Abstract
Butyrate is one of the most important intermediates during anaerobic digestion of protein wastewater, and its oxidization is considered as a rate-limiting step during methane production. However, information on syntrophic butyrate-oxidizing bacteria (SBOB) is limited due to the difficulty in isolation of pure cultures. In this study, two anaerobic chemostats fed with butyrate as the sole carbon source were operated at different dilution rates (0.01/day and 0.05/day). Butyrate- and acetate-oxidizing bacteria in both chemostats were investigated, combining DNA-Stable Isotope Probing (DNA-SIP) and 16S rRNA gene high-throughput sequencing. The results showed that, in addition to known SBOB, Syntrophomonas, other species of unclassified Syntrophomonadaceae were putative butyrate-oxidizing bacteria. Species of Mesotoga, Aminivibrio, Acetivibrio, Desulfovibrio, Petrimonas, Sedimentibacter, unclassified Anaerolineae, unclassified Synergistaceae, unclassified Spirochaetaceae, and unclassified bacteria may contribute to acetate oxidation from butyrate metabolism. Among them, the ability of butyrate oxidation was unclear for species of Sedimentibacter, unclassified Synergistaceae, unclassified Spirochaetaceae, and unclassified bacteria. These results suggested that more unknown species participated in the degradation of butyrate. However, the corresponding function and pathway for butyrate or acetate oxidization of these labeled species need to be further investigated.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Anaerobic digestion (AD) is an effective and environment-friendly technology for organic waste/wastewater treatment. Butyrate accounts for 17–20% of the total volatile fatty acid (VFA) intermediates during anaerobic digestion of protein wastewater, such as dairy (casein) and meat-processing (beef offal, flesh, pork flesh) wastewater [1,2,3]. Generally, the anaerobic oxidization of butyrate to H2 and acetate (ΔG0′ = + 48.1 kJ/mol) is not spontaneous under standard conditions, unless it can be overcome by the collaborative interaction between syntrophic butyrate-oxidizing bacteria (SBOB) and hydrogenotrophic methanogens, which could keep a much lower H2 partial pressure [4]. Due to the thermodynamic barrier, the accumulation of butyrate occurs easily, causes a further decrease in pH, and even leads to failure of the AD process [5]. Therefore, butyrate oxidization performed by SBOB is considered as a limited step for methane production.
Owing to the symbiotic relationships with hydrogenotrophic methanogens, pure cultures of only a few SBOB strains have been successfully obtained so far. These isolates include 11 mesophilic species and/or subspecies of the genus Syntrophomonas [6,7,8,9,10,11,12,13,14,15], two thermophilic species of the genera Thermosyntropha and Syntrophothermus [16, 17], one mesophilic species of the genus Syntrophus [18], and a psychrotolerant species of the genus Algorimarina [19]. The analysis of two SBOB genomes (Syntrophomonas wolfei and Syntrophus aciditrophicus) showed that they have multiple copies of genes related to β-oxidization for butyrate degradation [20, 21]. However, little is known about SBOB diversity in the anaerobic digester [22,23,24]. Therefore, the roles and contributions of SBOB during the AD process for biogas production need to be further investigated.
Considering the difficulty in isolating SBOB, it is necessary to explore the frontiers of technology for identifying novel SBOB in anaerobic digesters. The recent development of the stable isotope probing (SIP) technique, based on DNA or RNA, enables linking of metabolic function and taxonomic identity, especially for exploring the function of uncultured microorganisms [25]. Several reports have proven that SIP is an effective tool for identifying potential butyrate degraders in different methanogenic habitats [26,27,28,29]. More non-Syntrophomonas species, such as Tepidanaerobacter, Clostridium, Syntrophospora, Syntrophomonadaceae, Syntrophaceae, and Actinobacteria, were found to be possibly responsible for syntrophic butyrate oxidization. However, to date, only one study has investigated the diversity of butyrate-oxidizing bacteria (BOB) using SIP in methanogenic sludge [27]. Therefore, research on SBOB is of great importance for learning and regulating the AD process for protein wastewater treatment.
There are complex microbial communities and a relatively low abundance of SBOB in AD reactors for protein waste/wastewater treatment, which could reduce the chances of identifying SBOB using 13C-butyrate SIP. Moreover, our previous studies demonstrated that the dilution rate (the reciprocal of hydraulic retention time, HRT) could seriously affect the community structure of VFA-degrading bacteria in a continuous stirred tank reactor (CSTR) for AD [30, 31]. Therefore, in this study, we constructed two butyrate-fed mesophilic anaerobic CSTRs and operated them at different dilution rates (0.01/day and 0.05/day) for SBOB enrichment. Then, the potential SBOB in both reactors were investigated using DNA-SIP combined 16S rRNA high-throughput sequencing. Simultaneously, acetate-oxidizing bacteria (AOB) were also analyzed, considering that acetate is an important intermediate (butyrate + 2H2O ⟶ 2acetate + 2H2) during butyrate metabolism.
Materials and Methods
Construction and Operation of the Chemostats Fed with Butyrate as the Sole Carbon Source
Two anaerobic chemostats were constructed using two CSTRs, each with a working volume of 1.8 L as described previously [31]. The seed sludge was obtained from an anaerobic reactor treating kitchen waste. The chemostats, designated as BL and BH reactors, were fed with synthetic wastewater, containing butyrate as the sole carbon source (TOC 8000 mg/L) at dilution rates of 0.01/day and 0.05/day, respectively. BL and BH reactors operated at 37 °C for approximately 400 and 300 days, respectively, during which, parameters, including pH, suspended solids (SS), volatile suspended solid (VSS), total organic carbon (TOC), VFAs, were measured regularly, as described previously [32]. After the reactor reached a steady state at each dilution rate, the sludge was used for microbial community analysis and DNA-SIP experiments.
SIP Incubation with 13C-Butyrate and 13C-Acetate
Considering the co-existence of acetate and butyrate degraders during butyrate metabolism, the sludge was incubated separately with 13C-butyrate and 13C-acetate to aid SBOB identification. Incubations of 12C-butyrate and 12C-acetate were used as controls. Sludge was collected directly from the BL and BH reactors on days 372 and 290, respectively. Microcosms were set up individually in 50-mL serum bottles amended with 15 mL BL or BH digester sludge. After sealing with rubber stoppers and aluminum seal, and purged with nitrogen gas, cysteine-HCl (final concentration of 0.5 g/L) and resazurin (final concentration of 1 mg/L) were added as reducing agent and anaerobic condition indicator, respectively. The serum bottles were supplemented once every two days with 13C or 12C-substrate via a gas-tight syringe, and eight different treatments, each in duplicates, were established (Table 1). All microcosms were incubated at 37 °C and 150 rpm on a shaker. The incubation time had previously been optimized by a pretest (data not shown). Gas production was measured, with a syringe, every two days. After incubation, the remaining VFAs in each bottle were determined and sludge from each bottle was collected for DNA extraction. The [1-13C] sodium butyrate (98 atom% 13C) used in this study was purchased from Shanghai Engineering Research Center of Stable Isotope in China, while [2-13C] sodium acetate (99 atom% 13C) was purchased from Cambridge Isotope Laboratories, USA.
