Abstract
Pooled sequencing-based fitness assays are a powerful and widely used approach to quantifying fitness of thousands of genetic variants in parallel. Despite the throughput of such assays, they are prone to biases in fitness estimates, and errors in measurements are typically larger for deleterious fitness effects, relative to neutral effects. In practice, designing pooled fitness assays involves tradeoffs between the number of timepoints, the sequencing depth, and other parameters to gain as much information as possible within a feasible experiment. Here, we combined simulations and reanalysis of an existing experimental dataset to explore how assay parameters impact measurements of near-neutral and deleterious fitness effects using a standard fitness estimator. We found that sequencing multiple timepoints at relatively modest depth improved estimates of near-neutral fitness effects, but systematically biased measurements of deleterious effects. We showed that a fixed total number of reads, deeper sequencing at fewer timepoints improved resolution of deleterious fitness effects. Our results highlight a tradeoff between measurement of deleterious and near-neutral effect sizes for a fixed amount of data and suggest that fitness assay design should be tuned for fitness effects that are relevant to the specific biological question.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Accurate fitness measurements are central to questions in experimental evolution, quantitative genetics, and functional genomics. Traditional methods include approaches such as estimating maximum growth rate from growth curves (Hall et al. 2014) and quantifying colony sizes from spot assays (Baryshnikova et al. 2010). An alternative, and increasingly used, approach is a competitive fitness assay in which a reference strain with known fitness is competed directly with a test strain. The relative fitness of the test strain can be inferred from its change in frequency compared to the reference, with either colony counts (Lenski et al. 1991) or fluorescence as a readout (Breslow et al. 2008; Thompson et al. 2006). However, pairwise competition assays are lower throughput and challenging to scale to thousands of measurements.
Competitive fitness assays can be adapted from measuring fitness of a single test strain per assay to measuring fitnesses of several thousand strains in parallel. This typically involves uniquely tagging each strain with a DNA barcode and tracking changes in the frequency of the barcodes over time using deep sequencing (Smith et al. 2009, 2010). Applications of such sequencing based fitness measurements and phenotyping range from CRISPR (Shalem et al. 2014; Wang et al. 2014) and transposon mutagenesis screening for essential genes (van Opijnen and Camilli 2013; Wetmore et al. 2015), genetic interaction screens (Du et al. 2017; Jaffe et al. 2017), deep mutational scanning of proteins (Fowler and Fields 2014; Fowler et al. 2010; Stiffler et al. 2015), codon usage (Kelsic et al. 2016), fitness measurements of thousands of adaptive mutations from evolution experiments (Venkataram et al. 2016), genetic crosses (Nguyen Ba et al. 2022), and natural variants (Carrasquilla et al. 2022).
Despite their scalability, highly parallel sequencing-based fitness assays are prone to biases in estimation of fitness. Li et al. demonstrated that fold enrichment based fitness metrics cannot be quantitatively compared across pools of strains with different underlying distributions of fitness effects, and developed FitSeq, a fitness estimation method that accounts for changes in the mean population fitness over time (Li et al. 2018, 2023). Their simulations also showed that measuring multiple timepoints makes fitness estimates more robust to changes in the distribution. However, uncertainty in fitness measurements depend on the true fitness and are typically worse for more deleterious fitness effects. Consequently, it is not evident if parameter regimes improving resolution of fitness measurements are the same regardless of the true fitness effect under investigation.
Here, we combined simulated fitness assays for a range of experimental regimes and reanalysis of a deeply sequenced transposon mutagenesis dataset to explore how experimental parameters impact uncertainty in fitness measurements across a wide range of fitness effects. Some of the results presented here have appeared in other work and are cited when appropriate (Li et al. 2018; Robinson et al. 2014); the purpose of this paper is to combine both our findings and these existing insights to derive recommendations for designing pooled sequencing-based fitness assays.
