for fungi. However, both bacterial and fungal communities showed consistent elevational distance-decay relationships, i.e., the dissimilarity of assemblage composition increased with greater elevational differences. Bacterial alpha diversity patterns were mainly affected by chemical variables such as total nitrogen and phosphorus, while fungi were affected by physical variables such as riparian shading and stream width. Notably, climatic variables such as mean annual temperature strongly affected the elevational succession of bacterial and fungal community compositions. Our study is the first exploration of microbial biodiversity and their underlying driving mechanisms for stream ecosystems in tropical alpine regions. Our findings provide insights on the response patterns of tropical aquatic microbial community composition and diversity under climate change.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Exploring elevational patterns of biodiversity has been one of the central research topics in ecology and biogeography [1, 2]. High mountain areas are home to aggregations of small-ranged species [3] and support approximately one third of terrestrial and aquatic biodiversity [4]. They are also natural laboratories to examine biodiversity patterns given its dramatically changes in biotic and abiotic environments within a relatively small spatial area [5]. Unlike the diversity and distribution of macroorganisms that have been studied for centuries, elevational patterns of microbial community have only been explored for less than two decades and thus still remain poorly understood [2, 6]. Compared to terrestrial environments [7, 8], aquatic microbes such as bacteria, fungi, and diatoms are still relatively understudied [2]. Therefore, the general diversity patterns and driving mechanisms remain to be explored especially for aquatic microorganisms on mountainsides.
Microbes are an important component of ecosystem in response to global change [9], and their biodiversity usually differs across regional and global scales as driven by various environmental variables [10, 11]. Biodiversity consists of two complementary facets, i.e., alpha and beta diversity, with the former usually measured by species richness and species abundance distribution, and the latter by community composition variations [12, 13]. For alpha diversity, species richness measures the total number of species at local sites, whereas species abundance distribution, often examined as evenness, measures the similarity level of species abundances, being both linked to multiple ecosystem processes and functioning [13, 14]. Microbes may not always follow the decreasing or hump-shaped elevational patterns in diversity traditionally observed in plants and animals [7, 15], but may show unexpected monotonically increasing patterns [11]. For beta diversity, it is originally proposed as the variation in species richness among communities in a geographical region [16] with additional methods developed since then, such as to disentangle the similarity decay in species composition with spatial distance [17]. Beta diversity has been recognized as an important facet to reveal the biodiversity patterns, and the ecological processes in maintaining the biodiversity [14, 18].
Local and regional environmental variables typically shape species occurrence across various spatial and temporal scales and constrain the biodiversity and community composition in freshwater ecosystems [19]. Among local environmental variables, pH and nutrient concentrations are the main factors in shaping the alpha diversity of diatom and bacteria in streams [11] or limiting the spatial distribution of bacterioplankton in lakes [20]. For regional environmental variables, the mean temperature of the warmest season or the mean precipitation of the coldest season serving as climatic factors is the best predictor of microbial alpha diversity and composition in three mountainsides streams across Eurasia [21]. Considering multiple aspects of biodiversity and their environmental drivers simultaneously may be helpful to improve our comprehension of the general biodiversity patterns and underlying mechanisms.
Here, we explored the patterns of biofilm bacterial and fungal diversity and community composition in the streams along an elevational gradient of 745–1874 m on Mt. Kilimanjaro, Tanzania. Mt. Kilimanjaro is located in a seasonally dry tropical climate zone, where climatic conditions are dominated by the Indian Ocean monsoon and vary greatly with elevation [22, 23]. This mountain is one of the world’s biodiversity hotspots and refuges for endangered species, and offers an outstanding example to investigate microbial biodiversity in African tropical habitats along elevational gradients due to its unique topographical conditions [24]. The mountain is affected by climate change and anthropogenic activities in the past decades, such as the pronounced decrease in glaciers thickness, leaving only a small portion covered by ice cap [25,26,27]. The unique microbial communities may be gradually compromised and lost with the continued recede of the glaciers on top of Mt. Kilimanjaro [28]. There is thus an urgent need to systematically study microbial diversity and ecosystem functions across the elevational gradients. In 2020, we sampled the bacteria and fungi of stream biofilm on Mt. Kilimanjaro and focused on three key objectives: (i) Are there general elevational patterns in bacterial and fungal species richness and evenness in the stream biofilm on Mt. Kilimanjaro? (ii) How do the community compositions change along the elevational gradients for bacteria and fungi? (iii) What are the key drivers among environmental factors underlying biodiversity patterns for bacteria and fungi?
Materials and Methods
Study Area and Field Sampling
Mt. Kilimanjaro, the Africa’s highest mountain, is located on the Kenya-Tanzania border, about 300 km south of the equator [28]. The nature diversity of this site is exceptional: from lowland savannah to upper icecap, a massive range of ecosystems is presented in its slopes [29]. Our study area (3° 10′ 27′′–3° 26′ 30′′ S, 37°12′ 19′′–37° 34′ 40′′ E) is located in the south slope of Mt. Kilimanjaro with an elevational range of 745–1874 m. The whole forest zone above 1700 m has been part of Kilimanjaro National Park since 2005 [30]. The region is classified as a seasonal dry tropical climate zone, with two distinct rainy seasons: long rains from March to May and short rains around November [31]. The southern slope of Mt. Kilimanjaro can be distinguished into seven vegetation zones: natural savanna woodlands (700 – 1100 m), traditional home gardens (1100 – 1700 m), lower montane forests (1700 – 2100 m), montane zone that consists of a mixture of dense natural and disturbed Ocotea forests (2100 – 2800 m), upper montane zone that consists of natural and disturbed Podocarpus forest (2800 – 3200 m), subalpine zone that consists of natural Erica trimera forest and degraded bushland (3500 – 4000 m), alpine zone dominated by several species of Helichrysum dwarf shrubs, and grasses (above 4000 m) [23, 30].
In February and March 2020, we sampled stream water to determine environmental variables, and scraped stream biofilm from stone surface for microbial analyses at each sampling location. Forty locations were selected along elevations, and eventually a total of 40 water and biofilm samples were separately collected. Water samples and stream biofilm were acquired according to the methods reported in previous literature [32]. Briefly, each site was divided into five or ten transects depending on the width of streams. Ten stones were randomly selected from the shallow/stream habitats along the transects, and biofilms were obtained by scarping the upside surfaces (~9 cm2 per stone) of each stone with a sterile sponge. The biofilms were collected into a composite sample as the microbial sample from each site. Water and biofilm samples were immediately stored in mobile refrigerator at −20 °C on site and were kept at −20 °C in the laboratory until the analyses of chemical variables and microbial communities.
Physicochemical and Climatic Variables
The latitude, longitude, and elevation of each sampling point were recorded using a GPS device. Physical factors of streams, including water depth and width, velocity, shading (% canopy coverage), temperature, and stone size, were determined at each sample site as shown in previous literature [32]. Briefly, water electrical conductivity and pH were determined on site by using mobile pH meters (Sanxin, China). Nitrogen oxides (NOx−-N), nitrite (NO2−-N), ammonium (NH4+-N), and phosphate (PO43−-P) were measured using a flow injection analyzer (Skalar SA1000). Total nitrogen (TN) and total phosphorus (TP) were measured using peroxodisulfate oxidation with the spectrophotometric method [33]. Dissolved organic matter (DOM) is a diverse and complex blend of labile and persistent ingredient that actively governs the biogeochemical cycles of ecosystems [34]. The availability of resources, especially nutrients and carbon, shapes the structure and function of microbial communities. UV-vis absorption spectroscopy is a widely used tool to effectively investigate the quantity and quality of light absorbing compounds in DOM (i.e., chromophoric DOM, cDOM) [35]. Therefore, the water samples were scanned in the wavelength ranging from 200 to 800 nm with a UV-visible spectrophotometer (UV2700, Shimadzu) to obtain the absorbance data for cDOM. Mili-Q water was used as the blank. The fluorescence excitation-emission matrices (EEMs) offer a plethora of information about various fluorophores. The EEMs fluorescence spectroscopy was generated with Mili-Q water as the background using a fluorescence spectrophotometer (Fluorolog⁃3, Horiba). Scans were conducted at excitation wavelengths in the range of 250–450 nm at 5 nm intervals and emission wavelengths in 250–580 nm at 1 nm intervals [36].