DNA Extraction, Density-Gradient Centrifugation, and Fractionation
Total genomic DNA of each sludge sample was extracted by CTAB method [33]. Purified DNA was prepared for density-gradient centrifugation and fractionation as described by Lueders et al. [34]. Briefly, total DNA (∼ 2.5 μg) was added to Quick-Seal polyallomer tubes (6.3 mL, Beckman Coulter, Australia), along with 1.2 mL gradient buffer (GB) (containing 0.1 M Tris-HCl (pH 8.0), 0.1 M KCl, and 1 mM EDTA) and 4.8 mL CsCl solution (final buoyant density of 1.9 g/mL). Then, the tubes were sealed and centrifuged at 177,000×g for 40 h at 20 °C in a Beckman ultracentrifuge with Ti90 fixed angle rotor (Beckman, USA). Following centrifugation, 15 density fractions each with 400 μL, numbered fractions 1–15 from the bottom (high density) to the top (low density), were collected from each tube using a fraction recovery system (Beckman Coulter, USA). The buoyant density of each fraction was determined by a digital refractometer (AR200, Reichert, USA) [34], and DNA was recovered from each fraction by PEG6000 precipitation with glycogen [35].
In order to profile the DNA gradient distribution, bacterial and archaeal 16S rRNA genes in each fraction were quantified by qPCR, using the EcoTM real-time PCR system (Illumina, USA) with primer sets Eu27f/Eu518r [36] and Arch349f/Arch806r [37], respectively. Reaction mixtures (20 μL) were prepared with 2 μL template DNA (~ 2 g/μL and ~ 7 g/μL DNA used for amplification of bacterial and archaeal 16S rRNA gene, respectively) and 0.8 μL of each primer. qPCR conditions for bacterial 16S rRNA gene were as follows: denaturation at 95 °C for 10 s, annealing at 55 °C for 5 s, and extension at 72 °C for 40 s for a total of 40 cycles. qPCR conditions for archaeal 16S rRNA gene were as follows: denaturation at 95 °C for 40 s, annealing at 53 °C for 45 s, and extension at 72 °C for 45 s for a total of 40 cycles. Based on qPCR results, DNA samples from several heavy density fractions and total DNA for each treatment were used for sequencing, as shown in Table 1.
High-Throughput Sequencing and Phylogenetic Analysis
The V4–V5 regions of the bacterial and archaeal 16S rRNA genes of the DNA samples were amplified using the primers, 515F (5′-GTGCCAGCMGCCGCGGTAA-3′) and 909R (5′-CCCCGYCAATTCMTTTRAGT-3′) [38]. Sequencing was performed on the Illumina MiSeq platform using the MiSeq v2 reagent kit (2 × 250 bp) by Chengdu Institute of Biology. Raw FASTQ files were quality-processed using the QIIME Pipeline-Version 1.7.0 (http://qiime.org/). Chimeric sequences were removed using the Uchime algorithm [39]. Operational taxonomic units (OTUs) were defined by clustering at 97% similarity. Final OTUs were taxonomically classified using Ribosomal Database Project classifier and NCBI blast [40]. Phylogenetic analysis was performed using the MEGA4 software package (http://megasoftware.net/mega4/) after multiple sequence alignment by Clustalx1.8 (http://www.clustal.org/). Phylogenetic trees were constructed using neighbor-joining method.
Nucleotide Sequence Accession Numbers
The original sequencing data is available at the National Center for Biotechnology Information database (accession no. PRJNA475621).
Results
Microbial Community of Butyrate-Fed Chemostats Operated at Different Dilution Rates
BL (0.01/day) and BH (0.05/day) reactors operated for approximately 400 and 300 days, respectively, and both retained the steady state. The performance of both reactors is displayed in Fig. S1. The biogas yield of the BL and BH reactors was stable, at approximately 80 and 900 mL/day during this period, respectively. TOC and pH in the two reactors were 40–50 mg/L and 8–8.3, respectively. No organic acid accumulation was observed, and butyrate fed into each reactor was completely mineralized. The average VSS concentration was 1.7 g/L for BL reactor and 1.5 g/L for BH reactor. The sludge from each chemostat was used for microbial community analyses.
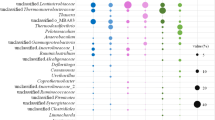
Based on the high-throughput sequencing of 16S rRNA gene, the composition of bacterial community in both reactors was obviously different (Fig. 1). In the BL reactor, 84.22% of bacterial OTUs were assigned to four phyla: Thermotogae (25.65%), Firmicutes (22.83%), Bacteroidetes (18.24%), and Proteobacteria (17.50%) (Fig. 1a). In the BH reactor, phylum Firmicutes accounted for 75.33% abundance (Fig. 1b). At the genus level, Defluviitoga, Mesotoga, Coprothermobacter, and Petrimonas were the dominant genera at the dilution rate of 0.01/day (BL reactor), accounting for 13.27%, 12.37%, 10.16%, and 9.52% of the total valid reads, respectively. Several other key genera, including Syntrophomonas (6.09%), Desulfovibrio (5.71%), Dechloromonas (4.3%), and Anaerobaculum (3.35%), also were detected. Among them, Syntrophomonas is known as a SBOB in association with H2-utilizing methanogens. It was observed following enrichment at the higher dilution rate (0.05/day) and became the most abundant genus (54.54%) in the BH reactor. In addition, one OTU, which could not be affiliated to any bacterial phylum, accounted for 15.19% of total bacterial reads and was the second most abundant genus in the BH reactor.
The archaeal communities of both reactors are listed in Table S1. In the BL reactor, acetoclastic Methanosaeta (53.62%) and hydrogenotrophic Methanobacterium (36.15%) and Methanothermobacter (7.14%) were the primary methanogens. Methanosaeta remained dominant in the BH reactor with an abundance of 78.60%. Methanoculleus (17.27%) became the most dominant hydrogenotrophic methanogen, followed by Methanobacterium (3.63%). Methanothermobacter sp. was not detected in the BH reactor.
DNA-SIP Analysis
Sludge from both reactors was incubated with 13C- and 12C-substrates for 30 and 20 days, respectively (Table 1). The VFA consumption and gas yield in each microcosm were measured. Two biological replicates of each treatment showed almost the same VFA consumption and biogas production. As shown in Table S2 and Fig. S2, 13C- and 12C-butyrate could be completely consumed by both methanogenic sludges, and the gas yields were approximate 74.18–92.40% of the theoretical value. For treatments with BL sludge, more 12C-acetate (100%) than 13C-acetate (about 80%) was utilized, and their gas yields were 60.15–66.9% of the theoretical value. BH sludge consumed 100% acetate, and the gas yields were around 56% of the theoretical value.
