Abstract
Geographic location and physicochemical properties are thought to represent major factors that shape soil bacterial community abundance and diversity. Crude oil contamination is becoming a notable concern with respect to soil property variation; however, the quantifiable influences of geographic location, physicochemical properties, and oil contamination are still poorly understood. In this study, the 16S ribosomal RNA genes of bacteria in the four oil fields in China were analyzed by using pyrosequencing. Results showed that physicochemical properties were the most dominant factor of bacterial community distribution, followed by geographical location. Oil contamination was a driving factor whose indirect influence was stronger than its direct influence. Under the impact of these three factors, different oil fields presented diversified and distinguishable bacterial community features. The soil of sites with the highest total petroleum hydrocarbon content (HB), nitrogen content (DQ), and phosphorus content (XJ) contained the largest proportion of functional groups participating in hydrocarbon degradation, nitrogen turnover, and phosphorus turnover, respectively. The first dominant phylum of the site with loam soil texture (HB) was Actinobacteria instead of Proteobacteria in other sites with sandy or sandy loam soil texture (DQ, SL, XJ). The site with the highest salinization and alkalization (SL) exhibited the largest proportion of unique local bacteria. The site that was located in the desert with extremely low precipitation (XJ) had the most diversified bacteria distribution. The bacterial community diversity was strongly influenced by soil physicochemical properties.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Bacteria are the most abundant organisms found in soils and support soil functions by nutrient cycling, organic material degradation, and other ecological services [1–3]. Such features as biodiversity and composition are greatly influenced by the location and physicochemical properties of the soil [4]. Numerous studies have investigated the relationship between bacterial communities and environmental factors [5–7]. Due to the high complexity and plasticity of bacterial community, different methods used to reveal bacterial diversity, frequency and magnitude of environmental disturbance, and limited knowledge on spatiotemporal scales of bacterial community in ecosystems, previous studies have shown inconsistent or even contradictory results [8]. It is necessary to clarify how environmental factors shape bacterial community, especially the one in contaminated setting.
As the demand of oil increases rapidly and certain toxic chemicals found in crude oil accumulate in ecosystems, oil contamination in soil is becoming one of the most severe global environmental problems [9–11]. When exogenous chemicals are released into soil, the environment for bacteria will be greatly changed, and the overall bacterial community structure will shift to adapt to the new habitat [12]. Therefore, many studies have been carried out to analyze bacterial communities in oil-contaminated fields [9–17]. However, most of the efforts to describe bacterial communities in oil-contaminated fields have been focused on phylogenetic composition. Few studies have quantifiably studied both of the influence of natural factors and human factors on its bacterial communities. Better understanding how indigenous bacterial communities are shaped by geographic location, soil physicochemical properties, and oil contamination is vital for soil remediation and the management of oil-contaminated fields.
Hence, this study aims to (1) clarify the bacterial community feature of oil-contaminated soil, (2) investigate the impact of oil contamination on soil physicochemical properties, and (3) quantify the influence of oil field geographic location, oil contamination, and physicochemical properties of soil on bacterial community using pyrosequencing. The understanding of bacterial diversity in oil-contaminated soil and its relationship with natural and anthropic environmental factors would be useful in soil bioremediation in response to oil contamination.
Material and Methods
Research Sites and Soil Sampling
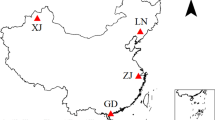
Four representative oil fields that produce approximately 50 % of the total crude oil of China were chosen as the research sites (Fig. S1). The Daqing (DQ) oil field (46° 35′ N, 125° 18′ E), in northeast China, is characterized by a temperate continental monsoon climate with a mean annual rainfall of 428 mm. The Huabei (HB) oil field (39° 10′ N, 116° 20′ E), in the Huabei Plain in northern China, is characterized by a temperate monsoon climate with a mean annual rainfall of 575 mm. The Shengli (SL) oil field (37° 28′ N, 118° 29′ E), in the Yellow River area in northern China, is characterized by a warm temperate continental semi-humid monsoon climate with a mean annual rainfall of 550 mm. The Karamay (XJ) oil field (45° 37′ N, 85° 01′ E), in western China, is characterized by a temperate continental arid climate with a mean annual rainfall of 109 mm. Field research was carried out in September to December 2012. Contaminated soils (DQ, HB, SL, XJ) were collected adjacent to crude oil pumping wells near surface (0–10 cm), while uncontaminated soils (DQ(BLK), HB(BLK), SL(BLK), XJ(BLK)) were collected 100 km from the wells. For each oil field, four subsamples of contaminated soil and four subsamples of uncontaminated soil were collected. Soils were sealed in sterile sampling bags, transported to the laboratory in an icebox, and stored in the dark at −20 °C until processing. Since all those fields are located in plain areas, no obvious soil variation was observed during field work. And further statistic analysis showed that there was no significant difference in soil physicochemical properties of subsamples for each oil field (p > 0.05). Thus, four subsamples for contaminated soil and uncontaminated soil were later respectively pooled prior to experiments to reduce the effects of fine scale spatial heterogeneity on bacterial community [18].