Climatic variables were extracted using the package “raster” V3.6-3 from “WordClim” website, which is a database of globally interpolated climate data at a high spatial resolution (approximately 1 km2) [37, 38]. These variables are derived from the monthly temperature and rainfall values, and represent annual trends, seasonality, and extreme or limiting environmental factors [39].
Bacteria and Fungi Community Analysis
Genomic DNA was extracted from stream biofilm using MoBio PowerSoil DNA Isolation Kit (MoBio, Carlsbad, USA). The 16S rRNA genes of bacteria were amplified using universal primers [515F, 5′-GTG YCA GCM GCC GCG GTA A-3′ and 806R, 5′-GGA CTA CNV GGG TWT CTA AT-3′] targeting the V4 region [40]. For fungi, amplicon of ITS2-2 region was targeted using the primers [gITS7F, 5′-GTG ART CAT CGA RTC TTT G-3′ and ITS4R, 5′-TCC TCC GCT TAT TGA TAT GC-3′] [14]. We obtained amplification products and normalized them at equal molar concentration, and then paired-end sequenced (2 × 250bp) on the Illumina NovaSeq 6000 sequencing platform. Before sequence analysis processes, quality control was performed to select high quality sequences for bioinformatic and statistical analyses. Briefly, any reads that are shorter than 200 bp were excluded, while those with over 40% unqualified bases were discarded. Sequencing data for each bacterial or fungal sample were included in an individual fastq file. The raw sequence data were obtained after removing the barcode sequence and divided it into paired-end read files. Raw sequence data were firstly removed potential primers from the obtained 16S rRNA and ITS2-2 using qiime2-cutadapt in QIIME2[41, 42], and then were denoised and quality control using DADA2 for obtaining high-quality amplicon sequence variants (ASVs). The taxonomic assignment of each ASV for both bacteria and fungi was performed using the q2-feature-classifier, by comparing to the databases of SILVA 138 and UNITE (99% threshold levels, release 10.05.2021) [43], respectively. The fragments that could not be mapped were defined as unknown sequences. ASV tables were then generated based on the mapped sequences. All above analyses were performed in QIIME2 software (version 2022.8) [44]. The mapped ASV results for each dataset in USEARCH tabular format were imported directly into the R software and merged into the integrated ASV tables for further analysis. Our study was currently focused on bacteria and fungi, and the ASVs designated as “mitochondria”, “chloroplasts”, “archaea”, and “eukaryotes” were removed from the bacterial communities before further analyses. The sequences were rarefied to 30,007 and 51,166 per sample for bacteria (30,007–73,717 sequences) and fungi (51,166–106,161 sequences), respectively, using the “rrarefy” function in R package “vegan” V 2.6-2.
Statistical Analyses
The stream chromophoric dissolved organic matter (cDOM) spectroscopic data of absorbance and fluorescence were analyzed using the R package “staRdom” V1.1.25. Briefly, we used “staRdom” to perform parallel factor analysis (PARAFAC) of excitation-emission matrices (EEMs) [45], including “peak picking” of EEM. Fluorescence indices, absorbance indices and absorbance slope indices were calculated from EEMs and absorbance spectra [46]. The indices of cDOM included classical peaks based on manual peak picking: B (tyrosine-like), T (tryptophan-like), A (humic-like), M (marine humic-like), C (humic-like), which are proxies for tyrosine-like, tryptophan-like, UV humic-like, microbial-derived and visible humic-like DOM, respectively [47, 48]. Other indices were also included to reflect various aspects of DOM: Humification index (HIX) ranges from 0 to 1, and higher values mean greater degree of DOM humification [49]. Autochthonous productivity index (BIX) has values < 1 suggesting the presence of autochthonous and fresh DOM, and < 0.6 indicating low or zero autochthonous DOM production [50]. Fluorescence index (FI) is proxy of the DOM origin, with high values suggesting the prevalence of stream-produced DOM [51]. Absorbance coefficient at 254 nm (a254) is used to indicate the concentration of cDOM [52]. Spectral slope (S275–295) is a reliable proxy of and negatively correlated with cDOM average molecular weight (MW) [53]. More details of these indices could be found in the Supporting Information (Tables S2 - S4). A four-component model was validated via split-half test and random initialization analysis along with the residuals analysis in our study, with the components being identified according to the shapes and locations of the spectral peaks [46, 54, 55]. The components identified by PARAFAC were compared with previous studies based on their Ex/Em maxima online at the “OpenFluor” database [35, 56].
We firstly examined the elevational patterns of environmental variables through linear or quadratic model based on lower value of Akaike’s information criterion (AIC) [57]. We grouped the environmental variables into four categories: that is, climatic, physical, chemical, and cDOM. The variables for these categories were: for climatic, including mean annual temperature and precipitation; for physical, including water temperature, speed, stream width, water depth, shading, and stone size; for chemical, including pH, conductivity, TN, TP, NOx−-N, NO2−-N, NH4+-N, and PO43−-P; for cDOM, including B, T, A, M, C, HIX, BIX, FI, a254, and S275–295. Pearson correlations were used to determine the covariation among environmental variables, and the highly correlated variables (|r| > 0.75) were excluded to minimize collinearity in the subsequent analyses.
The metrics of species richness and evenness were calculated to characterize the two aspects of bacterial or fungal alpha diversity using the R package “vegan” V 2.6-2 [58]. Chao1 is a good measure for species richness, and Pielou’s evenness for distribution of relative abundance in a community [59]. Pielou’s evenness was calculated by J = H/log (S), where H is the Shannon-Weaver diversity index and S is the number of species [60]. The relationship between elevation and Chao1 or evenness was explored with linear or quadratic model. The better fitting model was selected based on lower value of AIC. The elevational patterns of dominant phyla/genera were also explored using a similar approach. Nonmetric multidimensional scaling (nMDS) based on the pairwise Bray-Curtis dissimilarity was carried out using the “metaMDS” function to visualized the variation in the community composition of bacteria and fungi across elevations [61, 62]. The distance-decay relationships between Bray-Curtis dissimilarity and geographical distance were estimated for bacteria or fungi [63]. In addition, we calculated the Spearman correlation between environmental factors and the relative abundance of dominant phyla using “corr.test” function. The bacterial and fungal ASVs with the threshold of incidence > 50% were separately selected, and then Spearman correlations between these ASVs and the environmental variables were calculated using “corr.test” function.
