Abstract
Transcriptomic sequencing together with bioinformatic analyses and an automated annotation process led us to identify novel phytochelatin synthase (PCS) genes from two extremophilic green algae (Chlamydomonas acidophila and Dunaliella acidophila). These genes are of intermediate length compared to known PCS genes from eukaryotes and PCS-like genes from prokaryotes. A detailed phylogenetic analysis gives new insight into the complicated evolutionary history of PCS genes and provides evidence for multiple horizontal gene transfer events from bacteria to eukaryotes within the gene family. A separate subgroup containing PCS-like genes within the PCS gene family is not supported since the PCS genes are monophyletic only when the PCS-like genes are included. The presence and functionality of the novel genes in the organisms were verified by genomic sequencing and qRT-PCR. Furthermore, the novel PCS gene in Chlamydomonas acidophila showed very strong induction by cadmium. Cloning and expression of the gene in Escherichia coli clearly improves its cadmium resistance. The gene in Dunaliella was not induced, most likely due to gene duplication.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Contamination of soil and water by toxic heavy metals poses a major environmental hazard due to their toxicity and accumulation behavior [1]. Thus, metals are among the most studied polluting agents. Heavy metals are usually toxic because they displace endogenous metal cofactors from their cellular binding sites, undergo aberrant capping reactions with the thiol groups of proteins and thiol-containing coenzymes, and promote the formation of active oxygen species [2]. Of the various detoxification pathways activated in plants under heavy metal stress, increased synthesis of sulfur containing defense compounds, namely cysteine, glutathione (GSH), and phytochelatins (PCs), is considered to be of prime importance for their tolerance and survival [3, 4]. In addition to a variety of plants, their synthesis is also increased in algae and fungi [4, 5] upon metal exposure.
Phytochelatins (PCs) mediate high-affinity binding and contribute to detoxification of heavy metal ions and metalloids, such as cadmium or arsenic, by promoting the vacuolysosomal sequestration of heavy metals [6]. These compounds are enzymatically synthesized from reduced glutathione (GSH) and related thiols in a γ-glutamyl-cysteinyltranspeptidation reaction catalyzed by phytochelatin synthase (PCS). Genes encoding phytochelatin synthases have been found in all vascular plants as well as some algae, fungi, diatoms and invertebrates [4, 7–9]. However, studies regarding PCS gene expression levels have only been examined in a few vascular plants and, in many cases, the results have been contradictory [10–12]. In addition to the eukaryotic full-length phytochelatin synthase genes, PCS-like or half-size PCS proteins were detected initially in prokaryotes (reviewed by Rea et al. [2]). These proteins were described to have a similar N-terminal but lack the more sequence-variable C-terminal, being therefore shorter (220–237 compared to 421–506 amino acid residues). Since then, PCS-like genes have also been reported in some eukaryotes (e.g., fungi: Saccharomyces pombe, Phytophtora infestans, Dictyostelium discoideum; heterokonta: Tetrahymena thermophila; metazoan: Caenorhabditis elegans, Hydra magnipapillata, Strongylocentrotus purpuratus) with scattered distribution but the functionality of these transcripts has largely remained untested [13, 14].
Most studies concerning heavy metals and microorganism physiology are related to environments polluted by industrial and domestic wastes. Only little is known about the toxicity of these substances in environments with a natural, non-anthropogenic source of high levels of heavy metals, such as extreme acidic ecosystems. Acidic environments tend to contain unusually high concentrations of heavy metals, because their solubility increases markedly as the pH decreases [15]. Despite these extreme environmental conditions, a large number of prokaryotic and eukaryotic organisms have been identified to live in the presence of high concentrations of heavy metals [16, 17]. Acidophiles can survive in ≤1000-fold higher heavy metal amounts than neutrophilic microorganisms [18]. These highly metal-resistant organisms seem to have multiple and/or more efficient active resistance systems than neutrophiles [19]. Modern biotechnological methods like high-throughput sequencing of non-model organisms that live in extreme habitats can help to understand the metabolism that allows these organisms to live in their environment and also enables the discovery of novel chemical components.
In a recent study on genes affected by copper excess in a strain of the microalga Chlamydomonas acidophila isolated from Río Tinto (one of the largest extreme acidic environments described until now) and transcripts with a putative function related to stress and heavy metal detoxification were identified from a de novo transcriptomic assembly [20]. Several of these transcripts got a best blast hit in other organisms than green algae. One of these, a transcript annotated as a glutathione gamma-glutamyl-cysteinyl-transferase (synonymous to PCS), was hypothesized to represent a novel gene absent in other green algae. A putative PCS-like transcript was later also found in another microalga, in a Dunaliella acidophila strain isolated from the same environment [21]. The present study aims to (1) characterize these novel PCS genes by additional sequencing to recover the complete genes; (2) analyze their protein domains and transmembrane topology; and (3) study their functionality and induction under cadmium exposure. Cadmium was selected because its immense water solubility determines its wide distribution in aquatic systems, and its relatively high mobility in comparison to other metals, makes it a metal of major concern in environmental contamination [22–24]. A phylogenetic analysis was performed in order to uncover relationships between PCS genes in plants, bacteria, and green algae and to interpret their evolutionary history.
