Abstract
The black-legged tick, Ixodes scapularis, is a well-known vector for the Lyme disease-causing pathogen (Borrelia burgdorferi) but can also carry other disease-causing pathogens such as Rickettsia, Anaplasma, Bartonella, Ehrlichia, and Theileria. Hence, tick screening using highly specific protein signatures for specific pathogens will help assess the prevalence of infected ticks and understand the pathogen-tick interactions in a specific geographic area. In this study, we used data-dependent acquisition to key pathogen protein signatures in black-legged ticks collected from the Southern Tier New York. Bottom-up proteomic analysis of extract from five combined ticks identified 2,052 tick proteins and 41 pathogen proteins with high confidence (≥ 99% C.I.). Results show high peptide spectral match counts for Rickettsia species and Borrelia species and lower counts for other rarer pathogens such as Anaplasma phagocytophilum. Parallel reaction monitoring performed on protein extracts from individual ticks (n = 10) revealed that 8 out of the 10 screened ticks carried Rickettsia species, 5 carried Borrelia species, 3 carried both pathogens, and only 1 tick carried no detectable bacteria. Mass spectrometry-based proteomics is a highly specific way to define the expression of different types of pathogen proteins in infected ticks. This might bring insights into the tick-pathogen interactions at the molecular level and especially expression pathogen surface proteins in ticks.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ixodes scapularis, commonly known as the black-legged tick or a deer tick, is a vector for several disease-causing pathogens that can affect both animals and humans [1]. Lyme disease is one of the commonly reported tick-borne illnesses caused by the spirochete Borrelia burgdorferi that is transmitted to humans through a tick bite [2]. Its prevalence across New York State and the United States (USA) poses a serious public health threat as new cases increase annually [3]. Climate change and global warming encourage geographical expansion of ticks throughout the United States and will therefore incite more tick-borne diseases in the coming years [4]. Although B. burgdorferi is the most known tick-borne pathogen to cause Lyme disease, other pathogens such as Rickettsia, Babesia, and Anaplasma species are also prevalent among tick populations and cause other serious diseases such as babesiosis, ehrlichiosis, Rocky Mountain spotted fever and anaplasmosis [5,6,7].
The natural life cycle of the I. scapularis tick and B. burgdorferi bacteria involves a complex relationship between small animals (i.e., mice, rabbits, squirrels, birds) and larger mammals (i.e., dogs, horses, deer) [8]. The natural reservoir for B. burgdorferi is the white-footed mouse, which transmits the pathogen to feeding I. scapularis ticks in the larval or nymphal stages. It is then transmitted to other larger mammals such as dogs, deer, and humans during the nymphal or adult life stages of the tick [9,10,11,12].
I. scapularis ticks also carry other bacterial pathogens such as Rickettsia, Bartonella, and Anaplasma; protozoan pathogens such as Babesia microti and Theileria species; and viruses such as the Powassan virus, Heartland virus, and Bourbon virus [13]. Rickettsiae organisms are intracellular pathogens that infect endothelial cells in vertebrates and include Rickettsia and Ehrlichia species (spp.). Rickettsia spp. can cause spotted fevers with a mortality rate of up to 25% if not treated early and a typhus rash. Ehrlichia spp. can cause ehrlichiosis that is like spotted fever [14]. Anaplasma phagocytophilum is a gram-negative bacterium that can also be carried by the I. scapularis ticks. Anaplasma species is genetically similar to Rickettsia species and causes flu-like symptoms in early infection stages. If left untreated, it can lead to respiratory issues, organ failure, and even death [15]. In a genetic study, 97% of female I. scapularis ticks collected in Massachusetts tested positive for Rickettsia species and 10% tested positive for A. phagocytophilum [16]. Because of the low A. phagocytophilum infection rate in I. scapularis ticks, there is also a low infection rate of humans with a maximum yearly infection rate of 5,762 in 2017 according to the CDC [17]. Hence, tick surveillance and screening for pathogen species are needed for effective tick control and impact on human health. Pathogens carried by ticks can be extremely devastating and deadly if not correctly identified and quickly treated.
The commonly used method to screen ticks for potential pathogens is polymerase chain reaction (PCR). A recent study utilizing a PCR assay targeting pathogen-specific genes successfully screened 459 I. scapularis ticks to determine the prevalence of infection and co-infection in ticks [18]. The results showed prevalence of Borrelia burgdorferi (47%), Anaplasma phagocytophilum (6.9%), Babesia microti (4.7%), Borrelia miyamotoi (7.3%), and Rickettsial endosymbionts (93.3%) [18]. Although PCR is a highly sensitive method for tick surveillance, detection of these pathogens at the protein levels might bring insight into the pathogen’s antigen expression within a tick and perhaps develop future assays to screen for these same antigens in infected humans and other mammals.
