Abstract
Biological soil crust (biocrust) is a composite of mosses, lichens, and bacteria that performs many important soil system functions, including increasing soil stability, protecting against wind erosion, reducing nutrient loss, and mediating carbon and nitrogen fixation cycles. These cold desert and steppe ecosystems are expected to experience directional changes in both climate and disturbance. These include increased temperatures, precipitation phase changes, and increased disturbance from anthropogenic land use. In this study, we assessed how climate and grazing disturbance may affect the abundance and diversity of bacteria in biocrusts in cold steppe ecosystems located in southwestern Idaho, USA. To our knowledge, our study is the first to document how biocrust bacterial composition and diversity change along a cold steppe climatic gradient. Analyses based on 16S small subunit ribosomal RNA gene sequences identified the phylum Actinobacteria as the major bacterial component within study site biocrusts (relative abundance = 36–51%). The abundance of the phyla Actinobacteria and Firmicutes was higher at elevations experiencing cooler, wetter climates, while the abundance of Cyanobacteria, Proteobacteria, and Chloroflexi decreased. The abundance of the phyla Cyanobacteria and Proteobacteria showed no significant evidence of decline in grazed areas. Taken together, results from this study indicate that bacterial communities from rolling biocrusts found in cold steppe ecosystems are affected by climate regime and differ substantially from other cold desert ecosystems, resulting in potential differences in nutrient cycling and ecosystem dynamics.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Steppe ecosystems, characterized by cold desert and semi-arid conditions, are widespread in the Northern Hemisphere and are projected to experience increases in temperature and changes in amount and phase of precipitation [1]. Climate shifts are predicted to increase mean annual surface temperatures by about 3–5 °C by 2100 [1]. Potentially more important to these ecosystems is the projection of increased variability in precipitation and phase shift, with change in averages being season-specific [2–4]. For high latitude regions of western North America, for example, recent models predict increases in winter precipitation, accompanied by decreases in summer precipitation [5]. Along with warming trends, changes in phase of precipitation will likely cause shifts from snow-dominated to rain-dominated winters [6].
Biological soil crusts, or biocrusts, are complex microbial communities of mosses, lichens, algae, fungi, and bacteria that commonly colonize the soil surface of cold desert and steppe ecosystems. Biocrusts range in structure from flat to rolling to pinnacled in morphology depending on soil texture and climatic conditions such as the amount of precipitation and frost heaving, and can vary in color from white to black hues depending on successional stage [7]. The majority of research has focused on the structure and function of rugose and pinnacled biocrusts associated with cold desert ecosystems, where mean annual precipitation ranges from 15 to 26 cm [8]; less is known about the composition and bacterial diversity associated with rolling biocrusts, typically found in cold steppe deserts, where precipitation is 50–100% of potential evapotranspiration (see Belnap 2003, Fig. 5 for biocrust morphologies) [7]. Rugose and pinnacled biocrusts in unproductive cold desert ecosystems have been shown to increase soil stability and moisture holding capacity [9]. Some bacteria found within these biocrust communities can produce polysaccharides that hold soil particles together, providing physical structure, which reduces wind erosion of soil [9] and nutrient loss by water runoff [10]. Additionally, biocrust communities have been shown to be important mediators of biogeochemical cycling processes like carbon (C) and nitrogen (N2) fixation, which increase soil fertility [9, 11], and help in the germination and survival of vascular plants [7, 9].
Changes in climate are occurring in cold steppe ecosystems, and these changes are likely to alter the bacterial communities associated with biocrusts, as biocrust morphology is influenced by precipitation and frost heaving. Thus, altered precipitation and temperature regimes may result in changes over time in community structure and composition. Studies monitoring long-term patterns in climate at the Reynolds Creek Experimental Watershed (now the Reynolds Creek Critical Zone Observatory (RC CZO)) in the Intermountain West Region of the US have reported directional trends in climate [12]. Specifically, minimum temperatures have increased and the proportion of precipitation due to snow had decreased, particularly at mid-elevations [12].