Mantel test with Spearman’s rank correlation [64] was deployed to determine the impacts of environmental variables on the bacterial and fungal community. We also calculated the Spearman’s rank correlation between environmental variables and microbial biodiversity by considering species richness and the community composition represented by nMDS1, that is, the first axis of the nonmetric multidimensional scaling (nMDS). Random Forest modeling was then used to predict the importance of the climatic and physicochemical factors on the species richness, evenness, and community composition of bacteria and fungi. Model cross-validation for random forests was performed using 75% of the dataset for training, with the remainder validated at each resampling iteration. We trained the model using 500 trees and sampled 2 predictors per node for splitting selected to minimize the root-mean-squares (measures of variance, RMSE) [65]. The R packages included “vegan” V2.6-4, “raster” V3.6-23 [66], “FactoMineR” V2.7 [67], “caret” V6.0-93 [68], and “randomForest” V4.7-1.1 [69]. Finally, the variation of microbial biodiversity was partitioned into pure effects of the four environmental categories (that is, climatic, physical, chemical, and cDOM), and their joint effects using variation partitioning analysis [70]. We used forward selection model to identify a best explained subset of environmental variables to avoid overloading the model, by using “ordiR2step” function in R package “vegan” V 2.6-2 [71]. Variation partitioning was conducted using the “varpart” function in the R package “vegan” V 2.6-2. All statistical analyses were performed in R version 4.2.1 unless otherwise indicated.
Results
Climate and Environmental Variables along Elevation Gradients
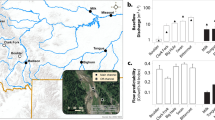
There were considerably variations of the measured environmental variables across the 40 sampling sites, such as temperature, stream morphology, and nutrient conditions (Table S1). For instance, water temperature ranged from 16.50 to 28.20 °C (SD = 2.83), conductivity from 16 to 251 mS m−1 (SD = 47.68), total nitrogen from 0.23 to 1.68 mg L−1 (SD = 0.44), and total phosphorus from 0.03 to 0.15 (SD = 0.03). There were generally significant (P < 0.05) elevational patterns for more than half of the climate and environmental variables (Figs. 1, S1). For instance, phosphate and nitrite showed a significant decreasing pattern towards higher elevations, whereas total nitrogen and nitrogen oxides first increased slightly then decreased towards higher elevations (Figs. 1, S1).
Map of stream sampling sites on the Mt. Kilimanjaro, Africa, and the elevational trends of partly main climatic and local variables The lines were fitted with linear or quadratic models. The model was chosen based on the lower value of Akaike’s information criterion. The significant (P < 0.05) models are shown as solid lines, and nonsignificant trends (P > 0.05) are shown as dotted lines
For cDOM, we found that humic substances (A, C), tyrosine (B) and absorption coefficient (a254) showed significantly decreasing elevational patterns (Figs. 1, S1). Across the 40 sampling sites, HIX value ranged from 0.09 to 0.45, and only five samples had BIX values larger than 1, which indicates that most of DOM was input from surrounding terrestrial lands and showed a lower degree of humification. This is because HIX values ranged from 0 to 1 with increasing degree of humification [49], and high BIX values (>1) suggest the presence of autochthonous and fresh DOM [72] (Table S2).
The Structure and Composition of Bacteria and Fungi
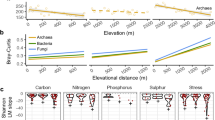
The bacterial and fungal groups dominant at different taxonomic levels, i.e., phylum and genus, demonstrated multiple elevational patterns which illustrated the strong influence of elevation on community (Figs. 2, S2). For bacteria, the most dominant phylum was Proteobacteria, which accounted for 56.9% of the bacterial community, followed by Cyanobacteria (25.61%) and Deinococcota (4.16%). The relative abundance of Proteobacteria varied from 28.15 to 66.67% in the bacterial communities across the 40 sampling sites, but showed a nonsignificant elevational pattern (P > 0.05). Four dominant phyla showed significant elevational trends (P < 0.01, Fig. S2), such as a decreasing trend for Deinococcota, while U-shaped trends for Acidobacteriota, Planctomycetota, and Actinobacteriota. Although Proteobacteria was the most dominant bacterial phylum, the genera belonging to the Cyanobacteria occupied the top four positions in the dominant genera. The genera Chroococcopsis showed a significantly hump-shaped elevational pattern (P < 0.05), while Pleurocapsa_PCC−7319, Chamaesiphon_PCC−7430, and SpeB-3 were nonsignificant (P > 0.05, Fig. 2). For fungi, the most dominant phylum was Ascomycota (34.86%), followed by Basidiomycota (4.02%). The proportion of Ascomycota fluctuated between 14.49% and 66.01% in fungal communities across 40 sampling sites, with a nonsignificant elevational pattern (Fig. S2). Except for Rozellomycota and Mortierellomycota, the other dominant fungal phyla showed non-significant elevational trends. The most dominant genera for fungi all belong to the members of Ascomycota (Fig. 2). The drivers for the dominant phyla of bacteria and fungi differed greatly, such as Ascomycota primarily driven by water depth, whereas Cyanobacteria by total phosphorus (Fig. S4).
The elevational trends of the relative abundance of the top 10 genera for bacteria (a) and fungi (b) The lines were fitted with linear or quadratic models. The model was chosen based on the lower value of Akaike’s information criterion. The significant (P < 0.05) models are shown as solid lines, and nonsignificant trends (P > 0.05) are shown as dotted lines
At the ASV level, the Spearman correlations between the relative abundance of microbial ASVs with incidence > 50% and the environmental variables were mainly positive, which was particularly true for fungi with over 62.30% ASV showing positive correlations (Figs. S5, S6). Further analysis revealed that negative correlations were prevalent between elevation and the dominant ASVs for bacteria and fungi, e.g., ASVs belonging to the genera Chroococcopsis and Truepera in the bacterial community, and Penicillium and Pyrenochaetopsis in the fungal community, whereas positive correlations prevailing with other environmental factors (Fig. S7).
Bacteria and Fungi Diversity Patterns
The elevational trends of species richness and evenness showed different elevational trends between bacteria and fungi (Fig. 3). The bacteria had significant, monotonically increasing elevational patterns in both species richness and evenness (R2 = 0.16, P < 0.01; R2 = 0.09, P < 0.05), while non-significant elevational patterns for fungi. There were significant correlations between species richness and evenness for both bacteria and fungi (Fig. S7).
Elevational alpha diversity patterns for bacteria (a–b) and fungi (c–d) For each group, we considered two indicators: Chao1 (a, c) and Evenness (b, d). The relationships between elevation and alpha diversity were modeled by linear models. The significant (P < 0.05) models are shown as solid lines, and nonsignificant trends (P > 0.05) are shown as dotted lines
The bacterial or fungal communities were more similar at lower elevations than those at higher elevations indicated by Bray-Curtis dissimilarity (Fig. 4). Specifically, both microbial groups showed more pronounced separation toward higher elevations in nMDS plots, implying higher microbial community dissimilarity. The community compositional dissimilarity significantly increased with larger elevational distance for bacteria and fungi with Mantel’s r of 0.35 and 0.23, respectively (P = 0.0001 and 0.0004), indicating there were predictable turnover patterns of microbial assemblage composition along the elevational gradients (that is, elevational distance-decay relationship).
The variation of community composition along the elevational gradient The nMDS of bacterial (a) and fungal (c) communities and the elevations of the sample points were distinguished by color. The relationships between community dissimilarity and elevational change for bacteria (b) and fungi (d). The regression slope of the linear relationship based on the Gauss generalized linear model is expressed by a solid line. According to the Mantel test (9999 permutations, P < 0.05), the relationships of bacteria and fungi and elevation were statistically significant
Drivers of Bacterial and Fungal Biodiversity
The Spearman correlations between fungal and bacterial diversity indices and environmental variables were mainly negative, with only four positive correlations (Fig. S8). For instance, the Chao1 for bacteria negatively correlated with Comp.1 (rho = 0.47, P < 0.01) a254 (rho = −0.46, P < 0.01), and PO43−-P (rho = −0.45, P < 0.01). The bacterial and fungal assemblage compositions represented by nMDS1 were strongly correlated with mean annual temperature with rho values of −0.86 and −0.90, respectively (P < 0.001). It was worth noting that bacterial and fungal nMDS1 had positive Spearman correlation with mean annual precipitation with rho values of 0.42 and 0.40 (P < 0.05).
