Abstract
Electrospray ionization mass spectrometry (ESI-MS) experiments, including ion mobility spectrometry mass spectrometry (ESI-IMS-MS) and electron capture dissociation (ECD) of proteins ionized from aqueous solutions, have been used for the study of solution-like structures of intact proteins. By mixing aqueous proteins with denaturants online before ESI, the amount of protein unfolding can be precisely controlled and rapidly analyzed, permitting the characterization of protein folding intermediates in protein folding pathways. Herein, we mixed various pH solutions online with aqueous cytochrome C for unfolding and characterizing its unfolding intermediates with ESI-MS charge state distribution measurements, IMS, and ECD. The presence of folding intermediates and unfolded cytochrome c structures were detected from changes in charge states, arrival time distributions (ATDs), and ECD. We also compared structures from nondenaturing and denaturing solution mixtures measured under “gentle” (i.e., low energy) ion transmission conditions with structures measured under “harsh” (i.e., higher energy) transmission. This work confirms that when using “gentle” instrument conditions, the gas-phase cytochrome c ions reflect attributes of the various solution-phase structures. However, “harsh” conditions that maximize ion transmission produce extended structures that no longer correlate with changes in solution structure.
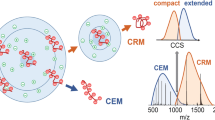
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mass spectrometry (MS) is a leading tool for studying protein dynamics and structure [1,2,3,4]. When studying protein function, it is important to characterize the conformational ensembles of functional, non-functional, and dysfunctional forms, gaining an understanding of what roles primary, secondary, tertiary, and quaternary structure play in the cellular milieu [4,5,6]. Biophysical approaches like NMR and X-ray crystallography heavily depend on protein stability and purity [7, 8]. X-ray and cryo-electron microscopy are limited to giving static snapshots of protein structure [9]. Although these biophysical tools have been proven to yield incredible amounts and quality of information, mass spectrometry has unique advantages compared to other methods due to its ability to rapidly probe heterogeneous proteins and protein complexes as well as being able to confirm protein identity, binding, and interactions [10, 11].
Cytochrome c is a well-studied, heme-containing protein that aids in the electron transport chain [12, 13]. Cytochrome c’s folding mechanism, as measured by techniques such as tryptophan fluorescence, UV circular dichroism, and hydrogen exchange [14, 15], has multiple folding phases, resulting in many different short-lived (ms time range) intermediates [16]. Methods like hydrogen exchange labeling have helped to characterize some of these folding intermediates [17]. Previous trends show that the amide groups on the N and C termini of folded structures are the first regions to show protection against hydrogen exchange, indicating the development of secondary structure [17]. Native (i.e., nondenaturing) electrospray (ESI) mass spectrometry [18] has also been applied to the study of cytochrome c structure but in the gas phase. Zhang et al. sequenced ubiquitin and cytochrome c native-like charge states from ESI from 100mM ammonium acetate solutions by electron capture dissociation (ECD) and determined that the native-like charge states fragment only in regions of higher structural dynamicity (i.e., regions with high crystallographic B factors) [19, 20] due to the tendency of ECD to not disrupt intramolecular noncovalent bonding such as hydrogen bonding and salt bridges or fragile covalent bonds [21, 22]. Additionally, mass spectrometric studies utilizing online rapid mixing devices [23,24,25] have been used to control the timescales and extent of protein unfolding and refolding to study folding intermediate conformations and kinetics [24]. However, instead of changing the mixing time scale, varying the concentration of denaturant can also be used to direct protein unfolding. When paired with ion mobility spectrometry (IMS) and ECD, such a method of protein unfolding could give a localized picture of protein unfolding and refolding structural details.
IM and ECD paired with ESI-MS have been used to evaluate to what extent gas-phase ions retain their native solution-phase structures [26,27,28,29,30]. ESI, as a soft ionization technique, does not impart excess energy into ions as they transfer into the gas phase, allowing noncovalent interactions to be preserved [26, 31, 32]. However, there are still questions about to what extent, once in the instrument, the gas-phase ions retain native structure, especially on various instrumentation platforms that may or may not, by their nature, impart internal energy into protein ions [33]. Here, we use a rapid mixing device coupled with IMS and ECD for controlled protein unfolding to characterize the transfer of native-like, partially folded, and unfolded structures into the gas phase. We also measured the IMS arrival time distributions (ATDs) and ECD patterns of ions being transferred under lower energy or higher energy conditions through the instrument to assess the effects of gas-phase activation on folded, unfolded, and intermediate structures.