Geographical, Contaminant, and Physicochemical Indices
Latitude and longitude coordinates of all the samples were measured by the Global Positioning System (GPS) during field work. Precipitation data came from the China Meteorological Data Sharing Service System [19]. The concentration of total petroleum hydrocarbons (TPH) was determined using the Ultrasonic-Soxhlet extraction gravimetric method [20]. The four components of TPH (saturates, naphthene aromatics, polar aromatics, and asphaltenes) were separated using the Carhett method according to ASTM D4124 [21]. The following indices of soil physicochemical properties were measured according to the recommended soil testing procedures [22]: soil texture, density, water content (WC), pH, salinity, total nitrogen (TN), olsen-nitrogen (Olsen-N), total phosphorus (TP), olsen-phosphorus (Olsen-P), total potassium (TK), olsen-potassium (Olsen-K), and organic matter (OM) (Table 1).
DNA Extraction and PCR Amplification
Microbial DNA was extracted from eight samples from four research sites using a PowerSoil DNA Isolation kit (MO BIO Laboratories, Inc., West Carlsbad, CA) according to the manufacturers’ protocols. The V1–V3 region of the bacteria 16S ribosomal RNA gene was amplified by polymerase chain reaction (95 °C for 2 min, followed by 25 cycles at 95 °C for 30 s, 55 °C for 30 s, 72 °C for 30 s, and a final extension at 72 °C for 5 min) using primers 27 F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 533R (5′-TTACCGCGGCTGCTGGCAC-3′). PCR reactions were performed in a 20-μL mixture containing 4 μL of 5× FastPfu buffer, 2 μL of 2.5 mM dNTPs, 0.8 μL of each primer (5 μM), 0.4 μL of FastPfu polymerase, and 10 ng of template DNA. The DNAs of SL(BLK) and XJ(BLK) were not detected even after multiplying DNA extractions and pooling in a spin column to concentrate, and increasing PCR cycle. Low abundance of microbial community in these two samples might result from the high salinity of SL(BLK) and XJ(BLK) and the lack of water in XJ(BLK) (Table 1), since high salinities limit the microbial access to hydrocarbon and the availability of oxygen [23].
454 Pyrosequencing
After purification using the AxyPrep DNA Gel Extraction Kit (Axygen Biosciences, Union City, CA, USA) and quantification using QuantiFluor™ -ST (Promega, USA), a mixture of amplicons was used for pyrosequencing on a Roche 454 GS FLX + Titanium platform (Roche 454 Life Sciences, Branford, CT, USA) according to standard protocols [24]. The raw reads were deposited into the NCBI Sequence Read Archive (SRA) database (accession number SRP041085).
Bioinformatic Analysis
Pyrosequencing flowgrams were converted to sequence reads by using Mothur [25]. Trimmed sequences were produced by removing low-quality sequences (quality scores <25, sequence lengths <200 bp) and ambiguous reads (ambiguous base >0) using QIIME v. 1.3.0 [26]. Sequences were aligned against the Silva database (SSU111 version: http://www.arb-silva.de/) using k-mer searching (http://www.mothur.org/wiki/Align.seqs) [27]. Potential chimeric sequences were detected using UCHIME (http://drive5.com/uchime) and removed [28]. The high-quality sequences were pre-clustered (http://www.mothur.org/wiki/Pre.cluster) and then clustered using an uncorrected pairwise algorithm. Operational taxonomic units (OTUs) were defined as sharing >97 % sequence identity using the average neighbor method (http://www.mothur.org/wiki/Cluster).
Statistical Analysis
Univariate analysis of variance (UNIANOVA) testing contaminant and physicochemical index difference and Pearson’s correlation analysis were performed by SPSS version 17 (SPSS, Inc., Chicago, IL, USA). The contributions of geographic location (G), oil contamination (C), and physicochemical properties (P) to bacterial community variation and the contributions of geographic location (G) and oil contamination (C) to soil physicochemical property variation were evaluated with variance partitioning analysis using (partial) canonical correspondence analysis (CCA) and redundancy analysis (RDA), respectively, by CANOCO for Windows Version 4.5 [29] (Plant Research International, Wageningen, The Netherlands). The significance test was carried out by Monte Carlo permutation (999 times). Weighted principal component analysis (PCA) was generated by using CANOCO for Windows Version 4.5 to demonstrate the clustering of different sites. Pairwise similarities between individual sites were calculated based on the patterns of OTUs0.03 using Jaccard’s similarity index (Cjaccard = SAB / (SA + SB − SAB), where SAB is the number common OTUs0.03 in sites A and B, SA is the number of OTUs0.03 in the sample A, and SB is the number of OTUs0.03 in the sample B [30]. Good’s coverage index, abundance-based coverage estimator (ACE), Chao1 richness estimator, and Shannon-Wiener diversity index were calculated by Mothur 1.15.0 [25]. Venn diagrams and Heatmap figure were implemented by the R (http://www.R-project.org) packages VennDiagram [31] and pheatmap [32], respectively.