Advances in mass spectrometry and proteomic methods have started shedding light on tick-borne pathogens, their molecular signatures, and their host-dependent protein expression [19,20,21]. Matrix-assisted laser desorption-time of flight mass spectrometry (MLADI-TOF–MS) was successfully used to detect pathogens in ticks based on pathogen-specific polypeptide signatures [19]. Although the technique is rapid, further characterization and sequencing of the pathogen’s polypeptide might be more informative and might bring insights into types of pathogen protein expressed in infected ticks.
Single tick screening for potential pathogens in a given geographic area is highly important to human and animal health within that area. It would also aid in understanding the life cycle of tick-borne pathogens and their interaction with the tick vector, and eventually aid in developing direct diagnostic biomarkers for different pathogens and treatment decision-making based on the presence of specific pathogen antigens in bodily fluids of acutely infected patients [22]. In this study, we focused on surveying tick-borne pathogens in I. scapularis ticks collected from Chenango Valley State Park located in Broome County, New York, an area where Lyme disease is highly prevalent and remains challenging to diagnose and properly treat. Subsequently, we developed a parallel reaction monitoring method to screen single ticks and determine their pathogen load status. This might raise awareness about tick-borne disease among the community in the area and broader scientific community.
Materials and methods
Tick collection and processing
Ixodes scapularis ticks were collected at Chenango Valley State Park in the spring and summer of 2018 and 2019. A 1-m-squared corduroy drag was used on commonly travelled nature paths in the park to collect questing ticks. Latched ticks were carefully picked using tweezers and individually transferred into 1.5-mL Eppendorf tubes containing 1 mL of methanol. Storage in alcohol was reported to maintain the integrity of the tick for a long period if stored refrigerated for subsequent pathogen screening using PCR methods which is more vulnerable to degradation than proteins [23,24,25]. Only female, adult I. scapularis ticks were used for analysis as they are larger and more abundant during dragging durations and have the probability of carrying more diseases due to an additional blood meal compared to larval and nymphal ticks.
For tick processing and protein extraction, excess methanol was removed from each tube followed by a very brief drying at 95 °C (< 1 min) on a heat block to evaporate residual methanol and facilitate tick cuticula crushing. Individual ticks were grinded using a handheld mortar and pestle, and 3 × 70µL of ice-cold radioimmunoprecipitation assay (RIPA) buffer with protease inhibitor cocktail (Halt Protease Inhibitor Cocktail, Thermo Fisher Scientific) was added to collect the tick dust. The lysed tick samples were transferred into a clean Eppendorf and placed on ice for 15 min with occasional vortexing before being sonicated for another 2 min. Samples were then centrifuged at 16,000 × g for 30 s and supernatants were collected and transferred into clean 1.5-mL Eppendorf tubes and placed on ice. Protein concentration in each sample was determined using bicinchoninic acid (BCA) assay (Thermo Fisher Scientific). Protein extracts were then flash frozen in liquid nitrogen and stored at − 80 °C until further use.
Sample processing for mass spectrometry analysis
For global proteome profiling and pathogen identification, extracts from 5 ticks were pooled together and a sample aliquot containing 100 µg total protein was transferred to a clean Eppendorf tube. 5 µL of 1% SDS was added to the sample followed by 5 µL of 200 mM dithiothreitol (DTT) to reduce any disulfide bonds and denature the proteins. Then, 5 µL of lithium dodecyl sulfate (LDS) sample buffer was added and the sample was vortexed and placed on a heating block at 95 °C for 5 min. The sample was loaded onto a 3–8% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and separation was performed at 150 V until the samples ran to the bottom of the gel (between 41 and 86 min, depending on the gel type). The gel was then washed with water and stained for 4 to 24 h using G250 Coomassie stain (Bio-Rad), and then de-stained in water for 4 to 24 h before being imaged. The gel lane containing fractionated proteins was sliced into 2- to 3-mm gel slices using a razor blade and a glass marked guide with 4 mm lengths to ensure correct band cut sizes. Each slice was placed in a clean Eppendorf tube; cysteine residues were reduced and alkylated before in-gel digestion and processed for in-gel digestion using an established protocol for Gel-LC–MS/MS-based method [26]. A 50-µL trypsin solution (Promega, Madi-son, WI) was added to each gel slice at a concentration of 12.5 ng/µL in 50 mM ammonium bicarbonate and incubated at 37 °C overnight. The resulting peptides were extracted, dried by vacuum centrifugation, and analyzed by LC–MS/MS as described below.
