Abstract
Microorganisms are the driver of petroleum hydrocarbon degradation in soil micro-ecological systems. However, the distribution characteristics of microbial communities and hydrocarbon degradation dynamics during the remediation of petroleum-contaminated soil by enhancing moisture content are not clear. In this study, polymerase chain reaction and high-throughput sequencing of soil microbial DNA were applied to investigate the compositions of microorganisms and alpha diversity in the oil-polluted soil, and the hydrocarbon removal also being analyzed using ultrasonic extraction and gravimetric method in a laboratory simulated ex-situ experiment. Results showed the distribution of petroleum hydrocarbon degrading microorganisms in the petroleum-contaminated loessal soil mainly was Proteobacteria phylum (96.26%)—Gamma-proteobacteria class (90.03%)—Pseudomonadales order (89.98%)—Pseudomonadaceae family (89.96%)—Pseudomonas sp. (87.22%). After 15% moisture content treatment, Actinobacteria, Proteobacteria, and Firmicutes still were the predominant phyla, but their relative abundances changed greatly. Also Bacillus sp. and Promicromonospora sp. became the predominant genera. Maintaining 15% moisture content increased the relative abundance of Firmicutes phylum and Bacillus sp. As the moisture-treated time increases, the uniformity and the richness of the soil bacterial community were decreased and increased respectively; the relative abundance of Pseudomonas sp. increased. Petroleum hydrocarbon degradation by enhancing soil moisture accorded with the pseudo-first-order reaction kinetic model (correlation coefficient of 0.81; half-life of 56 weeks). The richness of Firmicutes phylum and Bacillus sp. may be a main reason for promoting the removal of 18% petroleum hydrocarbons responded to 15% moisture treatment. Our results provided some beneficial microbiological information of oil-contaminated soil and will promote the exploration of remediation by changing soil moisture content for increasing petroleum hydrocarbon degradation efficiency.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Increasingly serious oil-contaminated soil concern was initially realized deeply in several well-known oil spills accidents, which induced permanent threats to humans, animals, and plants on a global scale due to their toxicity brought by the harmful hydrocarbons such as long-chain alkanes and polycyclic aromatic hydrocarbons [1,2,3,4]. The area of soil contaminated by petroleum was expanding in north of Shaanxi province, which was the important petrochemical base of China [5, 6]. In recent years, annual oil production exceeded 200 million tons, causing more than 100,000 tons of newly contaminated soil each year [7]. The promising technologies, natural attenuation, biostimulation, and bioaugmentation techniques, based on the little disturbance to the environment, were employed for remediation of oil-contaminated soil [3, 8, 9]. Generally, the stability and efficiency of remediation of oil-contaminated soil were relied on greatly the bioavailability and activity of soil microorganisms and the synergy between these microbial and environmental conditions like moisture contents and temperatures [10, 11]. Therefore, exploring the petroleum-contaminated soil remediation effects primly calls for a synthetic evaluation of not just which forms of program implementations are devised; the taxonomy in soil microorganisms are also requisite [12, 13].
Some studies have demonstrated the powerful advances of high throughput sequence technology in microbial biogeography for analysis of the soil microbial compositions and structures [14]. Many microorganisms belonging to different taxonomy levels have the ability to degrade petroleum components including hydrocarbons and non-hydrocarbons [15, 16]. A metagenomics and deep sequencing research reported by Shahi et al. [3] revealed that the relative abundance of Firmicutes and Bacteroides, two kinds of petroleum-degrading bacteria, can change with the adjustment of the ratio of carbon, nitrogen, and phosphorus in the oil-contaminated soil. Now, with the highly accurate visual species analysis of the soil microbial populations, more and more potentially hydrocarbon-degrading microbes will probably be found and applied.
Great changes of soil microbial population and diversity across the oil-polluted sites occurred compared to clean, uncontaminated soil, and their growth, reproduction, and metabolic activities were susceptible to sensitivity by the external environment [17, 18]. The distribution of microbes in harsh and restrictive environments such as sand, clay, loam, desert, polar, and frozen soil has been well interpreted [12, 19, 20]. Previous studies have shown that psychrophilic microorganisms were emerged adapted to low temperature and heavy oil pollution with large molecular weight and high viscosity under the cold environment [21]. Essentially, the petroleum mixture in the soil were transferred, absorbed, and degraded in a parallel manner with extracellular transport and intracellular degradation by microbial communities [22]. Humidity, one of the vital factors for the survival of soil microbes, affects the activity of indigenous microorganisms in oil-contaminated soil, which has low water holding capacity and is a strong water repellent [5]. Wang et al. confirmed that 33% of the water increased the community diversity of microbes in diesel and lubricant-contaminated soils by sodium azide and mercuric chloride as experimental controls and nucleotide sequence analysis [23]. A large number of studies have proved that the degradation of petroleum hydrocarbons is related to the soil moisture content, and found the optimum soil moisture content range which was beneficial to remediate different oil-contaminated soil sites [24, 25]. Ali et al. investigated the performance of total petroleum hydrocarbons in different soil moisture content, and found that maintaining a moisture content of 20% for sand soil for 270 days would result in a TPH removal rate of 70% [12]. However, to the best of our knowledge, the information of the effects of moisture content on the soil microbial community and hydrocarbon degrading microorganisms in petroleum-contaminated soil is still not clear.