Total DNA from all microcosms was used for density-gradient centrifugation and fractionation. The DNA distribution profiles for different density fractions are illustrated by relative copies of bacterial and archaeal 16S rRNA genes (Fig. 2; Fig. S3). As shown in Fig. 2a, heavy density fractions, 1.71 to 1.72 g/mL (fraction 8), contained more bacterial 16S rRNA genes (29.11% and 17.85%) in the 13C-butyrate treatments with BL sludge than in the 12C-controls (8.91% and 7.48%). For 13C-acetate treatments with BL sludge (Fig. 2b), 13.7% and 10.5% of bacterial 16S rRNA genes were enriched at a density of 1.73 g/mL (fraction 7). Figure S3a shows that the abundance of archaeal 16S rRNA genes increased from 14.8% and 8.05% (12C-controls) to 52.52% and 36.33% (13C-butyrate treatments) in the heavy fractions. More archaeal 16S rRNA genes (37.4% and 20.5%) were obtained in the heavy density DNA fractions from 13C-acetate treatments with BL sludge (Fig. S3b). Similarly, about 18.66% and 16.91% bacterial 16S rRNA genes were enriched at density fractions ranging from 1.72 to 1.73 g/mL (fraction 8) in the 13C-butyrate treatments with BH sludge (Fig. 2c). At the same density fractions, archaeal 16S rRNA genes presented with 3.77% and 6.38% abundance (Fig. S3c). For the 13C-acetate treatments with BH sludge, 13.99% and 14.59% of 16S rRNA genes (Fig. 2d) were found at a density of 1.73 g/mL (fraction 7), and more archaeal 16S rRNA genes (14.23% and 13.19%) were obtained in the heavy density fractions (Fig. S3d). These results suggested that bacterial and archaeal species were labeled successfully by 13C-substrates. In order to be closer to the potential BOB and AOB, the 16S rRNA genes in the heavy density fractions (fraction 8–10 for butyrate of BL and BH, fraction 7–10 for acetate of BL and fraction 6–9 for acetate of BH) were used for sequencing analyses. Whole DNA for 16 samples from eight different treatments (Table S2) were sequenced.
Phylogenetic Identification of the Labeled Bacterial and Archaeal Species
Bacterial Community at the Phylum Level
Microbial community data obtained from total DNA from 12C- and 13C-treatments were compared at the phylum level, using 16S rRNA gene sequences. The results showed that the composition and abundances of phyla were similar in the total DNA from BL or BH sludge treated with 12C- and 13C-substrates (data not shown). Phyla Proteobacteria, Thermotogae, Firmicutes, Bacteroidetes, Chloroflexi, and Synergistetes were predominant in BL sludge treatments, while Firmicutes, Synergistetes, Spirochaetae, Thermotogae, Bacteroidetes, and unclassified bacteria presented in BH sludge treatments.
For 13C-butyrate-treated BL sludge (Fig. 3a), Firmicutes and Synergistetes (two dominant phyla), accounted for 11.17% and 6.47% of the total reads in fraction 8 (1.718 g/mL), respectively, while there were only 1% of Firmicutes and Synergistetes in the corresponding 12C-control. In addition, the abundance of Bacteroidetes increased nearly three times, both in fraction 8 (from 0.14% in 12C-butyrate treatment to 0.44% in 13C-butyrate treatment) and fraction 9 (from 2.83% in 12C-butyrate treatment to 7.52% in 13C-butyrate treatment). More phyla related to acetate oxidation were observed in BL sludge, especially in fraction 7 (1.727 g/mL) and fraction 8 (1.714 g/mL) (Fig. 3a). Firmicutes was enriched in fraction 7 and 8 with an abundance of 5.44% and 9.48%, respectively, while it was only 0.21% and 1.16% in the corresponding fraction of 12C-control. Synergistetes was enriched in fraction 7 (from 0.09% in 12C-acetate treatment to 3.48% in 13C-acetate treatment) and in fraction 8 (from 1.23% to 3.02%). Moreover, phyla Proteobacteria, Bacteroidetes, Chloroflexi, and Thermotogae slightly increased in abundance in fraction 8 of DNA treated with 13C-acetate, compared with the 12C-control.
For 13C-butyrate-treated BH sludge (Fig. 3b), four key phyla were significantly enriched, largely in the DNA of fraction 8 (1.727 g/mL), including Firmicutes (5.21%), Synergistetes (6.81%), Spirochaetae (1.88%), and unclassified bacteria (2.42%), while phylum Thermotogae (0.27%) slightly increased. These phyla accounted for 0.44%, 0.21%, 0.15%, 0.39%, and 0.04% in fraction 8 of DNA from the 12C-butyrate treatment, respectively. For acetate-treated BH sludge, Synergistetes was enriched, largely in fractions 6–8 (1.749 to 1.722 g/mL) of DNA from the 13C-acetate treatment, with an abundance of 5.75%, 10.91%, and 10.01%, respectively, compared with the abundances from the 12C-acetate treatment (0.17%, 0.31%, and 2.08%, respectively). The abundance of two other phyla, Firmicutes and unclassified bacteria, was 2–3 times higher in fractions 7 and fraction 8 of DNA from 13C-acetate treatment than those in the 12C-control. In addition, small quantities of Spirochaetae, Thermotogae, and Bacteroidetes were enriched in the heavy density DNA fraction from 13C-acetate treatments. The results above suggested that much more phyla involved in the butyrate and acetate oxidation.
Bacterial Community at the Genus Level
There were more apparent differences between the bacterial communities in heavy density fractions of 13C- and 12C-treatments at the genus level (Figs. 4, 5, and 6). The abundance of several represented bacteria increased in the heavy density 13C-treated DNA fractions, especially focusing on fraction 8 for both butyrate and acetate.
Compared with 12C-butyrate-treated BL sludge, Syntrophomonas (BL-OTU281, 7.61%), unclassified Synergistaceae (BL-OTU29, 4.43%), Sedimentibacter (BL-OTU152, 0.59%), and unclassified Syntrophomonadaceae (BL-OTU7514, 0.63%) populations increased separately, ten-fold in fraction 8 of DNA from the 13C-butyrate treatments (Figs. 4a, b, e and 6). In acetate-treated BL sludge, Sedimentibacter (BL-OTU152) and unclassified Synergistaceae (BL-OTU29) were greatly enriched. Sedimentibacter populations increased from 0.05% in 12C-acetate treatments to 5.25% in 13C-acetate treatments (105-fold increase) in fraction 8, while unclassified Synergistaceae increased from 0.02 to 1.77% (88.5-fold increase) in fraction 7 (Figs. 4c–e and 6). In addition, the abundance of Mesotoga (BL-OTU23, 1.92%), unclassified Anaerolineae_1 (BL-OTU45, 1.79%), Desulfovibrio (BL-OTU146, 1.25%), and Petrimonas (BL-OTU95, 0.9%) in the eighth density 13C-acetate-treated DNA fractions were higher than those from 12C-controls, which suggested that these bacteria were enriched by 13C-acetate.