Compared to fungi, the community of bacteria generally exhibited stronger correlations with various environmental variables with rho values ranging from 0.15 to 0.41 in Mantel test (P < 0.05, Fig. 5). Specifically, the bacterial communities were more correlated with several chemical and climatic variables such as conductivity and mean annual temperature (Mantel’s r = 0.41 and 0.35, P = 0.0001), while the fungal communities were with the physicochemical variables, i.e., water depth (Mantel’s r = 0.27, P = 0.0001) and conductivity (Mantel’s r = 0.31, P = 0.002). In addition, bacterial and fungal communities were both significantly related to cDOM, such as a254 (Bacteria: Mantel’s r = 0.28, P = 0.0001; Fungi: Mantel’s r = 0.16, P = 0.026) and FI (Bacteria: Mantel’s r = 0.17, P = 0.022; Fungi: Mantel’s r = 0.26, P = 0.001).
Relationship between environmental factors and bacterial and fungal community The color gradient is used to represent Spearman’s correlation coefficient of pairwise comparisons of environmental variables. Spearman’s correlation was used to analyze the correlation of bacterial and fungal community with each environmental factor. EC, electrical conductivity; TN, total nitrogen; TP, total phosphorus; PO43−-P, orthophosphate; NOx−-N, nitrogen oxides; Depth, water depth; Shading, riparian shading; Speed, stream flow speed; MAP, mean annual precipitation; and MAT, mean annual temperature. BIX, autochthonous productivity index; HIX, humification index; FI, fluorescence index; a254, absorbance at 254 nm; S275–295, spectral slope; Comp.1, microbial humic-like; Comp.2, Dark-incubation peak
Bacterial and fungal alpha diversity were mainly driven by chemical and physical variables, respectively, while community compositions by climatic factors, according to random forest analysis (Fig. 6). Regarding bacteria, total nitrogen was the most important factors governing species richness and evenness (Fig. 6a, b). Further research revealed a significantly negative correlation between total nitrogen and both bacterial species richness and evenness (Fig. S7). Regarding fungi, shading was the best driver for species richness, and stream width for evenness, with shading showing a positive significant correlation with both indices (Figs. 6d, e; S7). Different from the diversity indices, the microbial community compositions were mainly driven by climatic variables with mean annual temperature being the strongest driver for both bacterial and fungal community compositions (Fig. 6c, f).
Relative contribution of climate, physical, chemical, and cDOM variables in shaping the Chao1 diversity, Evenness diversity, and assemblage composition of bacteria and fungi The independent effects of the selected variables on the biological indices for bacteria (a)–(c) and fungi (d)–(f) examined by random forest. The variances in Chao1 diversity, Evenness diversity, and assemblage composition for bacteria (g)–(i) and fungi (j)–(l) associated with the climatic and local non-climatic variables were obtained using variation partitioning. The values in the diagrams refer to the coefficient of determination. Negative variance components were not displayed. The assemblage composition was estimated using first axis coordinates from nonmetric multidimensional scaling (nMDS) based on Bray-Curtis dissimilarity matrices. Purple, green, yellow, and blue colors represent climatic, physical, chemical, and cDOM factors, respectively
The alpha diversity were explained by different categories of explanatory variables between bacteria and fungi, while variations in community composition were mainly explained by the climatic variables, as revealed by the variation partitioning analysis (Fig. 6). Regarding alpha diversity, the physical (9.3%) and climatic variables (5.2%), and the joint effect of the them (8.7%) accounted for most of the variation in bacterial species richness, and the climatic variables (13.4%) explained most in bacterial evenness (Fig. 6g, h). For fungi, physical variables explained most of the variation in species richness (27.0%) and evenness (27.6%) (Fig. 6j, k). Regarding community composition, the climatic variables explained the largest variation in both bacteria and fungi with 23.2% and 34.6%, respectively (Fig. 6i, l).
Discussion
Despite the increasing number of studies on the distribution, characteristics, and diversity of microorganisms along elevational gradients, studies on freshwater ecosystems are still scarce, but are an important part of understanding the global microbial geographic distribution patterns. Here, by examining the elevational biodiversity patterns and drivers of stream bacteria and fungi on the southern slope of Mt. Kilimanjaro, we had three main findings: (1) The species richness and evenness of bacteria increased significantly towards higher elevations, whereas fungi showed non-significant trends. (2) The dissimilarity of assemblage composition increased significantly with greater elevational distances for both bacteria and fungi. (3) Bacterial and fungal alpha diversity were predominantly driven by chemical and physical factors, respectively, whereas community compositions by climatic factors.
Microbial Community Structure in the Streams
For bacterial groups, we found that the most dominant phylum in the streams biofilms was Proteobacteria, with the prevalence of class Alphaproteobacteria (Figs. S2, S3), while Betaproteobacteria was not detected in our stream samples. Generally, Alpha-, Beta-, and Gammaproteobacteria, as members of Proteobacteria, are typically the main classes in the freshwater ecosystems [73, 74]. Our unusual findings are partly consistent with previous study in the Cloghoge River, County Wicklow, Ireland, where Alphaproteobacteria was the key component of freshwater biofilms , while Betaproteobacteria also occurred [75]. In addition, the relative abundance of Cyanobacteria, as a dominant phylum second to Proteobacteria, was significantly correlated with total phosphorus and PO43−. This may be attributed to the large phosphorus accumulation capacity of Cyanobacteria at low phosphorus concentrations [76].
For fungi groups, Ascomycota had the broadest distribution in the stream ecosystems (Figs. S2, S3). This finding is consistent with Hulun lake that the Ascomycota was the most dominant phyla in the lake water and sediment [77]. Ascomycota is probably a potentially critical structuring element of stream biofilms that facilitates the decomposition of submerged organic matter [78, 79], hence typically plays an important role in aquatic ecosystems. Previous studies in soil have demonstrated that the Ascomycota are mostly saprophytes, which breakdown difficult-to-degrade organic matter and play an important role in nutrient cycling [80]. The relative abundance of Ascomycota was significantly negatively correlated with water depth (P < 0.001, Fig. S4), which could be attributed to the fact that Ascomycota are aerobic fungi [81], while the oxygen availability gradually may decrease with depth. It should be noted that our findings are different from a global survey of terrestrial ecosystems showing the dominance of Basidiomycota [82].
The most abundant ASVs in the subset of ASVs with > 50% incidence in bacterial communities in stream biofilm were mainly belong to the genus Pleurocapsa_PCC−7319, followed by the genus Chroococcopsis. Significant correlations (P < 0.001) were observed between NH4+ and two prominent ASVs belonging to the genus Pleurocapsa_PCC−7319, which could be the potential nitrogen-fixing bacteria that fix atmospheric nitrogen gas to ammonia serving nitrogen in ecosystem [83]. Furthermore, four of the ten ASVs with the highest relative abundance in the subset of ASVs with >50% incidence in fungal communities were belonged to the genus Plectosphaerella and correlated negatively with PO43−, while several ASVs from the genera Cladosporium, Pyrenochaetopsis, and Penicillium showed positive correlations with PO43−. This phenomenon indicates that some taxa of fungi were sensitive to phosphorus conditions, which is likely relevant to human activities with phosphorus inputs. This highlights the importance of the further research on functional fungal genes such as those governing the phosphorus cycle.