In the past few years, the alpha diversity index (Chao1, ACE, Simpson, Shannon) reflected richness and uniformity of the species in an ecosystem and was commonly used for microbial diversity analysis [26]. The activity and number of petroleum hydrocarbon degrading microorganisms were measured indirectly by soil headspace carbon dioxide emissions and most probable number procedure to evaluate the removal ability of petroleum hydrocarbons [27]. Recently, Li et al. [28] found that biostimulation remediation promoted the degradation of petroleum hydrocarbons and affected the distribution and metabolic activity of bacteria in the soil by phospholipid fatty acid (PLFA) analysis, but less information about the specific microbial species was acceptable due to the imperfect analytical technique.
This paper carried out a study to weaken the stress of petroleum hydrocarbons on the loessal soil by enhancing soil moisture content. The compositions and the α-diversity of bacterial community in petroleum-contaminated soil were discussed in detail. The relative abundances of petroleum hydrocarbon-degrading bacteria were analyzed simultaneously. We also investigated the reaction kinetic and the correlation between petroleum hydrocarbon removal rate and the hydrocarbon degrading populations in the polluted soil. The results will provide microbiological information of the dominant petroleum-degrading microorganisms in order to obtain improved petroleum hydrocarbon removal rates for the rehabilitation of petroleum-contaminated soils.
Materials and Methods
Soil Sampling and Analysis
The long-term oil-contaminated loess soil samples which were loose, soft and a light yellowish soil were obtained from an oil well located in the north of Shaanxi province, China. The methods of sampling, collection and transportation were according to the description of Wu et al. [29]. The granulometric compositions of soil are silt (28.64%), fine sand (62.17%), medium sand (5.44%), and medium sand (3.75%) (types II); and the physical, chemical, and biological properties of the soil are shown in Table 1.
15% Moisture Treatment
A microcosm experiment for bioremediation oil-contaminated soils was performed at 24 °C for up to 12 weeks. A 0.8 kg of soil was placed in pots in triplicate, maintaining 15% moisture content with distilled water and periodically agitated artificially to obtain oxygen. The expression of S0, S1, and S12 represented soil samples without distilled water addition, 15% moisture treatment for 1 week, and 15% moisture treatment for 12 weeks, respectively.
Enrichment and Analysis of Soil Total Microorganisms and the Hydrocarbon Degrading Community
For enrichment of soil total microorganisms, 5 g of oil-contaminated soil was added to 50 mL of PBS buffer, vibrating for 2 h with 150 rpm in a water bath shaker at room temperature. After standing for 30 min, the soil total microorganisms were obtained by dumping out the supernatant.
The screening and enrichment of TPH-degrading microbial cells from petroleum-contaminated soil has been previously introduced by Wu et al. [29], and the process is further modified. A 6% inoculum of the total microorganisms in 100 mL of PBS buffer with 1% petroleum hydrocarbon as the sole carbon and energy source cultured for 1 week at room temperature and subjected to three consecutive transfer cultures. After that, the petroleum hydrocarbon-degrading flora was obtained by centrifugation from the last cultures.
Both soil total bacterial and the petroleum hydrocarbon degrading flora were analyzed using high-throughput sequencing technology by Sangon Biotech Co., Ltd. China (ftp://ftp.sangon.com:21148) and the details were as follows.
Genomic DNA Extraction, Illumina Sequencing
The DNA of the total microbial microorganisms and hydrocarbon degrading community in initial and 15% moisture content soil samples was extracted and quantified using Power Soil DNA extraction kit (MoBio Laboratories, USA). The integrity of the extracted DNA was examined by agarose gel electrophoresis. The sequencing mode was Miseq PE 300 with the paired-end. The primers 341F (CCCTACACGACGCTCTTCCGATCTGCCTACGGGNGGCWGCAG) and 805R (GACTGGAGTTCCTTGGCACCCGAGAATTCCAGACTACHVGGGTATCTAATCC) were used to complete the PCR reaction [30].
The temperature parameter was (1) repetition 5 cycles of pre-denaturation at 94 °C for 3 min, denaturation at 94 °C for 30 s, annealing at 45 °C for 20 s, extension at 65 °C, and stretching for 30 s; (2) repetition 20 cycles of denaturation at 94 °C for 20 s, annealing at 55 °C for 20 s, and extension at 72 °C for 30 s; (3) repetition for 5 cycles of pre-denaturation at 95 °C for 30 s, denaturation at 95 °C for 15 s, annealing at 55 °C for 15 s, and extension at 72 °C for 30 s after introduction of Illumina bridge PCR compatible primers set during the PCR reaction.