In addition to climate changes, land-use changes are also occurring in cold-desert and steppe ecosystems worldwide, resulting in increased disturbance by animal grazing, agriculture, fire, and recreational activities [9]. These activities often physically disturb the soil surface and biocrust structure. Reductions in biocrust cover from disturbance can lead to lowered water retention, decreased soil fertility, and increased erosion with up to 35 times more dust production [8, 11]. Pinnacled biocrust communities have been shown to be susceptible to a variety of anthropogenic disturbances, with recovery potentially taking over 1000 years depending on the severity of disturbance experienced, soil characteristics, and climate [11, 13]. Disturbance has also been found to alter the community composition of pinnacled biocrusts, with particularly negative effects to Cyanobacteria species [9]. Little is known, however, about the potential responses of rolling biocrusts to climate change and disturbance.
In this study, we examined the variation in composition and diversity of bacterial communities in rolling biocrusts in cold steppe ecosystems with respect to climate (elevation gradient) and disturbance (grazing). We hypothesized that altered relative bacterial abundances and community composition in biocrusts would occur along the climatic gradient due to changes in temperature and precipitation. Further, we hypothesized that disturbance form grazing would alter the bacterial abundance and community composition of biocrusts, and predicted that bacterial phyla sensitive to disturbance, such as Cyanobacteria, would be replaced with higher relative abundances of more disturbance-tolerant types of bacteria. Thus, we posit that biocrust community composition can be used as a potential bioindicator of both climate change and the severity of impacts from land use.
Materials and Methods
Study Area and Experimental Design
All biocrust samples were collected from the Reynolds Creek Critical Zone Observatory, a large (239 mile2) watershed located in the Owyhee Range in southwestern Idaho, USA. Aside from watershed research, the area is primarily managed for cattle grazing by the United States Department of Agriculture Agricultural Research Service (USDA-ARS). Vegetation within the CZO is common to cold steppe ecosystems of the Intermountain West region. Four sites along a 1000 m elevational gradient at the CZO were selected for use in this study. The four sites, increasing in elevation, are designated as Flats (F), Wyoming Big Sage (WBS), Low Sage (LS), and Mountain Big Sage (MBS).
Big sagebrush (Artemisia tridentata) is the dominant vegetation at all sites, though A. tridentata subspecies vary as a function of climate (Table 1). As elevation increases, mean annual precipitation (MAP) increases from 235 to 803 mm, and mean annual temperature (MAT) decreases from 9.1 to 5.4 °C (Table 1). Rain is the main form of annual precipitation at the lower elevation sites, whereas snow is the dominant form of annual precipitation at the highest elevation site. Variation in other factors such as topography and parent material was minimized by insuring that all sites were on relatively flat surfaces with a common (volcanic/basalt) parent material.
Biocrust and Soil Sampling
All biocrust and soil samples were collected in October 2014. For DNA analysis, interspace and shrub-canopy biocrust samples were collected from five randomly selected shrubs in both grazed and ungrazed (exclosure ˃40 year) areas at each of the elevational sites as described by Schwabedissen (SG Schwabedissen, 2016. M.S. Thesis, Idaho State University). The experimental layout reflected a replicated randomized block design, with samples obtained randomly from within both grazed and ungrazed levels, within each elevation (block). Shrub-interspace vs. shrub-canopy effects are not explicitly considered here. Each sample area was defined with a 10 × 10 cm metal frame, excised intact with a spatula and soil knife to a depth of 2.5 cm, and placed in an ethanol-sterilized storage container. All samples were transported and stored on ice until processing. A total of 5 × 2 shrub/interspace levels × 2 grazed/ungrazed levels × 4 elevation levels = 80 samples were obtained.
Soil Characterization
For geochemical analysis, soil was collected to a depth of 5 cm below each biocrust sample and transported on ice to the laboratory for processing. Soils were passed through a 2-mm sieve, and subsamples were used for determination of pH, electrical conductivity (EC), extractable inorganic nitrogen (N) as ammonium-N and nitrate-N, extractable soluble phosphate (ortho-P), and gravimetric water content (GWC). Subsamples of the biocrust were taken for determination of total C (TC) and total N (TN). Samples were sieved through a 2-mm sieve and then dried at 55 °C for 24–48 h before being placed in a ball mill grinder and ground for approximately 2 min until a fine powder was reached. Samples were packed into 5 × 9-mm tin capsules and sent to the ISU Center for Archeology, Materials, and Applied Spectroscopy (CAMAS) where the samples were analyzed on an Elemental Combustion System 4010 (Costech Analytical Tech, Inc., USA). Soil moisture was determined as percent gravimetric water content (% GWC) by weighing 10.0 ± 1 g of homogenized soil into an aluminum tin and drying at 105 °C for 24–48 h before weighing again to determine water loss. Sample pH and EC were determined using a 1:1 ratio of water to soil except at the highest elevation site, MBS, where a 2:1 ratio was needed due to high organics. After swirling the soil and water mixture for 1 min, samples rested for 1 h before measuring pH in triplicate on a Dual Channel pH/Ion/Conductivity Meter XL50 (Fisher Scientific, Pittsburg, PA).