Mass spectrometry analysis
Data-dependent acquisition and protein identification
The dried peptide samples were resuspended in 10 µL of 0.1% formic acid, and 7µL of this peptide solution was injected onto a Dionex UltiMate 3000 RS UPLC system connected to a Q Exactive™ HF-X Hybrid Quadrupole-Orbitrap™ Mass Spectrometer. Because of the complexity of the tick proteome and the low abundance of the pathogens, a 165-min gradient was used to survey for proteins belonging to a potential pathogen. The mobile phase A consisted of 0.1% formic acid in water, and a mobile phase B consisted of 0.1% formic acid in 80% acetonitrile. An EASY-Spray™ LC Column (75 µm diameter, 50 cm length, pore size 100 Å, particle size 2 μm; Thermo Scientific) column was first equilibrated with 98% mobile phase A and 2% mobile phase B for 8 min, then 5–30% gradient of mobile phase B for 87 min, 30–100% B for 5 min, 100% B for 5 min, 100–2% B for 5 mi, and 2% B for 10 min. Data-dependent acquisition (DDA) was performed at 120,000 resolution for a scan mass range of 375–1600 m/z, with one full MS scan followed by 20 MS/MS scans. DDA was done at 27 eV collision energy, 300 V ion spray, and 35 °C column temperature. Blanks consisting of 0.1% formic acid were run in the beginning and between each sample analysis.
Raw mass spectrometry data files were collected and processed with a specific workflow designed in Proteome Discoverer 2.2 (Thermo Fisher Scientific). Searches were performed against the following FASTA databases (Homo sapiens_SwissProt TaxID = 9606, Ixodes scapularis_Black-legged tick_Deer tick, Borreliella group_64895, spotted fever group_114277, Anaplasma phagocytophilum strain HZ, Ehrlichia muris, Ehrlichia chaffeensis, Theileria annulata UP000001950, Theileria parva UP000001949, Bartonella henselae, Bartonella schoenbuchensis, and Babesia microti UP000002899) downloaded from www.uniprot.org using Sequest HT (Thermo Fisher Scientific) with precursor and fragment mass tolerance set at ± 10 ppm and ± 0.05 Da respectively, partial trypsin digestion, and the following dynamic modifications: carbamidomethyl on cysteine, acetyl on protein N terminus, and oxidation of the methionine residue. A complete Proteome Discoverer workflow and parameter settings are provided under supplemental material S1.
Parallel reaction monitoring
Single ticks were processed for total protein extraction the same as above, except for total protein loaded into the SDS-PAGE which depended on the size of the tick. On average, single ticks yielded total protein extracts ranging from 15 to 50 µg. A total of 8 bands, where the targeted pathogen’s proteins are expected to migrate, were excised from SDS-PAGE of protein extract from a single tick and in-gel digested for LC–MS/MS as described above. Then, 3 µL of each tryptic peptide sample was analyzed in PRM mode using the same collision energy and resolution as used in the DDA analysis. The specific charge, m/z, and retention time window for each peptide were used to build the method (Supplemental material S2). Peptides were targeted with an isolation window of ± 1.2 m/z and a retention time window of ± 5 min. Database search was performed as described above and peptides of interest were targeted and confirmed using Skyline data analysis on the raw MS/MS data files to ensure confidence in selected transition ions and peptide identification. All detectable transition ions for each targeted peptide were used to add confidence in peptide identification.
Western blot analysis of Borrelia burgdorferi’s outer surface protein A
To confirm the presence of B. burgdorferi’s outer surface protein A (OspA) in potentially infected ticks and confirm its approximate molecular mass on the SDS-PAGE gel, total protein extract (about 50 µg per tick) from single ticks (n = 17) was separated by SDPS-PAGE as described above. Gel separated proteins were then transferred to Novex™ nitrocellulose membrane overnight using NuPAGE™ transfer buffer at 0.17Amp for 18 h. The membrane was then incubated overnight with polyclonal rabbit antibody against Borrelia OspA (ab106081, Abcam, MA, USA) at 1:10,000 dilution. The membrane was washed three times for 10 min with phosphate buffer saline + tween (PBST) and incubated for 10 min in blocking buffer (PBST + 5% nonfat milk) before incubation with the donkey monoclonal anti-rabbit secondary antibody at 1:4000 dilution for 30 min. Processed membranes were then washed 3 times for 10 min with PBST and imaging was performed using enhanced chemiluminescence (ECL) reagent.