The PCR products were analyzed by agarose gel electrophoresis and purified to recover using 0.6 times of magnetic beads. The amount of DNA per sample was 10 ng, and the final loaded sequencing concentration was 20 pmol.
Sequencing Data Analysis
To perform some quality control processing on the original sequence, such as de-joining and mass-cutting, Prinseq software (version 0.20.4) were used. After removing the non-amplified region portion in the pre-processed sequence, Usearch (version 5.2.236) was put to use to correct all sequence errors and clustered according to the distance between sequences, i.e., operational taxonomic units (OTUs) [26]. The database sequence of Blast was used to compare against the measured sequences. The RDP classifier (version 2.12) divided the OTU with a sequence similarity threshold of 0.97 into the same genus, and did it as the same species with the value of 0.99. The alpha diversity index reflecting the richness and uniformity of the microbes in soil was calculated by Mothur (version 1.30.1) [26].
TPH Removal Performance and Microbial Population
In order to investigate the effect of 15% moisture content on hydrocarbons degradation in petroleum-contaminated soil, TPH concentrations during the maintenance of 15% moisture content were detected. The TPH were extracted and determined by ultrasonic extraction and gravimetric methods respectively [31]. Firstly, 1 g of air-dried, ground soil sample from each pot and 15 mL of mixed extract (V (n-hexane: methylene chloride) = 1:1) were placed in a 50 mL polyethylene centrifuge tube to extract TPH weekly using an ultrasonic cell disrupter at a power of 180 W by repeating three times for 15 min each time. Then, the three extracts were centrifuged at − 4 °C, 8000 r/min and filtered in a 30-mL weighing bottle of known weight. Finally, the weighing bottle was placed in a fume hood to evaporate the organic extract, air dried, and weighed. The TPH removal performance was obtained from the difference between the two weights. Also, according the description of Wu et al. [31], the standard petroleum hydrocarbons and a modified most probable number (MPN) procedure were used to count the TPH-degrading microbial populations. Briefly, 1 g of soil sample was uniformly dispersed in 9 mL PBS buffer solution, and 0.2 ml of the suspension was transferred to 1.8 mL Bushnell-Haas medium containing standard petroleum hydrocarbon and 2% NaCl. After the transferred suspension was incubated at room temperature for 1 week, iodonitrotetrazolium violet (INT) and MPN table were applied to count TPH degrading bacteria by 1 mL of soil microbial extract with dilution gradients of 10−1, 10−2, 10−3, 10−4, 10−5, 10−6, 10−7, 10−8, 10−9, and 10 μL TPH, in five replicates per gradient..
Statistical Analysis
Data for all TPH concentrations was represented by a combination of mean and standard deviation (mean ± SD). For the study of petroleum hydrocarbon degradation and microbial community, analysis of relationship was conducted by origin software (version 9.0, China).
Results and Discussion
Total Bacterial Community Compositions in the Petroleum-Contaminated Soil
The information of total microorganisms in petroleum-contaminated soil was obtained by high-throughput sequencing analysis of DNA directly extracted from microorganisms in soils. Figure 1 a showed the relative abundance of the top 10 dominant phyla. Actinobacteria, Proteobacteria, and Firmicutes were the three dominant phyla of which the relative abundances were 47.34%, 37.44%, and 9.16% in the petroleum-contaminated loessal soil. The three bacterial phyla were ubiquitous in oil-contaminated soils reported in previous literatures [3, 9], and most of them belonged to Gram-positive bacteria, confirming their universality and potential for habitation in oil-contaminated sites.
Figure 1 b displayed Promicromonospora sp. which was the top dominant genus with the relative abundance of 18.96% in the contaminated soil. Exiguobacterium sp. was the subordinate genus and the relative abundance of 8.49% among all microorganisms. In addition, genera of Nocardioides sp., Citrobacter, Mycobacterium. sp., Acinetobacter sp., and Leifsonia sp. were also prevalent in the oil-contaminated soil. Members of Promicromonospora sp., Exiguobacterium sp., and Nocardioides were detected in previous studies [32, 33]. It was reported that n-alkanes with 9 to 26 carbon and aliphatic hydrocarbons can be degraded by them in diesel and oil pollution environments [34].
