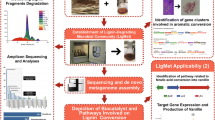
Abstract
An exploration of the ligninolytic potential of lignocellulolytic microbial consortia can improve our understanding of the eco-enzymology of lignin conversion in nature. In this study, we aimed to detect enriched lignin-transforming enzymes on metagenomes from three soil-derived microbial consortia that were cultivated on “pre-digested” plant biomass (wheat straw, WS1-M; switchgrass, SG-M; and corn stover, CS-M). Of 60 selected enzyme-encoding genes putatively involved in lignin catabolism, 20 genes were significantly abundant in WS1-M, CS-M, and/or SG-M consortia compared with the initial forest soil inoculum metagenome (FS1). These genes could be involved in lignin oxidation (e.g., superoxide dismutases), oxidative stress responses (e.g., catalase/peroxidases), generation of protocatechuate (e.g., vanAB genes), catabolism of gentisate, catechol and 3-phenylpropionic acid (e.g., gentisate 1,2-dioxygenases, muconate cycloisomerases, and hcaAB genes), the beta-ketoadipate pathway (e.g., pcaIJ genes), and tolerance to lignocellulose-derived inhibitors (e.g., thymidylate synthases). The taxonomic affiliation of 22 selected lignin-transforming enzymes from WS1-M and CS-M consortia metagenomes revealed that Pseudomonadaceae, Alcaligenaceae, Sphingomonadaceae, Caulobacteraceae, Comamonadaceae, and Xanthomonadaceae are the key bacterial families in the catabolism of lignin. A predictive “model” was sketched out, where each microbial population has the potential to metabolize an array of aromatic compounds through different pathways, suggesting that lignin catabolism can follow a “task division” strategy. Here, we have established an association between functions and taxonomy, allowing a better understanding of lignin transformations in soil-derived lignocellulolytic microbial consortia, and pinpointing some bacterial taxa and catabolic genes as ligninolytic trait-markers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Agricultural residues are an abundant source of sugars used for production of commodity chemicals (e.g., biofuels). However, the high complexity of plant biomass decreases the saccharification (i.e., release of monosaccharides) rates in biorefineries [1]. This drawback has been attributed to the recalcitrant nature of plant cell walls and to strong linkages (e.g., ester linkage) between its main components (lignin, cellulose, and xylan) [2]. Lignin is a highly heterogeneous aromatic polymer network formed via radical coupling reactions involving the three major monolignols: p-coumaryl, coniferyl, and sinapyl alcohol, linked by C–C and C–O bonds [3]. Delignification of agricultural residues could increase enzymatic access to plant polysaccharides, enhancing the saccharification rate [4]. Although lignin does not contain sugars, valorization of its aromatic constituents is a relevant topic for bioenergy, food, and cosmetics industries [5,6,7]. Lignin mineralization requires two main steps: (1) extracellular lignin depolymerization (carried out mainly by peroxidases, multicopper oxidases, and/or laccases) to yield mono, di, and oligomers; and (2) ring fission of the resulting aromatic compounds. Lignin-derived aromatic compounds are metabolized intracellularly by different type of microbes with specific metabolic capabilities. Here, a niche partitioning of lignin-degrading microbes might be a common feature. During lignin catabolism, aromatic compounds are typically shunted through a number of reactions referred to as “funneling pathways.” Eventually, they converge on a couple of conserved “ring fission pathways” (e.g., beta-ketoadipate) where aromatic rings of the intermediate compounds (e.g., protocatechuate, gentisate, and/or catechol) are cleaved. The subsequent catabolic products (e.g., pyruvate, acetyl-coA, oxaloacetate, fumarate, and/or succinate) can enter the tricarboxylic acid cycle (TCA) to generate energy [8,9,10,11,12].
In terrestrial and aquatic ecosystems, different types of microbes transform lignin. These are typically white rot fungi (e.g., Phanerochaete chrysosporium), filamentous fungi, yeast, and bacteria mostly belonging to Actinobacteria (e.g., Streptomyces, Arthrobacter, and Rhodococcus species) and Proteobacteria (e.g., Pseudomonas, Sphingobium, Enterobacter, and Sphingobacterium species) [13, 14]. Apparently, bacterial populations, instead of fungal ones, contribute significantly to degradation of native forms of lignin in forest soils and seawater [15,16,17]. However, scientific knowledge about the eco-enzymology (defined here as the study of enzymes and their role in microbial interactions and the modification of surrounding environments) of lignin degradation is very limited. Here, we provide evidence that a complete catabolism of lignin and its derived aromatic compounds is a complex process carried out by microbial communities (and their enzymes) instead of a single species, similar to (hemi)cellulose bioconversion [16, 18]. Lignocellulolytic microbial consortia have been constructed using the dilution-to-stimulation approach in which natural communities (e.g., from soils) are selected based on a unique carbon source (e.g., agricultural residues) [19]. This strategy allows enrichment of different microbial populations that have the capability to deconstruct plant polymers [20, 21]. In recent years, multi-omic strategies (i.e., metasecretomics and metagenomics) have been used to identify enzymatic mechanisms for polysaccharide degradation functioning in these lignocellulolytic microbial consortia [22,23,24,25]. However, a comprehensive understanding of lignin transformation in these systems is still missing. Recently, Moraes et al. [18] explored the ligninolytic potential of a soil-derived microbial consortium cultivated on soluble lignin. This lignin was obtained after the acidification of black liquor generated from delignification of steam-exploded sugarcane bagasse. Other environments, such as mangrove sediments, redwood compost, decay wood, and Mediterranean Sea, have also been used as initial inoculum to enrich lignin-adapted microbial populations [17, 26,27,28]. In these latter studies, bacterial 16S rRNA amplicon sequencing and shotgun metagenomic analyses were performed to evaluate the microbial composition and ligninolytic potential within the consortia. A metagenomic exploration on lignin-enriched and low diversity systems (i.e., lignocellulolytic microbial consortia) can help to improve our understanding of lignin catabolism in nature. These approaches can be useful to identify novel pathways, key microbes, genes, and/or enzymes as ligninolytic trait-markers. In addition, such studies could inform further lignin valorization strategies.
In this study, we explored the lignin-transforming potential of three lignocellulolytic microbial consortia obtained from forest soil and cultivated on “pre-digested” plant biomass (wheat straw, switchgrass, and corn stover) under aerobic and mesophilic conditions. We hypothesized that the use of biological-pretreated plant biomass can favor the presence of bacteria able to degrade recalcitrant plant polymers (e.g., lignin) [23]. In order to evaluate the ligninolytic potential of these engineered microbial consortia, a metagenome “gene-centric” approach was set up, selecting 60 enzyme-encoding genes that are putative involved in lignin depolymerization and metabolism of its derived aromatic compounds. Moreover, the taxonomic affiliation of some genes allowed us to link specific functions with particular taxa. Our results suggest that the catabolism of lignin in these microbial systems is a specialized process in which each taxon has its own niche and can perform a specific job, following the “task division” strategy.