Inorganic N extractions as ammonium (NH4 +) and nitrate (NO3 −) were conducted with a 1:5 ratio of soil to 2 M potassium chloride (KCl, CAS: 7447–40-7, Fisher Scientific, Pittsburg, PA). After addition of KCl, the samples were shaken for 1 h and then poured through Whatman #1 filters (CAT:1001–110, Fisher Scientific, Pittsburg, PA), pre-leeched with 2 M KCl. The inorganic N extracts were stored at 4 °C until analysis on the SmartChem 200 Discrete Analyzer Auto-Spectrophotometer (Westco Scientific Instruments, Inc., Brookfield, CT). A salicylate method (AMM-003-A) was used for NH4 + as N and a nitrate reduction method with a cadmium metal column (NO3–001-A) was used for NO3 − as N. Ortho-P extractions were performed using a 1:20 ratio of soil to 0.5 M sodium bicarbonate (NaHCO3, CAS: 144–55-8, Fisher Scientific, Pittsburg, PA). After addition of NaHCO3, the samples were shaken for 1 h and then poured through Whatman 40 filters (CAT: 1440–110, Fisher Scientific, Pittsburg, PA), pre-leeched with 0.5 M NaHCO3. The ortho-P extracts were stored at 4 °C until colorimetric analysis on the SmartChem 200 Discrete Analyzer Auto-Spectrophotometer using the PHO-001-A method and reported as ortho-P.
Microbial Community Structure
Biocrusts were sub-sampled for DNA extractions and bacterial community analysis from the original biocrust samples (80 samples) on the same day as field collection in October 2014. The samples were collected using an ethanol and flame-sterilized cork borer inserted 1 cm into each biocrust sample to collect a volume of approximately 1.75 mL, which was placed immediately into an autoclaved 2-mL tube and stored at −20 °C preceding extraction. DNA extraction was performed following the manufacturer’s protocol using the PowerSoil DNA Isolation Kit (CAT: 12,888–100, MoBio Laboratories, Inc., Carlsbad, CA). DNA was quantified using a NanoDrop ND-1000 Spectrophotometer at the Molecular Research Core Facility (MRCF) at Idaho State University.
The DNA extractions were pooled into 16 samples representing ungrazed and shrub-canopy, ungrazed and interplant space, grazed and shrub-canopy, and grazed and interplant space, within each of the four elevations. Thus, two independent DNA extractions were obtained for consideration of grazed and ungrazed effects within each elevation. We conducted PCR of the 16S SSU rRNA V3-V4 region to generate amplicons for next-generation sequencing on the MiSeq Illumina platform (Illumina San Diego, CA) using the primer sequences V3-Forward (5′-CCTACGGGNGGCWGCAG-3′) and V4-Reverse (5′-GACTACHVGGGTATCTAATCC-3′) [14] (Integrated DNA Technologies, Coralville, IA). The 50 μL reaction consisted of the primers (10 μM), Vent polymerase (CAT: M0258S, New England Biolabs, Ipswich, MA), 1× ThermoPol Buffer (New England Biolabs, Ipswich, MA), 400 μM deoxynucleotide triphosphates (CAT: BP2564–1, Fisher BioReagents, Fair Lawn, NJ), nuclease-free water, and 200 ng of pooled template DNA. Thermocycling was conducted using a PTC-200 gradient thermocycler (MJ Research, Waltham, Massachusetts). Amplification conditions were an initial incubation at 95 °C for 10 min, followed by a cycling regime (35 cycles total) of 95 °C for 1 min, 41.5 °C for 2 min, and 72 °C for 4 min, and followed by a final extension of 72 °C for 10 min and a hold at 4 °C until analysis on a 1% agarose gel. PCR amplicons were pair-end sequenced at the MRCF and the raw data obtained were uploaded to the Galaxy server [15–17]. Using Galaxy, the files were groomed, joined by paired ends, and converted to FASTA format before uploading to the program Metagenomics Rapid Annotation using Subsystems Technology (MG-RAST) for identification [18, 19]. Abundance values were normalized to the highest abundance for comparison between samples. Sequences have been submitted to the NCBI Sequence Repository. Shannon diversity index, richness, and evenness were calculated at both the phylum and genus taxonomic levels by site and by grazing disturbance.