PCR screening for Rickettsia peptidoglycan-associated protein gene and B. burgdorferi ospA gene in crude lysate from single ticks
Total DNA was extracted from four leftover single tick lysates above using the Qiagen DNeasy Blood & Tissue Kit according to the instructions provided in the manufacturer’s Supplementary Protocol for purification of total DNA from crude lysates (Qiagen, Hilden, Germany). PCR amplification was performed using forward (GCATTATGTGTGCTTGCAGGGTG) and reverse (GTGGTGTACAACTGTTACAGCTCTACG) primers targeting the gene coding for Rickettsia peptidoglycan-associated protein and forward (GGATCTGGAGTACTTGAAGGCG) and reverse (CAACTGCTGACCCCTCTAATTTGGTG) primers targeting the gene coding for B. burgdorferi OspA protein. The PCR reaction consisted of 1.5 mM MgCl2, 0.2 mM dNTP mix, 0.4 µM forward and reverse primers, and 2 units Platinum™ Taq DNA polymerase (Thermofisher Scientific). The cycling parameters consisted of an initial 2-min denaturation at 95 °C followed by 40 cycles of denaturation at 95 °C for 30 s, annealing at 55 °C for 30 s, and extension at 72 °C for 15 s, with a final 2-min extension at 72 °C. The presence of PCR product was assessed by gel electrophoresis, and amplicon identity was confirmed by Sanger sequencing as follows: PCR products were purified using the MultiScreen®PCRμ96 filter plate (Millipore Corporation, Billerica, MA), and were sequenced in both directions with the BigDye Terminator Kit v3.1 on an ABI 3730xl 96-capillary DNA Analyzer (Applied Biosystems, Foster City, CA). Sequence bases were called, and their quality was checked using Sequencing Analysis v5.1 software (Applied Biosystems, Foster City, CA).
Results
Data-dependent acquisition on combined tick samples
Five samples consisting of protein extract from five pooled ticks in each sample were used to survey tick-borne pathogens. The 5 ticks in each sample were selected randomly from the total collected ticks. Figure 1 shows pie charts visualizing the number of identified proteins in pooled tick extracts. Of the total identified proteins, 2052 proteins belonged to I. scapularis tick, and 41 proteins belonged to tick-borne pathogens. These proteins were identified by at least two peptides or more and with high confidence (≥ 99% C.I.), except for few pathogen proteins of interest that were identified with one single peptide but with high confidence XCorr score, and specificity based on blast search was retained. The list of all identified proteins including tick and pathogen proteins are listed in supplemental Table S1 with their Uniprot accession number, number of peptides per proteins, peptide sequence, change state, and XCorr scores. The identified proteins for each specific pathogen, along with their identified peptide sequences, m/z values, DeltaM (ppm), and Xcorr scores, are listed under separate Excel sheets in the same supplemental Table S1. Each of these identified pathogen peptide sequences was manually checked for specificity using blast search against the entire Uniprot knowledge database (UniProtKB). Figure 1 shows a pie chart distribution of the number of tick and pathogen identified proteins. The most abundant pathogen found in the black-legged tick was Rickettsia spp. (33 unique proteins) followed by B. burgdorferi (5 unique proteins), and A. phagocytophilum (3 proteins).
Observed total tick and pathogen proteome. I. scapularis specific proteins consisted of 2,052 proteins and pathogens’ specific proteins consisted of 41 unique proteins out of total identified proteins of 2,093. This is cumulative data from 5 replicate experiments analyzing pooled extract from 5 ticks in each experiment. Number denotes number of proteins identified for each detected pathogen
Although some of the detected pathogens, such as Rickettsia spp., were represented by several proteins (e.g., 33 proteins), in this study, we will only focus on membrane and outer surface proteins owing to their potential involvement in host cell adhesion and immunogenicity. Table 1 lists key pathogen surface proteins that were identified by unique peptide sequences along with peptide counts, sequence coverage, and detection frequency across the 5 analyzed sample pools. Each peptide sequence was manually checked for specificity using Blast search against the entire Uniprot database.
Rickettsia spp. were consistently identified by multiple surface proteins across all 5 analyzed samples. These included the putative adhesin (Pa) identified by 9 unique peptides covering 53% of the protein sequence, the peptidoglycan-associated lipoprotein (Pal) that was identified by 4 unique peptides covering 35% of the protein sequence, and the 120-kDa outer membrane protein (OmpB) that was identified by 5 unique peptides covering 5% of the protein sequence. As shown in sequence alignments (Supplemental material S3), of the nine peptide sequences identified for Pa, the peptide with the sequence [AELAYSWIYDGR] mapped with 100% sequence homology the Pa tryptic peptide belonging to four Rickettsia species (e.g., R. endosymbiont, R. buchneri, R. monacensis, and R. akari). Furthermore, the specificity of the peptide sequences [IDAGAAMFNK] and [NVIYQGTSVPTGGMR] eliminated R. akari and kept R. endosymbiont, R. buchneri, and R. monacensis. The 120-kDa OmpB was identified by 5 unique peptides that mapped to several Rickettsia species but the peptide with the sequence [SSDENYKETSTTVANK] narrowed the mapping to OmpB belonging to R. endosymbiont and R. buchneri. Similarly, POps was identified by 4 unique peptide sequences belonging to the Rickettsia spp. and the peptide with the sequence [LNAGGMIFDK] was unique to POsp that belongs to R. buchneri. The 4 unique Pal peptides on the other hand although specific to Rickettsia mapped to Pal protein belonging to several Rickettsia spp. Overall, these data point to R. buchneri as the likely Rickettsia species carried by the black-legged tick collected from the Southern Tier of New York State.