Hydrocarbon-Degrading Bacterial Compositions in the Petroleum-Contaminated Soil
Analysis of petroleum hydrocarbon degrading bacterial community enriched using petroleum as sole carbon and energy source from petroleum-contaminated soil was performed by high throughput sequencing method. The top 30 dominant bacterial taxonomies based on species including phylum, class, order, family, and genus levels were shown in Fig. 2. The top 30 bacterial species involved to two phyla, three classes, five orders, eight families, and 16 genera. Proteobacteria and Firmicutes were the dominant phyla with the relative abundance of 92.26% and 3.71%, respectively. Although Actinobacteria was dominant bacterial phylum in the oil-polluted soil, it was not petroleum-degrading phylum. Proteobacteria was considered as the most easily cultivated bacteria in oil-contaminated soils reported by previous study [35]. Members of Proteobacteria and Firmicutes phyla have been proven to degrade TPH or PAHs in oil-polluted soil [9, 35]. Kim et al. [36] enriched the microorganisms that produced biosurfactants with the potential of degrading hydrocarbons in oil-contaminated soil in Kuwait and found that the relative abundance of Proteobacteria was the most; Firmicutes were the second, and Actinobacteria was the lowest, and the relative abundance of Actinobacteria only accounted for 0.1% [36]. These similar results have once again confirmed the classification homology of microorganisms from oil-contaminated sites from different regions [9]. Gamma-proteobacteria was the first among three classes in terms of their relative abundance, which was 90.03%, 6.22%, and 3.71% attached to Gamma-proteobacteria, Beta-proteobacteria, and Bacilli respectively. Gamma-proteobacteria, Beta-proteobacteria, and Bacilli were also recognized as hydrocarbon-degrading strains in oil-contaminated soil, but the relative abundance was different from this study due to the soil texture and the time of oil pollution [36]. A previous study also reported Bacilli, one of the important hydrocarbon degrading bacteria, preferentially used more toxic aromatic hydrocarbons as their energy source and carbon source in petroleum-contaminated soils [37]. Among the 16 dominant genera, the relative abundance of Pseudomonas (87.22%) was the highest, while other 15 genera had a total relative abundance of less than 7%. Of the hydrocarbon-feeding bacteria, Pseudomonas is widely known for the ability to degrade hydrocarbons by producing a variety of glycolipid surfactants [38]. Even, Ramadass et al. found Pseudomonas sp. increased the removal of weathered hydrocarbons by about 20% compared to natural attenuation in engine oil-contaminated soil [39], Based on the information presented in Fig. 2, the distribution of petroleum hydrocarbon-degrading bacteria in the petroleum-contaminated soil was Proteobacteria phylum (96.26%)—Gamma-proteobacteria class (90.03%)—Pseudomonadales order (89.98%)—Pseudomonadaceae family (89.96%)—Pseudomonas sp. (87.22%).
Impacts of 15% Moisture Treatment on Total Bacterial Community
In our study, initial polluted soil (S0) and 15% moisture-treated soil samples at the first week (S1) and the 12th week (S12) were collected for MiSeq sequencing analysis. The average number of sequences for the three soil samples was more than 45,000. After deleted the sequence outside the target region and the chimera by Usearch software, their effective sequence numbers were 41,369; 45,917; and 45,132, respectively. This sequencing was successful with an optimization rate of over 88% and a sequencing depth of 98% coverage. Across the three soil samples, the order of OTUs number obtained by classifying effective sequences was S1 (1888)>S12 (1865)>S0 (1669) (Table 2). The details of community alpha diversity index including the richness index (ACE and Chao 1) and the uniformity index (Shannon and Simpson) [26] varied in different soil samples. It can be seen that the Shannon and Ace indices of the S0 sample were the highest, while the Chao1 index of the S1 soil sample was the highest. In all the samples, the order was S0>S12>S1 for Shannon index, S0>S1>S12 for Ace index and S1>S12>S0 for Chao1 index. Therefore, compared with the initial soil, the uniformity of soil bacterial community was reduced, and the richness increased after 1 week of 15% moisture treatment. After 12 weeks of 15% moisture remediation, the soil diversity was lower than that of the initial contaminated soil generally.
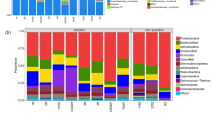
Figure 3 a showed the relative abundance of the top 10 dominant bacterial phyla in the S0, S1, and S12 samples. Actinobacteria, Proteobacteria, and Firmicutes were the first three dominant phyla in all samples, and together of which accounted for over 90% for each soil sample. Compared to the initial contaminated soil, the relative abundances of Actinobacteria and Proteobacteria decreased while Firmicutes, which was one of the hydrocarbon degradation bacterial phyla (shown in Fig. 2), increased by 15% moisture treatment. In the S1 sample, Firmicutes became the most important dominant phylum with the relative abundances of 62.29%, while the relative abundances of Actinobacteria and Proteobacteria decreased to 19.44% and 14.72%, respectively. In addition, phyla of Planctomycetes and Verrucomicrobia increased slightly after 1 week of 15% moisture treatment. Compared to S1, the relative abundance of Firmicutes reduced to 40.81%, and Actinobacteria and Proteobacteria increased to 34.43% and 19.70% in the S12 sample. The relative abundance of Bacteroidetes made a discernible increase while Planctomycetes and Verrucomicrobia increased slightly after 12 weeks of 15% moisture treatment.