Results
Basic Characteristic of Geographic Location and Physicochemical Property
The geographical and physicochemical data was listed in Table 1. The latitude of all the samples were approximately 40°, while longitude varied from 84° to 125°. XJ had lowest precipitation (109 mm/a), while HB had the highest (575 mm/a). The TPH concentration varied substantially among the contaminated sites from 25,560 to 183,760 mg/kg, which far exceeds the threshold of TPH concentration in environment (200 mg/kg) set by Organization for Economic Cooperation and Development (OECD). The TPH concentration of uncontaminated soils was 0.00 mg/kg, confirming that they were “uncontaminated”. UNIANOVA showed that the TPH concentrations of the four oil fields were significantly different (p < 0.01), and the concentrations of TPH’s four main components (saturates, naphthene aromatics, polar aromatics, and asphaltenes) were significantly different (p < 0.01). Saturates were the most dominant component. Other tested soil parameters (soil texture, density, WC, pH, salinity, TN, Olsen-N, TP, Olsen-P, TK, Olsen-K, and OM) were different between contaminated and uncontaminated soil. Physicochemical properties of the uncontaminated soil in four oil fields were not significantly different (p > 0.05).
Diversity of Bacterial Community
There were 125,654 (70.1 %) high-quality sequences that could be grouped into 4679 OTUs0.03. Different numbers of OTUs0.03 were assigned in different oil fields (Table 2). Good’s coverage ranged from 89.8 to 99.2 %, except for the bacterial 16S ribosomal RNA (rRNA) library from DQ(BLK) (79.3 %), indicating that most of the libraries could well reflect the majority of the bacterial community. According to the Shannon-Wiener index values, the highest level of biodiversity was found in the XJ (Table 2). Each oil field possessed a bacterial community feature that differed significantly from others based on the OTUs0.03 abundance (p < 0.01). Venn diagrams and PCA were used to visualize the overall spatial heterogeneity in bacterial community patterns. The Venn diagram showed that only 229 OTUs0.03 out of 4355 were shared by all sites. DQ had the largest number of shared communities (72.14 %), while SL had the largest number of unique OTUs0.03 (49.17 %) (Fig. S2). The shared portion between HB and SL was the lowest. In the PCA plot, only DQ and HB clustered close, while the other two showed a clear separation, with one quadrant composed entirely of one site, indicating that the bacterial communities from SL and XJ were quite different from the other three (Fig. 1).
Taxonomic Composition of the Bacterial Community
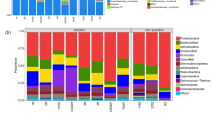
In addition to OTU-based analysis, the individual sequences were assigned to phylotypes and were grouped into 35 phyla, 92 classes, and 606 genera in oil-contaminated soil. At the phylum level, soil was dominated by Proteobacteria (38.33 %), Actinobacteria (23.30 %), Bacteroidetes (11.13 %), and Firmicutes (5.33 %). Unequal distribution among the sites was detected (Fig. 2). In HB, the largest dominant phylum was Actinobacteria, at approximately 51 % of all sequences, instead of Proteobacteria, as in other sites. The percentage of Bacteroidetes was relatively low in HB (2.88 %) and XJ (5.89 %), while it reached 24.29 % in SL.
At the class level, the first three largest dominant bacteria were Gammaproteobacteria (22.21 %), Actinobacteria (17.69 %), and Alphaproteobacteria (11.88 %). A hierarchal clustering double dendogram was performed based upon the relative percentage of the top 40 abundant bacteria at the class level (Fig. S3). Gammaproteobacteria, Actinobacteria, and Alphaproteobacteria were dominant in DQ and HB. Bacilli was far more abundant (13.79 %) in XJ than in other sites, so was Cytophagia in SL (19.74 %). A prevalent group in three of the sites, Actinobacteria was to be found rare in SL (2.88 %). Many of the classes were present in low proportions in all sites.