Borrelia spp., the second frequently detected pathogens in black-legged ticks used in this study, were identified by three unique surface proteins: the OspA that was identified by 7 unique peptides covering 33% of the protein sequence, flagellin (FlaB) that was identified by 3 unique peptides covering up to 22% of the sequence coverage, and Bmp domain-containing protein (Bmp) that was identified by one peptide. OspA was detected in all 5 analyzed sample pools and was represented by 7 tryptic peptide sequences with 100% sequence homology to OspA belonging to B. burgdorferi (see sequence alignment in supplemental material S2). FlaB was detected in 1 out of the 5 samples analyzed and was represented by 3 tryptic peptides whose sequences mapped to FlaB protein belonging to several Borrelia spp. Similarly, Bmp was detected in 2 out of the 5 samples analyzed and was represented by 1 tryptic peptide [KIEEVSEKGIK] whose sequence was unique to Bmp protein belonging to Borrelia bissettiae.
Additional pathogens detected by mass spectrometry proteome profiling in pooled tick extracts included rarer pathogens such as Babesia microti, Bartonella henselae, Bartonella schoenbuchensis and multiple Theileria spp. and A. phagocytophilum. These were represented by only few proteins and most of them were not specific when manually checked using blast search except for A. phagocytophilum that was represented by one surface protein, the p44-14 outer membrane protein (p44-14) which was identified by one unique peptide [VELEIGYER] but good MS/MS probability score.
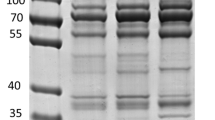
All these outer surface pathogen proteins were detected in gel bands corresponding to their expected molecular weight, except for the 120 kDa OmpB which was detected as a fragment of the C-terminal part of the protein (Fig. 2). Representative MS/MS spectra of peptides belonging to OspA, FlaB, and Pal are shown in Fig. 3 with good signal to noise ratios for the “y” and “b” ion series.
Fragmentation spectra of signature tryptic peptides from proteins of tick-borne pathogens MS/MS data of representative tryptic peptides belonging to specific proteins from the tick-borne pathogens. Top panel, MS/MS of a specific peptide of OspA from B. burgdorferi pathogen; middle panel, MS/MS of a specific peptide from flagellin from B. burgdorferi; and bottom panel, MS/MS of a specific peptide to Pal from R. parkeri. Identified peptides for these three targeted proteins are shown in Fig. 2
Western blot analysis confirmed the presence of OspA around its expected molecular weight of 30 kDa, in an independent set of ticks (Fig. 4). Eight out of seventeen analyzed ticks were positive. The presence of Rickettsia pal gene in ticks was confirmed by PCR analysis of leftover crude extracts from single ticks used in the PRM assay below (Fig. 5).
Western blot confirming the presence of OspA in protein extracts from single ticks. Individual protein extracts from 17 ticks were separated by SDS-PAGE and processed for western blot analysis to reveal OspA protein from B. burgdorferi pathogen. OspA was detected around its expected molecular weight of 30 kDa in 8 out of the 17 analyzed ticks
PCR assay targeting Rickettsia pal gene and Borrelia ospA gene in crude extracts from single ticks. Top panel shows the gel of PCR products and their migration pattern. Bottom panel shows the sequencing of the PCR products and the identified genes. Forward and reverse primers are indicated in red in each identified sequence
Pathogens survey in a single tick using PRM
Pathogen peptides that were easily detected by LC–MS/MS for Rickettsia spp. and B. burgdorferi and for rarer infectious pathogen such as A. phagocytophilum were selected to create an inclusion list for targeted mass spectrometry analysis using PRM on extracts from individual ticks. Additional peptides belonging to tick proteins were added to the list as internal reference. The full inclusion list including peptide sequences, m/z, and retention time are provided in supplemental Table S2. PRM analysis was performed on protein extracts from 10 individual ticks. Eight gel bands corresponding to the protein of interest were excised and processed for targeted mass spectrometry based on the peptide m/z and retention time determined from DDA analysis above.