Compared with 12C-butyrate-treated BH sludge, unclassified Synergistaceae (BH-OTU209, 6.10%) and Syntrophomonas (BH-OTU324, 2.75%) populations increased by 47- and 18-fold, respectively, in the 8th density DNA fractions from the 13C-butyrate treatments (Figs. 5a, b, e and 6). Syntrophomonas (BH-OTU80, 2.01%), unclassified Spirochaetaceae (BH-OTU142, 1.80%), unclassified bacteria (BH-OTU344, 2.42%), and Mesotoga (BH-OTU48, 0.27%) were enriched 5–9-fold in the same DNA fraction. Compared with 12C-acetate-treated BH sludge, unclassified Synergistaceae (BH-OTU209, 10.61% in fraction 7), Acetivibrio (BH-OTU287, 1.23% in fraction 8), and unclassified Spirochaetaceae (BH-OTU142, 3.02% in fraction 8) were enriched 40-, 17-, and 5-fold, respectively, in DNA from 13C-acetate treatments (Figs. 5c–e and 6). The abundance of three other genera increased slightly in heavy density DNA fractions from 13C-acetate treatments compared with the 12C-control, including unclassified bacteria (BH-OTU344, 1.79%), Mesotoga (BH-OTU48, 0.6%), and Aminivibrio (BH-OTU82, 0.71%).
Archaeal Community at the Genus Level
Archaeal community studies at the genus level showed that both acetotrophic and hydrogenotrophic methanogens were enriched by the 13C-substrate (Figs. S4 and S5). Compared with the 12C-butyrate-treated BL sludge microcosm, Methanosaeta (21.17%), Methanoculleus (14.63%), Methanobacterium (4.70%), and Methanothermobacter (3.62%) were enriched 3.3–6.7-fold in fraction 8 of 13C-butyrate treatments (Fig. S5). Four genera, including Methanosaeta, Methanothermobacter, Methanobacterium, and unclassified Methanomassiliicoccaceae, were largely enriched following 13C-acetate treatments. Unclassified Methanomassiliicoccaceae and Methanosaeta particularly presented with an abundance of 4.97% and 20.13% in the 7th density DNA fractions from 13C-acetate treatments, while there were only 0.01% and 0.35% in the corresponding 12C-controls, respectively. In the BH sludge microcosm (Fig. S5), Methanosaeta, Methanoculleus, and Methanobacterium increased in the heavy fractions of 13C-butyrate treatments compared with 12C-control treatments, while the enrichment of Methanosaeta and Methanoculleus was observed in the heavy fractions of 13C-acetate treatments.
Discussion
In this study, two mesophilic chemostats fed with butyrate as the sole carbon source were constructed and operated at different dilution rates. The potential butyrate-oxidizing bacteria (BOB), acetate-oxidizing bacteria (AOB), and methanogens in both chemostats were investigated by DNA-SIP and 16S rRNA high-throughput sequencing. The results showed that different species were responsible for butyrate and acetate oxidation in both chemostats.
Among the species enriched in the heavy density DNA fractions from 13C-butyrate and 13C-acetate-treated BL sludge, only two Syntrophomonadaceae OTUs (BL-OTU281 and BL-OTU7514) were enriched in the heavy density DNA fractions from 13C-butyrate treatment. Syntrophomonas (BL-OTU281) and unclassified Syntrophomonadaceae (BL-OTU7514) shared 98% and 93% sequence identity with Syntrophomonas wolfei, respectively. The 10-fold enrichment observed for both OTUs suggests that they played key roles in butyrate oxidation. Four genera were enriched in the heavy density DNA fractions from 13C-acetate treatment, including Mesotoga (BL-OTU23), unclassified Anaerolineae (BL-OTU45), Desulfovibrio (BL-OTU146), and Petrimonas (BL-OTU95). This suggested that these OTUs may be related to acetate oxidation. BL-OTU23 was affiliated to Mesotoga infera VNs100 (97% similarity), a members of the phylum Thermotogae. According to metagenomic analysis results, Nobu et al. found that Mesotoga may syntrophically oxidize acetate through a previously uncharacterized pathway [41]. BL-OTU45 showed 98% similarity with function-unknown uncultured clone 22 (MH040197) and clone B146 (HQ640609), belonging to class Anaerolineae of phylum Chloroflexi. Several reports have suggested that syntrophic metabolism of butyrate and propionate could occur by filamentous Anaerolineaceae and Methanosaeta via direct interspecies electron transfer (DIET) during the AD process [42, 43]. Therefore, the syntrophic oxidation of acetate via DIET may have occurred between Anaerolineae and Methanosaeta in this study, considering the high abundance of Methanosaeta in BL sludge; however, this requires further investigation. BL-OTU146 and BL-OTU95 showed highest similarity with Desulfovibrio oryzae PETROMIC B02 (AY664600) (96%) and Petrimonas sulfuriphila BN3 (97%), respectively. Desulfovibrio spp. are known as sulfate-reducing bacteria [44], and to date, only the butyrate oxidation ability of Desulfovibrio butyratiphilus sp. nov. BSYT has been described [45]. Recently, several bacteria in the sulfate-reducing sediment were labeled by 13C-acetate, including unclassified Desulfobacteraceae and Desulfovibrio [46]. Here, we inferred that these sulfate reducers may utilize acetate for energy production or growth, as a building block of biosynthesis. Petrimonas sulfuriphila is a strictly anaerobic bacteria, which is able to utilize sugars as carbon and energy sources, and reduce sulfur to sulfide with hydrogen [47]. Therefore, the roles of Petrimonas in BL sludge remain unclear; however, the labeling of this bacterium suggests that it may be related to acetate oxidation. BL-OTU152 and BL-OTU29 were largely enriched in the heavy density DNA fractions from both 13C-butyrate and 13C-acetate treatments, suggesting that both OTUs may be mostly involved in acetate oxidation. BL-OTU152 was closely related to Sedimentibacter (96%) of the family Synergistaceae, and species in this genus are often identified as amino acid-utilizing bacteria [48, 49]. Recently, several fermentative microorganisms including Sedimentibacter were found in acetate-fed microbial fuel cells, although acetate was a non-fermentative substrate [50, 51]. Regueiro et al. believed that Sedimentibacter played an important role in the degradation of accumulated VFAs, considering that Sedimentibacter appeared or increased in population and remained until VFA levels decreased in a temperature-changed AD system [52]. Thus, Sedimentibacter labeled by 13C-acetate (105-fold increase) in our study suggests that it is an acetate-oxidizing bacterium. BL-OTU29 also showed high similarity (98%) with another amino acid-fermenting bacterium, Synergistaceae DZ-S4 (MF185666), isolated from a municipal anaerobic sewage sludge digester. Although most cultured microbes belonging to family Synergistaceae have the ability to degrade amino acids into VFAs [53, 54], some species from Synergistaceae may also ferment VFAs via syntrophic relationships with methanogens during anaerobic digestion [55]. Hence, the roles of this OTU in BL sludge may be related to acetate oxidation. The simultaneous labeling of BL-OTU152 and BL-OTU29 by 13C-butyrate may be due to cross-feeding.