Methods/experimental
Reagents and materials
Cytochrome c from equine heart (purchased from Sigma-Aldrich St. Louis, MO) solutions was prepared at 10μM in Milli-Q water (approximately pH 6.98) and stored at 20°C. Acidic solutions were made at 1, 2.5, 5, and 10 volume percent glacial acetic acid (purchased from Fischer Scientific Fairmont, NJ) in Milli-Q water corresponding to pH values 2.57, 2.35, 2.19, and 1.96, respectively, for online mixing with aqueous cytochrome c solutions.
Experimental design
The experimental arrangement was adapted from the time-resolved ESI setup developed by Konermann and coworkers [25, 34, 35]. Aqueous cytochrome c and the various acidic solutions were infused by two separate syringe pumps at 5 μL/min into a three-way low pressure mixing tee (Idex P-714, Oak Harbor, WA) using 70 cm fused silica lines (75.5 μm ID and 357.4 μm OD purchased from Polymicro Technologies, Phoenix, AZ) as depicted in Fig. 1, with a metal union in the acidic solution line for the application of high voltage. After mixing, the solution traversed a third 7 cm line pulled at the distal end with a butane torch into an emitter for ESI with an applied potential of ~ +3.2 kV. The overall mixing time can be roughly estimated as 1.9 s (0.31 μL volume in the third capillary). IMS and mass analysis were performed with a Waters Synapt G2-Si IMS-MS (Wilmslow, UK) using helium in the trap cell and He cell, nitrogen in the mobility cell, and argon in the transfer cell. To collisionally heat cytochrome c in the gas phase to extended structures, the gas flow rates were lowered from 200 ml/min in the helium cell and 40 ml/min in the mobility cell to 25 ml/min and 7 ml/min, respectively, while keeping the traveling wave amplitude in the mobility cell the same (i.e., 40 V), dramatically increasing the ratio of the electric field to gas number density (E/N), and thus, the ion temperatures [36]. Collisional heating was observed by the detection of a single narrow feature in the mobility cell, suggesting an extended structure. We refer to these lower-pressure, higher E/N conditions as “harsh” conditions, and the higher-pressure, lower E/N conditions as “gentle” conditions. Under each different condition, each cytochrome c charge state was m/z selected before mobility separation and then subjected to ECD (eMSion Corvallis, OR) after mobility selection. ECD settings were optimized for maximum electron capture and minimum collisional heating. In the ECD cell, we observed that increasing the LM5 voltage decreases collisional heating, observed by the elimination of the protonated heme peak presumably produced by collision induced dissociation (CID). Therefore, we increased the voltage difference between the LM5 and L6 lenses in the ECD cell to slow the ions. The ECD settings were also optimized for each different instrument setting. Under gentle conditions, the ECD lenses used much lower voltages of L1 0.0, L2 10.0, LM3 3.0, L4 7.0, FB 3.5, LM5 5.0, L6 15.0, and L7 15.0, while harsh instrument conditions used lens voltages of L1 −23.3, L2 −17.0, LM3 −0.4, FB 8.6, LM5 13.0, L6 27, and L7 27.
Data analysis
ECD data for each charge state under each different condition were collected in triplicate. After fragmentation, c and z ions were annotated by LCMS Spectator using a 20 ppm mass error [37] and then manually verified for each replicate. Fragments observed in less than two spectra were not counted. The filtered sequence fragments for each charge state under each condition were visualized in Prosight Lite [38].
