Abstract
Fungi provide essential ecosystem services and engage in a variety of symbiotic relationships with trees. In this study, we investigate the spatial relationship of trees and fungi at a community level. We characterized the spatial dynamics for above- and belowground fungi using a series of forest monitoring plots, at nested spatial scales, located in the tropical South Pacific, in Vanuatu. Fungal communities from different habitats were sampled using metagenomic analysis of the nuclear ribosomal ITS1 region. Fungal communities exhibited strong distance–decay of similarity across our entire sampling range (3–110,000 m) and also at small spatial scales (< 50 m). Unexpectedly, this pattern was inverted at an intermediate scale (3.7–26 km). At large scales (80–110 km), belowground and aboveground fungal communities responded inversely to increasing geographic distance. Aboveground fungal community turnover (beta diversity) was best explained, at all scales, by geographic distance. In contrast, belowground fungal community turnover was best explained by geographic distance at small scales and tree community composition at large scales. Fungal communities from various habitats respond differently to the influences of habitat and geographic distance. At large geographic distances (80–110 km), community turnover for aboveground fungi is better explained by spatial distance, whereas community turnover for belowground fungi is better explained by plant community turnover. Future syntheses of spatial dynamics among fungal communities must explicitly consider geographic scale to appropriately contextualize community turnover.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fungi and plants have coexisted for millions of years, often forming important symbiotic relationships. Fungi influence forest communities [1, 2] both through antagonism (by causing negative density-dependent growth and mortality [3]) and through mutualisms (as with mycorrhizae [4]). Similarly, plants impact fungal communities by modifying the local environment or engaging in specificity [5]. Bipartite relationships between host plants and fungi are important for understanding population-level dynamics, but a more complete picture of complex ecosystems might be drawn at the community level. Estimates of fungal diversity range from 1.5 to 3.8 million species [6]. Plant species richness is predicted to be around 450,000 species, two-thirds of which are found in the tropics [7]. A framework for examining interactions between co-occurring communities is to compare their compositional turnover along gradients.
The distance–decay of similarity (DDS) is a recurrent phenomenon that describes how the relationship between two entities changes over geographic space, a pattern consistent with Tobler’s first law of geography [8]. Tobler’s law intuitively states that nearby things have a tendency to be more similar than distant things. Within community ecology, DDS describes a pattern of community-membership turnover (beta diversity) with increasing geographic distance and is used to uncover patterns of species distribution and aggregation [9]. The distance–decay relationship is a powerful tool in spatial ecology because the slope of this relationship reflects a combination of environmental and biotic variables [10]. Ecologists have used the DDS to infer the relative importance of such diverse topics as dispersal limitation in trees [11] and in the niche partitioning of diatoms [12]. For communities of bacteria, in general, there is recurrent evidence of a distance–decay relationship predicting community composition divergence positively correlated with geographic distance [13,14,15]. Distance–decay of similarity is not a constant in community ecology [16, 17] which raises questions about the circumstances that lead exceptions of distance–decay relationships [18].
Distance–decay relationships are both scale and system dependent, so recognizing scale dependency is an important step to revealing insights into processes driving patterns of biodiversity. Scale-dependent patterns have been documented in microbes, indicating that the relative importance of mechanisms generating spatial structure, such as dispersal limitation and environmental filtering, varies by geographic scale [19]. Aboveground, the fungal phyllosphere community may [20, 21] or may not [22] exhibit scale-dependent DDS relationships, sometimes showing variable results within the same study [23]. Belowground, soil fungi exhibit distance–decay relationships that vary by soil horizon [24]. The spatial structure among ectomycorrhizal communities is related to host density at a local scale, but climate seems to be more important at a global scale [25]. The relative importance of mechanisms generating spatial structure, such as dispersal limitation and environmental filtering, is scale and habitat dependent. Past studies have typically addressed a single habitat type or scale, making the synthesis of results across studies difficult due to the high variability in sampling methodology [26]. There is a clear need for the consideration of multiple habitats and spatial scales, simultaneously, to allow for an ecosystem-wide perspective within complex forest systems.
Characterization of the spatial dynamics for both fungi and plants from the same environment gives us a window into the ecology of complex forest systems [27]. Plant communities have been studied for centuries under the lens of spatial ecology [28]. By comparison, descriptions of the distribution of fungal communities remain relatively less common. Hawksworth and Lücking [6] estimated less than 10% of all fungal species have been formally described, leaving our understanding of fundamental ecological tenants for fungi behind those of animals and plants. Recent studies have highlighted how little we know of the natural distribution patterns of fungi [29, 30], and ongoing research has yet to establish the extent to which fungi and plants exhibit similar biogeographic patterns [12, 31].