Among the species enriched in the heavy density DNA fractions from 13C-butyrate and 13C-acetate-treated BH sludge, two syntrophic BOB, Syntrophomonas BH-OTU80 and BH-OTU324, were only enriched from 13C-butyrate treatment. They showed 98% and 96% similarity with Syntrophomonas wolfei Goettingen G311 and Syntrophomonas zehnderi OL-4, respectively. Aminivibrio BH-OTU82 and Acetivibrio BH-OTU287 were only enriched in the heavy density DNA fractions from 13C-acetate treatment, suggesting that they play key roles in acetate oxidation. BH-OTU82 was most closely related to Aminivibrio pyruvatiphilus 4F6E (99% similarity), an amino acid-degrading bacterium when in co-culture with the hydrogen-utilizing methanogen Methanobacterium formicicum JCM 10132(T) [54]. According to the phylogenetic tree, BH-OTU82 belonged to Synergistes group 4 (clone RSg13-6 and clone 13Cpro-5 in the Fig. 6), which was recently identified by RNA-SIP and MAR-FISH as the only predominant acetate-utilizing bacteria in anaerobic digester sludge [56]. In addition, Synergistes group 4 was found to have maximum utilization rate and high Km for acetate, and they are more competitive than acetoclastic Methanosaeta at high acetate concentrations. Taken together, BH-OTU82 may play a role in syntrophic oxidation of acetate in BH sludge. BH-OTU287 had 96% similarity with Acetivibrio cellulolyticus HL-2, a well-known anaerobic cellulolytic microorganism [57]. However, no report referred to its function in acetate oxidation, and this needs to be further confirmed. Four common OTUs were enriched in the heavy density DNA fractions from both the 13C-butyrate and 13C-acetate treatments, including BH-OTU209, BH-OTU142, BH-OTU48, and BH-OTU 344. These were most likely involved in acetate oxidation. BH-OTU209 was most closely related to Synergistaceae clone VHW_D_R9 (JQ085712, 99% similarity) from a two-stage digester treating solid wastes. And, it had 95% similarity with Thermovirga lienii DSM 17291, an amino-acid-degrading bacterium [53]. In other studies in our lab, the same OTU was also largely enriched in different mesophilic digester sludges when incubated with 13C-acetate or 13C-propionate (data not shown). Xu et al. [58] reported that one OTU (99% similarity with BH-OTU209) accounted for nearly half of the total reads in the anaerobic reactor fed with acetate. Therefore, BH-OTU209 may have the ability for syntrophic acetate oxidation. BH-OTU142 only had 94% similarity with the pure-culture bacterium, Rectinema cohabitans HM (NR_156915). It was affiliated to uncultured Spirochaetaceae clone F3 (MG674678, 99% similarity), which was obtained from an anaerobic butyrate oxidation system. Selective enrichment of Spirochaetes was observed during the AD process, accepting VFAs, especially acetate, as substrate, and suggesting the possible role of Spirochaetes in syntrophic acetate oxidation [59]. BH-OTU48 was affiliated to Mesotoga infera VNs100 (99% similarity). BH-OTU344 was highly abundant (15.19%) in the original BH reactor sludge (Fig. 1b). It showed 99% similarity with uncultured clone QEDR1AF11 obtained from an anaerobic digestion system treating sludge [60], but could not be affiliated to any bacterial phylum (Fig. 5). Taken together, the four OTUs may be putative AOB, and cross-feeding may lead to the labeling of them in the 13C-butyrate treatments.
The community analysis above showed that different species were retrieved from different sludges. From the DNA-SIP results, species from Syntrophomonadaceae may be putative BOB in both the BL and BH sludges. Liu et al. determined, using DNA-SIP, that Syntrophomonadaceae, together with the methanogens, Methanosarcinaceae and Methanocellales, were responsible for syntrophic oxidation of butyrate in paddy soil [29]. But, in other environments, more non-Syntrophomonadaceae bacteria were labeled by 13C-butyrate [26,27,28]. This may be because Syntrophomonadaceae-affiliated species were more easily enriched in chemostats during long-term incubation using butyrate as the sole carbon source. The diversity of bacteria labeled by 13C-acetate was higher than expected in this study. Species from Mesotoga, unclassified Anaerolineae, Desulfovibrio, Petrimonas, Sedimentibacter, and unclassified Synergistaceae may be the AOB in the BL sludge, while species from Aminivibrio, Acetivibrio, Mesotoga, unclassified Synergistaceae, unclassified Spirochaetaceae, and unclassified bacteria may be related to acetate oxidation in the BH sludge. Similar to our results, several reports, using DNA- or RNA-SIP identified some species belonging to Desulfovibrionaceae, Synergistaceae, Firmicutes, Bacteroidetes, and Proteobacteria as acetate utilizers [46, 56, 61]. Hao et al. also observed that species from the classes Clostridia, Thermotogae, and Spirochaetes were labeled by 13C-acetate in thermophilic methanogenic reactors with high ammonia levels [62].
Regarding archaea in 13C-butyrate-treated BL or BH sludge, nearly all methanogens, except Methanomassiliicoccus in BL sludge, existing in the original chemostats were labeled, which suggested that they were all involved in methane production from butyrate. Acetotrophic Methanosaeta and hydrogenotrophic Methanoculleus were particularly largely concentrated in the heavy fractions. Tang et al. also found that these two genera dominated in a mesophilic butyrate-degrading methanogenic reactor, at low dilution rates [31]. Lower H2 partial pressures of Methanoculleus than other hydrogenotrophic methanogens may have permitted its dominant position in both chemostats [63]. In fact, hydrogenotrophic methanogens, including Methanoculleus, Methanobacterium, and Methanothermobacter, should compete for hydrogen with each other due to their different hydrogen affinities [64]. This may allow the enrichment and labelling of only Methanothermobacter in the BL sludge. Similarly, different hydrogenotrophic methanogens were enriched in 13C-acetate treatments with BL (Methanobacterium and Methanothermobacter) and BH (Methanoculleus) sludges. Detection of hydrogenotrophic methanogens in our study suggests that syntrophic acetate oxidation could occur [65].