Statistics
All analyses acknowledged the replicated randomized block design of the experiment. A significance level of α = 0.05 was used for all null hypothesis tests. To examine univariate site and grazing differences in phyla (e.g., evenness, richness, etc.), we conducted analysis of variance (ANOVA) and post hoc Tukey tests after transforming data, if necessary, to meet linear model assumptions. We graphically depicted multivariate relationships of site bacterial communities within species space using Nonmetric Multi-Dimensional Scaling (NMDS) [20]. Bray-Curtis (Steinhaus) distance was used in NMDS and other resemblance metric requiring multivariate procedures [21]. We formally tested for differences in biocrust bacterial communities using Permutational Multivariate Analysis of Variance (PERMANOVA) [22]. We used the R statistical environment (version 3.2.0; R core team [https://www.R-project.org]) for all statistical procedures, with heavy reliance on the “vegan” package [23].
Results
Soil Characteristics
Soil characteristics varied with climate (Table 2). Soil pH became more acidic as elevation increased from the lowest elevation, Flats (F), at 8.2 ± 0.54 (mean ± SD) to the highest elevation site, Mountain Big Sage (MBS), at 6.2 ± 0.32. Extractable ammonium (NH4 +) as N, soil moisture (GWC), total nitrogen (TN), and total carbon (TC) all increased with elevation. Electrical conductivity (EC), extractable nitrate (NO3 −), and extractable soluble phosphate (ortho-P) varied among elevations with no clear pattern (Table 2). The lowest elevation site, F, had the highest NO3 − at 2.9 ± 1.92 μg N g−1 dry soil−1. The middle elevation site, Wyoming Big Sage (WBS), had the highest EC at 212.7 ± 161.96 μS cm−1. Concentration of ortho-P was highest at the middle elevation site Low Sage (LS) at 10.1 ± 10.63 μg P g−1 dry soil−1 (Table 2). In areas of grazing, the soil characteristics of EC and NO3 − were consistently higher in samples from grazed areas than in samples from ungrazed exclosure areas (Table 3). Measurements of pH, NH4 +, ortho-P, GWC, TN, and TC did not demonstrate clear patterns of variation with grazing disturbance (Table 3).
Biocrust Community Variation
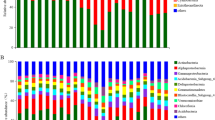
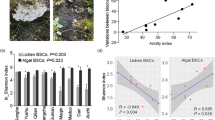
Actinobacteria was the most abundant phylum at each site, ranging from 35.85 to 51.07% of total bacterial abundance (Table 4). Other dominant phyla included Bacteroidetes, ranging in relative abundance from 6.24 to 16.71%, Proteobacteria, ranging from 9.46 to 11.77%, and Cyanobacteria, ranging from 0.12 to 10.58% (Table 4). Quantities of the two gram-positive bacterial phyla Actinobacteria and Firmicutes both increased with elevation, while Cyanobacteria, Chloroflexi, and Proteobacteria decreased. The abundance of the phyla Bacteroidetes and Verrucomicrobia was highest at the mid-elevation sites, WBS and LS. There were no clear patterns with elevation for the phyla Acidobacteria and Gemmatimonadetes. Unclassified bacteria made up about 21 to 26% of total bacteria sequences, and the minor phyla combined were <1% of the total bacterial sequences (Table 4). Minor phyla included Aquificae, Chlamydiae, Chlorobi, Deferribacteres, Deinococcus-Thermus, Dictyoglomi, Lentisphaerae, Nitrospirae, Planctomycetes, Poribacteria, Spirochaetes, Synergistetes, Tenericutes, and Thermotogae (Fig. 1).