In this study, we describe the spatial distribution patterns of fungal communities from multiple habitat types (soil, understory, and Selaginella) and multiple spatial scales. We then integrated plant community data, collected in parallel with the fungal data, to link distribution patterns of trees and fungi. The aim of this study was to assess whether fungal community beta diversity, derived from three different habitats, varied over three geographical scales ranging from local (3.33–37.23 m), to within islands (3.7–26 km), to between islands (80–110 km), and to what extent fungal spatial dynamics are synchronous with tree communities. We characterized the tree and fungal communities from tropical forests found on the nearby islands of Aneityum and Tanna, in the South Pacific archipelago of Vanuatu, to address the following questions: (i) do fungal communities show distance–decay patterns at multiple geographic scales? If so, (ii) does the strength of decay vary with the geographic scale of investigation? And, (iii) to what extent is fungal community beta diversity attributable to plant community diversity and distribution? We predicted that the rate of community turnover would increase with geographic scale and that these scale-dependent relationships would vary depending on whether the fungal community was sampled from either soil, understory, or Selaginella. Furthermore, we expected a collinear relationship between plant and fungal beta diversity, such that each would display similar patterns of correlated scale-dependent community turnover. Evidence of scale-dependent patterns among plants and fungal communities provides us with new insights into the factors governing biodiversity in tropical forests.
Materials and Methods
Study Site and Sampling
The Republic of Vanuatu is an archipelago of more than 80 islands located in the Southwestern Pacific. The island nation is of volcanic origin and is relatively young, where 90% of the nation is less than 1.8 million years old [32]. Sampling for this project occurred in the southernmost province of Tafea on the islands of Tanna and Aneityum (Fig. 1). Recent efforts to document the flora and vegetation of Tafea have led to a network of sampling transects that were used in this study. Sampling efforts on Tanna and Aneityum occurred in habitats classified as low- to mid-elevation rain forest [33]. Fieldwork occurred over the span of two trips during August 2017 (Aneityum) and December 2017 (Tanna). The two islands are separated by 86 km of open ocean and share a similar climate. Aneityum receives 2322 mm of rain annually and has an average annual temperature of 19.7 °C, although there is a large seasonal and interannual variation [34]. On small islands, topography can have an important impact on local climate typically producing rain shadows. For Tanna and Aneityum, there is a rain shadow on the northwest side of each island.
Location of the 6 transects sampled in this study. Three transects were each sampled on the islands of Tanna and Aneityum, Vanuatu (a). Each transect contained 36 sampling sites, split equidistant among 4 contiguous plots (b). At each sampling site, one fungal community was harvested from each of three habitats; soil, Selaginella, and understory (c, Online Resource 2)
Three long-term vegetation monitoring transects were sampled on each island. The average distances between transects within islands are 5.45 km (Aneityum) and 21.94 km (Tanna) (Fig. 1a). Transect dimensions were 10 m by 40 m. The vegetation in each transect was cataloged and all trees greater than 5 cm (diameter at breast height) were mapped and identified. Common tree genera within transects included Syzygium, Dysoxylum, and Hedycarya (Online Resource 1). Although plant species composition differed among transects, Selaginella (Selaginellaceae) dominated the understory throughout. Selaginella is a small spikemoss in the class Lycopodiopsida and is commonly found among the forests of Tanna and Aneityum. We sampled Selaginella spp., specifically targeting S. firmula to reduce the effect of host phylogeny on microbiome variation.
To characterize the fungal communities occurring within the transects, 36 sampling sites were established in a grid in each transect (Fig. 1b). At each sample site, three separate samples were taken: soil, Selaginella spp., and understory (swabbed angiosperm leaf surface) (Fig. 1c). The details of the sampling methodology for each sample source are outlined in Online Resource 2. Selaginella voucher specimens have been accessioned in the Joseph Rock Herbarium at the University of Hawai‘i. In total, 6 transects, each containing 36 sampling sites, at which there were 3 sampling events (soil, Selaginella, and understory), resulted in 648 fungal community samples. Samples were transported to the laboratory at the University of Hawai‘i at Mānoa. Desiccated Selaginella samples were pulverized in a biosafety cabinet using sterile mortars and pestles and liquid nitrogen. Both Selaginella powder and CTAB swabs were stored at − 20 °C until DNA extraction.
Fungal DNA Extraction, PCR Amplification, and Sequencing
DNA extraction was performed using the Qiagen DNeasy PowerSoil DNA isolation Kit (Qiagen, Cat No./ID: 12888). All extraction steps were performed in a biosafety cabinet to minimize environmental contamination. Negative control DNA extractions were performed using sterile swabs and CTAB solution that traveled to the sampling site but did not come in contact with organic material.
Amplicon libraries were prepared in a single PCR reaction using Illumina-barcoded fungal-specific primers. Primers ITS1F and ITS2 were used to target the hypervariable nuclear ribosomal ITS1 region which is flanked by the 18S and 5.8S nrDNA regions. The ITS1 region was amplified using 8-base pair indexed primers (0.2 μM), gDNA (∼ 5 ng), and Phusion Master Mix (New England Biolabs, Massachusetts), using thermocycler parameters recommended by Phusion. Fungal libraries were purified and normalized using Just-a-plate Purification and Normalization Kit (Charm Biotech, San Diego, California) and quantified on a Qubit fluorometer (Invitrogen, Carlsbad, California). Samples were pooled and submitted for sequencing at the Genomics Core Facility at the University of California–Riverside, Institute for Integrative Genome Biology. The Core Facility conducted quality control using a Bioanalyzer and qPCR to optimize cluster density. Sequencing occurred on the Illumina MiSeq platform (NextSeq500 Sequencer) using V3 chemistry (Illumina Inc., San Diego, CA), allowing for 300-bp paired-end reads.