Material and Methods
Algae Cultivation and Exposure to Cadmium
Chlamydomonas acidophila strain RT46 was collected from water samples taken in 2010 at the CEM station of Río Tinto (SW Spain) [17] and isolated to grow in the presence of antibiotics, vancomycin 50 μg/mL, cefotaxime 100 μg/mL, and chloramphenicol 15 μg/mL (Sigma Aldrich, MO, US), on agar plates made with 0.22 μM-filtered river water. Clonal cultures were grown in 50-ml flasks in K medium [25] at pH 2 under an illumination of ca. 70 microE s-1 m-2 irradiance provided by day-light fluorescent tubes, 16:8 h LD cycle at 20 °C.
Dunaliella acidophila RT5 was also collected in 2010 at RICHI sampling station and grown in ARTII medium (developed in our laboratory, see Online Resource 1) at pH 2. When the cultivations reached stationary stage, the cultures were exposed to a Cd solution CdCl2 2 ½ H2O (Panreac Química, Barcelona, Spain). Cultures were maintained during 1, 3, and 24 h with a final concentration of 1 μM. Cells were collected in 50-ml Falcon tubes and centrifuged for 5 min at 5000 rpm. Pellets were frozen with liquid nitrogen and stored at −80 °C until RNA extraction.
Nucleic Acid Extraction
Total RNA was extracted with TRI Reagent® Solution (Thermo Fisher Scientific, MA, US) following manufacturer’s protocol. To remove possible DNA contamination, the samples were treated with DNase I (RNase free) (Thermo Fisher Scientific). DNA was extracted from cultures not exposed to cadmium using the Wizard® Genomic DNA Purification Kit (Promega Corporation, WI, US) or PurelinkTM Genomic DNA Mini Kit (Thermo Fisher Scientific). DNA and RNA quality were determined spectrometrically (A260/A280 > 1.8) using a NanoDrop ND-1000 spectrometer (NanoDrop, DE, US).
Primer Design
Primers for amplification of the full-length genes (CaPCS1, CaPCS2, and DaPCS1) as well as qRT-PCR, and cloning and expression of CaPCS2 were designed using the program Primer3 (http://frodo.wi.mit.edu/cgi-bin/primer3/primer3_www.cgi) with default settings. Due to high variability in the PCS transcripts, species specific primers were needed. In quantitative reverse transcription, PCR (qRT-PCR) 18S rRNA [26] and actin were both used as housekeeping genes to normalize gene expression. All primers used in this study together with optimal annealing temperatures are listed in Table 1.
PCR Amplification, Cloning, and Sequencing
PCR amplifications (Bio-Rad Tetrad 2, Bio-Rad, CA, US) of DNA were performed in 25 μl-reactions following the recommendation for GoTaq® DNA Polymerase (Promega Corporation). PCR settings were as follows: 2 min 95 °C, 35 cycles (1 min 95 °C, 1 min primer specific annealing temperature, 1 min 72 °C), 10 min 72 °C. The number of cycles to 40 was frequently used for obtaining stronger products. The PCR products were separated on a 1 % agarose gel and visualized with ethidium bromide using 50 bp DNA ladder (Thermo Fisher Scientific) for size determination. The amplification specificity and efficiency were tested, and one primer pair was discarded due to unspecific amplification. The amplified products for each primer pair were cloned with the TOPO TA Cloning kit (Thermo Fisher Scientific) and sequenced on an Applied Biosystems 3730xl DNA Analyzer (Thermo Fisher Scientific) using M13pUC primers and BigDye Terminator v3.1 Cycle Sequencing Kit (Thermo Fisher Scientific). The sequences were edited manually with PhyDE® v1.0 [27] and primer sequences eliminated before submission to GenBank (accession numbers KT348512-KT348515).
Occasional non-specific amplification of the target region in Dunaliella acidophila was detected. Strong amplification of the non-specific products was detected using a gradient annealing temperature program (1 min 95 °C, 40 cycles (1 min 95 °C, 1 min 53 °C–57 °C, 1 min 72 °C, 10 min 72 °C).
Quantitative Reverse Transcription PCR (qRT-PCR)
For qRT-PCR, protocols established by Díaz et al. [28] were followed, except that the annealing temperature was 55 °C and actin (ACT1) was used in addition to 18S as endogenous control gene. ACT1 primers were modified from the Tetrahymena gene TTHERM_00190950 available on the Tetrahymena Genome Database (TGD) [29]. The cDNA synthesis was carried out using 3 μg RNA in 20 μl-reaction, according to the protocol supplied by first Strand cDNA Synthesis kit (AMV, Roche, Basel, Switzerland). All qRT-PCR reactions were carried out in an iQTM5 multicolor Real-Time PCR detection System (Bio-Rad) apparatus with the following cycling conditions (1) 5 min at 95 °C to denature reverse transcriptase, (2) 40 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C after 20 s. Both NTC (no template control) and RT minus control were negative. The real-time dissociation curve was used to check primer specificity and to confirm the presence of a unique PCR product. Standard curves were obtained using 10-fold serial cDNA dilutions and determining the Ct (cycle threshold) values. The standard line parameters (amplification efficiency, slope and correlation coefficient) are reported in Online Resource 2. Analysis of relative gene expression was carried out according to the Standard-curve quantification method [30] from at least four independent experiments (each performed in duplicates). Only CaPCS2 was amplified with qRT-PCR, since CaPCS1 was not in the focus of this study and reliable results would not be possible to obtain for DaPCS2 due to assumed gene duplication.