Figure 3 b displayed the bacterial community composition at the genus level, and a more pronounced difference occurred in the dominant bacterial genera among the three samples. Compared to the initial contaminated soil (S0), 15% moisture treatment caused greatly change in the soil bacterial community structure at the genus level. Promicromonospora sp. became the secondary genus while Bacillus sp. which was one type of hydrocarbon-degrading bacteria (shown in Fig. 2) turned into the first dominant in the 15% moisture soil. After 1 week of 15% moisture treatment, the relative abundance of Bacillus sp. drastically increased from 0.29 to 38.20%. The relative abundance of Promicromonospora sp. and Exiguobacterium sp. became the sub-dominant genera which reduced to 19.44% and 5.06%, respectively. Some new genera such as Fictibacillus sp. and Paenisporosarcina sp. appeared in the soil. The flora structure in S12 sample was similar with the 1 week of moisture treatment (S1), and the relative abundances of dominant genera slightly changed. Nocardioides sp. became the subordinate dominant genus and the relative abundance increased to 6.94%. The relative abundance of Fictibacillus sp., Pseudomonas sp., and several other genera increased in the S12 compared to the S1 soil.
There was a significant difference between the soil total bacterial community (Fig. 1) and the petroleum hydrocarbon-degrading bacterial community (Fig. 2). The most prevalent bacteria community in the oil-contaminated soils were Actinobacteria (47.34%), Proteobacteria (37.44%), and Firmicutes (9.16%) phyla, Promicromonospora sp. (18.96%), and Exiguobacterium sp. (8.49%) genus. While, petroleum hydrocarbon-degrading bacterial compositions mainly included the phyla of Proteobacteria (92.26%) and Firmicutes (3.71%) and the genus of Pseudomonas (87.22%), Achromobacter (6.12%), and Bacillus (3.51%). Figure 3 showed that Actinobacteria was the most dominant bacterial phylum in the initial contaminated soil (47.34%) and 15% moisture-treated soils (19.44~34.43%), but which was not dominant petroleum-degrading bacteria according to Fig. 2. At the genus level, Promicromonospora sp. was the most abundant both in the initial contaminated soil (18.96%) and 15% moisture-treated soil (19.44%), while the relative abundance of Promicromonospora sp. was little in the petroleum-degrading bacterial community. Pseudomonas sp. was the predominant in petroleum-degrading bacterial compositions which accounted for 87.22%; however, enhanced soil humidity did not promote Pseudomonas sp. growth. The reason may be that both environmental factors and nutrients have the significant effects on the growth of Pseudomonas sp. [40]. Enhanced humidity just provided a single element of water, but the deficiency of nitrogen and phosphorus or other growth factors may limit to the growth of Pseudomonas sp.
Effects of 15% Moisture Content on Petroleum Degradation and Hydrocarbon-Degrading Microbial Population
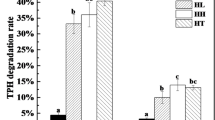
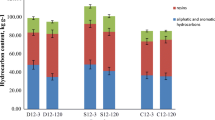
The petroleum hydrocarbon concentration and the petroleum hydrocarbon-degrading microbial populations in the soils by 15% moisture treatment were shown in Fig. 4a. After 12 weeks of 15% moisture treatment, the TPH concentration decreased from 18,800 to 15,411 mg kg−1 soil. It represented 18.0% of petroleum hydrocarbons removal by 15% moisture treatment while only 3% in the control. The microbial population in soil without humidity treatment was 6.3 × 102 cells g−1 to 3.4 × 103 cells g−1, while it significantly increased from 3.3 × 103 to 1.5 × 106 cells g−1 corresponding to 15% moisture content, which indicated that 15% moisture content was beneficial to promote the petroleum degrading microorganism population.
Petroleum hydrocarbons in soil were subject to various spontaneous migration and transformation process such as volatilization, photo-oxidation, chemical oxidation, and microbial degradation under relatively sufficient water and temperature conditions [41,42,43]. It was considered that microbial metabolism was the most important way for dissipation of petroleum hydrocarbons [44]. There were more than 200 functional microorganisms accompanied with higher occurrence frequency of bacteria, actinomycetes, fungi yeast, and mold, which were giving play to the utmost importance effect in different soil terrestrial ecosystems [45]. Among the heterotrophic groups, Pseudomonas (bacteria), Streptomyces (actinomycetes), Candida (fungi)-producing yeast proteins, and even some cyanobacteria used petroleum hydrocarbons as their sole source of carbon and energy to perform biological oxidation [46]. The suitable soil moisture formed an equilibrium between the water-vapor-solid three phases, which can not only accelerate the transfer of oxygen and petroleum hydrocarbons in the soil, but also promoted the absorption and utilization of petroleum hydrocarbons by microorganisms [24]. In this study, distilled water was added to soil to supply the essential moisture for the growth and development of microorganisms in the soil, and a small portion of petroleum hydrocarbons was removed, which was similar to previous results [31].