Elevational Patterns of Microbial Diversity
Our results showed the contrasting elevational patterns in species richness or evenness of bacteria and fungi in the alpine stream ecosystems (Fig. 3). Regarding species richness, the monotonically increasing patterns towards higher elevations for bacteria are consistent with the previous observations for stream bacteria in the Pyrenees Mountain of Spain and the Haba Mountain of China [11], and contrary to biofilm microbial communities in the glacier-fed streams in Austria [84]. This increasing pattern does not follow the traditional ecological theory that the relationship between species richness and elevation generally shows a hump-shaped or decreasing pattern with increasing elevation for plants and animals in terrestrial ecosystems [85, 86]. The elevation-based increase in species richness observed may be linked to the sampling scale effect, as the extent of the geographical area or gradient of sampling has been confirmed to significantly affect the resulting species richness patterns [85, 87].
As for evenness, an index directly linked to ecosystem stability [88], there were also an increasing trend for bacteria towards higher elevations (Fig. 3). This may be largely because high species evenness usually contributes to ecosystem stability, while the ecosystem at high altitudes is more vulnerable, and therefore they require high evenness for the maintenance of related functions [89]. Furthermore, we found that there was a significantly positive correlation between evenness and species richness for both bacteria and fungi (Fig. S7), which is opposite to the typically negative correlation for macroorganisms in stream ecosystems [90]. This could be explained by the lower extinctions of microorganisms, arising from the high dispersal capacity and large population, compared to macroorganisms [11, 91].
We found that the bacterial and fungal communities at the lower elevations exhibited greater similarity compared to those at higher elevations and showed a significant elevational distance-decay pattern (Fig. 4). This result suggested that elevation had crucial effects on structuring the composition of microbial communities, because elevation as a geographic factor has always been a complex and direct driving force for different climatic gradients, leading to the succession of habitats along gradients [85, 92]. The habitats at lower elevations are more benign that could allow many similar species to survive, while relatively harsher environments at higher elevations may be causing specialized species surviving in our study streams. For elevational distance-decay pattern, one possible explanation is that communities in close spatial proximity are likely to experience more similar environmental and climate condition and share more species through localized dispersal, resulting in more similar community structure than those located further apart in space [93].
Environment Effects on Microorganisms
The diversity patterns of bacterial community were primarily influenced by chemical factors, such as total nitrogen and phosphorus (Fig. 6), which are fundamental nutrient in aquatic ecosystems. For instance, the species richness and evenness of bacteria were both strongly driven by the total nitrogen, with a negative correlation observed between them (Figs. 6, S8). This may be related to the different strategies that the dominant groups in bacterial communities respond to environmental nutrient status. For example, Gammaproteobacteria prefer to high nutrient concentrations because they are typical R-strategists and are able to use the extra nutrients quickly (Shafi et al. 2021). In contrast, Alphaproteobacteria likely thrive in more oligotrophic environments as most of its species prefer to grow in oligotrophic environments [75]. At the ASV level, the ASVs sharing the same phylum also exhibited varying correlation to the environmental nutrient variables (Figs. S5, S6), further supporting that various dominant bacterial groups differently responded to environmental nutrient status. Furthermore, the productive activities of microorganisms in the region may be limited by nitrogen levels, as the nutrient inputs East Africa are generally lower than in the densely populated subtropical and temperate zones of the Northern Hemisphere [94].
Bacterial evenness was also strongly driven by total phosphorus and showed negative correlations with total phosphorus, the concentrations of which typically correlate with human influence and are stronger at lower elevations [11]. The intensity of human activities, such as agricultural cultivation and domestic wastewater discharges, positively influences nitrogen and phosphorus contents in the aquatic environments, leading to changes in the diversity of microbial communities. On the slopes of Mt. Kilimanjaro, most of savanna woodlands have been converted to agriculture and thousands of hectares of forest were destroyed by slashing and burning, driven by human activity in search of resources [95]. Our sampling area gets closer to Kilimanjaro National Park as elevation increases; a gradual reduction in human impact may happen.
For fungi, both species richness and evenness were strongly driven by physical variables, such as riparian shading, with a significant positive correlation between both indices and riparian shading (Figs. 6, S8). Fungi are the primary decomposers of the leaf litter, therefore, more sensitive to variation in the litter environment [96]. Riparian shading could have important effects on stream organic inputs by directly affecting the amount of litter entering [97, 98], accordingly becoming a crucial driver of fungal species richness. The fungal species richness and evenness were also driven by other physical factors, such as flow speed and width, which characterized the hydrological changes (Fig. 6). This finding was consistent with previous researches that have demonstrated the significant influence of hydrological conditions in shaping bacterial diversity [99, 100]. This is because regional hydrological transport can directly influences the community, with microorganisms being actively or passively dispersing along hydrological flow paths, thereby reaching downstream habitats [101]. Stream width may also reflect human influence, as wider streams have larger catchments and more complex human impacts or nutrient conditions [17].
It should be noted that the community assemblages for bacteria and fungi were strongly driven by climatic factors (Figs. 6, S8). For instance, the mean annual temperature was the strongest driver for the dissimilarities of bacterial and fungal community compositions, which could be attributed to the effect of temperature on microbial enzyme activity and its role as a primary driver of biological metabolic processes [102, 103]. In addition, mean annual precipitation was also a predictor of fungal community composition in the streams, although its relative importance is relatively lower than that observed in other ecosystems such as soils [82]. Precipitation could arouse higher stream flow, which causes increasing riparian soil input and sediment resuspension in rivers, eventually acting on microbial communities [17]. Generally, mean annual temperature and precipitation could directly regulate bacterial diversity and composition through speciation, competition, and dispersal of microbial communities, or indirectly through mediating other environmental factors [104].
Conclusion
In conclusion, we conducted a comprehensive study on the elevational patterns of stream bacteria and fungi in the Mt. Kilimanjaro region, and revealed the potential mechanisms underlying the observed patterns. We found that there were no consistent elevational patterns in species richness and evenness for the two microbial taxonomic groups bacteria and fungi, with the former showing a rarely monotonically increasing trend while the latter a nonsignificant trend. However, the beta diversity between the two taxonomic groups showed a consistent trend, i.e., significant elevational distance decay in community similarity. The inconsistency of species richness or evenness patterns between bacteria and fungi may be attributed to their different underlying drivers, with bacteria mainly by chemical factors such as total nitrogen and phosphorus, while fungi by physical factors such as riparian shading. For beta diversity, the consistent patterns may be related to the drivers of regional factors such as mean annual temperature and precipitation. Our study shows a new genetic data collection for better understanding the effects of multiple environmental stressors on stream microbes in this rarely studied region in Africa, and further provides novel insights on the response patterns of tropical aquatic microbial community composition and diversity under climate change.
Data Availability
The raw sequencing data for bacterial and fungal communities have been submitted to the National Omics Data Encyclopedia (NODE: https://www.biosino.org/node) database under the accession number OEP004246.