Materials and Methods
Construction of Plant Biomass-Degrading Microbial Consortia
The soil-derived lignocellulolytic microbial consortia were developed following a dilution-to-stimulation approach [23, 29]. Briefly, soil suspension was added to triplicate flasks containing mineral salt medium with 1% of plant biomass, trace mineral, and vitamin solutions. Flasks were incubated at 28 °C in oxic conditions (with shaking at 150 rpm). Once systems reached high bacterial cell density (7–8 log cells/ml, between 5 and 6 days and determined by microscopical cell counting), aliquots of microbial suspension were transferred to fresh medium (diluted 1000-fold). These procedures were repeated ten times. Five different consortia were analyzed in this study: (1) cultivated on fresh and heat-treated wheat straw (10-RWS and 10-TWS; transfer 10) [22], and (2) cultivated on “once used” or “pre-digested” (highly recalcitrant) wheat straw, corn stover, and switchgrass (WS1-M, CS-M and SG-M, respectively). The “pre-digested” substrates were obtained after a biological pretreatment with different soil-derived lignocellulolytic microbial consortia [23].
Soil-Derived Lignocellulolytic Microbial Consortia Metagenomes
Total DNA from the lignocellulolytic microbial consortia (10-RWS, 10-TWS, WS1-M, CS-M, and SG-M) was extracted using the UltraClean Microbial DNA Isolation Kit (MoBio Laboratories Inc., Carlsbad, CA, USA). Metagenome sequencing was performed using the Illumina MiSeq v2 (2 × 250 bp paired-end reads) at LGC Genomics (Berlin, Germany). Forest soil (initial inoculum) metagenome (FS1) was sequenced using the same platform [22, 23]. For the purpose of comparison, we have also used a dataset retrieved from a ligninolytic microbial consortium (denoted LigMet). This metagenome was sequenced using the Illumina HiSeq 2500 (2 × 100 bp paired-end reads) [18]. The total metagenomic information retrieved was approximately 136 Mb for FS1; 112 Mb for 10-RWS; 198 Mb for 10-TWS; 1.5 Gb for WS1-M; 1.6 Gb for SG-M; 1.8 Gb for CS-M; and 18 Gb for LigMet.
Metagenomic Analysis (“Gene-Centric” Approach)
All metagenomes (unassembled sequences reads) were uploaded to the MG-RAST v3.1.2 server [30]. Overlapping sequence pairs were matched, and non-overlapping reads retained as individual reads, after which, dereplication was performed. Duplicate read-based inferred sequencing error estimation and quality trimming (phred score < 20) used default settings. Gene predictions were done using the FragGeneScan software, and subsequently, the proteins were annotated based on BLASTX searches against the RefSeq and KEGG databases using an e-value cutoff of 1e-5, a minimum alignment length of 50 amino acids, and a minimum identity of 50%. Data from MG-RAST annotation were statistically analyzed using the STAMP package [31]. This software was used also to obtain correlation values of the taxonomic (genus level) and functional (KO level) profiles across the metagenomes. All metagenome datasets are publicly accessible on the MG-RAST server IDs 4547279.3 (10-TWS); 4547280.3 (10-RWS); 4547285.3 (FS1); 4790808.3 (LigMet); 4579477.3 to 4579479.3 (CS-M); 4579485.3 to 4579487.3 (SG-M); and 4579476.3, 4579480.3 and 4579481.3 (WS1-M).
Selection and Analysis of Genes Encoding Lignin-Metabolizing Enzymes
Based on an in-depth comprehensive bibliographic search [7, 9,10,11, 27, 32,33,34,35], we have selected 60 enzyme-encoding genes (KO IDs) that could be involved in lignin depolymerization, oxidative stress response, catabolism of lignin-derived aromatic compounds (e.g., funneling and fission pathways), and tolerance to lignocellulose-derived inhibitors. For each enzyme, EC numbers, KO IDs, names, and functions are shown in supplementary information Table S1. A matrix of read counts per gene in all the metagenomes was analyzed using the STAMP software [31]. To evaluate the relative abundance of reads per selected enzyme-encoding gene, the counts were normalized to hits, or unique matches, per million reads, thereby accounting for differences in metagenome sizes [36]. In order to evaluate significant differences in gene abundance within each consortium, an ANOVA and post hoc Tukey-Kramer test were performed using the software R with a confidence interval of 95% (α = 0.05). Heat maps were constructed in the web server Heatmapper using row Z score for each enzyme [37]. Differential abundance analysis was used to determine which lignin-metabolizing enzymes were highly enriched in WS1-M, CS-M, and SG-M consortia compared with forest soil inoculum (FS1). This analysis was conducted using the R package DESeq2 with a log fold change > 1 [17, 38]. Unassembled sequence reads (in FASTA format) belonging to the selected 60 lignin-metabolizing enzymes were extracted from WS1-M and CS-M using the MG-RAST webserver. The taxonomic assignment was performed using the lowest common ancestor (LCA) algorithm within the Kaiju web server [39]. These sequences were also clustered at 97 and 99% of nucleotide identity using the CD-HIT software [40]. Using the taxonomic affiliation and read counts in each gene, we have built a PCA in the software R.
Results and Discussion
In this study, we aimed to unveil the ligninolytic potential of different soil-derived lignocellulolytic microbial consortia. For this purpose, 60 catabolic genes that are putative involved in lignin depolymerization, oxidative stress response, and catabolism of lignin-derived aromatic compounds were evaluated (supplementary information Table S1). The selected gene list (i.e., KO IDs) could be a useful input to explore the lignin-degrading potential in other meta-omic surveys. Here, we have used a metagenomic “gene-centric” approach, instead “genome-centric” approach, as this is known to be useful to evaluate gene differential abundance profiles and cause minimal disturbance with respect to representation of sequences within the abundant taxa [41]. It is important to note that our study has a drawback inherent to any metagenomic study, and unfortunately, we cannot claim enzymatic activity based on similarity and/or functional annotation of DNA sequences. However, our approach allowed us to link potential functions with particular taxa, improving our understanding of the eco-enzymology of the lignin transformation process in these engineered microbial systems.