At the genera level, the 20 most prevalent bacteria were counted, of which Streptomyces, belonging to the Actinobacteria phylum, was the largest dominant genus (Table S1). The top 20 genera in each site were quite different (Table 3). The maximum number of genera (407) was detected in XJ, with Marinobacter (6.22 %), Halomonas (6.04 %), and Alcanivorax (4.84 %) as the three most prevalent genera. In particular, Halothiobacillus (1.01 %) and Sediminibacter (0.94 %) were the two unique but abundant genera in SL, while Haloactinosporathe (1.07 %) was only found in XJ.
Functional Group Distribution
In this study, bacteria known to perform a particular function and appeared in our study were identified according to the literature and were summarized in Table S2 at a genus level to analyze their distinct distribution in the oil-contaminated soil [17, 33–48]. The largest percentage of functional genera related to nitrogen metabolism was detected in DQ, while HB possessed the largest proportion of bacteria able to utilize oil as a carbon source, followed by XJ (Fig. 3). XJ also contained the largest percentage of phosphorus metabolism bacterium, Gemmatimonadetes. The sulfur and sulfate-reducing bacterium Desulfuromonas was found to be prevalent in SL. Significant numbers of the hydrocarbon-degrading bacterium Mycobacterium and the saturate-degrading bacterium Alcanivorax were detected in HB and SL, respectively. HB had no sulfur and sulfate-reducing bacterium (Desulfuromonas) being detected, while there was no polyphosphate-accumulating bacterium (Gemmatimonas) found in XJ.
Influence Factor Analysis of Geographic Location and Oil Contamination on Physicochemical Properties
As the research sites shared similar latitudes, longitude converted into projected coordinates was selected as geographic location indices. Alternative indices of contamination factor were the concentration of TPH and the percentages of its four components. To avoid the arching effect in RDA and CCA, Pearson’s correlation analyses were conducted among alternative indices of contamination factors. The results showed that TPH, saturates, naphthene aromatics, polar aromatics, and asphaltenes were mutually significantly correlated (p < 0.05) (Table S3). TPH, as the quantum of the four components, was thus selected as the contamination factor index.
Variance partitioning analysis was performed to determine the quantifiable contributions of the geographic location factor (G) and the contamination factor (C) to physicochemical property variation. The result showed that the geographic location contributed the most (28.40 %) to physicochemical properties (Fig. 4a). The interaction between geographical location and oil contamination could be ignored, and 65.80 % of the physicochemical properties could not be explained by these two components. Oil contamination had less individual impact (5.70 %) on physicochemical properties. This 5.70 % contribution led to changes in physicochemical properties under contamination interference.
Comparing the physicochemical properties of contaminated soil and uncontaminated soil, regular changes were found in density, WC, TN, TK, and OM. The density, WC, and TK obviously decreased, while the TN and OM obviously increased (Table 1). Oil contamination transformed most soil texture from loam to sandy loam. In addition, Pearson’s correlation analysis was also carried out between the soil physicochemical index and the oil contamination index to clarify the contribution of oil contamination to specific changes in physicochemical properties (Table S4). The result showed that OM significantly positively correlated to oil pollutant contents (r polar aromatics = 0.589, p < 0.01; r saturates = 0.495, p < 0.01). WC and pollutants had a significantly negative correlation, among which WC and saturates had the highest correlation coefficient (r saturates = 0.621, p < 0.01). In the PCA plot, samples of uncontaminated soil clustered much closer, while samples of contaminated soil had a much dispersed distribution (Fig. S4). HB had the farthest distance between its contaminated soil and uncontaminated soil. Therefore, oil contamination had changed the soil physicochemical properties and enhanced its heterogeneity.
Influence Factor Analysis of Geographic Location, Oil Contamination, and Physicochemical Properties on Bacterial Communities
Pearson’s correlation analysis was also conducted among physicochemical factors (WC, density, pH, salinity, OM, TN, TP, TK, Olsen-N, Olsen-P, and Olsen-K). The results showed that TN and Olsen-N were significantly correlated (r = 0.891, p < 0.01), as were Olsen-P and Olsen-K (r = 0.521, p < 0.01) (Table S5). TN, TP, and TK were excluded as influence factor indices because of their low interactions with microbes, while Olsen-N, Olsen-P, and Olsen-K were retained for their different roles in microbial metabolism. Thus, WC, density, pH, salinity, OM, Olsen-N, Olsen-P, and Olsen K were selected as the physicochemical factor indices.