In conclusion, microbial community analyses showed that DNA-SIP successfully identified butyrate and acetate oxidizers. Different species were retrieved from two chemostats operated at different dilution rates, suggesting that more uncultured bacteria played roles in butyrate degradation during AD. However, some functional bacteria, which have slower growth rates and lower abundance in the sludge, may be difficult to identify using DNA-SIP. In addition, the presence of these species identified by DNA-SIP does not mean activity, which should be further confirmed by culture-dependent technology and RNA approach (such as RNA-SIP and metatranscriptomics).
References
Ramsay IR, Pullammanappallil PC (2001) Protein degradation during anaerobic wastewater treatment: derivation of stoichiometry. Biodegradation 12:247–256. https://doi.org/10.1023/a:1013116728817
Batstone DJ, Pind PF, Angelidaki I (2003) Kinetics of thermophilic, anaerobic oxidation of straight and branched chain butyrate and valerate. Biotechnol Bioeng 84:195–204. https://doi.org/10.1002/bit.10753
Tang Y, Shigematsu T, Morimura S, Kida K (2005) Microbial community analysis of mesophilic anaerobic protein degradation process using bovine serum albumin (BSA)-fed continuous cultivation. J Biosci Bioeng 99:150–164. https://doi.org/10.1263/jbb.99.150
Schink B (1997) Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev 61:262–280. https://doi.org/10.1016/j.ijpharm.2004.07.010
Schink B, Stams AJM (2013) Syntrophism among prokaryotes. In: Rosenberg E, Delong E, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes. Springer, New York, pp 471–493
Mcinerney MJ, Bryant MP, Hespell RB, Costerton JW (1981) Syntrophomonas wolfei gen. Nov. sp. nov., an anaerobic, syntrophic, fatty acid-oxidizing bacterium. Appl Environ Microbiol 41:1029–1039
Wu C, Liu X, Dong X (2006) Syntrophomonas cellicola sp. nov., a spore-forming syntrophic bacterium isolated from a distilled-spirit-fermenting cellar, and assignment of Syntrophospora bryantii to Syntrophomonas bryantii comb. nov. Int J Syst Evol Microbiol 56:2331–2335. https://doi.org/10.1099/ijs.0.64377-0
Lorowitz WH, Zhao H, Bryant MP (1989) Syntrophomonas wolfei subsp. saponavida subsp. nov., a long-chain fatty-acid-degrading, anaerobic, syntrophic bacterium; Syntrophomonas wolfei subsp. wolfei subsp. nov.; and emended descriptions of the genus and species. Int J Syst Bacteriol 39:122–126. https://doi.org/10.1099/00207713-39-2-122
Wu C, Dong X, Liu X (2007) Syntrophomonas wolfei subsp. methylbutyratica subsp. nov., and assignment of Syntrophomonas wolfei subsp. saponavida to Syntrophomonas saponavida sp. nov. comb. nov. Syst Appl Microbiol 30:376–380. https://doi.org/10.1016/j.syapm.2006.12.001
Wu C, Liu X, Dong X (2006) Syntrophomonas erecta subsp. sporosyntropha subsp. nov., a spore-forming bacterium that degrades short chain fatty acids in co-culture with methanogens. Syst Appl Microbiol 29:457–462. https://doi.org/10.1016/j.syapm.2006.01.003
Zhang C, Liu X, Dong X (2005) Syntrophomonas erecta sp. nov., a novel anaerobe that syntrophically degrades short-chain fatty acids. Int J Syst Evol Microbiol 55:799–803. https://doi.org/10.1099/ijs.0.63372-0
Zhao HX, Yang DC, Woese CR, Bryant MP (1990) Assignment of Clostridium bryantii to Syntrophospora bryantii gen. Nov., comb. nov. on the basis of a 16S rRNA sequence analysis of its crotonate-grown pure culture. Int J Syst Bacteriol 40:40–44. https://doi.org/10.1099/00207713-40-1-40
Zhang C, Liu X, Dong X (2004) Syntrophomonas curvata sp. nov., an anaerobe that degrades fatty acids in co-culture with methanogens. Int J Syst Evol Microbiol 54:969–973. https://doi.org/10.1099/ijs.0.02903-0
Sousa DZ, Smidt H, Alves MM, Stams AJ (2007) Syntrophomonas zehnderi sp. nov., an anaerobe that degrades long-chain fatty acids in co-culture with Methanobacterium formicicum. Int J Syst Evol Microbiol 57:609–615. https://doi.org/10.1099/ijs.0.64734-0
Hatamoto M, Imachi H, Fukayo S, Ohashi A, Harada H (2007) Syntrophomonas palmitatica sp. nov., an anaerobic, syntrophic, long-chain fatty-acid-oxidizing bacterium isolated from methanogenic sludge. Int J Syst Evol Microbiol 57:2137–2142. https://doi.org/10.1099/ijs.0.64981-0
Svetlitshnyi V, Rainey F, Wiegel J (1996) Thermosyntropha lipolytica gen. Nov., sp. nov., a lipolytic, anaerobic, alkalitolerant, thermophilic bacterium utilizing short- and long-chain fatty acids in syntrophic coculture with a methanogenic archaeum. Int J Syst Bacteriol 46:1131–1137. https://doi.org/10.1099/00207713-46-4-1131
Sekiguchi Y, Kamagata Y, Nakamura K, Ohashi A, Harada H (2000) Syntrophothermus lipocalidus gen. Nov., sp. nov., a novel thermophilic, syntrophic, fatty-acid-oxidizing anaerobe which utilizes isobutyrate. Int J Syst Evol Microbiol 50(Pt 2):771–779. https://doi.org/10.1099/00207713-50-2-771
Jackson BE, Bhupathiraju VK, Tanner RS, Woese CR, Mcinerney MJ (1999) Syntrophus aciditrophicus sp. nov., a new anaerobic bacterium that degrades fatty acids and benzoate in syntrophic association with hydrogen-using microorganisms. Arch Microbiol 171:107–114. https://doi.org/10.1007/s002030050685
Kendall M, Liu Y (2006) Butyrate- and propionate-degrading syntrophs from permanently cold marine sediments in Skan Bay, Alaska, and description of Algorimarina butyrica gen. Nov., sp nov. FEMS Microbiol Lett 262:107–114. https://doi.org/10.1111/j.1574-6968.2006.00380.x
Mcinerney MJ, Rohlin L, Mouttaki H, Kim UM, Krupp RS, Rios-Hernandez L, Sieber J, Struchtemeyer CG, Bhattacharyya A, Campbell JW (2007) The genome of Syntrophus aciditrophicus: life at the thermodynamic limit of microbial growth. Proc Natl Acad Sci U S A 104:7600–7605. https://doi.org/10.1073/pnas.0610456104