Clustering of Lignocellulolytic Microbial Consortia Based on Their Taxonomic/Functional Profile
In a previous study, we selected and characterized three soil-derived lignocellulolytic microbial consortia, denoted WS1-M (cultivated on wheat straw), CS-M (cultivated on corn stover), and SG-M (cultivated on switchgrass). These consortia were obtained using a new approach in which the plant biomass used to cultivate them was first partially degraded (“pre-digested”) by another lignocellulolytic microbial consortium [21, 23]. This strategy allows microbes to be selected with high capacities to degrade highly complex plant polysaccharides, as well as lignin. The compositional profile obtained by Fourier transform infrared (FTIR) spectroscopy showed that the “pre-digested” wheat straw contained approximately 16% lignin, while switchgrass and corn stover only 11% [23]. Previous metagenomic sequence annotation showed that bacteria, rather than fungal communities, dominate in the WS1-M, CS-M, and SG-M consortia [23]. Here, based on a PCA using RefSeq (taxonomic) annotation profile, we observed that SG-M and CS-M consortia clustered together (R2 > 0.88) (Fig. 1a). A similar result was observed using the KEGG database (Fig. 1b). These findings were already reported in the previous study [23]. In terms of taxonomic composition, SG-M and CS-M are very similar, showing high abundance of sequences affiliated to species belonging to Pseudomonadaceae and Caulobacteraceae families. In contrast, Bacteriodetes species were preferentially selected in WS1-M [23]. Wheat straw-degrading consortia (WS1-M, 10-RWS and 10-TWS) showed a similar functional profile based on total KEGG annotation (R2 > 0.75) (Fig. 1b). It is important to highlight that all consortia evaluated in this study were retrieved from the same forest soil inoculum, except LigMet. Interestingly, SG-M and CS-M showed the same functional profile when lignin-transforming enzymes were used to build the clustering (R2 > 0.97). However, this ligninolytic profile was highly dissimilar compared with the LigMet consortium and the forest soil inoculum (FS1 metagenome) (Fig. 1c). The consortium LigMet was retrieved from agricultural tropical soils and cultivated on soluble lignin [18]. Thus, these conditions can explain its dissimilar taxonomic and functional profile compared with the other forest soil-derived lignocellulolytic microbial consortia. The results indicated that the substrates used to develop the consortia are key factors to shape the taxonomic and ligninolytic potential in these engineered microbial systems.
Taxonomic and functional clustering of lignocellulolytic microbial consortia based on metagenome annotation. a Clustering based on taxonomic assignment (RefSeq database) of annotated sequences at genus level. b Clustering based on functional assignment (KEGG databases) of annotated sequences at KOs level. c Functional clustering based on the 60 selected enzyme-encoding genes involved in transformation of lignin and its derived aromatic compounds. Black squares correspond to initial forest soil inoculum (FS1) metagenome used to build the consortia WS1-M, CS-M, SG-M, 10-RWS, and 10-TWS
Abundance of Lignin-Transforming Enzyme-Encoding Genes
Although the WS1-M consortium showed a different functional profile compared with CS-M and SG-M, some lignin-transforming enzymes were highly abundant in all three consortia (p ≤ 0.05), for instance, alcohol dehydrogenases, glutathione S-transferases (GST), and catalases (katE) (Fig. 2). Proteins involved in tolerance to lignocellulose-derived inhibitors (S-(hydroxymethyl)glutathione dehydrogenase - frmA) and catabolism of gentisate (gentisate 1,2-dioxygenase) were highly abundant in CS-M and SG-M compared with WS1-M. In contrast, a vanillate monooxygenase (vanB) was highly abundant in WS1-M compared with CS-M and SG-M. Some specialized types of GST (e.g., beta-etherase) are involved in lignin depolymerization by cleaving beta-aryl ether linkages [42]. Its high abundance in the consortia can be correlated with the number of gene copies in Pseudomonas species. For instance, 14 GST genes have been identified in Pseudomonas putida KT2440 [14]. However, not all GST proteins have beta-etherase activity, and in-depth functional analysis of these sequences would be worthwhile. Enzyme-encoding genes involved in lignin depolymerization and/or oxidative stress responses (e.g., GST, katG, superoxide dismutase - SOD2, glutathione peroxidase, and glycolate oxidase - glcD) were highly abundant on the three consortia (Fig. 2). Manganese superoxide dismutases from Sphingobacterium sp. T2 have been identified as novel lignin-oxidizing enzymes, which are able to solubilize organosolv and kraft lignin to generate a mixture of polymeric and monocyclic aromatic products [43]. Our results showed that GST, glcD, and (S)-2-hydroxy-acid oxidase (HAO) were also abundant in the forest soil inoculum (Fig. 3). In particular, glcD and HAO could be accessory enzymes for lignin degradation, by generating hydrogen peroxide for peroxidase enzymes, and potentially detoxifying aldehyde by-products [44]. These findings can be an indication of the large enzymatic potential to breakdown lignin in the soil-derived consortia. In fact, WS1-M, CS-M, and SG-M showed values of lignin degradation of 25.3 ± 1.8%, 24.7 ± 1.2%, and 58.6 ± 1.0% over 6 days, respectively, obtained by FTIR spectroscopy [23]. The high value observed on SG-M can be explained by the lower lignin recalcitrance in switchgrass, compared with wheat straw and corn stover. Similarly, Wang et al. [45] have reported a bacterial consortium that could break down 60.9% of lignin in reeds (i.e., grass-like plants of wetlands) over 15 days.
Number of normalized sequences (per million) that were annotated within the 60 enzyme-encoding genes involved in transformation of lignin and its derived aromatic compounds. The top 16 ligninolytic enzyme-encoding genes that were significantly abundant (p ≤ 0.05) in WS1-M, CS-M, or SG-M are shown at the bottom. Statistical differences in gene abundance within each consortium are indicated with lowercase letters (ANOVA, p ≤ 0.05). vanB (vanillate monooxygenase, in bold) was significant abundant only in WS1-M, whereas aryl-alcohol dehydrogenase (in bold) was significant abundant only in CS-M
Heat map of normalized abundance values (Row Z score) obtained using the number of sequences annotated within the 60 enzyme-encoding genes involved in transformation of lignin and its derived aromatic compounds in each microbial consortium. Genes differentially and significantly enriched (padj-value ≤ 0.05, Wald test; and Log2 FC ≥ 1) in WS1-M, CS-M, and/or SG-M compared with FS1 are labeled in blue
Unfortunately, multicopper oxidases and dye-decolorizing peroxidases (DyPs) (e.g., K15733; EC, 1.11.1.19) involved in bacterial lignin oxidation processes were not identified in the metagenomic annotation, suggesting a low copy numbers within bacterial genomes and/or due to the relative low number of representative sequences available in the KEGG database. In fact, only three genes encoding for DyPs have been found in P. fluorescens [46], and around 100 DyPs sequences from bacterial origin are available in a specialized database (http://peroxibase.toulouse.inra.fr/) [14]. As was mentioned, the extracellular depolymerization of lignin releases a mixture of aromatic monomers that can be converted into metabolic intermediates via catechol and protocatechuate pathways. These routes can be divided in three blocks: (1) the branch of catechol intermediate (ortho-cleavage), which involves the following enzyme-encoding genes: catA, catB, and catC; (2) the branch of protocatechuate (meta-cleavage), which involves pcaG, pcaH, pcaB, and pcaC; and finally, (3) the reactions common for both branches, catalyzed by pcaD, pcaI, pcaJ, and pcaF [11, 18]. In this regard, we found that the metagenomes from the consortia 10-RWS and 10-TWS contained a high abundance of pca genes (e.g., pcaBCGHIJ) compared with the other consortia (Fig. 3), suggesting a high proportion of low molecular weight aromatic compounds in these systems. These microbial consortia were cultivated on untreated and heat-treated wheat straw [22], where lignin proportion is higher, but probably with lower recalcitrance compared with substrates used on the selection of WS1-M, SG-M, and CS-M. In a recent proteomic-based study, Park et al. [47] demonstrated that P. putida KT2440 significant induces the production of pca-derived proteins after growth on plant-derived lignolysate.