Cloning and Expression of the Novel PCS Gene (CaPCS2) from C. acidophila in E. coli
The CaPCS2 gene was amplified and cloned in the plasmid pdr111 using the following primers: FQUpHIN and FQLoNHE, which contain the restriction sites HindIII and NheI required for subsequent subcloning. Plasmid pdr111 was a gift from D. Rudner (Harvard Medical School) and contains the hyper-SPANK promoter, upstream of the cloning site, which is inducible by IPTG. Primer FQUpHIN contains a bacterial ribosomal binding sequence to promote the expression the CaPCS2 gene in E. coli. PCR reaction was carried out using cDNA as DNA template in 20 μl reaction containing: 1× PCR buffer 1 (Expand Long Template PCR System, Roche) with 17.5 mM MgCl2, 350 μM each dNTP, 300 nM each primer, 3.75 U (0.75 μl) Expand Long Template enzyme mix (Roche), and 1 μl of cDNA. Amplification was carried out with the following cycling conditions: 2 min 94 °C, 39 cycles (1 min 94 °C, 1.5 min 56 °C, 1.5 min 72 °C), 10 min 72 °C.
A unique PCR fragment of approximately 850 bp corresponding to the complete ORF of CaPCS2 gene was gel purified with the QIAquick extraction gel (Qiagen, Hilden, Germany), digested with the restriction endonucleases HindIII (Roche) and NheI (Roche), and ligated into pdr111 plasmid. Chemically competent Escherichia coli DH5α were transformed with pdr111 plasmid containing the cloned CaPCS2 gene. Bacterial plasmids were isolated with QIAprep Spin Miniprep kit (Qiagen). The growth medium for transformed bacteria was supplemented with 50 μg ml-1 ampicillin to prevent loss of the plasmid in E. coli.
Analysis of Cadmium Resistance in E. coli Clones
The cadmium resistance was analyzed by the minimum inhibitory concentration (MIC) using serial dilutions 0–10 of E. coli clones in LB solid medium (Luria Bertani medium, Laboratorios Conda, Madrid, Spain) and with growth curves by measured the optical density at 600 nm (OD600). Bacteria carrying the plasmid pdr111 without the gene CaPCS2 cloned were used as negative control. The MIC was determinate pipetting 4 μl of each dilution (0–10−4) onto LB solid medium containing 200–500 μM CdCl2 2 ½ H2O (Panreac Química). The cultures were maintained during 16–24 h at 37 °C. For the growth curves, OD600 was measured every 60 min by using a microplate reader (Tecan Genios, Mannedorf, Switzerland) at 37 °C for 50 cycles (49 h) as described in Mirete et al. [31]. At least three different clones in LB (non-treated) and LB supplemented with 300 or 500 μM CdCl2 (treated clones) were analyzed. The pdr111 vector contains a hyper-SPANK promoter, which makes it inducible by IPTG. For this reason, MIC determination and growth curves were carried out in the presence or in the absence of 1 mM IPTG.
Characterization of Novel Genes
To form a complete picture of the genes, both genomic and transcriptomic sequences were used. Transcripts were detected from de novo assemblies of C. acidophila [20] and D. acidophila [21] and in addition the corresponding sequences from another assembly for C. acidophila (Olsson et al. unpubl.) were retrieved. Sanger sequencing on genomic DNA was used to complement the information. A flowchart (Online Resource 3) outlines the process from culture to extraction to analyses and shows which assemblies are used in the current study.
Protein Characterization
The nucleotide sequences were translated to amino acids using ExPASy translation tool [32]. The novel predicted proteins were further characterized using the web-server version of HMMER [33] to identify protein domains and the Phobius web-server [34] to predict transmembrane topology and signal peptides.
Identification of Similar PCS Coding Sequences and Phylogenetic Analyses
The novel PCS transcripts were used as queries in BLASTx searches [35] against all protein sequences available in GenBank (nr database downloaded in April 2013), and the most similar sequences were downloaded. The number of species and different isoforms in closely related species of well represented groups like land plants were limited. Sequences were aligned with Mafft [36] using alignment option FFT-NS-I. The alignment was manually edited in PhyDE® v1.0 [27] by excluding ends of the alignments which could not be confidently aligned due to length differences and ambiguities in homology assessment, including the variable C-terminal domain. Bayesian analyses were performed with MrBayes v3.2.1 [37], applying search strategies suggested by [37, 38]. Calculations of the consensus tree and of the posterior probability of clades were performed based upon the trees sampled after the chains converged. Consensus topologies and support values from the different methodological approaches were compiled and drawn using TreeGraph2 [39].