References
Teittinen A, Wang JJ, Stromgard S, Soininen J (2017) Local and geographical factors jointly drive elevational patterns in three microbial groups across subarctic ponds. Glob Ecol Biogeogr 26:973–982
Wang J, Hu A, Meng F, Zhao W, Yang Y, Soininen J, Shen J, Zhou J (2022) Embracing mountain microbiome and ecosystem functions under global change. New Phytol 234:1987–2002
Rahbek C, Borregaard MK, Antonelli A, Colwell RK, Holt BG, Nogues-Bravo D, Rasmussen C, Richardson K, Rosing MT, Whittaker RJ, Fjeldså J (2019) Building mountain biodiversity: Geological and evolutionary processes. Science 365:1114–1119
Larigauderie A, Prieur-Richard A-H, Mace G, Lonsdale M, Mooney H, Brussaard L, Cooper H, Cramer W, Daszak P, Diaz S, Duraiappah A, Elmqvist T, Faith D, Jackson L, Krug C, Leadley P, Le Prestre P, Matsuda H, Palmer M, Yahara T (2012) Biodiversity and ecosystem services science for a sustainable planet: the DIVERSITAS vision for 2012-20. Curr Opin Environ Sustain 4:101–105
Gilbert KJ, Bittleston LS, Naive MAK, Kiszewski AE, Buenavente PAC, Lohman DJ, Pierce NE (2020) Investigation of an elevational gradient reveals strong differences between bacterial and eukaryotic communities coinhabiting Nepenthes Phytotelmata. Microb Ecol 80:334–349
Yasir M, Azhar EI, Khan I, Bibi F, Baabdullah R, Al-Zahrani IA, Al-Ghamdi AK (2015) Composition of soil microbiome along elevation gradients in southwestern highlands of Saudi Arabia. BMC Microbiol 15:65
Shen C, Gunina A, Luo Y, Wang J, He J-Z, Kuzyakov Y, Hemp A, Classen AT, Ge Y (2020) Contrasting patterns and drivers of soil bacterial and fungal diversity across a mountain gradient. Environ Microbiol 22:3287–3301
Siles JA, Margesin R (2016) Abundance and diversity of bacterial, archaeal, and fungal communities along an altitudinal gradient in Alpine forest soils: what are the driving factors? Microb Ecol 72:207–220
Cavicchioli R, Ripple WJ, Timmis KN, Azam F, Bakken LR, Baylis M, Behrenfeld MJ, Boetius A, Boyd PW, Classen AT, Crowther TW, Danovaro R, Foreman CM, Huisman J, Hutchins DA, Jansson JK, Karl DM, Koskella B, Mark Welch DB et al (2019) Scientists’ warning to humanity: microorganisms and climate change. Nat Rev Microbiol 17:569–586
Bello F, Lavorel S, Lavergne S, Albert CH, Boulangeat I, Mazel F, Thuiller W (2013) Hierarchical effects of environmental filters on the functional structure of plant communities: a case study in the French Alps. Ecography 36:393–402
Wang J, Meier S, Soininen J, Casamayor EO, Pan F, Tang X, Yang X, Zhang Y, Wu Q, Zhou J, Shen J (2017) Regional and global elevational patterns of microbial species richness and evenness. Ecography 40:393–402
Legendre P, De Cáceres M (2013) Beta diversity as the variance of community data: dissimilarity coefficients and partitioning. Ecol Lett 16:951–963
Magurran AE (2021) Measuring biological diversity. Curr Biol 31:R1174–R1177
Yeh C-F, Soininen J, Teittinen A, Wang J (2019) Elevational patterns and hierarchical determinants of biodiversity across microbial taxonomic scales. Mol Ecol 28:86–99
Fierer N, McCain CM, Meir P, Zimmermann M, Rapp JM, Silman MR, Knight R (2011) Microbes do not follow the elevational diversity patterns of plants and animals. Ecology 92:797–804
Whittaker RH (1960) Vegetation of the Siskiyou Mountains, Oregon and California. Ecol Monogr 30:279–338
Liu J, Zhao W, Ren M, Liu Y, Xu Y, Wang J (2022) Contrasting elevational patterns and underlying drivers of stream bacteria and fungi at the regional scale on the Tibetan Plateau. FEMS Microbiol Ecol 98:fiac050
Maloufi S, Catherine A, Mouillot D, Louvard C, Couté A, Bernard C, Troussellier M (2016) Environmental heterogeneity among lakes promotes hyper β-diversity across phytoplankton communities. Freshw Biol 61:633–645
Benito X, Fritz SC, Steinitz-Kannan M, Tapia PM, Kelly MA, Lowell TV (2018) Geo-climatic factors drive diatom community distribution in tropical South American freshwaters. J Ecol 106:1660–1672
Zhang QQ, Sheng-Long J, Ke-Mao L, Zhen-Bing W, Hong-Tao G, Jin-Wen H, Shu-Yi W, Yao-Yao L, Guo-Jie W, Ai-Hua L (2021) Community structure of bacterioplankton and its relationship with environmental factors in the upper reaches of the Heihe River in Qinghai Plateau. Environ Microbiol 23:1210–1221
Picazo F, Vilmi A, Aalto J, Soininen J, Casamayor EO, Liu Y, Wu Q, Ren L, Zhou J, Shen J, Wang J (2020) Climate mediates continental scale patterns of stream microbial functional diversity. Microbiome 8:92
Gütlein A, Gerschlauer F, Kikoti I, Kiese R (2018) Impacts of climate and land use on N2O and CH4 fluxes from tropical ecosystems in the Mt. Kilimanjaro region Tanzania. Global Change Biol 24:1239–1255
Hemp A (2009) Climate change and its impact on the forests of Kilimanjaro. Afr J Ecol 47:3–10
Pabst H, Kühnel A, Kuzyakov Y (2013) Effect of land-use and elevation on microbial biomass and water extractable carbon in soils of Mt. Kilimanjaro. Ecosyst Appl Soil Ecol 67:10–19
Park S-H, Lee M-J, Jung H-S (2016) Spatiotemporal analysis of snow cover variations at Mt. Kilimanjaro using multi-temporal Landsat images during 27 years. J Atmos Sol Terr Phys 143-144:37–46
Stadelmann C, Fürst JJ, Mölg T, Braun M (2020) Brief communication: Glacier thickness reconstruction on Mt. Kilimanjaro. Cryosphere 14:3399–3406
Veettil BK, Kamp U (2019) Global disappearance of tropical mountain glaciers: observations, causes, and challenges. Geosci 9:196
Vimercati L, Darcy JL, Schmidt SK (2019) The disappearing periglacial ecosystem atop Mt. Kilimanjaro supports both cosmopolitan and endemic microbial communities. Sci Rep 9:10676
Sébastien L (2010) The Chagga people and environmental changes on Mount Kilimanjaro: lessons to learn. Clim Dev 2:364–377
Renner M, Rembold K, Hemp A, Fischer M (2022) Natural regeneration of woody plant species along an elevational and disturbance gradient at Mt. Kilimanjaro. Forest Ecol Manag 520:120404
Hemp A (2006) Continuum or zonation? Altitudinal gradients in the forest vegetation of Mt. Kilimanjaro. Plant Ecology 184:27–42
Wang J, Soininen J, Zhang Y, Wang B, Yang X, Shen J (2011) Contrasting patterns in elevational diversity between microorganisms and macroorganisms. J Biogeogr 38:595–603
Jin XC, Tu QY (1990) Standards of lake eutrophication survey of China. China Environmental Science Publishing House, Beijing, China (in Chinese)
Hu A, Jang K-S, Meng F, Stegen J, Tanentzap AJ, Choi M, Lennon JT, Soininen J, Wang J (2022) Microbial and environmental processes shape the link between organic matter functional traits and composition. Environ Sci Technol 56:10504–10516
Shi W, Zhuang W-E, Hur J, Yang L (2021) Monitoring dissolved organic matter in wastewater and drinking water treatments using spectroscopic analysis and ultra-high resolution mass spectrometry. Water Res 188:116406