Differential and Significant Enrichment of Lignin-Transforming Enzyme-Encoding Genes
Using relative abundance matrixes of the 60 enzyme-encoding genes (annotated by KEGG Orthology database) and the R package DESeq2, we identified which enzymes were significantly enriched in WS1-M, CS-M, and/or SG-M compared with the initial forest soil inoculum (FS1 metagenome) (Fig. 3). Moreover, in Fig. 4, we present a scatter plot that shows the most highly abundant genes in each microbial consortium compared with FS1. Based on the results, we found that 20 genes (~ 33%) were highly abundant (padj-value ≤ 0.05, Wald test; and Log2 FC ≥ 1) on WS1-M, CS-M, and/or SG-M compared with FS1 (Fig. 3). These genes could be involved in oxidative stress response (e.g., katE), lignin depolymerization/oxidation (e.g., superoxide dismutase - SOD1), generation of protocatechuate/gallate (e.g., vanillate monooxygenases - vanAB and methylenetetrahydrofolate reductase - metF), catabolism of catechol (e.g., muconate cycloisomerase - catB and muconolactone D-isomerase - catC), gentisate (e.g., gentisate 1,2-dioxygenase), and 3-phenylpropionic acid (e.g., hca genes). Additionally, genes involved in the beta-ketoadipate pathway (pcaI and pcaJ), thymidylate synthases (thyA), and aryl-alcohol dehydrogenases (K00055) were also highly abundant compared with FS1. Interesting, the pca genes in P. putida A514 are upregulated in response to lignin [48] and thymidylate synthases conferred tolerance to lignin monomers such as furfural, ferulic acid, vanillic acid, and syringic acid [49]. The vanA, vanB, katE, catC, and metF genes were highly abundant in the LigMet consortia. In addition, catC was highly abundant in the 10-RWS metagenome (Fig. 3). In particular, the O-demethylation of vanillate is catalyzed by the operon ligM-metF-ligH. This process can generate catecholic compounds, protocatechuate, and gallate. However, the O-demethylation steps are also important for the production of 5-methyl-H4folate, which is a C1-H4folate derivative in one-carbon (C1) metabolism [32]. Moreover, Ceballos et al. [27] have developed a lignin-degrading microbial consortium under high-solids and thermophilic conditions. In terms of enzyme-encoding genes that were identified by a predictive metagenomic approach, they found an enrichment of vanA, protocatechuate 3,4-dioxygenase (pcaG), catechol 1,2-dioxygenase (catA), catechol 2,3-dioxygenase (dmpB), catB, and aryl-alcohol dehydrogenase. In a study reported by Carlos et al. [50], eight filter papers and wood chip-degrading microbial consortia were “perturbed” in alkali lignin as the sole carbon source. They found a significant enrichment of enzyme-encoding genes involved in catechol ortho-cleavage, especially catA that cleaves the bond between the phenolic hydroxyl groups of catechol generating cis, cis-muconic acid. These type of catabolic genes (i.e., catA), and other genes involved in the phenylacetyl-CoA pathway, were highly abundant on a lignin-adapted consortia retrieved from Eastern Mediterranean seawater [17]. Although catA and dmpB were not significantly enriched in WS1-M, CS-M, and/or SG-M compared with FS1, they were highly abundant in the LigMet and 10-RWS consortia, suggesting that they could be key genes involved in lignin transformation through catechol catabolism. Notably, the gene dmpB has been found in a plasmid from a phenol-metabolizing Pseudomonas strain CF600. This enzyme-encoding gene catalyzes the conversion of catechol to 2-hydroxymuconic semialdehyde [51]. Moreover, alcohol dehydrogenases were highly abundant in the three consortia (e.g., adh) (Fig. 2), and one type of aryl-alcohol dehydrogenases was significant abundant in CS-M and SG-M compared with FS1 (Fig. 3). We suggest that these two proteins could be involved in oxidation of low molecular weight alcohols (e.g., coniferyl alcohol) that can be released after lignin depolymerization/oxidation [32, 52]. However, an in-depth functional analysis may be required to support this statement.
Comparison of ligninolytic profiles (percentage of relative abundance of genes involved in lignin transformation) between the soil inoculum (FS1) and metagenomes from the microbial consortia. Letters a to r indicate the most overrepresented gene functions in the observed metagenomes. In blue are the genes differentially and significantly abundant (padj-value ≤ 0.05, Wald test; and Log2 FC ≥ 1) in WS1-M, CS-M, and/or SG-M compared with FS1
Taxonomic Affiliation of Lignin-Transforming Enzyme-Encoding Genes
From the 60-ligninolytic genes selected, 22 were used for the taxonomic affiliation, in the WS1-M and CS-M metagenomes, using the LCA algorithm (Fig. 5). We have selected these consortia due to two main reasons: (1) CS-M and SG-M were highly similar between them, and (2) we wanted to explore the consortia cultivated on “pre-digested” agricultural residues, instead of switchgrass. Otherwise, enzyme-encoding gene selection was done taking in account three parameters: (1) genes that contained more than 20 assigned metagenomic reads; (2) genes that were differentially enriched compared with FS1 (Fig. 3); and (3) genes that are putative involved the catabolism of different lignin-derived aromatic intermediates (e.g., protocatechuate, gallate, gentisate, and catechol). Two genes involved in oxidative stress response (katG and katE) and one in lignin oxidation (superoxide dismutase - SOD1) were also selected. In a recent study, Rashid et al. [53] have found that extracellular MnSOD1 protein from Sphingobacterium sp. requires two mutations to have lignin demethylation activity. Apparently, these mutations are only found in Bacteroidetes phylum. In our two consortia, the SOD1-encoding genes were mostly affiliated to Pseudomonadaceae, Xanthomonadaceae, Alcaligenaceae, and Caulobacteraceae. Thus, we are doubtful whether these proteins can have ligninolytic activity. Moreover, in order to obtain a proxy of sequence diversity within each enzyme-encoding gene, we have used values of operational functional units (OFUs) per thousands of reads. The OFU values were obtained by clustering the sequences (with size average of 250 bp), affiliated within each catabolic gene, using a cutoff of > 97% nucleotide sequence similarity. These analyses were carried out using the CD-HIT software (http://weizhongli-lab.org/cd-hit/) [40]. It is important to clarify that we did not assigned taxa to the predicted OFUs. The results of OFUs/thousands of reads showed that genes katE and katG were highly diverse in WS1-M and CS-M consortia (≥ 0.5). This indicates that the metabolism of reactive oxygen species (by action of catalases/peroxidases), which could be correlated with lignin degradation, is a process where different taxa (mostly Pseudomonadaceae, Xanthomonadaceae, Alcaligenaceae, Caulobacteraceae, Flavobacteriaceae, Sphingobacteriaceae, and Sphingomonadaceae species) can be involved (Fig. 5). In contrast, low values of sequence diversity in catabolic genes involved in intracellular conversion of lignin-derived aromatics compounds were observed (< 0.2 OFUs/thousands of reads), except metF and thyA.