Along with the geographic location and contamination factor indices, variance partitioning analysis was further performed to quantify the contributions of the geographic location factor (G), the contamination factor (C), and the physicochemical factor (P) to the bacterial community variation represented by OTU0.03. The total variation was partitioned into the independent effects of G, C, and P, interactions between any two components (G-C, C-P, and G-P), common interactions of all three components (G-C-P), and any remaining unexplained data (Fig. 4b). A total of 49.28 % of the variation was significantly explained (p < 0.01) by these three components. Geographical location, oil contamination, and physicochemical properties could independently explain 2.65, 2.55, and 29.21 % of the total variation, respectively. The interactions between geographical location and physicochemical properties (11.22 %) had greater influence compared to the individual influence of the contamination factor (2.55 %). Besides, oil contamination altered soil physicochemical properties and soil physicochemical properties could contribute 29.21 % to the influence of bacterial community distribution. Thus, oil contamination had more intensive indirect impact on bacterial community compared to direct impact.
Significant difference was found between OTUs0.03 in contaminated samples and uncontaminated samples (p < 0.01). PCA was also carried out to visualize the differences in community structure. In the plot, DQ(BLK) and HB(BLK) clustered much closer compared to DQ and HB, showing that oil contamination altered the distribution pattern of the bacterial community and might strengthen its heterogeneity (Fig. 1). This was also confirmed by the Jaccard similarity index, with DQ and HB populations exhibiting a lower index (0.268) than that between DQ(BLK) and HB(BLK) (0.332) (Table S6). All these analyses suggested that oil contamination changed bacterial community features directly or through changing physicochemical properties of the soil and further strengthened the differences among the four oil fields, which were located in different geographic sites.
Discussion
Significant difference of physicochemical property of uncontaminated soil in four sites was not observed (p > 0.05). Oil contamination shifted the overall pattern of the soil physicochemical properties (Fig. S4), primarily causing a significant change in texture, a decrease in the water content, and an increase in the organic matter. Pignatello (1996) thought the sand particles might increase after the entrance of oil in soil, which resulted in the change of soil texture. This phenomenon might caused by the absorption of hydrophobic compounds by humic-coated clay, for the high viscosity of oil, and the change of soil sedimentation rate (Zhou et al. 1995; Pignatello and Xing 1996) [49, 50]. The significant decline in the water content may result from the strong hydrophobic characteristic of oil [12] and an increase in the water evaporation caused by the higher soil temperature after oil contamination [16]. Meanwhile, oil contamination could bring about a decrease in soil density, which is consistent with the result of Khamehchiyan’s study [12].
According to the results, we concluded that physicochemical properties were the most dominant factor influencing bacterial community distribution, followed by geographical location. Oil contamination had a stronger indirect influence and weaker direct influence on bacterial community, which significantly changed its distribution (p < 0.01, Fig. 1, Table S6). In the study of Liang et al., geographic location played a more essential role than the physicochemical environment to the microbe community [16]. As our study selected more integrated physicochemical parameters and added a specific analysis of the impact of geographic location and oil contamination on the physicochemical properties, the question of the bacterial community influence factor was better addressed. However, approximately 50.72 % of the bacterial composition still could not be explained by the geographic location, contaminant, and physicochemical indices, which was consistent with other studies [51, 52]. This might result from the incompleteness of the index setting. For instance, natural factors such as climate, temperature differences, and daylight duration would affect microbial metabolism [16]; the duration of contamination in the research site and the sampling process would also interfere with the microbial community [52, 53]; and animals and plants had an intimate relationship with microbes [52, 54, 55].
Geographical location, oil contamination, and physicochemical properties shaped the bacterial communities distribution and different environmental settings presented distinguished patterns. Proteobacteria, Actinobacteria, Bacteroidetes, and Acidobacteria, which play an important role in carbon, nitrogen and other nutrition cycling, were found to be prevalent in contaminated sites, which was in line with other research results [56, 57]. Many genera which play a major role in the fate of pollutants and the cycling of nutrients were detected in this study [58, 59] (Table S2). Although the distribution of functional groups had been described, whether these represented taxa were also functionally active in this study is still unknown. As stated, the physicochemical properties had the strongest influence on the uniqueness of the bacterial community composition and diversity. Daqing oil field had a large proportion of functional groups, especially those that take part in nitrogen turnover. This might be closely associated with the high nitrogen content of the soil in Daqing oil field (Table 1). A diversified bacterial community was found in the oil-contaminated soil of the Huabei oil field, particularly hydrocarbon-degrading bacteria, such as Mycobacterium. The highest TPH concentration in the Huabei oil field might be the cause of this site containing the highest proportion of hydrocarbon-degrading bacteria compared to other functional groups and sites (Table 1). Unlike other oil fields, the most dominant phylum in the Huabei oil field was Actinobacteria, which is an aerobic community that develops in soil with enough pores and gas, as well as effective gas exchange with the atmosphere [60]. The Huabei oil field is located in the Huabei plain, one of the grain-producing areas in China, and the agriculture soil provides an optimal habitat with loam for aerobic bacteria compared with other oil fields (Table 1). Thus, Actinobacteria predominated in the bacterial community. The Shengli oil field had Proteobacteria as the largest dominant phylum and possessed the largest proportion (49.17 %) of local genera such as RF3, and Actinobacteria, which were prevalent in other oil fields, was rarely detected in this oil field, which might be explained by its more severe soil salinization and alkalization (Table 1). The Karamay oil field is well known for its location in the Gobi desert, which experiences very low annual precipitation. The special geographic characteristics of the Karamay oil field might result in its unique bacterial community characteristics. The Karamay oil field had the most diversified soil bacterial community and a notable proportion of functional groups that are especially involved in oil utilization, indicating that in the barren Gobi desert, oil as a carbon source facilitated the development of hydrocarbon-degrading group. The largest proportion of bacteria involved in phosphorus turnover might be explained by its highest phosphorus content (Table 1). Actinomycetales, which are known to dominate alkaline and particularly arid soil, were only found in the Karamay oil field, and Bacilli, which adapt to arid environments, were far more abundant there (13.79 %) [61, 62].