Chen B, Zhao M, Liu C, Feng M, Ma S, Liu R, Chen K (2020) Comparison of copper binding properties of DOM derived from fresh and pyrolyzed biomaterials: insights from multi-spectroscopic investigation. Sci Total Environ 721:137827
Fick SE, Hijmans RJ (2017) WorldClim 2: new 1-km spatial resolution climate surfaces for global land areas. Int J Climatol 37:4302–4315
Hijmans RJ, van Etten J (2016) raster: geographic data analysis and modeling. R package version 2:5–8
Kumar P (2012) Assessment of impact of climate change on Rhododendrons in Sikkim Himalayas using Maxent modelling: limitations and challenges. Biodivers Conserv 21:1251–1266
Nelson AR, Narrowe AB, Rhoades CC, Fegel TS, Daly RA, Roth HK, Chu RK, Amundson KK, Young RB, Steindorff AS, Mondo SJ, Grigoriev IV, Salamov A, Borch T, Wilkins MJ (2022) Wildfire-dependent changes in soil microbiome diversity and function. Nat Microbiol 7:1419–1430
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583
Pearson T, Caporaso JG, Yellowhair M, Bokulich NA, Padi M, Roe DJ, Wertheim BC, Linhart M, Martinez JA, Bilagody C, Hornstra H, Alberts DS, Lance P, Thompson PA (2019) Effects of ursodeoxycholic acid on the gut microbiome and colorectal adenoma development. Cancer Med 8:617–628
Abarenkov K, Zirk A, Piirmann T, Pöhönen R, Ivanov F, Nilsson R, Kõljalg UJUC (2021) UNITE QIIME release for fungi version 10.05, vol 7, p 1264763
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y, Bisanz JE, Bittinger K, Brejnrod A, Brislawn CJ, Brown CT, Callahan BJ, Caraballo-Rodríguez AM, Chase J et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857
Harshman RA (1970) Foundations of the PARAFAC procedure: models and conditions for an "explanatory" multi-modal factor analysis. UCLA Work Pap Phon 16:1–84
Pucher M, Wünsch U, Weigelhofer G, Murphy K, Hein T, Graeber D (2019) staRdom: Versatile software for analyzing spectroscopic data of dissolved organic matter in R. Water 11:2366
Coble PG (1996) Characterization of marine and terrestrial DOM in seawater using excitation-emission matrix spectroscopy. Mar Chem 51:325–346
Ni M, Li S (2023) Ultraviolet humic-like component contributes to riverine dissolved organic matter biodegradation. J Environ Sci 124:165–175
Ohno T (2002) Fluorescence inner-filtering correction for determining the humification index of dissolved organic matter. Environ Sci Technol 36:742–746
Birdwell JE, Engel AS (2010) Characterization of dissolved organic matter in cave and spring waters using UV–Vis absorbance and fluorescence spectroscopy. Org Geochem 41:270–280
Granados V, Arias-Real R, Gutiérrez-Cánovas C, Obrador B, Butturini A (2022) Multiple drying aspects shape dissolved organic matter composition in intermittent streams. Sci Total Environ 852:158376
Fang Z, Wang W-X (2022) Dynamics of trace metals with different size species in the Pearl River Estuary, Southern China. Sci Total Environ 807:150712
Fichot CG, Benner R (2012) The spectral slope coefficient of chromophoric dissolved organic matter (S275–295) as a tracer of terrigenous dissolved organic carbon in river-influenced ocean margins. Limnol Oceanogr 57:1453–1466
Murphy KR, Stedmon CA, Graeber D, Bro R (2013) Fluorescence spectroscopy and multi-way techniques. PARAFAC. Anal Methods 5:6557–6566
Stedmon CA, Bro R (2008) Characterizing dissolved organic matter fluorescence with parallel factor analysis: a tutorial. Limnol Oceanogr Methods 6:572–579
Murphy KR, Stedmon CA, Wenig P, Bro R (2014) OpenFluor—an online spectral library of auto-fluorescence by organic compounds in the environment. Anal Methods 6:658–661
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Autom Control 19:716–723
Oksanen J, Simpson G, Blanchet FG, Kindt R, Legendre P, Minchin P, O`hara RB, Solymos P, Stevens H, Szöcs E, Wagner H, Barbour M, Bedward M, Bolker B, Borcard D, Carvalho G, Chirico M, De Cáceres M, Durand S, Weedon J (2022) Vegan community. Ecology package version 2:6
Jost L (2010) The relation between evenness and diversity. Diversity, pp 207–232
Pielou EC (1966) The measurement of diversity in different types of biological collections. J Theor Biol 13:131–144
Heger TJ, Giesbrecht IJW, Gustavsen J, del Campo J, Kellogg CTE, Hoffman KM, Lertzman K, Mohn WW, Keeling PJ (2018) High-throughput environmental sequencing reveals high diversity of litter and moss associated protist communities along a gradient of drainage and tree productivity. Environ Microbiol 20:1185–1203
Liao H, Hao X, Qin F, Delgado-Baquerizo M, Liu Y, Zhou J, Cai P, Chen W, Huang Q (2023) Microbial autotrophy explains large-scale soil CO2 fixation. Glob Chang Biol 29:231–242
Bell T (2010) Experimental tests of the bacterial distance–decay relationship. ISME J 4:1357–1365
Violle C, Nemergut DR, Pu Z, Jiang L (2011) Phylogenetic limiting similarity and competitive exclusion. Ecol Lett 14:782–787
Patoine G, Eisenhauer N, Cesarz S, Phillips HRP, Xu X, Zhang L, Guerra CA (2022) Drivers and trends of global soil microbial carbon over two decades. Nat Commun 13:4195
Hijmans RJ, Van Etten J, Cheng J, Mattiuzzi M, Sumner M, Greenberg JA, Lamigueiro OP, Bevan A, Racine EB, Shortridge AJR (2015). Package ‘raster’ 734:473
Lê S, Josse J, Husson F (2008) FactoMineR: an R package for multivariate analysis. J Stat Softw 25:1–18
Kuhn M (2008) Building predictive models in R using the caret package. J Stat Softw 28:1–26
Belgiu M, Drăguţ L (2016) Random forest in remote sensing: a review of applications and future directions. ISPRS J Photogramm Remote Sens 114:24–31
Borcard D, Legendre P, Drapeau P (1992) Partialling out the spatial component of ecological variation. Ecology 73:1045–1055
Bracewell SA, Barros TL, Mayer-Pinto M, Dafforn KA, Simpson SL, Johnston EL (2023) Contaminant pulse following wildfire is associated with shifts in estuarine benthic communities. Environ Pollut 316:120533
Huguet A, Vacher L, Relexans S, Saubusse S, Froidefond JM, Parlanti E (2009) Properties of fluorescent dissolved organic matter in the Gironde Estuary. Org Geochem 40:706–719
Gabriel Z, Byron CC, Kamst-van Agterveld MP, Hagen F, Han SK (2002) Typical freshwater bacteria: an analysis of available 16S rRNA gene sequences from plankton of lakes and rivers. Aquat Microb Ecol 28:141–155
Newton RJ, Jones SE, Eiler A, McMahon KD, Bertilsson S (2011) A guide to the natural history of freshwater lake bacteria. Microbiol Mol Biol Reviews : MMBR 75:14–49
Anderson-Glenna MJ, Bakkestuen V, Clipson NJW (2008) Spatial and temporal variability in epilithic biofilm bacterial communities along an upland river gradient. FEMS Microbiol Ecol 64:407–418