a Average percent abundance of phyla at each elevation. Minor phyla of Aquificae, Chloroflexi, and Deinococcus-Thermus were also significantly different with elevation. b Average percent abundance of phyla by grazing disturbance at each elevation. Site names increasing in elevation are Flats (F), Wyoming Big Sage (WBS), Low Sage (LS), and Mountain Big Sage (MBS)
The NMDS projection based on bacteria phyla demonstrated strong community differences between the highest site, MBS, and the other three elevations, F, WBS and LS, with greatest variation along the first dimension (Fig. 2). Significant abiotic correlates of the NMDS projection were soil pH (R 2 = 0.549; P = 0.003), NH4 + (R 2 = 0.613; P = 0.004), GWC (R 2 = 0.531; P = 0.005), TC (R 2 = 0.596; P = 0.004), and TN (R 2 = 0.674; P = 0.002) (Fig. 2). A projection of weighted averages of phyla scores from the NMDS ordination of sites in community shown in Fig. 2 is given in Fig. 3. Figure 3 reveals that the first NMDS dimension was strongly affected by the phylum Poribacteria, a positive correlate of soil NH4. Variation along the second dimension is primarily due to the phyla Chlorobi, Tenericutes, and Dictyoglomi, which were positively correlated to ortho-P and EC, and the phylum Cyanobacteria which was positively correlated with NO3 and soil alkalinity. Bacteria within the phylum Poribacteria were more abundant at the highest elevation site, MBS, while those belonging to the phylum Cyanobacteria were more abundant at the lowest elevation site, F (Figs. 2 and 3).
A projection of weighted averages of phyla scores (generated using the vegan function wascores) from the NMDS ordination of sites in phyla space shown in Fig. 2.Vector fits of the environmental soil characteristics are overlaid. Site names increasing in elevation are Flats (F), Wyoming Big Sage (WBS), Low Sage (LS), and Mountain Big Sage (MBS)
Biocrust Communities with Elevation
PERMANOVA demonstrated strong integrated phyla (community) differences with elevation (P < 0.001). Further, a large number of individual phyla varied in abundance by elevation (P ≤ 0.05) including Acidobacteria, Actinobacteria, Aquificae, Bacteroidetes, Cyanobacteria, Deinococcus-Thermus, and Firmicutes (Table 6). Respecting Actinobacteria abundance, the three lower elevations were not significantly different from each other but were significantly different from the highest elevation site, MBS after adjustment for simultaneous inference using Tukey’s method (Table 6). The abundance of Cyanobacteria was significantly different between each of the F, LS, and MBS sites (Table 6). For both the Acidobacteria and Firmicutes phyla, F and MBS were significantly different from one another, while neither was significantly different from either of the middle elevation sites WBS and LS (Table 6).
When calculated at the phylum level, Shannon diversity and evenness were both significantly higher in the F, WBS, and LS sites, compared to the highest elevation site, MBS (Table 5a). At the genus level, however, Shannon diversity, richness, and evenness were statistically indistinguishable (Table 5a).
Biocrust Communities with Grazing
We did not detect differences in overall phyla-level composition with grazing using PERMANOVA (P = 0.393). Nonetheless, several individual phyla, including Proteobacteria (P = 0.098), Cyanobacteria (P = 0.079), and particularly Acidobacteria (P = 0.0095) showed evidence of differences with grazing disturbance across all four elevations (Fig. 1, Table 6). The relative abundance of the phylum Acidobacteria was 0.25 to 0.63% higher in grazed areas than ungrazed (Table 4). Similarly, the relative abundance of Proteobacteria was 1.27 to 2.88% higher in grazing samples (Table 4). Grazed areas at all elevations had lower levels of Cyanobacteria, with a decline of −0.03% at MBS to −11.2% at F (Table 4). Measures of alpha diversity, richness, and evenness did not vary significantly with grazing at either the phylum or genus levels (Table 5b).