ITS1 sequences were extracted from the flanking ribosomal subunit genes using ITSxpress (Rivers et al., 2018) then filtered by quality scores using the FASTX-Toolkit (Hannon, 2010). Reverse reads were discarded due to low-quality reads. Chimeras were detected and removed using vsearch (Rognes et al., 2016). Sequences were clustered at 97% identity and fungal taxonomy was assigned using the Python package constax (Gdanetz et al., 2017) and the unite database. Putative contaminants were identified based on their prevalence in the extraction and PCR negative controls and removed using the R package decontam (Davis et al., 2018). OTUs that could not be assigned to a fungal phylum were discarded. Differences in sampling depth between samples were normalized using a variance stabilizing transformation (VST) [35] in DESeq2 [36] within R [37]. All downstream analyses were performed in R and used the VST transformed dataset except in cases when measurements of alpha diversity were employed, in which case the OTU table was rarefied to a minimum sequence depth of 3000 sequence reads. OTU read abundance data, taxonomic assignments, sample metadata, and ancillary collection data were compiled with the R package phyloseq [38].
Data Analysis
Spatial autocorrelation for tree and fungal communities was assessed using Mantel tests. Bray–Curtis dissimilarity matrices were calculated for Hellinger-transformed fungal communities using vegan::vegdist. Geographic distance was calculated using fossil::earth.dist. The decay slope at each spatial scale was calculated using linear regression and significance was tested by permutation. Distance–decay rates were calculated for the entire study and at three subset spatial scales: within transects (3.33–37.23 m), between transects within islands (3.7–26 km), and between islands (80–110 km). The minimum grain size for fungal and tree communities differed in that fungal communities can be represented at the grain size of the individual sample site (i.e., one of the 36 per transect), but tree community grain size could not be reduced beyond that of the transect level. For this reason, we were able to calculate distance–decay within transects for the fungal dataset but not the tree dataset. To test whether fungal distance–decay relationships were related to tree community distance–decay relationships, we performed partial Mantel tests on Bray–Curtis tree community dissimilarity and Bray–Curtis fungal community dissimilarity while accounting for any collinearity associated with geographic distance.
Variation in fungal communities as a function of geographic distance and tree community was tested using Generalized Dissimilarity Modeling (GDM; R package gdm) [39]. GDM provides a non-linear perspective on the relative importance of geographic distance and tree community in structuring fungal communities at various scales. GDM coefficients (the maximum height of its spline [39, 40]) can be interpreted as the amount of variation explained by a predictor variable when all other model variables are held constant. Variation in the GDM spline slope is also informative of how the model varies over a range. The significance of each GDM variable was tested by permutation. We tested how dissimilarity of sample OTU composition between transects varied with differences in tree community and geographic distance (fungal community ~ tree community + geographic distance).
To directly compare the fungal communities from each habitat, we examined the compositional turnover of tree community as a function of the fungal community (tree community ~ Selaginella + Understory + Soil + geographic distance). The relative importance and significance of each GDM variable were tested by permutation.
Results
Fungal Species Identification Through Sequencing
The field sampling occurred in a grid-like pattern but not all pre-determined sampling sites contained Selaginella and not all soil or phylloplane (understory epiphyte) samples resulted in sequence data. The final number of fungal metagenome samples totaled 580 (soil = 216, understory = 213, and Selaginella = 151). Within these samples, we identified a total of 27,581 OTUs (clustered at 97% similarity) of which 10 OTUs were identified as contaminants and were removed from the dataset. We then culled OTUs that could not be identified as fungal at the phylum level reducing our dataset to 18,147. Each sample type varied significantly in the number of OTUs (Tukey’s test; p < 0.01). The average number of OTUs was highest for understory samples (\( \overline{n}=254\pm 82\Big) \), followed by Selaginella (\( \overline{n}=225\pm 79 \)), and soil (\( \overline{n}=192\pm 60 \)). Alpha diversity of fungi (measured as a total number of observed OTUs) also varied by plot, transect, and island (Online Resource 3). Ascomycota and Basidiomycota were the two most common phyla to be recovered from all sample types. Glomeromycota and Mortierellomycota sequences were commonly recovered from Soil samples but not from aboveground samples (Selaginella and understory)(Online Resource 4).