Andersen T, Hessen DO (1991) Carbon, nitrogen, and phosphorus content of freshwater zooplankton. Limnol Oceanogr 36:807–814
Shang Y, Wu X, Wang X, Dou H, Wei Q, Ma S, Sun G, Wang L, Sha W, Zhang H (2022) Environmental factors and stochasticity affect the fungal community structures in the water and sediments of Hulun Lake China. Ecol Evol 12:e9510
Besemer K (2015) Biodiversity, community structure and function of biofilms in stream ecosystems. Res Microbiol 166:774–781
Das M, Royer T, Leff LG (2007) Diversity of fungi, bacteria, and actinomycetes on leaves decomposing in a stream. Appl Environ Microbiol 73:756–767
Beimforde C, Feldberg K, Nylinder S, Rikkinen J, Tuovila H, Dörfelt H, Gube M, Jackson DJ, Reitner J, Seyfullah LJ, Schmidt AR (2014) Estimating the Phanerozoic history of the Ascomycota lineages: combining fossil and molecular data. Mol Phylogenet Evol 78:386–398
Young D, Dollhofer V, Callaghan TM, Reitberger S, Lebuhn M, Benz JP (2018) Isolation, identification and characterization of lignocellulolytic aerobic and anaerobic fungi in one- and two-phase biogas plants. Bioresour Technol 268:470–479
Tedersoo L, Bahram M, Põlme S, Kõljalg U, Yorou NS, Wijesundera R, Ruiz LV, Vasco-Palacios AM, Thu PQ, Suija A, Smith ME, Sharp C, Saluveer E, Saitta A, Rosas M, Riit T, Ratkowsky D, Pritsch K, Põldmaa K et al (2014) Global diversity and geography of soil fungi. Science 346:1256688
Patel ZZ, Kumar D, Puvar A, Joshi H, Joshi C, Tipre DR, Joshi M (2023) Exploring bacteriome diversity of coral Goniopora sp. and Favia favus from the Gulf of Kutch Gujarat. J Sea Res 192:102361
Wilhelm L, Singer GA, Fasching C, Battin TJ, Besemer K (2013) Microbial biodiversity in glacier-fed streams. ISME J 7:1651–1660
Rahbek C (2005) The role of spatial scale and the perception of large-scale species-richness patterns. Ecol Lett 8:224–239
Zeng J, Zhao D, Li H, Huang R, Wang J, Wu QL (2016) A monotonically declining elevational pattern of bacterial diversity in freshwater lake sediments. Environ Microbiol 18:5175–5186
Rowe RJ, Lidgard S (2009) Elevational gradients and species richness: do methods change pattern perception? Glob Ecol Biogeogr 18:163–177
Bernhard AE, Kelly JJ (2016) Editorial: Linking ecosystem function to microbial diversity. Front Microbiol 7:1041
Wittebolle L, Marzorati M, Clement L, Balloi A, Daffonchio D, Heylen K, De Vos P, Verstraete W, Boon N (2009) Initial community evenness favours functionality under selective stress. Nature 458:623–626
Soininen J, Passy S, Hillebrand H (2012) The relationship between species richness and evenness: a meta-analysis of studies across aquatic ecosystems. Oecologia 169:803–809
Finlay BJ (2002) Global dispersal of free-living microbial eukaryote species. Science 296:1061–1063
Zhang Y, Cong J, Lu H, Li G, Xue Y, Deng Y, Li H, Zhou J, Li D (2015) Soil bacterial diversity patterns and drivers along an elevational gradient on Shennongjia Mountain, China. Microb Biotechnol 8:739–746
Clark DR, Underwood GJC, McGenity TJ, Dumbrell AJ (2021) What drives study-dependent differences in distance–decay relationships of microbial communities? Glob Ecol Biogeogr 30:811–825
Yao X, Zhao Z, Wang J, Ding Q, Ren M, Kimirei IA, Zhang L (2022) Sediment organic matter properties facilitate understanding nitrogen transformation potentials in East African lakes. Sci Total Environ 841:156607
Said M, Komakech HC, Munishi LK, Muzuka ANN (2019) Evidence of climate change impacts on water, food and energy resources around Kilimanjaro, Tanzania. Reg Environ Chang 19:2521–2534
Heino J, Tolkkinen M, Pirttilä AM, Aisala H, Mykrä H (2014) Microbial diversity and community–environment relationships in boreal streams. J Biogeogr 41:2234–2244
Bartels P, Cucherousset J, Steger K, Eklöv P, Tranvik LJ, Hillebrand H (2012) Reciprocal subsidies between freshwater and terrestrial ecosystems structure consumer resource dynamics. Ecology 93:1173–1182
Young RG, Matthaei CD, Townsend CR (2008) Organic matter breakdown and ecosystem metabolism: functional indicators for assessing river ecosystem health. J N Am Benthol Soc 27:605–625
Niño-García JP, Ruiz-González C, del Giorgio PA (2016) Interactions between hydrology and water chemistry shape bacterioplankton biogeography across boreal freshwater networks. ISME J 10:1755–1766
Read DS, Gweon HS, Bowes MJ, Newbold LK, Field D, Bailey MJ, Griffiths RI (2015) Catchment-scale biogeography of riverine bacterioplankton. ISME J 9:516–526
Cousin S (2009) Flavobacterial community structure in a hardwater rivulet and adjacent forest soil, Harz Mountain, Germany. Curr Microbiol 58:409–415
Zhang Y, Ning D, Wu L, Yuan MM, Zhou X, Guo X, Hu Y, Jian S, Yang Z, Han S, Feng J, Kuang J, Cornell CR, Bates CT, Fan Y, Michael JP, Ouyang Y, Guo J, Gao Z et al (2023) Experimental warming leads to convergent succession of grassland archaeal community. Nat Clim Change 13:561–569
Zhou J, Deng Y, Shen L, Wen C, Yan Q, Ning D, Qin Y, Xue K, Wu L, He Z, Voordeckers JW, Nostrand JD, Buzzard V, Michaletz ST, Enquist BJ, Weiser MD, Kaspari M, Waide R, Yang Y, Brown JH (2016) Temperature mediates continental-scale diversity of microbes in forest soils. Nat Commun 7:12083
Singh D, Lee-Cruz L, Kim W-S, Kerfahi D, Chun J-H, Adams JM (2014) Strong elevational trends in soil bacterial community composition on Mt. Halla. South Korea Soil Biol Biochem 68:140–149
Acknowledgements
We appreciate anonymous reviewer for their comments and valuable suggestions. We appreciate Yaseen Mshuza, Molla Bundala, and Claudia Hemp for the assistance in field sample collection, and Hao Wu for valuable comments of the manuscript.
Funding
This study was supported by the Research Program of Sino-Africa Joint Research Center, Chinese Academy of Sciences (151542KYSB20210007) and the National Natural Science Foundation of China (42225708, 92251304).
Author information
Authors and Affiliations
Contributions
Y.Z.: data curation, formal analysis, writing—original draft, writing—review and editing. F.M.: data curation. B.O.: writing—review and editing. J.X.: data curation. L.Z.: resources. I.K.: resources. M.F.: resources. L.Z: writing—review and editing. J.W.: conceptualization, formal analysis, writing—review and editing. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing of Interest
The authors declare no competing interests.
Supplementary Information
ESM 1
(DOCX 3681 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, Y., Meng, F., Ochieng, B. et al. Climate and Environmental Variables Drive Stream Biofilm Bacterial and Fungal Diversity on Tropical Mountainsides. Microb Ecol 87, 28 (2024). https://doi.org/10.1007/s00248-023-02335-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-023-02335-2