Results and discussion
Charge state distributions
The ESI spectrum of cytochrome c from neutral (pH 6.98) water, for both harsh and gentle instrument conditions, displayed low charge states: 6+, 7+, 8+, and 9+ (Fig. 2c, f). Folded protein conformations are represented by the lowest charge states [39]. Therefore, the charge states from ESI of cytochrome c under neutral solution conditions correlate with compact, folded protein conformations. As expected, the observed charge state distribution was not dramatically changed between harsh and gentle instrument conditions since these are post-ionization changes. However, the signal-to-noise ratio was noticeably lower for the gentler ion transmission settings, as is typically observed when trying to reduce ion heating [40]. When the acidity of the mixing solution was 5% acetic acid (pH 2.19, pH of post-mixing water/acidic solution roughly 2.35), under both gentle and harsh instrument conditions, the charge state distribution shifts to include higher charge states of 10–18+ (Fig. 2b and e). In these low pH conditions, the unfolded protein is likely ionized by the chain ejection mechanism [41], resulting in the new distribution of higher charge states [42]. The lowest charge states, 6+ and 7+, substantially decrease, correlating with a decrease in the abundance of the native state. However, the presence of the abundant 9+ charge state indicates that a dominant compact species, though possibly a different structure than the native state, is present with the unfolded state. This specific bimodal distribution reflects a transition from the native state into at least two states, one more extended than the other, which correlates with other methods, such as Soret absorption, fluorescence, circular dichroism, and small-angle X-ray scattering [43], as well as other ESI measurements of charge state distribution changes upon decreasing pH [34, 44]. These other measurements show that the populations of acid-induced unfolded and molten globular (i.e., compact, non-native) structures are maximized at approximately pH 2. It is also apparent that the higher E/N of the harsher conditions results in significant CID of the highest charge states, evidenced by the reduction in the signal of the highest charge states and the appearance of many additional peaks that are not precursor ion charge states. The most acidic condition, 10% acetic acid (pH 1.96, pH of post-mixing water/acidic solution roughly 2.19) under both gentle and harsh conditions, shows a similar charge state distribution (Fig. 2a and d). However, the most abundant charge state shifts from the 9+ charge state to the 10+ charge state under the lower pressure conditions and the 15+ charge state in the higher-pressure condition, evidence of a shift in the overall population of solution structures to more unfolded structures. The differences in the harsh and gentle spectra with 10% acetic acid are again attributed to additional ion heating and CID at higher E/N. Mixing the protein with 1% and 2.5% acetic acid (pH 2.57 and 2.35, respectively, with pH of post-mixing water/acidic solution roughly 2.87 and 2.57), did not show major shifts in the charge state distribution. Therefore, experiments run in water and mixed with 5%, and 10% acetic acid are the only ones discussed in this work. We refer to the mixtures of the aqueous protein with 5% and 10% acid via the mixing tee by their final mixed pH values throughout the remainder of the text.
Ion mobility
The ATDs of proteins represent the overall shape/size of kinetically-trapped conformations if care is used to minimize collisional or thermal activation [4]. The various peaks in ATDs correlate with the different conformations present. Multiple broad, unresolved features indicate many conformations, while narrow peaks indicate only one conformational family. Cytochrome c 9+ is present in each solution and instrumental condition tested and thus can be used as a representative charge state for probing the effects of acid-induced unfolding on IMS ATDs. Under gentle conditions, the ion mobility peaks for each acidic condition tested show broad peaks representing multiple conformations. Since the 9+ charge state is one of the most abundant charge states for each solution condition, it is expected to show evidence of molten globular [44] or other intermediate states in equilibrium with the most compact states. For example, in Fig. 3, the mobility spectra for cytochrome c 9+ in water (f) show multiple broad features. However, the mobility peaks for pH 2.35 and pH 2.19 (Fig. 3e and d) have more abundant extended features (above 10 ms), due to the increase in more extended states. The many 9+ features correlate with solution measurements of cytochrome c refolding having multiple folding intermediates [45]. Under the harsh instrument conditions, the mobility peaks in Fig. 3a, b, and c mobility spectra for the 9+ charge state are all narrow peaks, showing only one major feature with a small shoulder. The narrow peaks for each of the solution conditions correlate with the harsh instrument conditions resulting in the collisional heating of the protein to fewer conformations. Since the ATDs from the different solution conditions are virtually identical under harsh conditions, these ATDs likely cannot inform on the solution distribution of structures.
The other charge states of cytochrome c also show similar trends in their ion mobility spectra. The 7+ charge state under gentle conditions has a single broad mobility peak when electrosprayed from neutral pH, indicating this lower charge state presents an ensemble of compact states in the gas phase (Fig. S1a). Cytochrome c 8+ ionized from pure water results in a similar ATD to the 7+ under gentle conditions (Fig. S1b). When the pH is reduced to 2.35, the 8+ ions shift mostly to more extended distributions, showing unfolding behavior (Fig. S1c). At pH 2.19, the ATD shows a high-intensity, broad peak at ~7.72 ms (Fig. S1d), indicating that a significant portion of the cytochrome c population refolds, likely into the molten globular state. Higher charge states (11+ and 15+) are only observed at pH 2.35 and pH 2.19. These charge states lead to narrower distributions under both solution conditions (Fig. S1e–h), indicative of extended cytochrome c structures. Overall, these ATDs are in excellent agreement with ATDs measured with similar instrumentation for acid-induced cytochrome c unfolding [25].