Taxonomic affiliation, using the lowest common ancestor algorithm (LCA), of 22 enzyme-encoding genes involved in transformation of lignin and its derived aromatic compounds in a WS1-M and b CS-M microbial consortia. Data in top panel are normalized functional diversity values (OFUs, operational functional units/thousands of annotated reads) obtained by clustering the sequences in each gene at 97% (line) and 99% (dashed line) similarity
Recently, Moraes et al. [18] have developed a lignin-degrading microbial consortium composed of 355 bacterial types. Based on 16S rRNA amplicon sequencing data, they demonstrated that around 50% of the total consortium comprised Achromobacter (Alcaligenaceae family), Paenarthrobacter (novel Actinobacteria able to transform lignin), Pseudaminobacter, and Paenibacillus species. These taxa were enriched in the consortium compared with sugarcane soil inoculum. Based on the metabolic functional profile unveiled by metagenomics, they showed that species from Actinobacteria and Proteobacteria contain putative novel enzyme-encoding genes involved in the metabolism of lignin. For instance, peroxidases and laccases were frequently found in Actinobacteria. In contrast, both taxa have the potential to metabolize lignin-derived phenolic compounds. Moreover, Granja-Travez et al. [14] state that exist a high diversity of mechanism for lignin oxidation process in different bacteria and not just one class of enzyme carry out this function in all microbial systems. These observations suggest that lignin transformation is a process in which multiple microbes are involved. However, the intracellular catabolism of its derived aromatic compounds could be a specialized job, in which particular taxa can metabolize specific lignin-derived intermediates.
Based on the taxonomic affiliation of sequences affiliated to genes hcaA1 (3-phenylpropionate/cinnamic acid dioxygenases) and hcaB (2,3-dihydroxy-2,3-dihydrophenylpropionate dehydrogenases), we suggest that, in both consortia, species from Caulobacteraceae, Alcaligenaceae, and Enterobacteriaceae could be involved in the catabolism of 3-phenylpropionic acid (Fig. 5, Fig. 6, and supplementary information Fig. S1). There are two branches at the start of the 3-phenylpropionate catabolic pathway: one branch is from phenylpropionic acid, which is dihydroxylated via the hcaAB genes to 2,3-dihydroxyphenylpropionic acid. In this regard, phenylpropionic acid has been found in anaerobic microbial systems treating lignocellulose [54, 55]. In this case, we hypothesized that 3-phenylpropionic acid could be generated from anaerobic lignin degradation processes, probably via dehydroxylation of hydroxycinnamic acids, occurring in the lignocellulolytic consortia (Fig. 6). Moreover, sequences assigned to catabolic genes vanA, vanB, catC, catB, pcaB (3-carboxy-cis, cis-muconate cycloisomerase), and benD-xylL (dihydroxycyclohexadiene carboxylate dehydrogenase) were mostly affiliated to Pseudomonadaceae (Fig. 5), suggesting that species within this family play key roles in the metabolism of protocatechuate, gallate, and catechol in both consortia. Similarly, Carlos et al. [50] found an enrichment of genes involved in catechol ortho-cleavage that were mostly affiliated to Pseudomonas species in a lignin-adapted consortium.
Schematic representation of transformation of lignin-derived aromatic compounds, showing genes/taxa (from WS1-M and CS-M microbial consortia) putatively involved in each catabolic pathway. This figure was build based on a figure reported by Brink et al. [11] that shows a schematic distribution of the known pathways for aromatic catabolism currently indexed in the eLignin database. In this figure, the 22 selected catabolic genes selected for taxonomic assignment were plotted based on their function within these pathways (bold arrows). Colored circles represent the taxa that could be involved in each catabolic step. The size of each circle is an estimation of the data obtained from Fig. 5 and supplementary information Fig. S1. Asterisks represent genes that were significantly abundant in WS1-M and/or CS-M compared with FS1
Interestingly, the taxonomic assignment of sequences affiliated to the enzyme gentisate 1,2-dioxigenase shows that the catabolism of gentisate is a process where Caulobacteraceae species can play a unique and pivotal role, especially in CS-M consortium (Fig. 5b and supplementary information Fig. S1). DeAngelis et al. [15] have reported enrichments of Caulobacteraceae bacterial types in lignin-amended tropical soils compared with unamended ones. Similarly, Woo and Hazen [17] reported an increase of Caulobacteraceae species (along with others) in a seawater-derived ligninolytic microbial consortium, compared with xylan and unamended microcosms. In an outstanding study, Wilhelm et al. [16] using SIP microcosm-based experiments with 13C-labeled lignin, coupled with shotgun metagenomics, have demonstrated that species from Caulobacteraceae and Comamonadaceae families are the most relevant microbes for lignin degradation on coniferous forest soils across North America. They found that members of Caulobacteraceae family could degrade all three lignocellulosic polymers, providing new evidence for their importance in plant biomass degradation. In addition, some of these species contained genes predicted to encode the entire beta-ketoadipate pathway. It is reported that this pathway appears to be the most common route for aromatic metabolism in lignin-degrading bacteria. However, some lignin-degrading bacteria such as Sphingobacterium and Paenibacillus apparently lack these gene clusters [14]. In the current study, we observed that, based on taxonomic affiliation of sequences within the gene pcaI (3-oxoadipate CoA-transferase), the protocatechuate catabolism via beta-ketoadipate pathway appears to be carried out primarily by species from Pseudomonadaceae and Alcaligenaceae families (Figs. 5 and 6). Observing the taxonomic affiliation of sequences within the gene desB (gallate dioxygenase), we suggest that the catabolism of gallate could be performed mostly by Yersiniaceae species in WS1-M and Sphingomonadaceae species in CS-M. For catabolism of catechol, species affiliated to Pseudomonadaceae and Alcaligenaceae families appear to be key players (Figs. 5 and 6). However, we observed a high proportion of sequences within the gene dmpB that were affiliated to Actinobacteria, indicating that this taxon could play an important role as well within the catechol catabolism, especially in the consortium CS-M (Fig. 5b and supplementary information Fig. S1). Overall, these findings suggest that particular bacterial taxa could do specific functional jobs in each microbial consortium.