Discussion
Biocrust Community Variation
The results from our study clearly support our hypothesis that the composition of bacterial communities in biocrusts varies with climate (elevation), and to a lesser degree with disturbance (grazing). The biocrusts located in RC CZO were predominantly composed of bacterial species from the phylum Actinobacteria. These findings were in stark contrast to several studies performed on pinnacled and rugose biocrusts located in the cold desert of the Colorado Plateau, USA [8, 9]. In accordance with our work, these studies also sampled biocrusts to a depth of 2.5 cm for DNA extraction; yet, these authors found that Cyanobacteria comprised the majority (32–59%) of the biocrust bacterial communities [9]. This difference may be a result of climatic differences in the Colorado Plateau, a cold desert that typically receives <25 cm precipitation [8], compared to our study area, a cold steppe desert that receives between 25 to 50 cm of precipitation depending on evapotranspiration (Table 1). More generally speaking, these differences also explain biocrust morphology shifts from pinnacled to rolling crusts, which have each been found to host different microbial communities [24]. These findings are supported in part by a study [25] which employed terminal restriction fragment length polymorphism (TRFLP) and reported the phylum Actinobacteria as the dominant bacterial phylum in soils (0–5 cm) in a hot desert in Israel along a precipitation gradient from 100 to 400 mm [25]. Another study on Northern Asia biocrusts reported the phylum Proteobacteria dominant (25–33%), while Cyanobacteria represented 9–24% and Actinobacteria only 7–9% of all phyla present [26]. Not only did the phylum Actinobacteria dominate the biocrust bacterial community within our rolling biocrusts, it was a strong correlate of elevation. Similarly, the Firmicutes lineage, which comprised much less of the overall abundance in the samples (2–7%), also showed a significant increase in relative abundance with elevation. Phyla including Cyanobacteria, Proteobacteria, and Chloroflexi decreased in relative abundance with elevation.
There remains a debate as to which factors are important for determining biocrust community composition and diversity. Biocrust studies performed in southern Africa [27] suggest that rainfall frequency is the best predictor of both biocrust composition and presence. However, a study in Israel [25] concluded that the bacterial diversity was independent of a precipitation gradient. Most studies agree, however, that soil pH (a correlate of soil water), along with precipitation and temperature regimes, have a strong influence on biocrust presence and the bacterial community composition and diversity. Our results support the premise that climate is the primary state factor controlling biocrust bacterial communities.
Overall, our findings indicate that biocrust diversity at the phylum level was quite similar within the range of 235 to 345 mm precipitation but dramatically declined with higher precipitation (>800 mm) and lower temperatures. The three lower elevation sites, F, WBS, and LS, have similar Shannon diversity indices and group together in the NMDS ordination, indicating community overlap (Fig. 2). Lower diversity and community distinctiveness at the MBS site was associated with increases in Actinobacteria and drastic decreases of Cyanobacteria observed in both grazed and ungrazed areas. Thus, as noted above, we posit that community differences are largely due to elevational effects on climate (lower temperatures and higher precipitation) and potentially concomitant edaphic factors (e.g., lower pH and higher NH4 +), rather than grazing disturbance. Due to the large increase in precipitation from LS to MBS, future studies at the RC CZO would benefit from sampling at higher spatial densities to examine changes in biocrust community structure and function.
Biocrust Community Variation with Grazing
Our work supported our hypothesis that grazing will alter some components of bacterial communities composition. Disturbance was correlated with a decline in the relative abundance of the Cyanobacteria phylum (P = 0.079). Lack of statistical significance was undoubtedly largely due to obfuscation from strong elevational effects. Conversely, the relative abundance of Proteobacteria (P = 0.098) and particularly Acidobacteria (P = 0.0095) was higher in grazed areas. These findings suggest that as bacteria from the phylum Cyanobacteria are lost from grazing, they may be replaced with bacteria from the Acidobacteria and Proteobacteria. Alternatively, the phyla Acidobacteria and Proteobacteria may be introduced from the microbial communities associated with the feces of grazing animals thereby increasing their abundance. The reduction in the quantity of Cyanobacteria species with disturbance is consistent with findings that show that chronic disturbance may alter biocrust communities [8], with particularly negative effects to Cyanobacteria [8, 9]. Previous work has also demonstrated that trampling may result in the increased presence of Acidobacteria and Proteobacteria [8, 9]. Acidobacterial communities, particularly those in subgroup Gp6, have previously been shown to thrive in grassland areas associated with heavy grazing [28].