Under harsh conditions for each different acidic condition, the other charge states (e.g., 7+, 8+, 11+) (Fig. S2) all have singular, narrow peaks, following the trend of the 9+. This means that each of the cytochrome c charge states is collisionally heated to more extended structures under harsh instrument conditions. These gas-phase conformers, as represented by their ATDs, no longer correlate with the ionization solution pH. The presence of singular, narrow peaks for gas-phase collisionally heated cytochrome c ions agrees with the ATDs at the highest collisional energies for other examples of the gas-phase collisional heating of cytochrome c ions in the literature [46, 47].
ECD under gentle conditions
While the ATDs reflect the overall gas-phase structures of cytochrome c, ECD is used as a higher structural resolution, albeit indirect, measurement of the changes in cytochrome c structure. Regional increases in fragmentation are suggestive of unfolding and refolding into structures that have lost native contacts in these areas. The cytochrome c heme is covalently bound to cysteines 14 and 17, and the heme iron is very weakly bound to methionine 80 [13, 48, 49]. Therefore, we expect that natively folded cytochrome c structures would show limited fragmentation between cysteine 14 and methionine 80, since even if the protein backbone is fragmented by ECD, the bonds to the heme and iron bind fragments in this region together, preventing the detection of fragment ions by their unique m/z. Under gentle instrument conditions, the 9+ charge state shows ECD almost exclusively N-terminal to cysteine 14 and C-terminal to methionine 80, correlating with a structure similar to the natively folded solution structure (Fig. 4). However, ECD sequence coverage increases from 10% sequence coverage to 35% as the acidity increases from water to pH 2.19. Figure 4b (pH 2.35) shows several fragments between cysteine 14 and methionine 80, whereas Fig. 4c (pH 2.19) shows extensive fragmentation throughout the protein. These results correlate with the loss of the methionine 80/iron ligand bond dissociation upon the addition of acid, supporting evidence from fluorescence spectroscopy that the acidified cytochrome c loses the iron to methionine 80 bond [15]. The increase in sequence coverage from pH 2.35 to pH 2.19 indicates the loss of native salt bridges, hydrogen bonds, or other intramolecular interactions with the additional decrease in pH. Specifically, there is extensive fragmentation observed between lysine 79 and aspartic acid 93. These residues do not directly interact in the cytochrome c crystal structure as they are distant from each other [50]. This likely indicates that this putative salt bridge [51] that is disrupted between pH 2.35 and pH 2.19 is a new intramolecular bond formed after unfolding and refolding into an intermediate structure in the pH 2.35 acid solution. Similarly, there is a marked increase in fragmentation for pH 2.19 versus pH 2.35 between lysine 53 and glutamic acid 69. These residues are much closer in the crystal structure. Further examination reveals that the lysine side chain is hydrogen bound to the backbone carbonyl oxygen of tyrosine 74 under native conditions. Therefore, it is likely that this intramolecular bond may be preserved at pH 2.35 but is lost upon acidifying to the pH 2.19 condition, allowing observation of fragments in this region.
The lower charge states in water, such as the 7+ charge state, had ATDs that showed evidence of compact gas-phase structures, which we expect would correlate with the retention of an increased number of native contacts. The ECD sequence coverage of 14% from the 7+ charge state, also covering only from the N-terminal to cysteine 14 and C-terminal to methionine 80 (Fig. S3), correlates to many stabilizing intramolecular interactions, similarly to what was observed for 9+. The most dominant charge state when electrosprayed from water, 8+, has a sequence coverage of 23% with fragmentation focused around both termini (Fig. S4). The terminal regions of the protein have the highest B-factor in the crystal structure (i.e., the most flexible regions) and are expected to fragment readily even for the native structure [19]. As the acidity, and therefore protein unfolding, increases, the 7+ charge state is no longer present and the 8+ charge state abundance decreases so significantly that ECD fragments are no longer detectable above the noise baseline. As the protein intramolecular bonds are disrupted more to produce unfolded higher charge states, the ECD expands from just the N-terminal and C-terminal fragments to between cysteine 14 and 80. The higher charge states such as 11+ and 15+ have extensive fragmentation in the interior region of the protein (Figs. S5 and S6). At pH 2.35 and pH 2.19, cytochrome c 15+ has ECD sequence coverage of 44% and 54%, respectively, with fragments throughout the protein, reflecting a trend of increasing ECD coverage as the acidity increases. As the protein unfolds in acetic acid, the higher charge states’ increased ECD sequence coverage correlates with the loss of native contacts and iron to methionine 80 binding (Figs. S5 and S6).