Results and Discussion
Transcript Coding for Novel PCS Genes
A full-length PCS from Chlamydomonas acidophila was detected and named as CaPCS1. Transcripts annotated as putative phytochelatin synthases or phytochelatin synthase-like genes in C. acidophila and Dunaliella acidophila were identified as novel genes and named CaPCS2 and DaPCS, respectively. Both of them had best BLAST hits in bacteria, but there are a few eukaryotic sequences among the most similar sequences as well (Blast results are provided in Online Resource 4). The nucleotide sequences have substantial differences between them and only the translated amino acid sequences can be aligned. The genes are illustrated in Fig. 1. Additional information on the introns is represented in Online Resource 5.
Structures of the predicted genes a) DaPCS1, b) CaPCS1, and c) CaPCS2. Composition of exons (black, numbered E1, E2, etc.), untranslated regions (gray) and introns (white) of the three genes is given in bp and drawn to scale. Predicted isoforms are depicted below each gene. An (asterisk) indicates that only transcriptomic data is available for the UTR
Characterization of the Full-Length PCS CaPCS1 from Chlamydomonas acidophila
The transcript that was similar to known PCS from plants and green alga from C. acidophila, CaPCS1, is 425 amino acids long and contains four introns. Seven different isoforms of the gene were detected. In the predicted 3’UTR (238 bp), two isoforms show a 66 bp long gap, indicating alternative polyadenylation of UTR. Alternative polyadenylation of UTR (APA-UTR) is a widespread phenomenon in eukaryotic cells [40] including Chlamydomonas reinhardtii [41] but has not been documented in C. acidophila.
There are some differences between the transcripts obtained from the different assemblies used in this study. From the de novo assembly by Olsson et al. [20], two different transcripts were obtained of which one (KT348511) is almost identical with the genomic sequence obtained by Sanger sequencing, while the other KT447466 has a unique 58 bp long insertion. Also the sequences from the other transcriptome assembly (Olsson et al. unpubl. corresponding sequences presented in Online resource 6) show several differences, most importantly in the 5’UTR of two of the sequences (comp17619_c0_seq4 and comp17619_c0_seq6 depicted in Fig. 2. Due to these differences, it is hypothesized that two copies of the gene exist. The transcript with the longest open reading frame, which was congruent with the genomic sequence (KT348511), was used for the characterization of the gene.
A partial sequence alignment of the predicted 3’UTR in CaPCS1 from Chlamydomonas acidophila. The sequences from different assemblies (2227_isotig04503 from the assembly by Olsson et al. [20] and comp17619_c0_seq4 and comp17619_c0_seq6 from a Trinity assembly based on merged data from 454- and Illumina sequencing) show differences in both nucleotide composition and length. Clustal format uses special characters to indicate that all residues in the column are identical (asterisk), conserved (colon), or semi-conserved substitutions have been observed (period)
Characterization of CaPCS2 from Chlamydomonas acidophila
CaPCS2 consists of four exons and three introns, with a predicted open reading frame of 285 amino acid residues. Alternative polyadenylation of the UTR is present also in this gene; in the eight isoforms covering a 1288 bp long stretch of predicted 5’UTR regions common to all isoforms alter with regions present in only some of the isoforms (Online Resource 7).
A deletion of a nucleotide 235 bp after the beginning of the last exon was detected in the transcript assembled for this study compared to an existing one [20]. Since the deletion occurs in the coding sequence, it affects the open reading frame of the gene. The genomic sequence was congruent with the transcript from our previous study (GBAH01012416), which was therefore used for the characterization of the gene for this inconsistent part (depicted in Fig. 1).
Characterization of DaPCS from Dunaliella acidophila
Two isoforms of DaPCS1 (comp2951_c3_seq1 and comp2951_c3_seq2) were predicted in the de novo transcriptome assembly of Dunaliella acidophila [21] corresponding accession numbers KT348509–KT348510). One of them has five and the other eight introns (Fig. 1).
The nucleotide sequence of contig comp2951_c3_seq1 is longer than comp2951_c3_seq2 but there is an alternative stop codon TAA (instead of CAA coding alanine), which ends the open reading frame. The open reading frame of contig comp2951_c3_seq2 has 390 amino acid residues including a complete phytochelatin synthase domain (pfam 05023) of 293 amino acids as well as a signal peptide and inwards orientated transmembrane domain. Contig comp2951_c3_seq1 has a shorter open reading frame of only 238 amino acids with a truncated phytochelatin synthase domain and neither signal peptides nor transmembrane domains are detected.
Amplification of DaPCS yielded in addition to the expected amplification product unspecific amplification products, observed by gel electrophoresis. Two different products differing about 650 bp in size were cloned and sequenced (see “Material and methods” and Online Resource 8). The sequences are otherwise identical for the overlapping parts except for two differing nucleotides, suggesting gene duplication in this haploid organism. The sequence of the larger product is congruent with the other genomic and transcriptomic sequences and thus probably represents the prevalent gene copy.