There was no correlation between petroleum hydrocarbon removal rate and the increasement of petroleum hydrocarbon degradation microbial populations (Fig. 4b). Fifteen percent of moisture content treatment indeed increased relative abundance of some petroleum hydrocarbon-degrading bacteria including Firmicutes phylum and Bacillus sp. (Fig. 3). It is therefore concluded that the removal of petroleum hydrocarbons did not depend on the total number of petroleum hydrocarbon degraders’ populations, but some specific hydrocarbon-degrading strains involving Firmicutes phylum and Bacillus sp. may be a promoting factor for the degradation of petroleum hydrocarbon pollutants by 15% moisture treatment.
Table 3 showed the reaction kinetic of petroleum hydrocarbon removal by 15% soil moisture treatment. Degradation of petroleum hydrocarbons accorded with the pseudo-first-order reaction kinetic model. The correlation coefficient was not high (R2 = 0.81) due to the complex degradation process affected by catalysis, volatilization, and biodegradation. Fifteen percent humidity treatment made a hydrocarbon degradation rate constant of 0.0126 per week, and the half-life of petroleum up to 56 weeks. Some biostimulation experiments by adding nitrogen, phosphorus, composting, and inoculating with microorganisms to oil-contaminated soil at low temperatures have similar results; and the hydrocarbon degradation rate constants were 0.004~0.016, 0.011~0.018, and 0.017~0.026 day−1 found by Gomez et al., Chang et al., and Paudyn et al. respectively [19,20,21].
In this study, only 18% hydrocarbon removed by 15% moisture treatment, and most of petroleum residue still remained in the soil. We had illustrated that Proteobacteria phylum and Pseudomonas genus were predominant petroleum-degrading bacteria in petroleum-contaminated soil (shown in Fig. 2). Fifteen percent moisture treatment could not promote these strains growth (Fig. 3 S1 and S2) and it maybe due to restrictive conditions about environmental factors such as pH, temperature, and nutrient concentration [40]. If these strains’ growth can be enhanced by further remediation strategy, petroleum hydrocarbon degradation efficiency may be promised to improve greatly.
Conclusions
High-throughput sequencing technology was used to analyze the diversity of total bacterial compositions and functional TPH degrading flora in the petroleum-contaminated soil. The composition of dominant hydrocarbon-degrading bacteria was different to soil total bacterial community, both at the phylum and genus level. The oil-contaminated soil microbial compositions were roughly the same, but their relative abundance changed upon to 15% moisture treatment. Fifteen percent of soil moisture content led to 18% of the hydrocarbon removal in the oil-contaminated soil, which may be attributed to the increment of some specific degrading bacteria belongs to Firmicutes phylum and Bacillus sp.
References
Cho E, Park M, Hur M, Kang GY, Kim S (2019) Molecular-level investigation of soils contaminated by oil spilled during the Gulf War. J. Hazard. Mater 373:271–277
Gao H, Zhang J, Lai HX, Xue QH (2017) Degradation of asphaltenes by two Pseudomonas aeruginosa strains and their effects on physicochemical properties of crude oil. Int. Biodeterior. Biodegradation 122:12–22
Shahi A, Aydin S, Ince B, Ince O (2016) Reconstruction of bacterial community structure and variation for enhanced petroleum hydrocarbons degradation through biostimulation of oil contaminated soil. Chem. Eng. J 306:60–66
Wang SY, Kuo YC, Hong A, Chang YM, Kao CM (2016) Bioremediation of diesel and lubricant oil-contaminated soils using enhanced landfarming system. Chemosphere 164:558–567
Wu ML, Ye XQ, Chen KL, Li W, Yuan J, Jiang X (2017) Bacterial community shift and hydrocarbon transformation during bioremediation of short-term petroleum-contaminated soil. Environ. Pollut 223:657–664
Xu JL, Zhao MH, Wang R, Du J, Zhang QJ (2019) Efficiently dedicated oxidation of long-chain crude oil in the soil by inactive SOM-Fe. Chem. Eng. J. https://doi.org/10.1016/j.cej.2019.121913
He J, Fan XR, Liu H, He XT, Wang QZ, Liu Y, Wei HF, Wang B (2019) The study on Suaeda heteroptera Kitag, Nereis succinea and bacteria's joint bioremediation of oil-contaminated soil. Microchem. J 147:872–878