ECD under harsh conditions
When the instrument is operated under lower pressure/harsh conditions, similar to the gentle conditions, there is an increased sequence coverage as the acidity of the denaturing conditions increases (Fig. 5). However, for the 9+ charge state, all solution conditions analyzed under harsh parameters exhibit much more fragmentation than any of the solution states fragmented by ECD with gentle instrument conditions. This is likely due to the disruption of intramolecular interactions due to extensive ion heating [52], which was also observed from the ATDs under these harsh conditions. Cytochrome c 9+ ECD sequence coverage increases from 54% in water to 72% at pH 2.35 and pH 2.19 as the ion heating further unfolds the protein compared to only solution unfolding before analysis with ion transmission under gentle conditions (10%, 15%, and 35%, respectively). Instead of exclusively fragmenting from the N-terminus to cysteine 14 and C-terminus to methionine 80 in the pH 6.98 mixtures, ECD occurs throughout the protein. This correlates with the ion heating under the harsh instrument conditions resulting in extensive unfolding since all three solvent conditions show extensive fragmentation. The 11+ charge state has ECD sequence coverage that is virtually identical in both acidic conditions, from 66% at pH 2.35 to 68% at pH 2.19 (Fig. S5). The fragmentation under harsh conditions for the lower charge states also does not follow the same fragmentation patterns as under gentle conditions. The more folded 7+ and 8+ charge states in water both show sequence coverage of 57% in neutral solutions (Figs. S3 and S4), indicating gas-phase unfolding. However, the 15+ charge state was not transferred through the instrument in high enough abundance to observe under harsh conditions.
Discussion of solution versus gas-phase unfolding
Our results suggest that solution and gas-phase unfolding pathways can be non-identical. Again, gas-phase collisional heating gives more extensive ECD than does solution unfolding by acidification with unfolded ions transmitted under gentle conditions. For example, the 7+ charge state, which is representative of the most compact solution states when electrosprayed from pure water, gives much higher sequence coverage upon ECD (85%) when transmitted under harsh conditions than the 11+ charge state at pH 2.19 (35%) transmitted under gentle conditions. This suggests that the acid-unfolded structures of the higher charge states are in fact not identical to the structures produced by collisional heating of even the most compact states.
Ruotolo et al. used collisional heating (collision induced unfolding (CIU)) of the lowest charge states to show that the number of foldable domains in solution is equal to the number of unfolding transitions by both solution and gas-phase unfolding [53]. Although our data is in general in agreement with this study, the gas-phase unfolded structures gave different ECD coverage than the solution-unfolded structures, except for the highest charge state we monitored (15+). This is likely due to processes that occur in the picoseconds after ESI during desolvation into gaseous protein ions as modeled by molecular dynamics [54]. One of the observations from molecular dynamics of proteins emerging from ESI droplets was that immediately after desolvation, charged side chains form new salt bridges and hydrogen bonds, giving additional stability to the nascent gas-phase structures. It is also reasonable to expect that acid-induced unfolded structures would become stabilized by these effects, resulting in less sequence coverage by ECD. This is likely especially true of more linear, extended structures produced by CEM [41], with many new charge/charge and charge/dipole interactions that can be formed from the large overall number of charged residues, in balance with coulombic repulsion. Though our results suggest that collisional activation might lead to greater unfolding than solution unfolding, this can alternatively be explained by the formation of new intramolecular interactions for solution-unfolded structures upon dehydration and the dissociation of these and native interactions by collisional heating, increasing ECD sequence coverage for collisionally activated ions compared to solution-unfolded, thermalized ions.
However, intermediate charge states, such as the 9+ charge state, are produced by ESI from pure water as well as by acidic conditions, which suggests that they are not necessarily produced by CEM. Even under unfolded conditions, the 9+ ions are part of a different, more compact distribution than the more extended, more highly charged ions (as in Fig. 2a, b, d, and e). The 9+ ions have unique ATDs when produced by ionization from acidic and neutral solutions (Fig. 3d–f). Therefore, if the ions are produced primarily by desolvation of the electrospray droplet until a bare ion remains (i.e., CRM) instead of CEM [41], then it is likely that 9+ ions electrosprayed from acidic conditions result in gaseous structures that may resemble acidic solution states. This is supported by ECD of the 9+ ions from the various pH solution conditions correlating with unfolding processes that are measured from solution methods. The 9+ ion unfolds to a much greater extent under collisional heating when ionized from pH 6.98 solutions than do the 9+ ions from low pH solutions when transmitted under gentle conditions. This result is significant as it confirms the work from Ruotolo et al. that gas-phase unfolding of any ions other than the most compact, lowest charged ions can lead to structures that do not represent solution-phase unfolded proteins and in fact may be gas-phase artifacts.