This study allowed us to shed light on the diversity and composition of bacterial communities thriving in oil-contaminated soil environments, where geographic location, oil contamination, and physicochemical properties lead to their significant differences among disparate oil fields across China.
References
Maier RM, Pepper IL, Gerba CP (eds) (2009) Environmental microbiology (Vol. 397). Academic Press, San Diego
Torsvik V, Øvreås L (2002) Microbial diversity and function in soil: from genes to ecosystems. Curr Opin Microbiol 5(3):240–245
Fulthorpe RR, Roesch LF, Riva A, Triplett EW (2008) Distantly sampled soils carry few species in common. ISME J 2(9):901–910
Gerdes B, Brinkmeyer R, Dieckmann G, Helmke E (2005) Influence of crude oil on changes of bacterial communities in Arctic sea‐ice. FEMS Microbiol Ecol 53(1):129–139
Fierer N, Nemergut D, Knight R, Craine JM (2010) Changes through time: integrating microorganisms into the study of succession. Res Microbiol 161:635–642
Banning NC, Gleeson DB, Grigg AH, Grant CD, Andersen GL et al (2011) Soil microbial community successional patterns during forest ecosystem restoration. Appl Environ Microbiol 77:6158–6164
Ding G, Piceno YM, Heuer H, Weinert N (2013) Changes of soil bacterial diversity as a consequence of agricultural land use in a semi-arid ecosystem. PLoS One 8:e59497
Zhang Y, Cong J, Lu H, Yang C, Yang Y, Zhou J, Li D (2014) An integrated study to analyze soil microbial community structure and metabolic potential in two forest types. PLoS One 9(4):e93773
Allen JP, Atekwana EA, Atekwana EA, Duris JW, Werkema DD, Rossbach S (2007) The microbial community structure in petroleum-contaminated sediments corresponds to geophysical signatures. Appl Environ Microbiol 73(9):2860–2870
Paissé S, Coulon F, Goñi‐Urriza M, Peperzak L, McGenity TJ, Duran R (2008) Structure of bacterial communities along a hydrocarbon contamination gradient in a coastal sediment. FEMS Microbiol Ecol 66(2):295–305
Yang YY, Wang J, Liao JQ, Huang Y, Xie SG (2014) Distribution of naphthalene dioxygenase genes in crude oil-contaminated soils. Microb Ecol. doi:10.1007/s00248-014-0457-7
Khamehchiyan M, Hossein Charkhabi A, Tajik M (2007) Effects of crude oil contamination on geotechnical properties of clayey and sandy soils. Eng Geol 89(3):220–229
Al Dousari A, Literathy P (2008) Evidence of hydrocarbon contamination from the Burgan oil field, Kuwait—interpretations from thermal remote sensing data. J Environ Manag 86(4):605–615
Gogoi BK, Dutta NN, Goswami P, Krishna Mohan TR (2003) A case study of bioremediation of petroleum-hydrocarbon contaminated soil at a crude oil spill site. Adv Environ Res 7(4):767–782
Hamid HRA, Kassim WM, El Hishir A, El-Jawashi SA (2008) Risk assessment and remediation suggestion of impacted soil by produced water associated with oil production. Environ Monit Assess 145(1–3):95–102
Liang Y, Van Nostrand JD, Deng Y, He Z, Wu L, Zhang X et al (2011) Functional gene diversity of soil microbial communities from five oil-contaminated fields in China. ISME J 5(3):403–413
Liu YJ, Chen YP, Jin PK, Wang XC (2009) Bacterial communities in a crude oil gathering and transferring system (China). Anaerobe 15(5):214–218
Berthrong ST, Buckley DH, Drinkwater LE (2013) Agricultural management and labile carbon additions affect soil microbial community structure and interact with carbon and nitrogen cycling. Microb Ecol 66(1):158–170
China Meteorological Data Sharing Service System (2012) http://jingyan.baidu.com/article/fa4125ac906aa328ac7092d6.html. Accessed 30 Dec 2012
Huesemann M (1995) Predictive model for estimating the extent of petroleum hydrocarbon biodegradation in contaminated soils. Environ Sci Technol 29:7–18
ASTM D4124–01 (2001) Standard test methods for separation of asphalt into four fractions. ASTM International, West Conshohocken
Lu RK (1999) Soil agricultural chemical analysis. China agricultural science and technology press, Nanjing