Note About Terminology
While we refer to fitness effects of mutations throughout the paper, the results can be extended to any collection of strains, for instance, derived from an evolution experiment, or from natural variation. For a mutant with a true fitness effect s (relative to a reference strain), the change in frequency of the mutant is given by:
where f1 and f2 are the frequencies of the mutants before and after selection, and t is the number of generations of selection. We note that this equation holds when the frequency of the reference lineage does not change. This assumption means that all the mutant (i.e., non-reference) frequencies are very small and do not impact mean fitness. In a sequencing-based fitness assay, we can estimate the fitness effect of the mutation as follows:
where n1 and n2 are the mutants counts before and after selection, N1 and N2 are total counts for those timepoints, and t is the number of generations of selection. Note that this equation also assumes that the frequency of the reference lineage does not change. While this estimator is biased for finite read depth and bottleneck size, we found in simulations that this bias was negligible compared to measurement error (Fig. S1). This definition can be readily generalized to multiple timepoints as the slope of the linear regression of ln(frequency) vs number of generations of selection in fitness assay. Under this definition, a neutral mutation has a fitness of 0, and an unviable mutation (say loss of an essential gene) has a fitness effect of -ln(2) = − 0.693 (Chevin 2011). Note that a fitness value of -ln(2) is in units of inverse generations, and is pertinent to microbes dividing by binary fission.
Simulating Sequencing-Based Fitness Assays
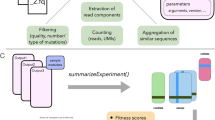
In our simulations, we decouple mutant abundances and the read counts from sequencing (Fig. 1A). We assume that the initial mutant abundances are Poisson distributed (with mean equal to the bottleneck size). Sequencing this mutant pool also leads to Poisson sampling, introducing additional noise, determined by the depth of sequencing. Each subsequent passaging in the fitness assay involves an additional bottleneck step.
We estimated fitness effects using linear regression of log(read count frequencies) vs number of generations, averaging over 5 replicates for each mutant (in practice typically done with redundant barcoding). Lastly, for mutant trajectories that disappeared (either due to demographic stochasticity, or due to deleterious fitness effect), we added a pseudocount, and restricted the regression to the first appearance of a zero read count. We made a few additional simplifying assumptions: we restricted our analysis to a focal mutant of interest, ignoring changes in the mean fitness of the population. We further ignored noise in measurement of reference strains, and overdispersion in the initial distribution of mutant abundances.
We examined the distribution of initial read counts prior to the start of the fitness assay (N(0)), involving Poisson sampling during the bottleneck and sequencing steps. We found that the read counts were overdispersed (Fig. 1B), and the variance was significantly higher than the mean regardless of sequencing depth (Fig. 1C). Due to the overdispersion, they are a more accurate proxy for real sequencing counts datasets than naïve Poisson distributed counts.
Results
First, we investigated the relative importance of bottleneck size and sequencing depth on measurement error. We calculated the uncertainty as the standard error of mean of fitness estimates of five replicates. We found that for neutral mutations, while errors decrease with bottleneck size and sequencing depth, we found that increasing sequencing depth much more beyond the bottleneck size has little impact on measurement error (Fig. 2). This pattern persisted for slightly deleterious mutations.
Diminishing returns of increasing sequencing depth well beyond the bottleneck size. Uncertainty in measurements is obtained as the standard errors of measurements obtained from simulated fitness assays (fixed parameters: number of timepoints = 2, number of generations between timepoints = log2(100))
Errors in Fitness Measurements Depend on the True Fitness of a Mutant
Next, we explored how the uncertainty in fitness estimates depends on the “true” fitness of a mutation. We observed that errors are consistently larger for more deleterious mutations (Fig. 3A). Next, we turned to a deeply sequenced transposon sequencing dataset of E. coli B REL606 from our previous work (Limdi et al. 2022). We found a statistically significant negative correlation between the estimated fitness of disrupting a gene and the error in the estimate (p value < 0.001, Fig. 3B); this pattern was more evident when we binned by mutant effect sizes (Fig. 3C). This was consistent with the result that FitSeq errors are larger for deleterious mutations (Li et al. 2018). Because errors were dependent on the effect size of the mutation, we decided to explore how experimental parameters impact both near-neutral fitness effects and deleterious fitness effects separately.