Conclusion
The combination of online mixing with ion mobility spectrometry and ECD was used to identify the structural details of steps in a protein unfolding process. By decreasing the ESI pH, the charge state distribution for cytochrome c shifted to higher charge states that indicate the presence of more extended structures. Ion mobility spectra of the various charge states under gentle instrument conditions showed evidence for multiple protein conformations in their ATDs. For the lowest charge states, broad peaks correlated to the coexistence of several compact, native-like states when ionized from water, but additional, more extended features were observed upon acidification. The 9+ ions, having intermediate charge, as well as the highest charge states, resulted in multiple peaks at various levels of acidification indicating various stages of protein unfolding. When using harsh instrument conditions, ion mobility spectra contained singular narrow peaks due to the protein’s unfolding into more extended conformers by collisional activation.
ECD of the various charge states of cytochrome c reflected differences in structure induced by acidification. The charge states observed from pure water showed fragmentation only at the flexible termini of the proteins but not within the interior of the protein, bound together by intramolecular interactions and from binding to the iron by methionine 80 and the heme by cystine 14 and 16. Upon acidification, ECD reflected the loss of the iron-methionine 80 bond as well as other native and non-native contacts. However, under harsh conditions, fragmentation was drastically different, showing extensive sequence coverage for all charge states regardless of the ESI solution pH.
Several important conclusions can be drawn from these studies. First, the structures at low pH probed by IMS/MS under gentle conditions are not completely unfolded structures, as neither the ATDs nor ECD fragments from harsh conditions reflect the unfolded protein ions examined under gentle conditions. Therefore, these experiments demonstrate the utility of native IMS/MS methods for characterizing protein folding intermediates under the timescale of the IMS measurement. Second, we note that our unfolding measurements, along with the measurements of others, show that tuning mass spectrometry conditions for maximum transmission leads to the unfolding of the protein in the gas phase into structures that likely have little biological relevance. Thus, echoing the work of others, we recommend that special care be given to the parameters used to study the protein unfolding and other dynamics by native IMS/MS methods. Additionally, gas-phase unfolding experiments should be performed only on the most compact, lowest charge state ions if correlations are being made with solution unfolding. Finally, though this study serves as a proof of concept with a model protein, we will apply our method to larger proteins, protein–protein complexes, and dynamic and disordered systems.
References
Lossl P, van de Waterbeemd M, Heck AJ. The diverse and expanding role of mass spectrometry in structural and molecular biology. EMBO J. 2016;35(24):2634–57.
Hodge EA, Benhaim MA, Lee KK. Bridging protein structure, dynamics, and function using hydrogen/deuterium-exchange mass spectrometry. Protein Sci. 2020;29(4):843–55.
Sinz A, Arlt C, Chorev D, Sharon M. Chemical cross-linking and native mass spectrometry: a fruitful combination for structural biology. Protein Sci. 2015;24(8):1193–209.
Clemmer DE, Russell DH, Williams ER. Characterizing the conformationome: toward a structural understanding of the proteome. Acc Chem Res. 2017;50(3):556–60.
Dill KA, MacCallum JL. The protein-folding problem, 50 years on. Science. 2012;338(6110):1042–6.
Shen H-B, Chou K-C. QuatIdent: a web server for identifying protein quaternary structural attribute by fusing functional domain and sequential evolution information. J Prot Res. 2009;8(3):1577–84.
Chavez JD, Bruce JE. Chemical cross-linking with mass spectrometry: a tool for systems structural biology. Curr Opin Chem Biol. 2019;48:8–18.
Yee AA, Savchenko A, Ignachenko A, Lukin J, Xu X, Skarina T, et al. NMR and X-ray crystallography, complementary tools in structural proteomics of small proteins. J Am Chem Soc. 2005;127(47):16512–7.
Wells JN, Marsh JA. Experimental characterization of protein complex structure, dynamics, and assembly. In: Marsh JA, editor. Protein Complex Assembly: Methods and Protocols. New York, NY: Springer New York; 2018. p. 3–27.
Kaltashov IA, Eyles SJ. Studies of biomolecular conformations and conformational dynamics by mass spectrometry. Mass Spectrom Rev. 2002;21(1):37–71.
Smith DP, Giles K, Bateman RH, Radford SE, Ashcroft AE. Monitoring copopulated conformational states during protein folding events using electrospray ionization-ion mobility spectrometry-mass spectrometry. J Am Soc Mass Spectrom. 2007;18(12):2180–90.
Ow Y-LP, Green DR, Hao Z, Mak TW. Cytochrome c: functions beyond respiration. Nat Rev Mol Cell Biol. 2008;9(7):532–42.