The genes pcaB, pcaH, ligB, ligC, ligI, and ligJ (supplementary information Table S1) were not significantly enriched in WS1-M, CS-M, and SG-M compared with FS1 (Fig. 3). However, their taxonomic affiliation shows that the catabolism of protocatechuate, by fission pathways (2,3-cleavage and 4,5-clevage), is a process probably carried out by species belonging to the taxa Pseudomonadaceae, Alcaligenaceae, Actinobacteria, Sphingomonadaceae, Xanthomonadaceae, and Comamonadaceae (Figs. 5 and 6). In a previously reported thermophilic lignin-degrading consortium, the family Xanthomonadaceae was highly selected along the enrichment, while species from family Alcaligenaceae survives in moderately lignin-rich environments [27]. In addition, Fang et al. [28] have reported that Stenotrophomas (Xanthomonadaceae member) is an abundant microbe in a consortium retrieved from decaying wood and cultivated in guaiacol and tree trimmings, suggesting that Xanthomonadaceae species could be associated with lignin-abundant environments. The ligAB genes are less common than the pca genes for protocatechuate degradation. They have been studied in Sphingobium SYK-6 (formerly known as Sphingomonas) [56]. Looking in detail the protocatechuate 4,5-cleavage fission pathway (Fig. 5), we observed that sequences from ligB were mostly affiliated to Sphingomonadaceae and Comamonadaceae in WS1-M. However, some sequences were affiliated to Actinobacteria in the consortia CS-M (Fig. 5). Regarding genes ligI and ligJ, the data showed that sequences were mostly affiliated to Xanthomonadaceae in the consortium CS-M, whereas in WS1-M, the sequences were mostly affiliated to Sphingomonadaceae and Comamonadaceae. Finally, we observed that Flavobacteriaceae family could be a key taxon in the WS1-M consortium, based on the high proportion of sequences of genes (katG, metF, pcaH, and thyA) affiliated to this family (Fig. 5 and supplementary information Fig. S1).
Conclusions
From examination of the results in this study, several conclusions can be made. Firstly, the SG-M and CS-M consortia are very similar, in terms of their taxonomical composition and ligninolytic gene profile, whereas the WS1-M consortium has different composition. Secondly, catabolic genes probably involved in lignin depolymerization or oxidative stress response (e.g., catalase/peroxidases and superoxide dismutases) were highly abundant in the three consortia, showing high sequence diversity. In contrast, we found less sequence diversity within catabolic genes involved in the intracellular metabolism of aromatic compounds, suggesting that these processes could have a degree of specialization. In general terms, we propose that lignin transformation follows a “task division” strategy, similar to that found for the degradation of cellulose and hemicellulose [57]. Thirdly, from the 60 enzyme-encoding genes involved in lignin catabolism, 20 were significantly abundant in the three soil-derived lignocellulolytic consortia (WS1-M, CS-M, and/or SG-M) compared with FS1, suggesting that these microbial communities have a large potential to transform lignin. As a perspective, the use of spectroscopic analyses to reveal the molecular structure of lignin [58] will be very valuable to correlate gene relative abundances and taxonomy with specific lignin linkages in each plant biomass-degrading consortia. Based on the enrichment of some catabolic genes found in our study and the results reported in other lignin-adapted microbial consortia, we suggest that genes vanA, pcaI, pcaJ, catA, catB, catC, and dmpB can be key gene-markers for lignin transformation, especially those involved in catechol and protocatechuate metabolism. Interestingly, hca genes involved in 3-phenylpropionic acid metabolism were significantly abundant in the lignocellulolytic consortia compared with FS1. We suggest that this aromatic compound can be released from the anaerobic lignin depolymerization and subsequently metabolized toward the TCA. Moreover, we conclude that the presence, in high abundance, of bacterial species belonged to Pseudomonadaceae, Caulobacteraceae, Xanthomonadaceae, Alcaligenaceae, and Comamonadaceae families could be a strong indication of the high potential to depolymerize lignin and metabolize its derived aromatic compounds in any microbial community. In this regard, we are in accordance with Wilhelm et al. [16] to state that the variation in lignin-degrading activity could be better explained by the catabolic gene content and community structure. Fourthly, we conclude that species belonged to Pseudomonadaceae family can be the most relevant ligninolytic members in these microbial consortia. They have the potential to participate in lignin depolymerization and in the metabolism aromatic compounds through the beta-ketoadipate pathway. Our predictive “model” (Fig. 6) allowed us to hypothesize that some bacterial populations could have a specific functional role within the lignin catabolism. For instance, some members have broad metabolic capacities (e.g., Pseudomonadaceae), while other ones could act as a specialist doing the catabolism of specific aromatic compounds (e.g., Caulobacteraceae and Sphingomonadaceae). Finally, we propose that by linking function and taxonomy, our metagenomic exploration has allowed us to better understand the lignin degradation process on soil-derived lignocellulolytic microbial consortia.