Whelan MJ, Coulon F, Hince G, Rayner J, McWatters R, Spedding T, Snape I (2015) Fate and transport of petroleum hydrocarbons in engineered biopiles in polar regions. Chemosphere 131:232–240
Wu ML, Wu JL, Zhang XH, Ye XQ (2019) Effect of bioaugmentation and biostimulation on hydrocarbon degradation and microbial community composition in petroleum-contaminated loessal soil. Chemosphere. https://doi.org/10.1016/j.chemosphere.2019.124456
Roy A, Dutta A, Pal S, Gupta A, Sarkar J, Chatterjee A, Saha A, Sarkar P, Sar P, Kazy SK (2018) Biostimulation and bioaugmentation of native microbial community accelerated bioremediation of oil refinery sludge. Bioresour. Technol 253:22–32
Abed RMM, Al-Kharusi S, Al-Hinai M (2015) Effect of biostimulation, temperature and salinity on respiration activities and bacterial community composition in an oil polluted desert soil. Int. Biodeterior. Biodegrad 98:43–52
Ali H, Mohammad HF, Mahin S (2016) The effect of soil type on the bioremediation of petroleum contaminated soils. J. Environ. Manag 180:197–201
Wu ML, Li W, Warren AD, Ye XQ, Chen LM (2017) Bioremediation of hydrocarbon degradation in a petroleum-contaminated soil and microbial population and activity determination. Chemosphere 169:124–130
Gao H, Zhang J, Lai HX, Xue QH (2017) Degradation of asphaltenes by two Pseudomonas aeruginosa strains and their effects on physicochemical properties of crude oil. Int. Biodeterior. Biodegradation 122:12–22
Xu JL, Zhang QJ, Li DY, Du J, Wang C, Qin JY (2019) Rapid degradation of long-chain crude oil in soil by indigenous bacteria using fermented food waste supernatant. Waste Manag 85:361–373
Whelan MJ, Coulon F, Hince G, Rayner J, McWatters R, Spedding T, Snape I (2015) Fate and transport of petroleum hydrocarbons in engineered biopiles in polar regions. Chemosphere 131:232–240
Trellu C, Mousseta E, Pechaud Y, Huguenot D, Hullebusch EDV, Esposito G, Oturan MA (2016) Removal of hydrophobic organic pollutants from soil washing/flushing solutions: a critical review. J. Hazard. Mater 306:149–174
Onotasamiderhi TI, Paola M, Russell JD, David W (2019) Impacts of activated carbon amendments, added from the start or after five months, on the microbiology and outcomes of crude oil bioremediation in soil. Int. Biodeterior. Biodegradation 142:1–10
Chang W, Dyen M, Spagnuolo L, Simon P, Whyte L, Ghoshal S (2010) Biodegradation of semi- and non-volatile petroleum hydrocarbons in aged, contaminated soils from a sub-arctic site: laboratory pilot-scale experiments at site temperatures. Chemosphere 80:319–326
Gomez F, Sartaj M (2013) Field scale ex-situ bioremediation of petroleum contaminated soil under cold climate conditions. Int. Biodeterior. Biodegradation 85:375–382
Paudyn K, Rutter A, Rowe RK, Poland JS (2008) Remediation of hydrocarbon contaminated soils in the Canadian Arctic by landfarming. Cold Reg. Sci. Technol 53:102–114
Karthick A, Roy B, Chattopadhyay P (2019) A review on the application of chemical surfactant and surfactant foam for remediation of petroleum oil contaminated soil. J. Environ. Manag 243:187–205
Wang SY, Kuo YC, Hong A, Chang YM, Kao CM (2016) Bioremediation of diesel and lubricant oil-contaminated soils using enhanced landfarming system. Chemosphere 164:558–567
Schjønning P, Thomsen IK, Petersen SO, Kristensen K, Christensen BT (2011) Relating soil microbial activity to water content and tillage-induced differences in soil structure. Geoderma 163:256–264
Sigouin MJP, Dyck M, Si BC, Hu W (2016) Monitoring soil water content at a heterogeneous oil sand reclamation site using a cosmic-ray soil moisture probe. J. Hydrol 543:510–522
Zhang HH, Feng J, Chen SN, Zhao ZF, Li BQ, Wang Y (2019) Geographical patterns of nirs gene abundance and nirs-type denitrifying bacterial community associated with activated sludge from different wastewater treatment plants. Microb. Ecol 77:304–316
Meynet P, Hale SE, Davenport RJ, Cornelissen G, Breedveld GD, Werner D (2012) Effect of activated carbon amendment on bacterial community structure and functions in a PAH impacted urban soil. Environ Sci Technol 46:5057–5066
Li XX, Fan FQ, Zhang BY, Zhang KD, Chen B (2018) Biosurfactant enhanced soil bioremediation of petroleum hydrocarbons: design of experiments (DOE) based system optimization and phospholipid fatty acid (PLFA) based microbial community analysis. Int. Biodeterior. Biodegradation 132:216–225