Errors in measurements depend on the true fitness of the mutation. a Uncertainty in measurements, defined as the standard error of mean of 5 replicates. Values plotted are the average of fitness assays for 1,000 mutations. Parameters: number of timepoints = 200, number of generations = log2(100), total sequencing depth per replicate = 200. b Error in fitness measurements (defined as standard error of mean) from a transposon sequencing dataset of E. coli B REL606, using two timepoints. c Same data as in b binned by fitness effects. Annotations above points indicate the number of genes in the bin
Near-Neutral Mutants Require More Time for Fitness Effects to Exceed Measurement Noise
Given fixed sequencing over the entire experiment, we investigated how varying the fitness assay design impacted the measurement errors of near-neutral mutations. First, we explored the impact of changing the number of timepoints in the fitness assay, keeping the number of generations between timepoints fixed. We found that in simulations, for a given sequencing depth, measurement errors were lower with more timepoints (and generations of selection) but with less sequencing per timepoint (Fig. 4A). We then tested this hypothesis in data from the transposon sequencing dataset, finding that over a 30-fold change in total sequencing depth over the experiment, errors were consistently smaller when sampling multiple timepoints and generations (Fig. 4B). Notably, an experiment with 3 × 105 reads spread out over five timepoints led to better estimates than 107 reads over two timepoints.
For fixed total sequencing depth over the experiment, increasing number of generations of selection leads to better resolution of near-neutral fitness effects despite less sequencing per time point. Schematic indicates design of the fitness assay, with circles in blue indicating which timepoints were sequenced. a Simulations: uncertainty in fitness estimates for a neutral mutation as a function of timepoints (and number of generations, interval = log2(100) generations) keeping total sequencing depth per mutant constant over the experiment. b Reanalysis of the TnSeq dataset: measurement uncertainty as a function of timepoints (and number of generations, interval = log2(100) generations), keeping total reads in the experiment constant. c Simulations: uncertainty in fitness estimates keeping total sequencing depth per mutant and total generations of selection constant (26.5 generations), while varying number of generations between sequencing. d Reanalysis of TnSeq dataset: measurement uncertainty in fitness estimates keeping total reads over the experiment and total generations of selection constant (26.5 generations), while varying number of generations between sequencing. Since there is no ground truth of neutrality, we averaged over the measurement errors for genes in the range of (-0.05, 0.05)
In the above analysis, both number of timepoints and number of generations of selection were varied simultaneously. We next probed the effect of changing the frequency of sequencing the mutant pools, keeping the total numbers of generations of selection constant. In both simulations and experimental data, we found that at low sequencing depths, sequencing only at two timepoints ~ 26.5 generations apart performed better than sequencing five timepoints ~ 6.6 generations apart (Fig. 4C, D). However, at high sequencing depths, the measurement errors were nearly independent of frequency of sampling. These results suggest that measurements of near-neutral mutations improve with longer durations of selection, while only weakly depending on frequency of sequencing.
For Deleterious Mutations, Increasing Timepoints and Generations of Selection at Fixed Total Sequencing Leads to Less Usable Data
Next, we investigated to what extent this intuition also held true for deleterious fitness estimates. For moderately deleterious mutations (s = − 0.1) we found that errors typically decreased with timepoints (and generations of selection). However, at low sequencing depth, (25 across the experiment), we found that going from 4 to 5 timepoints made measurements less reliable (Fig. 5A). To investigate this, we examined the average number of timepoints used in calculating fitness estimates. At low total depth, increasing timepoints (and therefore reducing sequencing reads per timepoint) measured in fact led to a reduction in usable data from the fitness assay, contributing to noisier measurements (Fig. 5B).
For fixed total sequencing, adding timepoints leads to less usable data for estimating deleterious fitness effects in simulated assays. Schematic indicates fitness assay designs, with blue circles indicating which timepoints were sequenced. a Uncertainty in fitness measurement for a mutant with true s = -0.1, as a function of timepoints measured (keeping number of generations between timepoints fixed: log2(100). b Average number of timepoints that were used to calculate fitness. c Uncertainty in fitness measurement for a mutant with true s = -0.25, as a function of timepoints measured (keeping number of generations between timepoints fixed: log2(100). d Average number of timepoints that were used to calculate fitness (Color figure online)
Similarly, for a strongly deleterious mutation (s = − 0.25), we found that errors increased from 3 to 4 timepoints for all but the highest sequencing depth (Fig. 5C). This trend corresponded to a decrease in the average number of timepoints used in fitness estimates (Fig. 5D). This suggests that for fixed sequencing depth, measuring earlier timepoints at greater depth provides better resolution of deleterious fitness effects, and that sampling additional timepoints can reduce the amount of useful data.