Sieber JR, Sims DR, Han C, Kim E, Lykidis A, Lapidus AL, Mcdonnald E, Rohlin L, Culley DE, Gunsalus R (2010) The genome of Syntrophomonas wolfei: new insights into syntrophic metabolism and biohydrogen production. Environ Microbiol 12:2289–2301. https://doi.org/10.1111/j.1462-2920.2010.02237.x
Hansen KH, Ahring BK, Raskin L (1999) Quantification of syntrophic fatty acid-β-oxidizing bacteria in a mesophilic biogas reactor by oligonucleotide probe hybridization. Appl Environ Microbiol 65:4767–4774. https://doi.org/10.1016/j.ydbio.2005.05.019
Menes RJ, Travers D (2006) Detection of fatty acid beta-oxidizing syntrophic bacteria by fluorescence in situ hybridization. Water Sci Technol 54:33–39. https://doi.org/10.2166/wst.2006.483
Zellner G, Macario AJL, Macario ECD (1997) A study of three anaerobic methanogenic bioreactors reveals that syntrophs are diverse and different from reference organisms 1. FEMS Microbiol Eco 22:295–301. https://doi.org/10.1111/j.1574-6941.1997.tb00381.x
Dumont MG, Murrell JC (2005) Stable isotope probing - linking microbial identity to function. Nat Rev Microbiol 3:499–504. https://doi.org/10.1038/nrmicro1162
Chauhan A, Ogram A (2006) Fatty acid-oxidizing consortia along a nutrient gradient in the Florida Everglades. Appl Environ Microbiol 72:2400–2406. https://doi.org/10.1128/AEM.72.4.2400-2406.2006
Hatamoto M, Imachi H, Yashiro Y, Ohashi A, Harada H (2008) Detection of active butyrate-degrading microorganisms in methanogenic sludges by RNA-based stable isotope probing. Appl Environ Microbiol 74:3610–3614. https://doi.org/10.1128/AEM.00045-08
Kristiansen A, Lindholst S, Feilberg A, Nielsen PH, Neufeld JD, Nielsen JL (2011) Butyric acid- and dimethyl disulfide-assimilating microorganisms in a biofilter treating air emissions from a livestock facility. Appl Environ Microbiol 77:8595–8604. https://doi.org/10.1128/AEM.06175-11
Liu P, Qiu Q, Lu Y (2011) Syntrophomonadaceae-affiliated species as active butyrate-utilizing syntrophs in paddy field soil. Appl Environ Microbiol 77:3884–3887. https://doi.org/10.1128/AEM.00190-11
Hori T, Haruta S, Ueno Y, Ishii M, Igarashi Y (2006) Dynamic transition of a methanogenic population in response to the concentration of volatile fatty acids in a thermophilic anaerobic digester. Appl Environ Microbiol 72:1623–1630. https://doi.org/10.1128/AEM.72.2.1623-1630.2006
Tang YQ, Shigematsu T, Morimura S, Kida K (2007) Effect of dilution rate on the microbial structure of a mesophilic butyrate-degrading methanogenic community during continuous cultivation. Appl Microbiol Biotechnol 75:451–465. https://doi.org/10.1007/s00253-006-0819-2
Jiang X, Hayashi J, Sun ZY, Yang L, Tang YQ, Oshibe H, Osaka N, Kida K (2013) Improving biogas production from protein-rich distillery wastewater by decreasing ammonia inhibition. Process Biochem 48:1778–1784. https://doi.org/10.1016/j.procbio.2013.08.014
Shigematsu T, Tang Y, Kawaguchi H, Ninomiya K, Kijima J, Kobayashi T, Morimura S, Kida K (2003) Effect of dilution rate on structure of a mesophilic acetate-degrading methanogenic community during continuous cultivation. J Biosci Bioeng 96:547–558. https://doi.org/10.1016/s1389-1723(04)70148-6
Lueders T, Manefield M, Friedrich MW (2010) Enhanced sensitivity of DNA- and rRNA-based stable isotope probing by fractionation and quantitative analysis of isopycnic centrifugation gradients. Environ Microbiol 6:73–78. https://doi.org/10.1046/j.1462-2920.2003.00536.x
Neufeld JD, Schäfer H, Cox MJ, Boden R, Mcdonald IR, Murrell JC (2007) Stable-isotope probing implicates Methylophaga spp and novel Gammaproteobacteria in marine methanol andmethylamine metabolism. ISME J 1:480–491. https://doi.org/10.1038/ismej.2007.65
Shigematsu T, Tang Y, Mizuno Y, Kawaguchi H, Morimura S, Kida K (2006) Microbial diversity of mesophilic methanogenic consortium that can degrade long-chain fatty acids in chemostat cultivation. J Biosci Bioeng 102:535–544. https://doi.org/10.1263/jbb.102.535
Takai K, Horikoshi K (2000) Rapid detection and quantification of members of the archaeal community by quantitative PCR using fluorogenic probes. Appl Environ Microbiol 66:5066–5072. https://doi.org/10.1128/AEM.66.11.5066-5072.2000
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M, Gormley N, Gilbert JA, Smith G, Knight R (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624. https://doi.org/10.1038/ismej.2012.8
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Nobu MK, Narihiro T, Rinke C, Kamagata Y, Tringe SG, Woyke T, Liu WT (2015) Microbial dark matter ecogenomics reveals complex synergistic networks in a methanogenic bioreactor. ISME J 9:1710–1722. https://doi.org/10.1038/ismej.2014.256
Xia Y, Wang Y, Wang Y, Chin FYL, Zhang T (2016) Cellular adhesiveness and cellulolytic capacity in Anaerolineae revealed by omics-based genome interpretation. Biotechnol Biofuels 9(111):111. https://doi.org/10.1186/s13068-016-0524-z
Wang G, Li Q, Gao X, Wang XC (2018) Synergetic promotion of syntrophic methane production from anaerobic digestion of complex organic wastes by biochar: performance and associated mechanisms. Bioresour Technol 250:812–820. https://doi.org/10.1016/j.biortech.2017.12.004
Ben AGZ, Thioye A, Cayol JL, Joseph M, Fauque G, Labat M (2018) Characterization of Desulfovibrio salinus sp. nov., a slightly halophilic sulfate-reducing bacterium isolated from a saline lake in Tunisia. Int J Syst Evol Microbiol 68:715–720. https://doi.org/10.1099/ijsem.0.002567
Daisuke S, Atsuko U, Toshiko S, Yoshimi O, Katsuji U (2010) Desulfovibrio butyratiphilus sp. nov., a gram-negative, butyrate-oxidizing, sulfate-reducing bacterium isolated from an anaerobic municipal sewage sludge digester. Int J Syst Evol Microbiol 60:595–602. https://doi.org/10.1099/ijs.0.013771-0