Conclusions
Our study shows that the community composition of rolling biocrusts in steppe ecosystems varies strongly with climatic (precipitation and temperature) and edaphic conditions, and that variation in community components may occur with land use changes. The presence and composition of biocrusts are linked to their ability to perform important functions in steppe ecosystems, such as increasing soil stability, aiding in establishment of vascular plants, protecting soil against wind erosion, reducing nutrient loss, and performing carbon and nitrogen fixation. This study is the first to examine the impacts of climate variation and grazing on bacterial diversity in rolling biocrusts utilizing a climatic gradient. Further studies monitoring changes in bacterial community composition of biocrusts may provide a means for predicting altered ecosystem stability, altered capacity to perform vital soil biogeochemical cycles, and other responses to climate and land use changes that are occurring in these ecosystems.
References
Stacker TF, Qin D, Plattner GK, Tignor M, Allen SK, Boschung J, Nauels A, Xia Y, Bex V, Midgley PM (eds) (2013) The physical science basis. Contribution of working group I to the fifth assessment report of the Intergovernmental Panel on Climate Change. Cambridge, United Kingdom and New York, USA
Dettinger MD, Cayan DR, Diaz HF, Meko DM (1998) North-south precipitation patterns in western North America on interannual-to-decadal timescales. J. Clim. 11:3095–3111. doi:10.1175/1520-0442(1998)011<3095:NSPPIW>2.0.CO;2
Groisman PY, Karl TR, Easterling DR, Knight RW, Jamason PF, Hennessy KJ, Suppiah R, Page CM, Wibig J, Fortuniak K, Razuvaev VN, Douglas A, Forland E, Zhai P (1999) Changes in the probability of heavy precipitation: important indicators of climate change. Clim. Chang. 42:243–283. doi:10.1023/A:1005432803188
Easterling DR, Karl TR, Gallo KP, Robinson DA, Trenberth KE, Dai A (2000) Observed climate variability and change of relevance to the biosphere. J. Geophys. Res. 105:20101–20114. doi:10.1029/2000JD900166
USGCRP, Karl TR, Melillo JM, Peterson TC (eds) (2009) Global climate change impacts in the United States. New York, USA
Klos PZ, Abatzoglou JT, Bean A, Blades J, Clark MA, Dodd M, Hall TE, Haruch A, Higuera PE, Holbrook JD, Jansen VS, Kemp K, Lankford A, Link TE, Magney T, Meddens AJH, Mitchell L, Moore B, Morgan P, Newingham BA, Niemeyer RJ, Soderquist B, Suazo AA, Vierling KT, Walden V, Walsh C (2015) Indicators of climate change in Idaho: an assessment framework for coupling biophysical change and social perception. Wea Climate Soc 7:238–254. doi:10.1175/WCAS-D-13-00070.1
Belnap J (2003) The world at your feet: desert biological soil crusts. Front. Ecol. Environ. 1:181–189. doi:10.2307/3868062
Steven B, Kuske CR, Gallegos-Graves LV, Reed SC, Belnap J (2015) Climate change and physical disturbance manipulations result in distinct biological soil crust communities. Appl. Environ. Microbiol. 81:7448–7459. doi:10.1128/AEM.01443-15
Kuske CR, Yeager CM, Johnson S, Ticknor LO, Belnap J (2012) Response and resilience of soil biocrust bacterial communities to chronic physical disturbance in arid shrublands. ISME J 6:886–897. doi:10.1038/ismej.2011.153
Barger NN, Harrick JE, Zee JV, Belnap J (2006) Impacts of biological soil crust disturbance and composition on C and N loss from water erosion. Biogeochemistry 77:247–263. doi:10.1007/s10533-005-1424-7
Eldridge DJ, Zaady E, Shachak M (2000) Infiltration through three contrasting biological soil crusts in patterned landscapes in the Negev, Isreal. Catena 40:323–326. doi:10.1016/S0341-8162(00)00082-5
Nayak A, Marks D, Chandler DG, Seyfried M (2010) Long-term snow, climate, and streamflow trends at the Reynolds Creek experimental watershed, Owyhee Mountains, Idaho, United States. Water Resour. Res. 46:W06519. doi:10.1029/2008WR007525
Belnap J (2002) Nitrogen fixation in biological soil crusts from southeast Utah, USA. Biol. Fertil. Soils 35:128–135. doi:10.1007/s00374-002-0452-x
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glockner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generations sequencing-based diversity studies. Nucleic Acids Res. 41:1. doi:10.1093/nar/gks808