Tuning Frequency of Sequencing to Detect Deleterious Fitness Effects
While increasing the duration between sequencing (for fixed total depth) can be helpful for resolving near-neutral fitness effects, it is not necessarily optimal for deleterious fitness effects. Over tens of generations of selection, if no intermediate timepoints are sampled, it is not possible to distinguish a slightly deleterious mutation from an unviable mutation, as the expected mutant abundance and read counts is nearly zero for both scenarios. Under this scenario, the most deleterious fitness effect detectable can be estimated as:
Assuming equal total sequencing depths at two timepoints, this approximation follows from Eq. (2) because when trajectories disappear to 0, n2 = 1 (from adding a pseudocount of 1). We verified this approximation in experimental data, finding that the average of the ten most deleterious fitness effects, a proxy for the most deleterious effect detectable, matched the theoretical predictions well (Fig. 6). This comparison shows that true fitnesses below the theoretical bound cannot be estimated from pooled fitness assays, and if calculated, will be systematically over-estimated.
Generations of selection between sequencing set a lower bound on fitness that can be inferred using pooled fitness assays. Schematic indicates fitness assay design, with blue circles indication timepoints that were sequenced. a Predicted lower bound as a function of number of generations, and sequencing depth, assuming that the mutant disappears after selection (see Eq. 3). b Average of the 10 most deleterious fitness effects estimated for a set of parameters (downsampling and number of generations of selection). We use this as a proxy for the resolution of deleterious effects in bulk fitness assays. Dotted line in indicates the fitness of an unviable mutation, -ln(2)
Discussion
We found that sequencing more timepoints (over more generations of selection) at relatively lower sequencing depth, as opposed to fewer timepoints (and generations of selection) at very high depth, improves resolution of near-neutral fitness estimates. Conversely, for deleterious fitness effects, with additional time points there is less new, usable information obtained, as these variants are depleted over time. Our results highlight that the timescale of sampling in fitness assays should be tuned to the timescale of change in mutant frequencies. Moreover, they suggest that there is no combination of experimental parameters that optimally resolves both ranges of fitness effects for a fixed amount of data.
A limitation of our simulations is that we make several simplifying assumptions in modeling fitness assays. We do not consider noise from PCR amplification and DNA extraction steps, which likely contribute to higher measurement noise. We also do not account for changes in the mean fitness of populations over the course of the fitness assay. For a detailed discussion of how the underlying distribution of fitness effects impacts estimates of mutant fitness using log-fold change metrics, and more generally inferring fitnesses from barcode frequencies, we recommend the following (Li et al. 2018, 2023; Ascensao et al. 2023). While changing mean fitness can be corrected for using neutral, reference strains, any measurement errors in these lineages will propagate to fitness estimates of all mutants and can introduce systematic biases. Initial mutant abundances are not typically perfectly Poisson distributed, as generating mutant libraries involves growth steps which can skew abundances toward mutants that have a fitness advantage. Conversely, mutants growing poorly in the growth media (prior to the fitness assay) will have noisier measurements by virtue of starting off with fewer cells, and therefore read counts.
Our simulations and reanalysis of transposon sequencing data, combined with previously published results, can be distilled into principles for experimental design:
Identify Fitness Effects that are Relevant for the Biological Question at Hand
Errors in measurements depend on the true fitness of the mutations, and there exists a tradeoff between resolution of near-neutral fitnesses and deleterious fitness effects.
For Measurements Near Neutrality and Fixed Total Sequencing Budget, Sequence Mutant Pools at More Timepoints, with Less Sequencing Depth
Our reanalysis of a deeply sequenced transposons sequencing data shows that sequencing more timepoints (over longer selection periods) at lower sequencing depth outperforms sequencing very deeply but for fewer generations of selection (Fig. 4B). While we find that there is no advantage to sequencing more timepoints if the period of selection is unchanged (Fig. 4C, D), it may provide additional robustness to fitness estimates.