Spatial Dynamics of Fungal and Tree Communities
Mantel tests showed significant distance–decay patterns. In general, as geographic distance increased, fungal communities became increasingly dissimilar for all sample types (Fig. 2). This general trend did not hold true when sub-setting among various spatial scales. At the smallest geographic scale, within transects (< 40 m), all fungal communities follow a distance–decay pattern similar to the general trend seen across the extent of the study. At an intermediate scale, between transects (3.7–26 km), fungal communities invert the general trend of distance–decay, such that fungal community composition increases in similarity with growing spatial distance. At the largest scale, between islands (80–110 km), fungal communities vary in the distance–decay response. Selaginella fungal communities decreased in similarity with distance, whereas soil fungal communities increased in similarity with distance. Linear regressions fit to pairwise sample distances indicated significant scale-dependent patterns present in all fungal communities at each scale, except for understory fungi between islands (80–110 km; Table 1). Tree communities (Mantel r = 0.357, p < 0.001) exhibited a distance–decay slope across the extent of the study (0–110 km) and was similar to what was observed among the fungal communities (Fig. 2; Table 1).
Distance–decay of similarity (DDS) relationships for fungal (soil, Selaginella, understory) and tree communities across various scales. In general, fungal communities become less similar with growing isolation. For fungal communities, there is an inversion of this trend occurring at intermediate scales. Regression lines denote the least-squares linear regression across the extent of the study. The three scales tested in this study are indicated with dotted lines and curly brackets: local (3.33–37.23 m), across islands (3.7–26 km), between islands (80–110 km). Geographic distances are the natural log of meters between sample sites. Dissimilarity between communities is calculated as the natural log of Bray–Curtis dissimilarity. Only the regression lines for significant Mantel tests (p < 0.05) are shown in this figure. Non-significant Mantel test results can be found in Table 1. Community similarity measured as 1-Bray–Curtis dissimilarity (1-BC)
The Relationship Between Tree and Fungi Community Composition
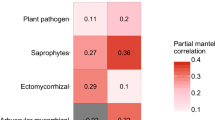
The relationship between tree community and fungal community composition was examined using Mantel tests, partial Mantel tests (Table 2), and GDM. Mantel tests showed a positive correlation between plant and fungal community Bray–Curtis dissimilarity, indicating synchronous beta diversity turnover. As plant community became dissimilar, so too did the fungal community for all three sample types (soil r = 0.612, Selaginella r = 0.632, understory r = 0.556; p = 0.001). This relationship held true when using partial Mantel tests that accounted for the collinearity of geographic distance (fungal community × plant community × geographic distance) (soil r = 0.535, Selaginella r = 0.548, understory r = 0.452; p = 0.001) (Online Resource 5).
GDM models testing the relative importance of both geographic space and tree community composition on structuring aboveground or belowground fungal community composition (fungal community ~ tree community + geographic distance) indicated that geographic distance, rather than tree community, better explained compositional dissimilarity for fungal communities aboveground; Selaginella (GDM coefficient; geography = 1.023, tree community = 0.301) and understory (GDM coefficient; geography = 0.682, tree community = 0.634) (Fig. 3; understory, Selaginella). By contrast, tree community, rather than geographic distance, explained more of the compositional dissimilarity for fungal communities belowground; soil (GDM coefficient; geography = 0.550, tree community = 0.721) (Fig. 3; soil).
Splines for generalized dissimilarity models (fungal community ~ tree community + geographic distance). Models quantify the relationship between tree community (dotted line) and geographic isolation (solid line) and their effect on structuring fungal communities of Selaginella, soil, and understory. The maximum height of a spline can be interpreted as the total contribution that a factor explains of the observed differences in fungal communities when all other variables in the model are held constant. The slopes of the GDM spines indicate the relative importance for a predictive variable across its range. Geographic distance best predicts fungal communities of Selaginella and understory across the entity study, whereas tree community is a better predictor of soil fungal communities at large distances (> 75 km). The top x-axis shows geographic distance between fungal samples, the bottom x-axis shows tree community distances.
The slopes of the GDM splines indicate the relative importance of a predictive variable across its range. For example, the slope of the geographic distance spline for soil fungi is steepest at intermediate ranges (Fig. 3; soil), indicating that geographic distance is most explanatory of soil fungal communities at an intermediate range (~ 30–80 km). By comparison, geographic distances around 100 km explain little of the variation observed in soil fungal community composition (Fig. 3; soil).
Fungal communities from different habitats differed in their strength of correlation with tree community composition. A second GDM model used tree community as a dependent variable and tested the relative influence of each fungal community (tree community ~ Selaginella + understory + soil + geographic distance). Soil fungal community had the largest maximal GDM coefficient (1.075) and explained the greatest cumulative compositional dissimilarity for tree community composition. Geographic distance had a coefficient of 0.758, and aboveground fungal communities (Selaginella and understory) had coefficients of 0.199 and 0.059, respectively (Online Resource 6). At short geographic distances, plant and fungal community turnover is best explained by spatial distances. At large geographic distances (~ 110 km), plant community turnover is best explained by soil fungal community turnover.