Na H, Lever MA, Kjeldsen KU, Schulz F, Jørgensen BB (2015) Uncultured Desulfobacteraceae and Crenarchaeotal group C3 incorporate (13) C-acetate in coastal marine sediment. Environ Microbiol Rep 7:614–622. https://doi.org/10.1111/1758-2229.12296
Grabowski A, Tindall BJ, Bardin V, Blanchet D, Jeanthon C (2005) Petrimonas sulfuriphila gen. Nov., sp. nov., a mesophilic fermentative bacterium isolated from a biodegraded oil reservoir. Int J Syst Evol Microbiol 55:1113–1121. https://doi.org/10.1099/ijs.0.63426-0
Imachi H, Sakai S, Kubota T, Miyazaki M, Saito Y, Takai K (2016) Sedimentibacter acidaminivorans sp. nov., an anaerobic, amino acids-utilizing bacterium isolated from marine subsurface sediment. Int J Syst Evol Microbiol 66:1293–1300. https://doi.org/10.1099/ijsem.0.000878
Obst M, Krug A, Luftmann H, Steinbüchel A (2005) Degradation of cyanophycin by Sedimentibacter hongkongensis strain KI and Citrobacter amalonaticus strain G isolated from an anaerobic bacterial consortium. Appl Environ Microbiol 71:3642–3652. https://doi.org/10.1128/AEM.71.7.3642-3652.2005
Hao LT, Zhang BG, Cheng M, Feng CP (2016) Effects of various organic carbon sources on simultaneous V(V) reduction and bioelectricity generation in single chamber microbial fuel cells. Bioresour Technol 201:105–110. https://doi.org/10.1016/j.biortech.2015.11.060
Lesnik KL, Liu H (2014) Establishing a core microbiome in acetate-fed microbial fuel cells. Appl Microbiol Biotechnol 98:4187–4196. https://doi.org/10.1007/s00253-013-5502-9
Regueiro L, Carballa M, Lema JM (2014) Outlining microbial community dynamics during temperature drop and subsequent recovery period in anaerobic co-digestion systems. J Biotechnol 192:179–186. https://doi.org/10.1016/j.jbiotec.2014.10.007
Dahle H, Birkeland N-K (2006) Thermovirga lienii gen. Nov., sp. nov., a novel moderately thermophilic, anaerobic, amino-acid-degrading bacterium isolated from a North Sea oil well. Int J Syst Evol Microbiol 56:1539–1545. https://doi.org/10.1099/ijs.0.63894-0
Honda T, Fujita T, Tonouchi A (2013) Aminivibrio pyruvatiphilus gen. Nov., sp. nov., an anaerobic, amino-acid-degrading bacterium from soil of a Japanese rice field. Int J Syst Evol Microbiol 63:3679–3686. https://doi.org/10.1099/ijs.0.052225-0
Meng X, Yuan X, Ren J, Wang X, Zhu W, Cui Z (2017) Methane production and characteristics of the microbial community in a two-stage fixed-bed anaerobic reactor using molasses. Bioresour Technol 241:1050–1059. https://doi.org/10.1016/j.biortech.2017.05.181
Ito T, Yoshiguchi K, Ariesyady HD, Okabe S (2011) Identification of a novel acetate-utilizing bacterium belonging to Synergistes group 4 in anaerobic digester sludge. ISME J 5:1844–1856. https://doi.org/10.1038/ismej.2011.59
Dassa B, Borovok I, Lamed R, Henrissat B, Coutinho P, Hemme CL, Yue H, Zhou J, Bayer EA (2012) Genome-wide analysis of Acetivibrio cellulolyticus provides a blueprint of an elaborate cellulosome system. BMC Genomics 13:210. https://doi.org/10.1186/1471-2164-13-210
Xu S, Han R, Zhang Y, He C, Liu H (2018) Differentiated stimulating effects of activated carbon on methanogenic degradation of acetate, propionate and butyrate. Waste Manag 76:394–403. https://doi.org/10.1016/j.wasman.2018.03.037
Lee SH, Park JH, Kang HJ, Lee YH, Lee TJ, Park HD (2013) Distribution and abundance of Spirochaetes in full-scale anaerobic digesters. Bioresour Technol 145:25–32. https://doi.org/10.1016/j.biortech.2013.02.070
Rivière D, Desvignes V, Pelletier E, Chaussonnerie S, Guermazi S, Weissenbach J, Li T, Camacho P, Sghir A (2009) Towards the definition of a core of microorganisms involved in anaerobic digestion of sludge. ISME J 3:700–714. https://doi.org/10.1038/ismej.2009.2
Wang HZ, Gou M, Yi Y, Xia ZY, Tang YQ (2018) Identification of novel potential acetate-oxidizing bacteria in an acetate-fed methanogenic chemostat based on DNA stable isotope probing. J Gen Appl Microbiol 64:221–231. https://doi.org/10.2323/jgam.2017.12.006
Hao L, Fan L, Mazéas L, Quéméner DL, Madigou C, Guenne A, Shao L, Bouchez T, He P (2015) Stable isotope probing of acetate fed anaerobic batch incubations shows a partial resistance of acetoclastic methanogenesis catalyzed by Methanosarcina to sudden increase of ammonia level. Water Res 69:90–99. https://doi.org/10.1016/j.watres.2014.11.010
Neubeck A, Sjöberg S, Price A, Callac N, Schnürer A (2016) Effect of nickel levels on hydrogen partial pressure and methane production in methanogens. PLoS One 11:e0168357. https://doi.org/10.1371/journal.pone.0168357
Zinder SH (1993) Physiological ecology of methanogens. Springer, US
Worm P, Koehorst JJ, Visser M, Sedanonúñez VT, Schaap PJ, Plugge CM, Sousa DZ, Stams AJ (2014) A genomic view on syntrophic versus non-syntrophic lifestyle in anaerobic fatty acid degrading communities. Biochim Biophys Acta 1837:2004–2016. https://doi.org/10.1016/j.bbabio.2014.06.005
Funding
This study was funded by the Ministry of Science and Technology of China (No. 2016YFE0127700) and by the National Natural Science Foundation of China (No. 51678378; No. 31200068).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(DOCX 2147 kb)
Rights and permissions
About this article
Cite this article
Yi, Y., Wang, H., Chen, Y. et al. Identification of Novel Butyrate- and Acetate-Oxidizing Bacteria in Butyrate-Fed Mesophilic Anaerobic Chemostats by DNA-Based Stable Isotope Probing. Microb Ecol 79, 285–298 (2020). https://doi.org/10.1007/s00248-019-01400-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-019-01400-z