Al-Mailem D, Sorkhoh N, Al-Awadhi H, Eliyas M, Radwan S (2010) Biodegradation of crude oil and pure hydrocarbons by extreme halophilic archaea from hypersaline coasts of the Arabian Gulf. Extremophiles 14:321–328
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA et al (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437(7057):376–380
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75(23):7537–7541
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J, Glöckner FO (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35(21):7188–7196
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19):2460–2461
ter Braak CJ and Smilauer P (1998) CANOCO reference manual and user’s guide to Canoco for Windows: software for canonical community ordination (version 4)
McCune B, Grace JB, Urban DL (2002) Analysis of ecological communities (Vol. 28). MjM software design, Gleneden Beach
Chen H (2011) VennDiagram: Generate high-resolution Venn and Euler plots. R package version, 113
Kolde R (2012) pheatmap: pretty heatmaps. R package version, 61
Chin KJ, Liesack W, Janssen PH (2001) Opitutus terrae gen. nov., sp. nov., to accommodate novel strains of the division ‘Verrucomicrobia’ isolated from rice paddy soil. Int J Syst Evol Microbiol 51(6):1965–1968
Camargo FAO, Bento FM, Okeke BC, Frankenberger WT (2004) Hexavalent chromium reduction by an actinomycete, Arthrobacter crystallopoietes ES 32. Biol Trace Elem Res 97(2):183–194
Zhang H, Sekiguchi Y, Hanada S, Hugenholtz P, Kim H, Kamagata Y, Nakamura K (2003) Gemmatimonas aurantiaca gen. nov., sp. nov., a Gram-negative, aerobic, polyphosphate-accumulating micro-organism, the first cultured representative of the new bacterial phylum Gemmatimonadetes phyl. nov. Int J Syst Evol Microbiol 53(4):1155–1163
DeBruyn JM, Nixon LT, Fawaz MN, Johnson AM, Radosevich M (2011) Global biogeography and quantitative seasonal dynamics of Gemmatimonadetes in soil. Appl Environ Microbiol 77(17):6295–6300
Li G, Huang W, Lerner DN, Zhang X (2000) Enrichment of degrading microbes and bioremediation of petrochemical contaminants in polluted soil. Water Res 34(15):3845–3853
Radwan SS, Al-Hasan RH, Ali N, Salamah S, Khanafer M (2005) Oil-consuming microbial consortia floating in the Arabian Gulf. Int Biodeterior Biodegrad 56(1):28–33
Sorkhoh N, Al-Hasan R, Radwan S (1992) Self-cleaning of the Gulf. Nature 359:109
Magot M, Ollivier B, Patel BK (2000) Microbiology of petroleum reservoirs. Antonie Van Leeuwenhoek 77(2):103–116
Hara A, Syutsubo K, Harayama S (2003) Alcanivorax which prevails in oil‐contaminated seawater exhibits broad substrate specificity for alkane degradation. Environ Microbiol 5(9):746–753
Yakimov MM, Denaro R, Genovese M, Cappello S, D’Auria G, Chernikova TN et al (2005) Natural microbial diversity in superficial sediments of Milazzo Harbor (Sicily) and community successions during microcosm enrichment with various hydrocarbons. Environ Microbiol 7(9):1426–1441
Gu J, Cai H, Yu SL, Qu R, Yin B, Guo YF et al (2007) Marinobacter gudaonensis sp. nov., isolated from an oil-polluted saline soil in a Chinese oilfield. Int J Syst Evol Microbiol 57(2):250–254
Bordoloi NK, Konwar BK (2009) Bacterial biosurfactant in enhancing solubility and metabolism of petroleum hydrocarbons. J Hazard Mater 170(1):495–505
Brito EMS, Guyoneaud R, Goñi-Urriza M, Ranchou-Peyruse A, Verbaere A, Crapez MA et al (2006) Characterization of hydrocarbonoclastic bacterial communities from mangrove sediments in Guanabara Bay. Brazil Res Microbiol 157(8):752–762
Saul DJ, Aislabie JM, Brown CE, Harris L, Foght JM (2005) Hydrocarbon contamination changes the bacterial diversity of soil from around Scott Base, Antarctica. FEMS Microbiol Ecol 53(1):141–155
Westerberg K, Elväng AM, Stackebrandt E, Jansson JK (2000) Arthrobacter chlorophenolicus sp. nov., a new species capable of degrading high concentrations of 4-chlorophenol. Int J Syst Evol Microbiol 50(6):2083–2092