Firstly, the mean fitness of the population can change over time depending on the underlying distribution of fitness effects. This can lead to biased fitness estimates; for instance, neutral mutations may appear deleterious without any correction. Quantifying mutant abundance over multiple timepoints allows for use of methods such as FitSeq to correct for this bias. Secondly, beneficial mutations can occur on otherwise neutral or even deleterious over the course of the fitness assay purely due to chance. Sequencing multiple timepoints can allow for identifying such outlier events and excluding them from downstream analysis.
For Measurements of Deleterious Mutations and Fixed Total Sequencing Budget, Sequence at Higher Depth for Fewer Timepoints and Fewer Generations
For fixed amount of sequencing, adding more timepoints does not add meaningful information as deleterious mutations will go extinct over time. As a starting point for parameters, we recommend using sequencing depth and number of generations of selection such fitness effects of interest are above this lower bound predicted in Eq. 3. Lastly, we recommend (from experience) always sequencing the mutant pools prior to any fitness assay, as deleterious mutations (or variants) disappear from the pool rapidly in a few generations.
Using Pilot Experiments and Simulations to Guide Fitness Assay Design
We present an approach for tuning fitness assay design; we suggest performing a pilot fitness assay and sequencing experiment, using simulations as a starting point for experimental parameters, and reanalyzing the data with subsampling (either fewer reads or fewer timepoints). If the errors or resolution of fitness effects of interest do not change with subsampling, it is possible to collect data for more strains/genetic backgrounds with the same total sequencing.
Data availability
Raw sequencing reads have been deposited in the NCBI BioProject database under accession number PRJNA814281. Processed data are deposited on Zenodo (https://doi.org/10.5281/zenodo.6547536), and source code for sequencing pipeline, downstream analyses, and figure generation are available at GitHub (https://github.com/baymlab/2022_Limdi_limits-pooled-fitness-assays).
References
Ascensao JA, Wetmore KM, Good BH, Arkin AP, Hallatschek O (2023) Quantifying the local adaptive landscape of a nascent bacterial community. Nat Commun 14:248
Baryshnikova A, Costanzo M, Kim Y, Ding H, Koh J, Toufighi K, Youn J-Y, Ou J, San Luis B-J, Bandyopadhyay S et al (2010) Quantitative analysis of fitness and genetic interactions in yeast on a genome scale. Nat Methods 7:1017–1024
Breslow DK, Cameron DM, Collins SR, Schuldiner M, Stewart-Ornstein J, Newman HW, Braun S, Madhani HD, Krogan NJ, Weissman JS (2008) A comprehensive strategy enabling high-resolution functional analysis of the yeast genome. Nat Methods 5:711–718
Carrasquilla, M., Drammeh, N.F., Rawat, M., Sanderson, T., Zenonos, Z., Rayner, J.C., and Lee, M.C.S. (2022). Barcoding genetically distinct Plasmodium falciparum strains for comparative assessment of fitness and antimalarial drug resistance. https://doi.org/10.1101/2022.04.05.487250
Chevin L-M (2011) On measuring selection in experimental evolution. Biol Lett 7:210–213
Du D, Roguev A, Gordon DE, Chen M, Chen S-H, Shales M, Shen JP, Ideker T, Mali P, Qi LS et al (2017) Genetic interaction mapping in mammalian cells using CRISPR interference. Nat Methods 14:577–580
Fowler DM, Fields S (2014) Deep mutational scanning: a new style of protein science. Nat Methods 11:801–807
Fowler DM, Araya CL, Fleishman SJ, Kellogg EH, Stephany JJ, Baker D, Fields S (2010) High-resolution mapping of protein sequence-function relationships. Nat Methods 7:741–746
Hall BG, Acar H, Nandipati A, Barlow M (2014) Growth rates made easy. Mol Biol Evol 31:232–238