References
Chundawat SP, Beckham GT, Himmel ME, Dale BE (2011) Deconstruction of lignocellulosic biomass to fuels and chemicals. Annu Rev Chem Biomol Eng 2:121–145
Himmel ME, Ding SY, Johnson DK, Adney WS, Nimlos MR, Brady JW, Foust TD (2007) Biomass recalcitrance: engineering plants and enzymes for biofuels production. Science. 315:804–807
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546
Deng Z, Xia A, Liao Q, Zhu X, Huang Y, Fu Q (2019) Laccase pretreatment of wheat straw: effects of the physicochemical characteristics and the kinetics of enzymatic hydrolysis. Biotechnol Biofuels 12:159
Ragauskas AJ, Beckham GT, Biddy MJ, Chandra R, Chen F, Davis MF, Davison BH, Dixon RA, Gilna P, Keller M, Langan P, Naskar AK, Saddler JN, Tschaplinski TJ, Tuskan GA, Wyman CE (2014) Lignin valorization: improving lignin processing in the biorefinery. Science 344:1246843
Cao L, Yu IKM, Liu Y, Ruan X, Tsang DCW, Hunt AJ, Ok YS, Song H, Zhang S (2018) Lignin valorization for the production of renewable chemicals: state-of-the-art review and future prospects. Bioresour Technol 269:465–475
Xu Z, Lei P, Zhai R, Wen Z, Jin M (2019) Recent advances in lignin valorization with bacterial cultures: microorganisms, metabolic pathways, and bio-products. Biotechnol Biofuels 12:32
Bugg TD, Ahmad M, Hardiman EM, Rahmanpour R (2011) Pathways for degradation of lignin in bacteria and fungi. Nat Prod Rep 28:1883–1896
de Gonzalo G, Colpa DI, Habib MH, Fraaije MW (2016) Bacterial enzymes involved in lignin degradation. J Biotechnol 236:110–119
Janusz G, Pawlik A, Sulej J, Swiderska-Burek U, Jarosz-Wilkolazka A, Paszczynski A (2017) Lignin degradation: microorganisms, enzymes involved, genomes analysis and evolution. FEMS Microbiol Rev 41:941–962
Brink DP, Ravi K, Lidén G, Gorwa-Grauslund MF (2019) Mapping the diversity of microbial lignin catabolism: experiences from the eLignin database. Appl Microbiol Biotechnol 103:3979–4002
Bugg TDH, Williamson JJ, Rashid GMM (2020) Bacterial enzymes for lignin depolymerisation: new biocatalysts for generation of renewable chemicals from biomass. Curr Opin Chem Biol 55:26–33
Lee S, Kang M, Bae JH, Sohn JH, Sung BH (2019) Bacterial valorization of lignin: strains, enzymes, conversion pathways, biosensors, and perspectives. Front Bioeng Biotechnol 7:209
Granja-Travez RS, Persinoti GF, Squina FM, Bugg TDH (2020) Functional genomic analysis of bacterial lignin degraders: diversity in mechanisms of lignin oxidation and metabolism. Appl Microbiol Biotechnol 104:3305–3320. https://doi.org/10.1007/s00253-019-10318-y
DeAngelis KM, Allgaier M, Chavarria Y, Fortney JL, Hugenholtz P, Simmons B, Sublette K, Silver WL, Hazen TC (2011) Characterization of trapped lignin-degrading microbes in tropical forest soil. PLoS One 6:e19306
Wilhelm RC, Singh R, Eltis LD, Mohn WW (2019) Bacterial contributions to delignification and lignocellulose degradation in forest soils with metagenomic and quantitative stable isotope probing. ISME J 13:413–429
Woo HL, Hazen TC (2018) Enrichment of bacteria from eastern Mediterranean Sea involved in lignin degradation via the phenylacetyl-CoA pathway. Front Microbiol 9:922
Moraes EC, Alvarez TM, Persinoti GF, Tomazetto G, Brenelli LB, Paixão DAA, Ematsu GC, Aricetti JA, Caldana C, Dixon N, Bugg TDH, Squina FM (2018) Lignolytic-consortium omics analyses reveal novel genomes and pathways involved in lignin modification and valorization. Biotechnol Biofuels 11:75
Jiménez DJ, Dini-Andreote F, DeAngelis KM, Singer SW, Salles JF, van Elsas JD (2017) Ecological insights into the dynamics of plant biomass-degrading microbial consortia. Trends Microbiol 25:788–796
Cortes-Tolalpa L, Jiménez DJ, de Lima Brossi MJ, Salles JF, van Elsas JD (2016) Different inocula produce distinctive microbial consortia with similar lignocellulose degradation capacity. Appl Microbiol Biotechnol 100:7713–7725
de Lima Brossi MJ, Jiménez DJ, Cortes-Tolalpa L, van Elsas JD (2016) Soil-derived microbial consortia enriched with different plant biomass reveal distinct players acting in lignocellulose degradation. Microb Ecol 71:616–627
Jiménez DJ, Chaves-Moreno D, van Elsas JD (2015) Unveiling the metabolic potential of two soil-derived microbial consortia selected on wheat straw. Sci Rep 5:13845
Jiménez DJ, de Lima Brossi MJ, Schückel J, Kračun SK, Willats WG, van Elsas JD (2016) Characterization of three plant biomass-degrading microbial consortia by metagenomics- and metasecretomics-based approaches. Appl Microbiol Biotechnol 100:10463–10477
Alessi AM, Bird SM, Bennett JP, Oates NC, Li Y, Dowle AA, Polikarpov I, Young JPW, McQueen-Mason SJ, Bruce NC (2017) Revealing the insoluble metasecretome of lignocellulose-degrading microbial communities. Sci Rep 7:2356
Alessi AM, Bird SM, Oates NC, Li Y, Dowle AA, Novotny EH, deAzevedo ER, Bennett JP, Polikarpov I, Young JPW, McQueen-Mason SJ, Bruce NC (2018) Defining functional diversity for lignocellulose degradation in a microbial community using multi-omics studies. Biotechnol Biofuels 11:166
Wu YR, He J (2013) Characterization of anaerobic consortia coupled lignin depolymerization with biomethane generation. Bioresour Technol 139:5–12
Ceballos SJ, Yu C, Claypool JT, Singer SW, Simmons BA, Thelen MP, Simmons CW, VanderGheynst JS (2017) Development and characterization of a thermophilic, lignin degrading microbiota. Process Biochem 63:193–203
Fang X, Li Q, Lin Y, Lin X, Dai Y, Guo Z, Pan D (2018) Screening of a microbial consortium for selective degradation of lignin from tree trimmings. Bioresour Technol 254:247–255
Jiménez DJ, Korenblum E, van Elsas JD (2014) Novel multispecies microbial consortia involved in lignocellulose and 5-hydroxymethylfurfural bioconversion. Appl Microbiol Biotechnol 98:2789–2803
Meyer F, Paarmann D, D’Souza M, Olson R, Glass EM, Kubal M, Paczian T, Rodriguez A, Stevens R, Wilke A, Wilkening J, Edwards RA (2008) The metagenomics RAST server—a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinf 9:386
Parks DH, Beiko RG (2010) Identifying biologically relevant differences between metagenomic communities. Bioinformatics 26:715–721
Kamimura N, Takahashi K, Mori K, Araki T, Fujita M, Higuchi Y, Masai E (2017) Bacterial catabolism of lignin-derived aromatics: new findings in a recent decade: update on bacterial lignin catabolism. Environ Microbiol Rep 9:679–705
Lin L, Wang X, Cao L, Xu M (2019) Lignin catabolic pathways reveal unique characteristics of dye-decolorizing peroxidases in Pseudomonas putida. Environ Microbiol 21:1847–1863