Table 2 shows peptides detected for specific pathogen proteins by PRM assay. Out of the ten individually screened ticks, eight tested positive for Rickettsia spp. based on the detection of the putative adhesin tryptic peptide [ADLTLGTIIGGK]; four tested positive for B. Burgdorferi based on the detection of three OspA tryptic peptides [LEGSAVEITK], [GYVLEGTLTAEK], and [LTISDDLGQTTLEVFKEDGK]; and one tick tested negative for all targeted pathogens. None of the single screened ticks tested positive for other rare pathogens (e.g., A. phagocytophilum, T. parva, B. microti, and Bartonella spp.) detected in pooled samples above. Interestingly, three out of the 10 screened ticks were co-infected with both Rickettsia spp. and B. burgdorferi. The transition ions of these PRM targeted pathogen peptides were further verified using Skyline software for consistency (Supplemental material S4). The intensity of transition ions of each targeted peptides matched very well to the theoretical transition ion intensities based on the dot-product correlation (Dotp) scores that was above 0.8 (Supplemental material S4).
To confirm the presence of Rickettsia pal gene and borrelia ospA gene, a PCR assay was performed on 4 leftover single tick sample extracts (18, 24, 25, and 26) analyzed by PRM above. Figure 5 shows the pal gene PCR product that was detected in 4 out of 6 screened samples. PCR and PRM data for Pal agree for single tick extracts 18, 25, and 26. In the case of tick extract 24, while PCR and sequencing confirmed the presence of pal gene, the PRM assay for the Pal protein was negative. The pal gene was found to be 100% homologous to the pal gene belonging to spotted fever group R. endosymbiont and R. buchneri (Fig. 5), in agreement with the proteomic data above. Phylogenetic tree supports the close identity of the pal gene and peptide sequences to R. buchneri and the R. endosymbiont group (Supplemental material 5). The partial ospA PCR product was 100% specific to ospA gene belonging to B. Burgdorferi.
Discussion
Proteome profiling to screen for tick-borne pathogens in the Ixodes scapularis population collected from the Chenango Valley State Park of the Broome County in NY revealed high positivity for Rickettsia spp. and B. burgdorferi. These two pathogens were readily identified by several proteins in both combined tick extracts and in individually screened ticks. Other rarer pathogens such as A. phagocytophilum, Theileria annulata, and Babesia microti were detected only in one or two pooled tick extracts and not in the individually screened ticks. Accordingly, the focus in this study is on the most prevalent pathogens detected in I. scapularis from the Southern Tier region of New York State (Rickettsia spp. and B. burgdorferi).
Rickettsia spp., spotted fever rickettsiosis causing pathogens [27, 28], were consistently detected by several proteins across all pooled tick extracts, as well as in single tick extracts screened by PRM assay. Of the 10 screened ticks, 8 tested positive for Rickettsia spp. and 5 tested positive to B. burgdorferi, in agreement with previous study conducted by PCR assay on I. scapularis ticks submitted by residents to the Connecticut agricultural experiment station tick testing laboratory [18].
Rickettsia was identified by 33 unique proteins in the five pooled tick extracts with several proteins consistently detected across all samples. Peptide sequences of these Rickettsia proteins mapped to several Rickettsia species. However, tree analysis of identified peptide sequences in combination with sequenced PCR product narrowed the species to the spotted fever group R. endosymbiont and R. bucheneri. Perhaps one of the most important classes of these pathogen proteins are outer surface proteins owing to their potential role in Rickettsia pathogenicity and immunogenicity. These included Pal, putative adhesin, and OmpB. OmpB is a surface protein that was demonstrated to play a role in the adherence of the bacteria to host cells [29]. Although it was not included in PRM screening of single tick extracts, due to its low abundance, OmpB was identified by 5 unique peptides in pooled tick extracts. Pal is an outer membrane protein that is part of the Tol-Pal system and is associated with the peptidoglycans in gram-negative bacteria. Pal is important for outer membrane integrity and is believed to be involved in the pathogenesis of certain gram-negative bacteria including B. burgdorferi [30]. While we did not detect Pal in B. burgdorferi in this study, it was readily detected in tick extracts that were positive for Rickettsia spp. Pal was confirmed by PCR assay and sequencing and was found to be conserved among the Rickettsia spp. A complete sequencing of the PCR product showed 100% identity to the pal gene belonging to R. endosymbiont and R. buchneri. Pal was recently identified by others from cultured R. acari using shotgun and gel-based proteomics approach and was found to be immunogenic after probing it with serum from clinically infected patients [31]. Putative adhesin was one of the most abundant outer surface proteins in infected ticks since it was detected by PRM in 8 out of 10 screened ticks. This same protein was also identified using a proteomic approach in cultured R. acari but was not found to be immunogenic [31]. R. buchneri is an obligate endosymbiont that established a symbiotic relationship with the tick. Although R. buchneri species is closely related to the pathogenic rickettsiae, it is only transmitted transversally from tick to tick [32]. Furthermore, a recent study has shown that R. buchneri inhibits growth of other pathogenic bacteria in tick cells [33] which could explain the low number of co-infected ticks in this and other studies.