Jaffe M, Sherlock G, Levy SF (2017) iSeq: a new double-barcode method for detecting dynamic genetic interactions in yeast. G3 7:143–153
Kelsic ED, Chung H, Cohen N, Park J, Wang HH, Kishony R (2016) RNA structural determinants of optimal codons revealed by MAGE-Seq. Cell Syst 3:563-571.e6
Lenski RE, Rose MR, Simpson SC, Tadler SC (1991) Long-term experimental evolution in Escherichia coli. I. adaptation and divergence during 2,000 generations. Am Nat 138:1315–1341
Li F, Salit ML, Levy SF (2018) Unbiased fitness estimation of pooled barcode or amplicon sequencing studies. Cell Syst 7:521-525.e4
Li F, Tarkington J, Sherlock G (2023) Fit-Seq2.0: an improved software for high-throughput fitness measurements using pooled competition assays. J Mol Evol. https://doi.org/10.1007/s00239-023-10098-0
Limdi A, Owen SV, Herren C, Lenski RE, Baym M (2022) Parallel changes in gene essentiality over 50,000 generations of evolution. bioRxiv. https://doi.org/10.1101/2022.05.17.492023
Nguyen Ba AN, Lawrence KR, Rego-Costa A, Gopalakrishnan S, Temko D, Michor F, Desai MM (2022) Barcoded bulk QTL mapping reveals highly polygenic and epistatic architecture of complex traits in yeast. Elife. https://doi.org/10.7554/eLife.73983
Robinson DG, Chen W, Storey JD, Gresham D (2014) Design and analysis of bar-seq experiments. G3 4:11–18
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelson T, Heckl D, Ebert BL, Root DE, Doench JG et al (2014) Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343:84–87
Smith AM, Heisler LE, Mellor J, Kaper F, Thompson MJ, Chee M, Roth FP, Giaever G, Nislow C (2009) Quantitative phenotyping via deep barcode sequencing. Genome Res 19:1836–1842
Smith AM, Heisler LE, St Onge RP, Farias-Hesson E, Wallace IM, Bodeau J, Harris AN, Perry KM, Giaever G, Pourmand N et al (2010) Highly-multiplexed barcode sequencing: an efficient method for parallel analysis of pooled samples. Nucleic Acids Res 38:e142
Stiffler MA, Hekstra DR, Ranganathan R (2015) Evolvability as a function of purifying selection in TEM-1 β-lactamase. Cell 160:882–892
Thompson DA, Desai MM, Murray AW (2006) Ploidy controls the success of mutators and nature of mutations during budding yeast evolution. Curr Biol 16:1581–1590
van Opijnen T, Camilli A (2013) Transposon insertion sequencing: a new tool for systems-level analysis of microorganisms. Nat Rev Microbiol 11:435–442
Venkataram S, Dunn B, Li Y, Agarwala A, Chang J, Ebel ER, Geiler-Samerotte K, Hérissant L, Blundell JR, Levy SF et al (2016) Development of a comprehensive genotype-to-fitness map of adaptation-driving mutations in yeast. Cell 166:1585-1596.e22
Wang T, Wei JJ, Sabatini DM, Lander ES (2014) Genetic screens in human cells using the CRISPR-Cas9 system. Science 343:80–84
Wetmore KM, Price MN, Waters RJ, Lamson JS, He J, Hoover CA, Blow MJ, Bristow J, Butland G, Arkin AP et al (2015) Rapid quantification of mutant fitness in diverse bacteria by sequencing randomly bar-coded transposons. Mbio 6:e00306–e00315
Acknowledgements
We thank Fernando Rossine, Eleanor Rand, and Indra Gonzalez Ojeda for feedback and discussion on analysis and figures. A.L. acknowledges support from the Molecules, Cells, and Organisms Graduate Program, Harvard University. M.B. acknowledges support from the NIGMS of the National Institutes of Health (R35GM133700), the David and Lucile Packard Foundation, the Pew Charitable Trusts, and the Alfred P. Sloan Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Additional information
Handling editor: Greg Lang.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Limdi, A., Baym, M. Resolving Deleterious and Near-Neutral Effects Requires Different Pooled Fitness Assay Designs. J Mol Evol 91, 325–333 (2023). https://doi.org/10.1007/s00239-023-10110-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-023-10110-7