Giardine B, Riemer C, Hardison RC, Burhans R, Elnitski L, Shah P, Zhang Y, Blankenberg D, Albert I, Taylor J, Miller W, Kent WJ, Nekrutenko A (2005) Galaxy: a platform for interactive large-scale genome analysis. Genome Res. 15:1451–1455. doi:10.1101/gr.4086505
Goecks J, Nekrutenko A, Taylor J, The Galaxy Team (2010) Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 11:R86. doi:10.1186/gb-2010-11-8-r86
Blankenberg D, Von Kuster G, Coraor N, Ananda G, Lazarus R, Mangan M, Nekrutenko A, Taylor J (2010) Galaxy: a web-based genome analysis tool for experimentalists. Curr Protoc Mol Biol Ch 19:Unit 19.10.1–21. doi: 10.1002/0471142727.mb1910s89
Meyer F, Paarmann D, D’Souza M, Olson R, Glass EM, Kubal M, Paczian T, Rodriguez A, Stevens R, Wilke A, Wilkening J, Edwards RA (2008) The metagenomics RAST server-a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinformatics 9:386. doi:10.1186/1471-2105-9-386
Wilke A, Bischof J, Harrison T, Brettin R, D’Souza M, Gerlach W, Matthews H, Paczian T, Wilkening J, Glass EM, Desai N, Meyer F (2015) A RESTful API for accessing microbial community data for MG-RAST. PLoS Comput. Biol. 11:1. doi:10.1371/journal.pcbi.1004008
Kruskal JB (1964) Nonmetric multidimensional scaling: a numerical method. Psychometrika 29:115–129. doi:10.1007/BF02289694
Bray JR, Curtis JT (1957) An ordination of the upland forest communities in southern Wisconsin. Ecol. Monogr. 27:325–349. doi:10.2307/1942268
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecology 26:32–46
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2015) vegan: Community Ecology Package. R package version 2.3–0. http://CRAN.R-project.org/package=vegan
Belnap J, Lange OL (eds) (2003) Biological soil crusts: structure, function, and management. Ecological Studies 150, Berlin, Germany
Bachar A, Al-Ashhab A, Soares MI, Sklarz MY, Angel R, Ungar ED, Gillor O (2010) Soil microbial abundance and diversity along a low precipitation gradient. Microb. Ecol. 60:435–461. doi:10.1007/s00248-010-9727-1
Kemmling A, Pfeiffer B, Daniel R, Hoppert M (2012) Bacterial diversity in biological soil crusts from extrazonal mountain dry steppes in northern Mongolia. Wiss Beitr Martin Luther Univ Halle Wittenberg 12:437–449
Budel B, Darienko T, Deutschewitz K, Dojani S, Friedl T, Mohr KI, Salisch M, Reisser W, Weber B (2009) Southern African biological soil crusts are ubiquitous and highly diverse in drylands, being restricted by rainfall frequency. Microb. Ecol. 57:229–247. doi:10.1007/s00248-008-9449-9
Naether A, Foesel BU, Naegele V, Wüst PK, Weinert J, Bonkowski M, Alt F, Oelmann Y, Polle A, Lohaus G, Gockel S (2012) Environmental factors affect acidobacterial communities below the subgroup level in grassland and forest soils. Appl. Environ. Microbiol. 78:7398–7406
Acknowledgments
This research was performed in collaboration with the United States Department of Agriculture Agricultural Research Service, and Northwest Watershed Research Center in Boise, Idaho, and the landowners within the Reynolds Creek Critical Zone Observatory. This work was supported by the National Science Foundation for Reynolds Creek Critical Zone Observatory Cooperative agreement under award #NSF EAR 1331872 and the Institutional Development Award (IDeA) from the National Institute of General Medical Sciences of the National Institutes of Health under grant #P20GM103408. Additional funding for sequencing was awarded to SG Schwabedissen by the Molecular Research Core Facility (MRCF) at Idaho State University. We appreciate the field assistance of E McCorkle, and sequencing help of E O’Leary-Jepsen, M Andrews, and L McDougall at MRCF.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Blay, E.S., Schwabedissen, S.G., Magnuson, T.S. et al. Variation in Biological Soil Crust Bacterial Abundance and Diversity as a Function of Climate in Cold Steppe Ecosystems in the Intermountain West, USA. Microb Ecol 74, 691–700 (2017). https://doi.org/10.1007/s00248-017-0981-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-017-0981-3