B. burgdorferi, a well-known Lyme disease-causing pathogen [34], was identified mainly by OspA which was confirmed by PCR assay and sequencing. OspA protein was relatively abundant compared to other Borrelia proteins in the positive ticks. It was represented by a maximum of 7 unique peptides and up to 61 PSMs in pooled tick extracts and was detected by three peptides in PRM screening of extracts from single ticks. This agrees with a previous study demonstrating that OspA and not OspC is overexpressed by B. burgdorferi when present in the tick [35]. The switch between OspA and OspC expression is believed to be involved in the Borrelia life cycle and OspA is downregulated when the pathogen is transmitted from the tick to the mammals. This switch might be important for Borrelia survival at different temperatures and different pH’s [36, 37]. Indeed, temperature and pH are different in an unfed tick’s midgut compared to mammalian blood and this might dictate which outer surface proteins are expressed depending on the environment [37]. Other studies reported that OspA might be involved in the interaction of the bacteria with ligands in the tick’s midgut [38]. However, further study comparing proteome profiles of Borrelia isolated from ticks to that of Borrelia isolated from mammals are needed to gain insight into Borrelia life cycle and pathogenesis.
This data could further be developed to identify changes between pathogen protein expressions within unfed, adult ticks compared to nymphal ticks, and fed ticks as well as pathogens within hosts such as dogs, horses, and humans. These subsequent experiments could bring insights into tick-pathogen interactions, at what stage ticks are more likely to encounter certain pathogens, and changes in protein expression that might occur during different tick life stages and between different hosts.
Interestingly, among the 10 individually screened ticks in this study, two were found to be positive for both Rickettsia spp. and B. burgdorferi in agreement with previous studies in which an increasing amount of co-infection rates has been detected in the past 20 years [39]. This further suggests that these two pathogens can be simultaneously transmitted to humans by co-infected ticks. However, the exact titer of R. parkeri relative to B. burgdorferi in co-infected ticks could only be estimated from peptide intensities and might not be accurate. Further studies using stable isotope-labeled peptides might be needed to determine the relative titer of pathogens in single ticks.
Some limitations in this study included the screening of a small number of ticks collected from one geographical area in Broome County, NY. Further screening of individual ticks might be required to ensure a true representation of I. scapularis populations and rate of positivity to tick-borne pathogens in Broome County as well as other surrounding areas. Furthermore, seasonal and annual testing could be performed to analyze the percentage of ticks carrying these pathogens to see how it changes in function of the seasons. In addition, a second method, such as PCR and DNA sequencing performed on half of the tick, might prove useful as a validation or confirmation of the data. The analysis time to screen a single tick was extremely long compared to the PCR method, as it required analysis of 8 gel bands per sample and a 90-min LC–MS/MS run per gel band, thus totaling 720 min for a single tick screening. The intent of this single tick screening was not to use rapid testing of ticks by any means but rather for understanding the pathogen content in single ticks and the type of pathogen proteins expressed or co-expressed in a tick, especially outer surface proteins that might play a role in pathogenicity and immunogenicity. Understanding the relationship between different microorganisms (pathogenic versus nonpathogenic) and their co-existence in the same tick might provide a way to control transmission of pathogen causing disease from ticks to mammals and humans. Indeed, recent network analysis study conducted on northern and southern ticks predicted an antagonistic interaction between R. buchneri and B. burgdorferi [40].
Conclusions
Mass spectrometry-based proteome profiling and parallel reaction monitoring are powerful methods to identify and screen for tick-borne pathogens for surveillance and awareness about tick-borne diseases in specific geographical areas. This might bring insight into tick-borne pathogens life cycles and their protein expression changes depending on the vector and might also help develop biomarkers for direct diagnosis and distinguishing between different tick-borne diseases, for which treatment remains empirical and often administered as prophylaxis.
Data availability
Data is available as supplementary material.
Code availability
NA.
References
Eisen RJ, Eisen L. The Blacklegged Tick, Ixodes scapularis: An Increasing Public Health Concern. Trends Parasitol. 2018;34(4):295–309. https://doi.org/10.1016/j.pt.2017.12.006.
Murray TS, Shapiro ED. Lyme disease. Clin Lab Med. 2010;30(1):311–28. https://doi.org/10.1016/j.cll.2010.01.003.