Stevens JM, Mavridou DA, Hamer R, Kritsiligkou P, Goddard AD, Ferguson SJ. Cytochrome c biogenesis System I. FEBS J. 2011;278(22):4170–8.
Colón W, Elöve GA, Wakem LP, Sherman F, Roder H. Side chain packing of the N- and C-terminal helices plays a critical role in the kinetics of cytochrome c folding. Biochem. 1996;35(17):5538–49.
Englander SW, Sosnick TR, Mayne LC, Shtilerman M, Qi PX, Bai Y. Fast and slow folding in cytochrome c. Accounts of Chemical Research. 1998;31(11):737–44.
Travaglini-Allocatelli C, Gianni S, Brunori M. A common folding mechanism in the cytochrome c family. Trends Biochem Sci. 2004;29(10):535–41.
Roder H, Elove GA, Englander SW. Structural characterization of folding intermediates in cytochrome c by H-exchange labelling and proton NMR. Nature. 1988;335(6192):700–4.
Leney AC, Heck AJR. Native mass spectrometry: what is in the name? J Am Soc Mass Spectrom. 2017;28(1):5–13.
Zhang H, Cui W, Gross ML. Native electrospray ionization and electron-capture dissociation for comparison of protein structure in solution and the gas phase. Int J Mass Spectrom. 2013;354-355:https://doi.org/10.1016/j.ijms.2013.06.019.
Fenn LS, Kliman M, Mahsut A, Zhao SR, McLean JA. Characterizing ion mobility-mass spectrometry conformation space for the analysis of complex biological samples. Anal Bioanal Chem. 2009;394(1):235–44.
Zubarev RA, Kelleher NL, McLafferty FW. Electron capture dissociation of multiply charged protein cations. A nonergodic process. J Am Chem Soc. 1998;120(13):3265–6.
Breuker K, Oh H, Lin C, Carpenter BK, McLafferty FW. Nonergodic and conformational control of the electron capture dissociation of protein cations. Proc Natl Acad Sci USA. 2004;101(39):14011–6.
Wilson DJ, Konermann L. A capillary mixer with adjustable reaction chamber volume for millisecond time-resolved studies by electrospray mass spectrometry. Anal Chem. 2003;75(23):6408–14.
Zinck N, Stark AK, Wilson DJ, Sharon M. An improved rapid mixing device for time-resolved electrospray mass spectrometry measurements. Chemistryopen. 2014;3(3):109–14.
Liuni P, Deng B, Wilson DJ. Comparing equilibrium and kinetic protein unfolding using time-resolved electrospray-coupled ion mobility mass spectrometry. Analyst. 2015;140(20):6973–9.
Wyttenbach T, Bowers MT. Structural stability from solution to the gas phase: native solution structure of ubiquitin survives analysis in a solvent-free ion mobility-mass spectrometry environment. J Phys Chem B. 2011;115(42):12266–75.
Breuker K, Oh H, Horn DM, Cerda BA, McLafferty FW. Detailed unfolding and folding of gaseous ubiquitin ions characterized by electron capture dissociation. J Am Chem Soc. 2002;124(22):6407–20.
Skinner OS, McLafferty FW, Breuker K. How ubiquitin unfolds after transfer into the gas phase. J Am Soc Mass Spectrom. 2012;23(6):1011–4.
Ruotolo BT, Giles K, Campuzano I, Sandercock AM, Bateman RH, Robinson CV. Evidence for macromolecular protein rings in the absence of bulk water. Science. 2005;310(5754):1658–61.
Ruotolo BT, Robinson CV. Aspects of native proteins are retained in vacuum. Curr Opin Chem Biol. 2006;10(5):402–8.
Robinson CV, Chung EW, Kragelund BB, Knudsen J, Aplin RT, Poulsen FM, et al. Probing the nature of noncovalent interactions by mass spectrometry. A study of protein-CoA ligand binding and assembly. J Am Chem Soc. 1996;118(36):8646–53.
Seo J, Hoffmann W, Warnke S, Bowers MT, Pagel K, von Helden G. Retention of native protein structures in the absence of solvent: a coupled ion mobility and spectroscopic study. Angew Chem Int Ed. 2016;55(45):14173–6.
Merenbloom SI, Flick TG, Williams ER. How hot are your ions in TWAVE ion mobility spectrometry? J Am Soc Mass Spectrom. 2012;23(3):553–62.
Konermann L, Collings BA, Douglas DJ. Cytochrome c folding kinetics studied by time-resolved electrospray ionization mass spectrometry. Biochemistry. 1997;36(18):5554–9.