Gene Expression of the Novel CaPCS2 Gene
Gene expression analysis of the CaPCS2 gene (Chlamydomonas acidophila) was carried out under cadmium treatment using qRT-PCR. The relative mRNA expression levels of target gene were normalized against the levels of actin and 18S rRNA. Statistical analysis for qRT-PCR fold induction indicates that there is a significant difference between the cadmium treatment and the control (Pair Wise Fixed Reallocation Randomisation test, p < 0.01). The gene showed a very strong induction (1275.3-fold induction, ±218.8 SD) at 3 h, which flatten out at 24 h (47.8-fold change, ±24.1 SD).
As discussed above, it is likely that DaPCS is duplicated, which explains the difficulties involved in the DNA amplification of this region. For this reason, qRT-PCR amplification was not performed on DaPCS. The focus of our study is on the novel PCS genes, and therefore, the function of the full-length CaPCS1 gene was not investigated in detail. Furthermore, different isoforms might be differently expressed thus introducing errors to the measurements. The results by Ramos et al. [10] showed that the regulatory mechanisms of PCS expression in plants is very complex including alternative splicing and gene duplication events.
Expression and Cadmium Resistance of CaPCS2 in E. coli Clones
The cadmium resistance analyzed by the minimum inhibitory concentration (MIC) as explained in “Material and methods” clearly shows that the expression of CaPCS2 gene in E. coli improves their resistance to Cd (Fig. 3). The effect is observed on LB solid medium with the highest tested concentration (500 μM) of Cd. In addition, the growth curves on liquid LB of the strain expressing CaPCS2 in the presence of Cd confirm this result. The effects of Cd in a liquid medium are different from solid medium, and the improved resistance to cadmium was better observed in LB broth with 300 μM added Cd (Fig. 4). In spite of the hyper-SPANK promoter in the used pdr111 plasmid, which controls the expression of the inserted gene CaPCS2 and is inducible by IPTG, addition of IPTG has no significant effect on the cell growth. This could be caused by the transcription of the gene from another constitutively promoter in the plasmid, which dims out the effect of IPTG.
a Drop assay using serial dilutions 0–10−4 of E. coli DH5 α negative control and E. coli DH5α with CaPCS2 gene cloned into pdr111 vector in LB platessupplemented with 50 μg ml−1 ampicillin. b E. coli DH5α negative control in LB with 500 μM Cd and 50 μg ml−1 ampicillin. c E. coli with CaPCS2 gene cloned into pdr111 vector in LB with 500 μM Cd and 50 μg ml−1 ampicillin
Phylogenetic Analyses of Phytochelatin Synthases
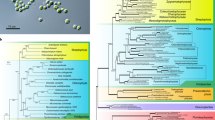
The results of the phylogenetic analyses show different topology from the expected one based on known species relationships. The only large clade reflecting phylogenetic relationships of organisms is the clade that contains known PCS genes from land plants and green algae. Even within this group, some species, such as lotus and Volvox carteri, have different variants or copies of the gene, which were not always monophyletic (Fig. 5). Indeed, many plant species are known to have at least two PCS genes [10, 13]. The PCS genes from other organisms do not form monophyletic groups congruent with species phylogenies, and many of the branches remain unresolved or with low support. Some smaller clades containing bacteria, metazoa, or fungi get statistical support, but none of them includes all the PCS genes of the group of organisms present in the analysis. Genes from cyanobacteria have a basal position in the phylogenetic tree and do not form a clade.
Phylogenetic relationships based on the novel PCS-like genes and the most similar genes obtained with a BLASTx search against NCBI’s databases. The tree represents the majority consensus of trees sampled after stationarity in the Bayesian analysis. Posterior probability values from the Bayesian inference equal or greater than 0.50 are shown above each branch. The tree is unrooted. The amino acid alignment was 383 aa long. The kingdoms of each clade are indicated. A miniature phylogram of the consensus tree is depicted to show the branch lengths
The full-length PCS (CaPCS1) present in Chlamydomonas acidophila is located in a clade containing PCS from other green algae. No similar transcripts from Dunaliella acidophila or other publicly available Dunaliella sequences were found. This could, however, be due to limited data available for the genus Dunaliella. The novel PCS transcripts (CaPCS2 from Chlamydomonas acidophila and DaPCS1 from Dunaliella acidophila) cluster together but are distinguished by long branches due to very different amino acid sequences between them (Fig. 5 and Online Resource 9).
The Scattered Distribution of PCS Genes in Nature
For a long time, researchers have been puzzled by the high constitutive expression of PCS genes in different plant tissues [2]. This is especially peculiar if we assume the presence of high heavy metal concentrations to be an exclusively recent phenomenon caused by human activity like mining. It might be, however, that high heavy metal concentration has been a much more severe problem for organisms early in the earth’s history and especially arsenic and cadmium comprised a major threat to incipient life forms [42, 43]. Indeed, all life forms have evolved strategies to cope with these heavy metals. The best-characterized metalloid resistances are encoded by the ars operons of Gram-negative and Gram-positive bacteria, which are non-specialized in the way that they share common organization and genes including repressors of resistance to Zn(II), Cd(II), and Pb(II) e.g., [44]. If life did emerge in waters rich in ionic metals, resistance to these compounds would probably be a very ancient trait, as suggested by Rosen [45]. Basal plants including mosses and other bryophyte lineages were for a long time supposed not to have PCS genes [13]. Recent results show the opposite, and the presence of constitutively expressed and functional phytochelatin synthases was demonstrated in all the bryophyte lineages by Petraglia et al. [46]. The presence of PCS genes in bryophytes was assayed by western blotting, in vitro activity and by measuring glutathione and phytochelatin contents, but the genes have not been sequenced and were therefore not included in our phylogenetic analysis. It is thus not clear if PCSs are key innovations to extreme habitats or ancestral traits and why they haven’t been downregulated or lost through evolution. One plausible explanation would be that PCS serve other physiological functions beyond cadmium or arsenic detoxification, which has been suggested by some authors [9, 14].