Discussion
For biological communities, the slope of a distance–decay relationship is a function of the environmental factors that act upon the community (exogenous factors) and organismal characteristics (endogenous factors) [10]. Our results show significant variability in the distance–decay relationship of tropical fungal communities and indicate that this variability is dependent on both scale and habitat. At the local scale (< 45 m), above- and belowground fungal communities respond similarly to geographic distance. At intermediate distances (3.7–26 km), fungal communities invert distance–decay relationships so that, as geographic distance increases fungal communities become more similar. Distance–decay patterns at the regional scale (~ 80–100 km) are mixed with negative spatial autocorrelation belowground and positive spatial autocorrelation aboveground (Fig. 2). The observed variation in the distance–decay relationships indicates that the relative influence of environmental and organismal characteristics on community dynamics is system and scale dependent. These results are supported by GDM analysis. Fungal communities are strongly influenced by geographic distance at local and intermediate scales. At large scales (~ 110 km), vegetation composition is more important in structuring belowground, but not aboveground fungal communities.
Identifying how dispersal and environment independently structure community composition is difficult in microbial systems. In this study, we observed positive spatial autocorrelation at short distances of less than 50 m, indicating a geographic distance between samples was a strong predictor of community similarity. Positive spatial autocorrelation is representative of an aggregating spatial dynamic (Fig. 4a), whereby fungal communities are clustered at a local scale. Similar results have been observed in other studies [25] looking at spatial autocorrelation of soil fungal communities over short distances of < 10 m [41], < 50 m [24], and < 1 km [20]. Dispersal limitation is an important factor at small scales [42], although we are unable to determine its specific contributions to our results. Fungi likely have a long-tailed dispersal distribution, where the vast majority of reproductive propagules travel over a distance of centimeters and meters, rather than kilometers, a process that would generate the fungal community aggregation patterns observed at small scales in Vanuatu.
Conceptual models of spatial autocorrelation. Positive spatial autocorrelation (a) is associated with a clustering, or aggregating, phenomenon. Positive spatial autocorrelation was observed for all fungal and tree communities at small scales and also across the extent of the study (Fig. 2). Negative spatial autocorrelation (b) was observed for all fungal communities at intermediate scales (3.7–26 km). No spatial autocorrelation (c) is associated with randomly spaced distribution patterns for fungal communities and is also the null hypothesis used in Mantel tests
The sharp inversion of distance–decay patterns at intermediate distances (3.7–26 km) was unexpected given our hypotheses. Mantel tests revealed a pattern of negative spatial autocorrelation at the island scale (3.7–26 km; Table 1), indicating that, at this scale, fungal communities exhibit some form of spatial patterning. The classic example of negative spatial autocorrelation is presented as a single species occurring in a checkerboard or interdigitating distribution pattern (Fig. 4b). While it is difficult to interpret a multivariate dataset, like fungal community composition, on a checkerboard distribution, the negative spatial autocorrelation observed in this study does indicate a pattern of recurrent fungal community composition at intermediate scales. This anomalous inversion of the DDS trend is an example of Simpson’s paradox, where observations at the island scale do not adhere to trends seen in the whole dataset. This same paradox was observed in a different study of continental tropical plants where, at the smallest distance interval (0–600 km) and across the entire study (0–1600 km), plants exhibited positive spatial autocorrelation but at the two intermediate geographic distances (800–1100 and 1200–1600 km), the trend inverted and plant similarity increased with geographic distance [43]. For the larger distance classes of that plant study, environmental factors were more important for explaining plant community structure than was geographic distance. We observed a similar pattern in our study for belowground fungi. At smaller scales, geographic distance was the more explanatory factor for fungal community dissimilarity, whereas, at larger scales (~ 100 km), environmental factors (i.e., tree community) were more important for explaining soil fungal community structure (Fig. 3; soil).
Spatial autocorrelation of fungal communities at a regional scale (between islands 80–110 km) differed depending on the habitat from which fungi were sampled (Fig. 2). Aboveground, Selaginella fungi demonstrated a distance–decay slope consistent with trends across the entire study; Selaginella fungal community composition became more dissimilar with geographic distance. By contrast, soil fungal communities showed an inverted trend. Soil communities became increasingly similar with increasing distance, a trend that was unexpected and implies factors other than geographic distance are structuring soil communities. Indeed, at large geographic scales (between islands 80–110 km), tree community composition explained more of the variability in soil fungal community composition than geographic distance (Fig. 3; soil), a hypothesis that had been previously demonstrated in ectomycorrhizal fungi of continental Europe [44].
The implication of this result is that exogenous factors, such as vegetation composition on separate islands, play a larger role in structuring soil fungal communities than do endogenous factors, such as dispersal limitation. At the largest scale (80–110 km), fungal communities isolated from soil and Selaginella showed opposite patterns of spatial autocorrelation. In previous research of culture-based fungal endophytes, aboveground endophytes exhibited a distance–decay relationship at the regional scale but belowground endophytes did not [45]. It is unclear why fungal communities from different habitats would present such variable results on a large scale. Martiny et al. [19] found that for bacteria, the relative importance of different environmental parameters varied by scale, such that moisture was important at the local scale but temperature and nitrate concentrations were important at regional and continental scales. Since variation in environmental parameters is also scale dependent, we might expect habitat-specific parameters, like edaphic characteristics, to exert differential influence on below- and aboveground fungal communities. A second possible explanation for the observed above/belowground spatial autocorrelation paradigm may be related to the physical spacing of habitats. Soil, as a habitat, forms a nearly continuous habitat coextensive with the forest, whereas Selaginella, as a habitat, is composed of individual plants dispersed in a patchwork across the same landscape. Perhaps Selaginella acts more like islands compared with soil resulting in a patchy habitat across the landscape.