Konermann L, Pan J, Wilson DJ. Protein folding mechanisms studied by time-resolved electrospray mass spectrometry. Biotechniques. 2006;40(2):135–41.
Revercomb HE, Mason EA. Theory of plasma chromatography/gaseous electrophoresis. Review. Anal Chem. 1975;47(7):970–83.
Park J, Piehowski PD, Wilkins C, Zhou M, Mendoza J, Fujimoto GM, et al. Informed-proteomics: open-source software package for top-down proteomics. Nat Methods. 2017;14(9):909–14.
Fellers RT, Greer JB, Early BP, Yu X, LeDuc RD, Kelleher NL, et al. ProSight Lite: graphical software to analyze top-down mass spectrometry data. Proteomics. 2015;15(7):1235–8.
Kaltashov IA, Mohimen A. Estimates of protein surface areas in solution by electrospray ionization mass spectrometry. Anal Chem. 2005;77(16):5370–9.
Chen S-H, Russell DH. How closely related are conformations of protein ions sampled by IM-MS to native solution structures? J Am Soc Mass Spectrom. 2015;26(9):1433–43.
Konermann L, Metwally H, Duez Q, Peters I. Charging and supercharging of proteins for mass spectrometry: recent insights into the mechanisms of electrospray ionization. Analyst. 2019;144(21):6157–71.
Konermann L, Douglas DJ. Acid-induced unfolding of cytochrome c at different methanol concentrations: electrospray ionization mass spectrometry specifically monitors changes in the tertiary structure. Biochem. 1997;36(40):12296–302.
Goto Y, Hagihara Y, Hamada D, Hoshino M, Nishii I. Acid-induced unfolding and refolding transitions of cytochrome c: a three-state mechanism in water and deuterium oxide. Biochem. 1993;32(44):11878–85.
Rob T, Wilson DJ. A versatile microfluidic chip for millisecond time-scale kinetic studies by electrospray mass spectrometry. J Am Soc Mass Spectrom. 2009;20(1):124–30.
Winkler JR. Cytochrome c folding dynamics. Curr Opin Chem Biol. 2004;8(2):169–74.
Yin V, Konermann L. Probing the effects of heterogeneous oxidative modifications on the stability of cytochrome c in solution and in the gas phase. J Am Soc Mass Spectrom. 2021;32(1):73–83.
Borotto NB, Osho KE, Richards TK, Graham KA. Collision-induced unfolding of native-like protein ions within a trapped ion mobility spectrometry device. J Am Soc Mass Spectrom. 2022;33(1):83–9.
Fisher WR, Taniuchi H, Anfinsen CB. On the role of heme in the formation of the structure of cytochrome c. J Biol Chem. 1973;248(9):3188–95.
Xu Y, Mayne L, Englander SW. Evidence for an unfolding and refolding pathway in cytochrome c. Nat Struct Biol. 1998;5(9):774–8.
Bushnell GW, Louie GV, Brayer GD. High-resolution three-dimensional structure of horse heart cytochrome c. J Mol Biol. 1990;214(2):585–95.
Zhang Z, Vachet RW. Gas-phase protein salt bridge stabilities from collisional activation and electron transfer dissociation. Int J Mass Spectrom. 2017;420:51–6.
Molano-Arevalo JC, Jeanne Dit Fouque K, Pham K, Miksovska J, Ridgeway ME, Park MA, et al. Characterization of intramolecular interactions of cytochrome c using hydrogen-deuterium exchange-trapped ion mobility spectrometry-mass spectrometry and molecular dynamics. Anal Chem. 2017;89(17):8757–65.
Zhong Y, Han L, Ruotolo BT. Collisional and coulombic unfolding of gas-phase proteins: high correlation to their domain structures in solution. Angew Chem Int Ed Engl. 2014;53(35):9209–12.
Breuker K, McLafferty FW. Stepwise evolution of protein native structure with electrospray into the gas phase, 10(-12) to 10(2) s. Proc Natl Acad Sci U S A. 2008;105(47):18145–52.
Acknowledgements
The authors would like to acknowledge Professor Derek Wilson and his research group for helpful discussions.
Funding
Portions of this work were funded by the National Science Foundation CAREER CHE-2143755.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
ABC Highlights: authored by Rising Stars and Top Experts.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 517 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cain, R.L., Webb, I.K. Online protein unfolding characterized by ion mobility electron capture dissociation mass spectrometry: cytochrome C from neutral and acidic solutions. Anal Bioanal Chem 415, 749–758 (2023). https://doi.org/10.1007/s00216-022-04501-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-022-04501-w