According to our results PCS genes probably have a bacterial origin and were subsequently inherited to different groups, in some cases multiple times. Bacterial origin is also supported by the domain structure and size of the PCS genes, the bacterial genes being smaller. It has been noted earlier that PCS genes have a non-monophyletic distribution across kingdoms [14] but a comprehensive phylogenetic analyses including PCS genes from both eukaryotes and prokaryotes has not been made earlier. It has been suggested earlier that there might be horizontal gene transfer events involved in the PCS genes in metazoa and fungi [14] and even among vascular plants [10]. Our results confirm that the evolutionary history of these genes is indeed hard to elucidate and show that prokaryotic genes have to be included to resolve their phylogeny. The complex evolution of the PCS genes involves several gene duplications and losses or independent insertions of the full-length PCS genes in plants and green algae.
The great differences in the amino acid composition and sequence length between the novel PCS genes DaPCS and CaPCS2 point to an ancient horizontal gene transfer event, although two independent events could also be possible. Horizontal gene transfer (HGT) from bacteria can be beneficial for eukaryotic organisms to survive in adverse conditions [47] and seems to be more common in algae than in vascular plants [48]. Recent studies have emphasized horizontally acquired key genes in adaptation to extreme and rapidly changing environments in other extremophilic algae [49, 50]. The novel PCS genes characterized in this study represent a likely example of genes that enable the organism to invade ecological niches otherwise unavailable, in this case an environment toxic for most other organisms.
Conclusions
Novel PCS genes from two extremophilic green algae are characterized in this study. The results of the phylogenetic analysis oppose the separation of prokaryotic PCS-like genes and suggest multiple horizontal gene transfer events from bacteria to eukaryotes within the PCS gene family. Further functional and structural analyses are needed to understand in detail the role of PCS both in heavy metal detoxification and other possible functions. The gene CaPCS2 increases cadmium tolerance in bacteria, but it remains to be seen if the novel genes are transferable to vascular plants since this could be of potential interest for crop science and phytoremediation.
References
Jain CK (2004) Metal fractionation study on bed sediments of river Yamuna, India. Water Res 38:569–578
Rea PA, Vatamaniuk OK, Rigden DJ (2004) Weeds, worms, and more. Papain’s Long-Lost Cousin, Phytochelatin Synthase. Plant Physiol 136:2463–2474
Rausch T, Wachter A (2005) Sulfur metabolism: a versatile platform for launching defence operations. Trends Plant Sci 10:503–509
Grill E, Winnacker E-L, Zenk MH (1987) Phytochelatins, a class of heavy-metal-binding peptides from plants, are functionally analogous to metallothioneins. Proc Natl Acad Sci USA 84:439–443
Mishra S, Srivastava S, Tripathi RD, Govindarajan R, Kuriakose SV, Prasad MN (2006) Phytochelatin synthesis and response of antioxidants during cadmium stress in Bacopamonnieri L. Plant Physiol Biochem 44:25–3
Cobbett CS (2000) Phytochelatin biosynthesis and function in heavy-metal detoxification. Curr Opin Plant Biol 3:211–216
Clemens S, Kim EJ, Neumann D, Schroeder JI (1999) Tolerance to toxic metals by a gene family of phytochelatin synthases from plants and yeast. EMBO J 18:3325–3333
Vivares D, Arnoux P, Pignol D (2005) A papain-like enzyme at work: native and acylenzyme intermediate structures in phytochelatin synthesis. Proc Natl Acad Sci U S A 102:18848–18853
Rea PA (2012) Phytochelatin synthase: of a protease a peptide polymerase made. Physiol Plant 145:154–164
Ramos J, Clemente MR, Naya L et al (2007) Phytochelatin synthases of the model legume Lotus japonicus. A small multigene family with differential response to cadmium and alternatively spliced variants. Plant Physiol 143:1110–1118
Liu Z, Gu C, Chen F, Yang D, Wu K, Chen S, Jiang J, Zhang Z (2012) Heterologous expression of a Nelumbo nucifera phytochelatin synthase gene enhances cadmium tolerance in Arabidopsis thaliana. Appl Biochem Biotechnol 166:722–734
Shukla D, Kesari R, Mishra S et al (2012) Expression of phytochelatin synthase from aquatic macrophyte Ceratophyllum demersum L. enhances cadmium and arsenic accumulation in tobacco. Plant Cell Rep 31:1687–1699
Clemens S (2006) Evolution and function of phytochelatin synthases. J Plant Physiol 163:319–332