The Relationship Between Plant and Fungal Communities
The effects of distance–decay have long been recognized in plants [11] and animals [46], and efforts have been made to relate biogeographic patterns of plants and animals with those of microbes [12, 31]. In this study, we show that tree communities and fungal communities share a distance–decay relationship, indicating a strong relationship between fungal and tree communities. Furthermore, a direct comparison of plant and fungal DDS demonstrates that the spatial turnover of trees is greater than for above- and belowground fungi (Fig. 2), a result previously reported for global soil fungi [29].
Past studies have indicated that host plant species identity drives an association with specific foliar fungal communities [47, 48], suggesting that beta diversity of plant and endophyte fungal communities is linked. Our results indicate similar patterns (Table 2), but in this case, geographic distance is consistently a stronger factor in structuring aboveground fungal communities (Fig. 3; understory, Selaginella). By contrast, at large geographic scales, tree community composition was better at explaining soil fungi (Fig. 3; soil). This leads us towards two non-mutually exclusive hypotheses. First, at regional scales, correlations between plant and soil fungal communities are best explained by their similar responses to climatic and edaphic variables [29], whereas dispersal limitation is more important in structuring aboveground fungi. Secondly, belowground plant–fungal symbioses are more impactful at structuring forests than are aboveground plant–fungal symbioses (Online Resource 6).
Conclusion
Understanding how environmental factors and those associated with fungal biology contribute to the spatial dynamics of fungal communities is challenging. Here, we contrasted the spatial dynamics of fungal communities from multiple habitats and compared them with turnover in the plant community. Our results show that fungal communities of tropical island forests exhibit strong spatial autocorrelation and that the strength and type of autocorrelation is scale dependent. Fungal communities at local scales (< 50 m) are aggregated and show positive spatial autocorrelation, a trend that is inverted at the island scale (3.7–26 km). The spatial dynamics of fungal communities at a regional scale (~ 80–100 km) are more varied and are dependent on fungal habitat (belowground or aboveground). Across the extent of our study, geographic distance is the dominating factor structuring aboveground fungal communities, whereas belowground, soil fungal community composition is better explained by forest community beta diversity. These results emphasize the importance of considering the scale and spatial autocorrelation in analyses of fungal communities. Furthermore, our findings support a growing body of evidence supporting the idea that fungi and plants exhibit similar biogeographic patterns.
Availability of Data and Material (Data Transparency)
The sequencing dataset analyzed during the current study is available in the NCBI Sequence Read Archive. (BioProject ID PRJNA634909). Tree community and transect data are available from Figshare (doi https://doi.org/10.6084/m9.figshare.12367475).
References
Van Der Heijden MGA, Bardgett RD, Van Straalen NM (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11:296–310. https://doi.org/10.1111/j.1461-0248.2007.01139.x
Schimann H, Bach C, Lengelle J, Louisanna E, Barantal S, Murat C, Buée M (2016) Diversity and structure of fungal communities in Neotropical rainforest soils: the effect of host recurrence. Microb Ecol 73:1–11. https://doi.org/10.1007/s00248-016-0839-0
Bagchi R, Gallery RE, Gripenberg S, Gurr SJ, Narayan L, Addis CE, Freckleton RP, Lewis OT (2014) Pathogens and insect herbivores drive rainforest plant diversity and composition. Nature 506:85–88. https://doi.org/10.1038/nature12911
Van Der Heijden MGA, Boller T, Sanders IR (1998) Different arbuscular mycorrhizal fungal species are potential. Ecology 79:2082–2091
Nacke H, Goldmann K, Schöning I, Pfeiffer B, Kaiser K, Castillo-Villamizar GA, Schrumpf M, Buscot F, Daniel R, Wubet T (2016) Fine spatial scale variation of soil microbial communities under European beech and Norway spruce. Front Microbiol 7. https://doi.org/10.3389/fmicb.2016.02067
Hawksworth DL, Lücking R (2017) Fungal Diversity Revisited : 2 . 2 to 3 . 8 Million Species:1–17. https://doi.org/10.1128/microbiolspec.FUNK-0052-2016.Correspondence
Pimm SL, Joppa LN (2015) How many plant species are there, where are they, and at what rate are they going extinct? Ann Missouri Bot Gard 100:170–176. https://doi.org/10.3417/2012018