Zhu D, Zhang P, Xie C, Zhang W, Sun J, Qian WJ, Yang B (2017) Biodegradation of alkaline lignin by Bacillus ligniniphilus L1. Biotechnol Biofuels 10:44
Zhu D, Si H, Zhang P, Geng A, Zhang W, Yang B, Qian WJ, Gabriel M, Sun J (2018) Genomics and biochemistry investigation on the metabolic pathway of milled wood and alkali lignin-derived aromatic metabolites of Comamonas serinivorans SP-35. Biotechnol Biofuels 11:338
Cardenas E, Kranabetter JM, Hope G, Maas KR, Hallam S, Mohn WW (2015) Forest harvesting reduces the soil metagenomic potential for biomass decomposition. ISME J 9:2465–2476
Babicki S, Arndt D, Marcu A, Liang Y, Grant JR, Maciejewski A, Wishart DS (2016) Heatmapper: web-enabled heat mapping for all. Nucleic Acids Res 44(W1):W147–W153
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550
Menzel P, Ng KL, Krogh A (2016) Fast and sensitive taxonomic classification for metagenomics with Kaiju. Nat Commun 7:11257
Fu L, Niu B, Zhu Z, Wu S, Li W (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152
Teeling H, Glöckner FO (2012) Current opportunities and challenges in microbial metagenome analysis—a bioinformatic perspective. Brief Bioinform 13:728–742
Masai E, Ichimura A, Sato Y, Miyauchi K, Katayama Y, Fukuda M (2003) Roles of the enantioselective glutathione S-transferases in cleavage of β-aryl ether. J Bacteriol 185:1768–1775
Rashid GMM, Taylor CR, Liu Y, Zhang X, Rea D, Fülöp V, Bugg TDH (2015) Identification of manganese superoxide dismutase from Sphingobacterium sp. T2 as a novel bacterial enzyme for lignin oxidation. ACS Chem Biol 10:2286–2294
Wei Z, Wilkinson RC, Rashid GMM, Brown D, Fülöp V, Bugg TDH (2019) Characterization of thiamine diphosphate-dependent 4-hydroxybenzoylformate decarboxylase enzymes from Rhodococcus jostii RHA1 and Pseudomonas fluorescens Pf-5 involved in degradation of Aryl C2 lignin degradation fragments. Biochemistry 58:5281–5293
Wang Y, Liu Q, Yan L, Gao Y, Wang Y, Wang W (2013) A novel lignin degradation bacterial consortium for efficient pulping. Bioresour Technol 139:113–119
Rahmanpour R, Bugg TD (2015) Characterisation of Dyp-type peroxidases from Pseudomonas fluorescens Pf-5: oxidation of Mn(II) and polymeric lignin by Dyp1B. Arch Biochem Biophys 574:93–98
Park MR, Chen Y, Thompson M, Benites V, Fong B, Petzold C, Baidoo E, Gladden J, Adams P, Keasling J, Simmons BA, Singer SW (2020) The response of Pseudomonas putida to complex aromatic-rich fractions from biomass. ChemSusChem. https://doi.org/10.1002/cssc.202000268
Lin L, Cheng Y, Pu Y, Sun S, Li X, Jin M, Pierson EA, Gross DC, Dale BE, Dai SY, Ragauskas AJ, Yuan JS (2016) Systems biology-guided biodesign of consolidated lignin conversion. Green Chem 18:5536–5547
Forsberg KJ, Patel S, Witt E, Wang B, Ellison TD, Dantas G (2015) Identification of genes conferring tolerance to lignocellulose-derived inhibitors by functional selections in soil metagenomes. Appl Environ Microbiol 82:528–537
Carlos C, Fan H, Currie CR (2018) Substrate shift reveals roles for members of bacterial consortia in degradation of plant cell wall polymers. Front Microbiol 9:364
Bartilson M, Shingler V (1989) Nucleotide sequence and expression of the catechol 2,3-dioxygenase-encoding gene of phenol-catabolizing Pseudomonas CF600. Gene 85:233–238
Nishimura M, Kohno K, Nishimura Y, Inagaki M, Davies J (2011) Characterization of two isozymes of coniferyl alcohol dehydrogenase from Streptomyces sp. NL15-2K. Biosci Biotechnol Biochem 75:1770–1777
Rashid GMM, Zhang X, Wilkinson RC, Fülöp V, Cottyn B, Baumberger S, Bugg TDH (2018) Sphingobacterium sp. T2 manganese superoxide dismutase catalyzes the oxidative demethylation of polymeric lignin via generation of hydroxyl radical. ACS Chem Biol 13:2920–2929
Glissmann K, Hammer E, Conrad R (2005) Production of aromatic compounds during methanogenic degradation of straw in rice field soil. FEMS Microbiol Ecol 52(1):43–48
Prem EM, Markt R, Lackner N, Illmer P, Wagner AO (2019) Microbial and phenyl acid dynamics during the start-up phase of anaerobic straw degradation in meso- and thermophilic batch reactors. Microorganisms 7:657
Masai E, Katayama Y, Fukuda M (2007) Genetic and biochemical investigations on bacterial catabolic pathways for lignin-derived aromatic compounds. Biosci Biotechnol Biochem 71:1–15
López-Mondéjar R, Zühlke D, Becher D, Riedel K, Baldrian P (2016) Cellulose and hemicellulose decomposition by forest soil bacteria proceeds by the action of structurally variable enzymatic systems. Sci Rep 6:25279
Lu Y, Lu YC, Hu HQ, Xie FJ, Wei XY, Fan X (2017) Structural characterization of lignin and its degradation products with spectroscopic methods. J Spectrosc 8951658:1–15
Acknowledgments
We thank Nancy N. Nichols for relevant comments and text corrections. Additionally, thanks to Alejandro Reyes, Alejandro Caro-Quintero, and Maryam Chaib De Mares for suggestions about the data analysis.
Funding
This work was supported by the FAPA project (Number PR.3.2018.5287) obtained by Diego Javier Jiménez at Universidad de los Andes (Bogotá, Colombia), and BBSRC research grant BB/P01738X/1 obtained by Timothy D.H. Bugg.
Author information
Authors and Affiliations
Contributions
LDG carried out all bioinformatic and statistical analysis. TDHB helped in the data discussion and drafting the text. DJJ conceived the study and drafted the manuscript.
Corresponding author
Ethics declarations
Conflicts of Interest
The authors declare that they have no conflict of interest.
Ethics Approval
Not applicable
Consent to Participate
Not applicable
Consent for Publication
All authors read and approved the final manuscript.
Electronic supplementary material
Figure S1
PCA using the abundance values and taxonomic affiliation of 22 selected ligninolytic catabolic genes in WS1-M (left) and CS-M (right). (PPTX 130 kb)
Table S1
The 60 enzyme-encoding genes involved in transformation of lignin and its derived aromatic compounds. Within the table, we showed the KOs number identifiers, gene or protein name, EC numbers and putative function or metabolic pathway within the lignin transformation processes. (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Díaz-García, L., Bugg, T.D.H. & Jiménez, D.J. Exploring the Lignin Catabolism Potential of Soil-Derived Lignocellulolytic Microbial Consortia by a Gene-Centric Metagenomic Approach. Microb Ecol 80, 885–896 (2020). https://doi.org/10.1007/s00248-020-01546-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-020-01546-1