Wu ML, Wu JL, Zhang XH, Ye XQ (2019) Effect of bioaugmentation and biostimulation on hydrocarbon degradation and microbial community composition in petroleum-contaminated loessal soil. Chemosphere. https://doi.org/10.1016/j.chemosphere.2019.124456
Qu YY, Zhang XW, Shen WL, Ma Q, You SN, Pei XF, Li SZ, Ma F, Zhou JT (2016) Illumina MiSeq sequencing reveals long-term impacts of single-walled carbon nanotubes on microbial communities of wastewater treatment systems. Bioresour. Technol 211:209–215
Wu ML, Dick WA, Li W, Wang XC, Yang Q, Wang TT, Xu LM, Zhang MH, Chen LM (2016) Bioaugmentation and biostimulation of hydrocarbon degradation and the microbial community in a petroleum-contaminated soil. Int. Biodeterior. Biodegradation 107:158–164
Hou JY, Liu WX, Wang BB, Wang QL, Franks AE (2015) PGPR enhanced phytoremediation of petroleum contaminated soil and rhizosphere microbial community response. Chemosphere 138:592–598
Vasileva-Tonkova E, Gesheva V (2005) Glycolipids produced by antarctic Nocardioides sp. during growth on n-paraffin. Process Biochem 40:2387–2391
Mohanty G, Mukherji S (2008) Biodegradation rate of diesel range n-alkanes by bacterial cultures exiguobacterium aurantiacum and burkholderia cepacia. Int. Biodeterior. Biodegradation 61:240–250
Zhang DC, Mörtelmaier C, Margesin R (2012) Characterization of the bacterial archaeal diversity in hydrocarbon-contaminated soil. Sci. Total Environ 421–422:184–196
Kim T, Hong JK, Jho EH, Kang GY, Lee SJ (2019) Sequential biowashing-biopile processes for remediation of crude oil contaminated soil in Kuwait. J Hazard Mater:378. https://doi.org/10.1016/j.jhazmat.2019.05.103
Bacosa H, Suto K, Inoue C (2010) Preferential degradation of aromatic hydrocarbons in kerosene by a microbial consortium. Int Biodeterior Biodegr 64:702–710
Das N, Chandran P (2011) Microbial degradation of petroleum hydrocarbon contaminants: an overview. Biotechnol. Res. Int. https://doi.org/10.4061/2011/941810
Ramadass K, Megharaj M, Venkateswarlu K, Naidu R (2018) Bioavailability of weathered hydrocarbons in engine oil-contaminated soil: impact of bioaugmentation mediated by pseudomonas, spp. on bioremediation. Sci. Total Environ 636:968–974
He SY, Ni YQ, Lu L, Chai QW, Yu T, Shen ZhQ Yang CP (2020) Simultaneous degradation of n-hexane and production of biosurfactants by Pseudomonas sp. strain NEE2 isolated from oil-contaminated soils. Chemosphere 242: https://doi.org/10.1016/j.chemosphere.2019.125237
Trellu C, Mousseta E, Pechaud Y, Huguenot D, Hullebusch EDV, Esposito G, Oturan MA (2016) Removal of hydrophobic organic pollutants from soil washing/flushing solutions: a critical review. J. Hazard. Mater 306:149–174
Almansoory AF, Hasan HA, Idris M, Abdullah SRS, Anuar N (2015) Potential application of a biosurfactant in phytoremediation technology for treatment of gasoline-contaminated soil. Ecol. Eng 84:113–120
Santos EVD, Sáez C, Cañizares P, Silva DRD, Rodrigo MA (2017) Treatment of ex-situ soil-washing fluids polluted with petroleum by anodic oxidation, photolysis, sonolysis and combined approaches. Chem. Eng. J 310:581–588
Marie TBA, Li TT, Shah MN, Zhong WH (2019) Biodegradation of total petroleum hydrocarbons (TPH) in highly contaminated soils by natural attenuation and bioaugmentation. Chemosphere 234:864–874
Varjani SJ (2017) Microbial degradation of petroleum hydrocarbons. Bioresour. Technol 223:277–286
Cristina MQ, Ana MTM, Leandro CPL (2019) Overview of bioremediation with technology assessment and emphasis on fungal bioremediation of oil contaminated soils. J. Environ. Manag 241:156–166
Funding
This work was supported by the National Natural Science Foundation of China (No. 21577109), the Program for Innovative Research Team in Shaanxi (PIRT) (Grant No. 2013KCT-13), the Natural Science Foundation of Shaanxi Province (2015JM5163), and the Key Laboratory Project of the Shaanxi Provincial Education Department (13JS048).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, H., Gao, H., Wu, M. et al. Distribution Characteristics of Bacterial Communities and Hydrocarbon Degradation Dynamics During the Remediation of Petroleum-Contaminated Soil by Enhancing Moisture Content. Microb Ecol 80, 202–211 (2020). https://doi.org/10.1007/s00248-019-01476-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-019-01476-7