Tobler W (1970) A computer movie simulating urban growth in the Detroit region. Econ Geogr 46:234–240
Morlon H, Chuyong G, Condit R, Hubbell S, Kenfack D, Thomas D, Valencia R, Green JL (2008) A general framework for the distance-decay of similarity in ecological communities. Ecol Lett 11:904–917. https://doi.org/10.1111/j.1461-0248.2008.01202.x
Nekola JC, White PS, Biogeography J et al (2007) Special Paper: The distance decay of similarity in biogeography and ecology the distance decay of similarity in biogeography and ecology. J Biogeogr 26:867–878
Condit R, Pitman N, Leigh EG et al (2002) Beta-diversity in tropical forest trees. Science 295(80):666–669. https://doi.org/10.1126/science.1066854
Astorga A, Oksanen J, Luoto M, Soininen J, Virtanen R, Muotka T (2012) Distance decay of similarity in freshwater communities: do macro- and microorganisms follow the same rules? Glob Ecol Biogeogr 21:365–375. https://doi.org/10.1111/j.1466-8238.2011.00681.x
Monroy F, van der Putten WH, Yergeau E, Mortimer SR, Duyts H, Bezemer TM (2012) Community patterns of soil bacteria and nematodes in relation to geographic distance. Soil Biol Biochem 45:1–7. https://doi.org/10.1016/j.soilbio.2011.10.006
Finkel OM, Burch AY, Elad T, Huse SM, Lindow SE, Post AF, Belkin S (2012) Distance-decay relationships partially determine diversity patterns of phyllosphere bacteria on Tamrix trees across the sonoran desert. Appl Environ Microbiol 78:6187–6193. https://doi.org/10.1128/AEM.00888-12
Qvit-Raz N, Finkel OM, Al-Deeb TM et al (2012) Biogeographical diversity of leaf-associated microbial communities from salt-secreting Tamarix trees of the Dead Sea region. Res Microbiol 163:142–150. https://doi.org/10.1016/j.resmic.2011.11.006
Bell T (2010) Experimental tests of the bacterial distance-decay relationship. ISME J 4:1357–1365. https://doi.org/10.1038/ismej.2010.77
Yavitt JB, Yashiro E, Cadillo-Quiroz H, Zinder SH (2012) Methanogen diversity and community composition in peatlands of the central to northern Appalachian Mountain region, North America. Biogeochemistry 109:117–131. https://doi.org/10.1007/s10533-011-9644-5
Saleem M, Pervaiz ZH, Traw MB (2015) Theories, mechanisms and patterns of microbiome species coexistence in an era of climate change. Microbiome community ecology: fundamentals and applications. Springer International Publishing, Cham, pp 13–53
Martiny JBH, Eisen JA, Penn K, Allison SD, Horner-Devine MC (2011) Drivers of bacterial beta-diversity depend on spatial scale. Proc Natl Acad Sci U S A 108:7850–7854. https://doi.org/10.1073/pnas.1016308108
Vaz ABM, Fontenla S, Rocha FS, Brandão LR, Vieira MLA, de Garcia V, Góes-Neto A, Rosa CA (2014) Fungal endophyte β-diversity associated with Myrtaceae species in an Andean Patagonian forest (Argentina) and an Atlantic forest (Brazil). Fungal Ecol 8:28–36. https://doi.org/10.1016/j.funeco.2013.12.008
Cordier T, Robin C, Capdevielle X, Desprez-Loustau ML, Vacher C (2012) Spatial variability of phyllosphere fungal assemblages: genetic distance predominates over geographic distance in a European beech stand (Fagus sylvatica). Fungal Ecol 5:509–520. https://doi.org/10.1016/j.funeco.2011.12.004
Vincent JB, Weiblen GD, May G (2016) Host associations and beta diversity of fungal endophyte communities in New Guinea rainforest trees. Mol Ecol 25:825–841. https://doi.org/10.1111/mec.13510
Oono R, Rasmussen A, Lefèvre E (2017) Distance decay relationships in foliar fungal endophytes are driven by rare taxa. Environ Microbiol 00:2794–2805. https://doi.org/10.1111/1462-2920.13799
Kadowaki K, Sato H, Yamamoto S, Tanabe AS, Hidaka A, Toju H (2014) Detection of the horizontal spatial structure of soil fungal communities in a natural forest. Popul Ecol 56:301–310. https://doi.org/10.1007/s10144-013-0424-z
Bahram M, Kõljalg U, Courty PE, Diédhiou AG, Kjøller R, Põlme S, Ryberg M, Veldre V, Tedersoo L (2013) The distance decay of similarity in communities of ectomycorrhizal fungi in different ecosystems and scales. J Ecol 101:1335–1344. https://doi.org/10.1111/1365-2745.12120
Baldrian P (2017) Forest microbiome: diversity, complexity and dynamics. FEMS Microbiol Rev 41:109–130. https://doi.org/10.1093/femsre/fuw040
Barberan A, Mcguire KL, Wolf JA et al (2015) Relating belowground microbial composition to the taxonomic, phylogenetic, and functional trait distributions of trees in a tropical forest. Ecol Lett 18:1397–1405. https://doi.org/10.1111/ele.12536
von Humboldt A, Bonpland A (1807) Essai sur la géographie des plantes
Tedersoo L, Bahram M, Põlme S et al (2014) Global diversity and geography of soil fungi. Science (80):346. https://doi.org/10.1126/science.1256688