Clemens S, Peršoh D (2009) Multi-tasking phytochelatin synthases. Plant Sci 177:266–271
Nordstrom DK, Alpers CN (1999) Negative pH, efflorescent mineralogy and consequences from environmental restoration at the Iron Mountain Superfund site, California. Proc Natl Acad Sci U S A 96:3455–3462
Nixdorf B, Wollmann K, Deneke R (1998) Ecological potentials for planktonic development and food web interactions in extremely acidic mining lakes in Lusatia. In: Geller J, Klapper H, Salomons W (eds) Acidic mining lakes. Springer, New York, pp 147–167
Aguilera A, Manrubia SC, Gómez F, Rodriguez N, Amils R (2006) Eukaryotic community distribution and their relationship with the water physicochemical parameters in an extreme acidic environment, Río Tinto (SW, Spain). Appl Environ Microbiol 72:5325–5330
Dopson M, Baker-Austin C, Koppineedi PR, Bond P (2003) Growth in sulfidic mineral environments: metal resistance mechanisms in acidophilic micro-organisms. Microbiol 149:1959–1970
Dopson M, Ossandon FJ, Lövgren L, Holmes D (2014) Metal resistance or tolerance? Acidophiles confront high metal loads via both abiotic and biotic mechanisms. Front Microbiol 5:1–4
Olsson S, Puente-Sánchez F, Gómez-Rodriguez M, Aguilera A (2015) Transcriptional response to copper excess and identification of genes involved in heavy metal tolerance in the extremophilic microalga Chlamydomonas acidophila. Extremophiles 19:657–672
Puente-Sánchez F, Olsson S, Aguilera A (2016) Comparative transcriptomic analysis of the response of Dunaliella acidophila (Chlorophyta) to short-term Cadmium and chronic natural metal-rich water exposures. Microb Ecol. doi:10.1007/s00248-016-0824-7
Guanzon NG, Nakahara H, Yoshida Y (1994) Inhibitory effects of heavy metals on growth and photosynthesis of three freshwater microalgae. Fish Sci 60:379–384
Worms IA, Boltzman J, Garcia M, Slaveykova VI (2012) Cell-wall-dependent effect of carboxyl-CdSe/ZnS quantum dots on lead and copper availability to green microalgae. Environ Pollut 167:27–33
de Abreu FC, da Costa PN, Brondi AM et al (2014) Effects of cadmium and copper biosorption on Chlorella vulgaris. Bull Environ Contam Toxicol 93:405–9
Keller MD, Selvin RC, Claus W, Guillard RRL (1987) Media for the culture of oceanic ultraphytoplankton. J Phycol 23:633–638
Fei X, Eriksson M, Li Y, Deng X (2010) A Novel Negative Fe-Deficiency-Responsive Element and a TGGCA-Type-Like FeRE Control the Expression of FTR1 in Chlamydomonas reinhardtii. J Biomed Biotechnol 2010:790247
Müller K, Quandt D, Müller J, Neinhuis C (2005) PhyDE ® 0.995: Phylogenetic Data Editor, http://www.phyde.de
Díaz S, Amaro F, Rico D et al (2007) Tetrahymena metallothioneins fall into two discrete subfamilies. PLoS ONE 2, e291
Stover NA, Krieger CJ, Binkley G et al (2005) Tetrahymena Genome Database (TGD): a new genomic resource for Tetrahymena thermophila research. Nucleic Acids Res 34:D500–D503
Larinov A, Krause A, Miller W (2005) Standard curve based method for relative real time PCR data processing. BMC Bioinform 6:62
Mirete S, Mora-Ruiz MR, Lamprecht-Grandío M, de Figueras CG, Rosselló-Móra R, González-Pastor JE (2015) Salt resistance genes revealed by functional metagenomics from brines and moderate-salinity rhizosphere within a hypersaline environment. Front Microbiol 6:1121
Artimo P, Jonnalagedda M, Arnold K et al (2012) ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res 40:W597–W603
Finn RD, Clements J, Eddy SR (2011) HMMER web server: interactive sequence similarity searching. Nucleic Acids Res 39:W29–W37
Käll L, Krogh A, Sonnhammer ELL (2007) Advantages of combined transmembrane topology and signal peptide prediction—the Phobius web server. Nucl Acids Res 35:W429–32
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: z new generation of protein database search programs. Nucl Acids Res 25:3389–3402
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucl Acids Res 30:3059–3066
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Huelsenbeck JP, Ronquist F, Nielsen R, Bollback JP (2001) Bayesian inference of phylogeny and its impact on evolutionary biology. Science 294:2310–2314
Stöver BC, Müller KF (2010) TreeGraph 2: Combining and visualizing evidence from different phylogenetic analyses. BMC Bioinform 11:7
Di Giammartino DC, Nishida K, Manley JL (2011) Mechanisms and consequences of alternative polyadenylation. Mol Cell 43:853–866
Shen Y, Liu Y, Liu L, Liang C, Li QQ (2008) Unique features of nuclear mRNA poly(A) signals and alternative polyadenylation in Chlamydomonas reinhardtii. Genetics 179:167–176