Liu J, Yang H, Zhao M, Zhang XH (2014) Spatial distribution patterns of benthic microbial communities along the Pearl Estuary, China. Syst Appl Microbiol 37:578–589
Pignatello JJ, Xing B (1996) Mechanisms of slow sorption of organic chemicals to natural particles. Environ Sci Technol 30(1):1–11
Zhou E, Crawford RL (1995) Effects of oxygen, nitrogen, and temperature on gasoline biodegradation in soil. Biodegradation 6(2):127–140
Zhou J, Kang S, Schadt CW, Garten CT (2008) Spatial scaling of functional gene diversity across various microbial taxa. Proc Natl Acad Sci U S A 105(22):7768–7773
Ramette A, Tiedje JM (2007) Multiscale responses of microbial life to spatial distance and environmental heterogeneity in a patchy ecosystem. Proc Natl Acad Sci U S A 104(8):2761–2766
Bhattacharya D, Sarma PM, Krishnan S, Mishra S, Lal B (2003) Evaluation of genetic diversity among Pseudomonas citronellolis strains isolated from oily sludge-contaminated sites. Appl Environ Microbiol 69(3):1435–1441
He J, Xu Z, Hughes J (2006) Molecular bacterial diversity of a forest soil under residue management regimes in subtropical Australia. FEMS Microbiol Ecol 55(1):38–47
Rooney-Varga JN, Giewat MW, Duddleston KN, Chanton JP, Hines ME (2007) Links between archaeal community structure, vegetation type and methanogenic pathway in Alaskan peatlands. FEMS Microbiol Ecol 60(2):240–251
Janssen PH (2006) Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes. Appl Environ Microbiol 72(3):1719–1728
Spain AM, Krumholz LR, Elshahed MS (2009) Abundance, composition, diversity and novelty of soil Proteobacteria. ISME J 3(8):992–1000
Paerl HW, Dyble J, Moisander PH, Noble RT, Piehler MF, Pinckney JL et al (2003) Microbial indicators of aquatic ecosystem change: current applications to eutrophication studies. FEMS Microbiol Ecol 46(3):233–246
Powell SM, Bowman JP, Snape I, Stark JS (2003) Microbial community variation in pristine and polluted nearshore Antarctic sediments. FEMS Microbiol Ecol 45(2):135–145
Akkermans AD, Van Elsas JD, De Bruijn FJ (1995) Molecular microbial ecology manual. Kluwer, Dordrecht, pp 1–8
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75(15):5111–5120
Bachar A, Al-Ashhab A, Soares MIM, Sklarz MY, Angel R, Ungar ED, Gillor O (2010) Soil microbial abundance and diversity along a low precipitation gradient. Microb Ecol 60(2):453–461
Acknowledgments
This work was supported by the Public Welfare Project of Ministry of Environmental Protection (No. 201309034).
Conflict of Interest
The authors declare that there is no conflict of interests regarding the publication of this article.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Most abundant 20 genera identified in the four oil fields (DOC 55 kb)
Table S2
Functional groups and their taxonomy (DOC 73 kb)
Table S3
Pearson’s correlation matrix of contamination factor alternative index (DOC 30 kb)
Table S4
Pearson’s correlation matrix of basic soil physicochemical index and contaminant (DOC 41 kb)
Table S5
Pearson’s correlation matrix of physicochemical factor alternative index (DOC 46 kb)
Table S6
The pair Jaccard similarity of bacterial community (DOC 30 kb)
Figure S1
Schematic map of sampling sites (DOC 1106 kb)
Figure S2
Venn diagram showing the unique and shared OTUs0.03 (DOC 422 kb)
Figure S3
Distribution of the top 40 most abundant classes of bacteria found in contaminated soil in research areas (DOC 196 kb)
Figure S4
Physicochemical PCA of contaminated and uncontaminated soil (DOC 46 kb)
Rights and permissions
About this article
Cite this article
Liao, J., Wang, J. & Huang, Y. Bacterial Community Features Are Shaped by Geographic Location, Physicochemical Properties, and Oil Contamination of Soil in Main Oil Fields of China. Microb Ecol 70, 380–389 (2015). https://doi.org/10.1007/s00248-015-0572-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-015-0572-0