Peay KG, Bidartondo MI, Elizabeth Arnold A (2010) Not every fungus is everywhere: scaling to the biogeography of fungal-plant interactions across roots, shoots and ecosystems. New Phytol 185:878–882
Meyer KM, Memiaghe H, Korte L, Kenfack D, Alonso A, Bohannan BJM (2018) Why do microbes exhibit weak biogeographic patterns? ISME J 12:1404–1413. https://doi.org/10.1038/s41396-018-0103-3
Mallick DIJ (1975) Development of the New Hebrides Archipelago. Philos Trans R Soc Lond Ser B Biol Sci 272:277–285. https://doi.org/10.1098/rstb.1975.0087
Mueller-Dombois D, Fosberg FR (2013) Vegetation of the tropical Pacific islands. Springer Science & Business Media
Vanuatu Meteorology and Geo-hazard Department, Australian Bureau of Meteorology, CSIRO (2015) Current and future climate of Vanuatu. Melbourne, Australia
McMurdie PJ, Holmes S (2014) Waste not, Want not: why rarefying microbiome data is inadmissible. PLoS Comput Biol 10:e1003531. https://doi.org/10.1371/journal.pcbi.1003531
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
R Core Team (2019) R: A language and environment for statistical computing. Accessed 1st April 2019
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PloS one 8(4):e61217
Ferrier S, Manion G, Elith J, Richardson K (2007) Using generalized dissimilarity modelling to analyse and predict patterns of beta diversity in regional biodiversity assessment. Divers Distrib 13:252–264. https://doi.org/10.1111/j.1472-4642.2007.00341.x
Fitzpatrick MC, Sanders NJ, Normand S, Svenning JC, Ferrier S, Gove AD, Dunn RR (2013) Environmental and historical imprints on beta diversity: insights from variation in rates of species turnover along gradients. Proc R Soc B Biol Sci 280:20131201–20131201. https://doi.org/10.1098/rspb.2013.1201
Robinson CH, Szaro TM, Izzo AD, Anderson IC, Parkin PI, Bruns TD (2009) Spatial distribution of fungal communities in a coastal grassland soil. Soil Biol Biochem 41:414–416. https://doi.org/10.1016/j.soilbio.2008.10.021
Peay KG, Kennedy PG, Talbot JM (2016) Dimensions of biodiversity in the earth mycobiome. Nat Rev Microbiol 14:434–447. https://doi.org/10.1038/nrmicro.2016.59
Tuomisto H, Ruokolainen K, Yli-Halla M (2003) Dispersal, environment, and floristic variation of Western Amazonian forests. Science 299(80):241–244. https://doi.org/10.1126/science.1078037
Van Der Linde S, Suz LM, Orme CDL et al (2018) Environment and host as large-scale controls of ectomycorrhizal fungi. Nature 558:243–248. https://doi.org/10.1038/s41586-018-0189-9
David AS, Seabloom EW, May G (2016) Plant host species and geographic distance affect the structure of aboveground fungal Symbiont communities, and environmental filtering affects belowground communities in a coastal dune ecosystem. Microb Ecol 71:912–926. https://doi.org/10.1007/s00248-015-0712-6
Poulin R (2003) The decay of similarity with geographical distance in parasite communities of vertebrate hosts. J Biogeogr 30:1609–1615. https://doi.org/10.1046/j.1365-2699.2003.00949.x
Kembel SW, Mueller RC (2014) Plant traits and taxonomy drive host associations in tropical Phyllosphere fungal communities. Botany. https://doi.org/10.1139/cjb-2013-0194
Thomas D, Vandegrift R, Roy BA, Hsieh HM, Ju YM (2019) Spatial patterns of fungal endophytes in a subtropical montane rainforest of northern Taiwan. Fungal Ecol 39:316–327. https://doi.org/10.1016/j.funeco.2018.12.012
Acknowledgments
The authors would like to thank the reviewers and Drs. Nicole Hynson, Tom Ranker, Nhu Nguyen, and Michael Kantar for improving this manuscript. We are also very appreciative of Presley Dovo and the Vanuatu Department of Forestry for logistical support. This project would not be possible without field support from the ever-growing network of people associated with Plants mo Pipol blong Vanuatu. We would also like to recognize the contributions of the late Philemon Ala who had been helping with Plants mo Pipol since its inception. Finally, we are grateful to the many communities of Aneityum and Tanna for their kindness, hospitality, and for sharing so much invaluable knowledge, tankyu tumas.
Funding
A.B., T.T., and A.S.A. were supported by a grant from the National Science Foundation (1555793). G.M.P. was supported by a National Science Foundation grant awarded to NYGB (1555657).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Boraks, A., Plunkett, G.M., Doro, T.M. et al. Scale-Dependent Influences of Distance and Vegetation on the Composition of Aboveground and Belowground Tropical Fungal Communities. Microb Ecol 81, 874–883 (2021). https://doi.org/10.1007/s00248-020-01608-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-020-01608-4