Oremland RS, Saltkov CW, Wolfe-Simon F, Stolz JF (2009) Arsenic in the evolution of earth and extraterrestrial ecosystems. Geomicrobiol J 26:522–536
Dani SU (2010) Gold, coal and oil. Med Hypotheses 74:534–541
Mendoza-Cózatl DG, Jobe TO, Hauser F, Schroeder JI (2011) Long-distance transport, vacuolar sequestration, tolerance, and transcriptional responses induced by cadmium and arsenic. Curr Opin Plant Biol 14:554–562
Rosen BP (1999) Families of arsenic transporters. Trends Microbiol 7:207–212
Petraglia A, De Benedicti M, Degola F et al (2014) The capability to synthesize phytochelatins and the presence of constitutive and functional phytochelatin synthases are ancestral (plesiomorphic) characters for basal land plants. J Exp Bot 65:1153–1163
Keeling PJ, Palmer JD (2008) Horizontal gene transfer in eukaryotic evolution. Nat Rev Genet 9:605–618
Richards TA, Soanes DM, Foster PG, Leonard G, Thornton CR, Tallbot NJ (2009) Phylogenomic analysis demonstrates a pattern of rare and ancient horizontal gene transfer between plants and fungi. Plant Cell 21:1897–1911
Blanc G, Agarkova I, Grimwood J, Kuo A, Brueggeman A, Dunigan DD et al (2012) The genome of the polar eukaryotic microalga Coccomyxa subellipsoidea reveals traits of cold adaptation. Genome Biol 13:R39
Schönknecht G, Chen WH, Ternes CM et al (2013) Gene transfer from bacteria and archaea facilitated evolution of an extremophilic eukaryote. Science 339:1207–10
Acknowledgments
This work was supported by the Spanish Ministry of Economy and Competitivity (MINECO) under Grant CGL-2011-22540 to AA. We acknowledge CSC-IT Center for Science Ltd. (Finland) for the allocation of computational resources. Dr. Salvador Mirete (Centro de Astrobiología CSIC-INTA) is acknowledged for assistance with the growth curves for the analysis of cadmium resistance in E. coli. FP-S was supported by a JAE-pre fellowship from the Spanish Consejo Superior de Investigaciones Científicas (CSIC).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1
ART II medium used for the cultivation of Dunaliella acidophila strain RT5. The volume was brought to 1 l with distilled water and the pH adjusted with 0.1 N H2SO4. (PDF 256 kb)
Online Resource 2
Quantitative reverse transcription PCR standard-curve parameters for target gene (novel phytochelatin synthase in Chlamydomonas acidophila CaPCS2) and the expression control (housekeeping) genes 18S rRNA and actin. E = amplification efficiency, S = slope, R2 = correlation coefficient. (PDF 393 kb)
Online Resource 3
Flow chart on the process from culture to extraction to analyses for the sequences used in the current study. (PDF 359 kb)
Online Resource 4
Results for BLASTx search using the open reading frame of transcripts coding for CaPCS2 (isotig12422) and DaPCS (comp2951_c3_seq1 and comp2951_c3_seq2) as query. (TXT 110 kb)
Online Resource 5
Introns in the predicted PCS genes from C. acidophila and D. acidophila. There are three spliceosomal introns in CaPCS2, four in CaPCS1 and three in DaPCS1. The introns are either the most common GT-AG splice site or variations of it. (PDF 19 kb)
Online Resource 6
Different contigs of component comp17619_c0 coding for the CaPCS1 gene in Chlamydomonas acidophila. (TXT 11 kb)
Online Resource 7
Different contigs of component comp11852_c0 coding for the CaPCS2 gene in Chlamydomonas acidophila. (TXT 18 kb)
Online Resource 8
PCR products amplified in Dunaliella acidophila strain RT5 using gradient annealing run on 1 % agarose gel (1X TBE). The gel electrophoresis shows amplification of products sized 1198 bp and 542 bp and several non-specific amplification products. The intensity of the non-specific bands depends on the used annealing temperature and PCR program. Wells 1: Ladder 1 kb (Promega Corporation), marker sizes from 250 kb to 10,000 kb, well 2: annealing 52-55 °C. Well 3: annealing 55-60 °C. (PDF 51 kb)
Online Resource 9
Phylogram based on the novel phytochelatin synthase genes and the most similar genes obtained with a BLASTx search against NCBI’s databases. The tree represents the majority consensus of trees sampled after stationarity in the Bayesian analysis. PP values from the Bayesian inference > 0.50 are indicated above each branch. The amino acid alignment was 383 aa long. (PDF 871 kb)
Rights and permissions
About this article
Cite this article
Olsson, S., Penacho, V., Puente-Sánchez, F. et al. Horizontal Gene Transfer of Phytochelatin Synthases from Bacteria to Extremophilic Green Algae. Microb Ecol 73, 50–60 (2017). https://doi.org/10.1007/s00248-016-0848-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-016-0848-z