Van Hout MC. The Controversies, Challenges and Complexities of Lyme Disease: A Narrative Review. J Pharm Pharm Sci. 2018; https://doi.org/10.18433/jpps30254.
El-Sayed A, Kamel M. Climatic changes and their role in emergence and re-emergence of diseases. Environ Sci Pollut Res Int. 2020;27(18):22336–52. https://doi.org/10.1007/s11356-020-08896-w.
Berrada ZL, Telford SR 3rd. Burden of tick-borne infections on American companion animals. Top Companion Anim Med. 2009;24(4):175–81. https://doi.org/10.1053/j.tcam.2009.06.005.
Sprong H, Azagi T, Hoornstra D, Nijhof AM, Knorr S, Baarsma ME, Hovius JW. Control of Lyme borreliosis and other Ixodes ricinus-borne diseases. Parasit Vectors. 2018;11(1):145. https://doi.org/10.1186/s13071-018-2744-5.
Földvári G, Široký P, Szekeres S, Majoros G, Sprong H. Dermacentor reticulatus: a vector on the rise. Parasit Vectors. 2016;9(1):314. https://doi.org/10.1186/s13071-016-1599-x.
Tilly K, Rosa PA, Stewart PE. Biology of infection with Borrelia burgdorferi. Infect Dis Clin North Am. 2008;22(2):217–v. https://doi.org/10.1016/j.idc.2007.12.013.
Kurokawa C, Lynn GE, Pedra JHF, Pal U, Narasimhan S, Fikrig E. Interactions between Borrelia burgdorferi and ticks. Nat Rev Microbiol. 2020;18(10):587–600. https://doi.org/10.1038/s41579-020-0400-5.
Roy-Dufresne E, Logan T, Simon JA, Chmura GL, Millien V. Poleward expansion of the white-footed mouse (Peromyscus leucopus) under climate change: implications for the spread of lyme disease. PLoS One. 2013;8(11):e80724. https://doi.org/10.1371/journal.pone.0080724.
Huang CI, Kay SC, Davis S, et al. High burdens of Ixodes scapularis larval ticks on white-tailed deer may limit Lyme disease risk in a low biodiversity setting. Ticks Tick Borne Dis. 2019;10(2):258–68. https://doi.org/10.1016/j.ttbdis.2018.10.013.
Liu Y, Nordone SK, Yabsley MJ, Lund RB, McMahan CS, Gettings JR. Quantifying the relationship between human Lyme disease and Borrelia burgdorferi exposure in domestic dogs. Geospat Health. 2019. https://doi.org/10.4081/gh.2019.750.
Moutailler S, Valiente Moro C, Vaumourin E, Michelet L, Tran FH, Devillers E, Cosson JF, Gasqui P, Van VT, Mavingui P, Vourc’h G, Vayssier-Taussat M. Co-infection of Ticks: The Rule Rather Than the Exception. PLoS Negl Trop Dis. 2016;10(3):e0004539. https://doi.org/10.1371/journal.pntd.0004539.
Walker DH. Rickettsiae. In: Baron S, editor. Medical Microbiology. 4th edition. Galveston (TX): University of Texas Medical Branch at Galveston; Chapter 38. 1996. Available from: https://www.ncbi.nlm.nih.gov/books/NBK7624/.
Centers for Disease Control and Prevention. Signs and symptoms. In: Centers for Disease Control and Prevention. 2019 https://www.cdc.gov/anaplasmosis/symptoms/index.html. Accessed 2021 August 6.
Guzman N, Yarrarapu SNS, Beidas SO. Anaplasma Phagocytophilum. In: StatPearls. Treasure Island (FL): StatPearls Publishing; 2021.
Centers for Disease Control and Prevention. In: Epidemiology and statistics. Centers for Disease Control and Prevention. 2021; https://www.cdc.gov/anaplasmosis/stats/index.html. Accessed 2021 August 6.
Pokutnaya D, Molaei G, Weinberger DM, Vossbrinck CR, Diaz AJ. Prevalence of Infection and Co-Infection and Presence of Rickettsial Endosymbionts in Ixodes Scapularis (Acari: Ixodidae) in Connecticut, USA. J Parasitol. 2020;106(1):30–7.
Fotso Fotso A, Mediannikov O, Diatta G, Almeras L, Flaudrops C, Parola P, Drancourt M. MALDI-TOF mass spectrometry detection of pathogens in vectors: the Borrelia crocidurae/Ornithodoros sonrai paradigm. PLoS Negl Trop Dis. 2014. https://doi.org/10.1371/journal.pntd.0002984.
Yssouf A, Almeras L, Terras J, Socolovschi C, Raoult D, Parola P. Detection of Rickettsia spp in ticks by MALDI-TOF MS. PLoS Negl Trop Dis. 2015. https://doi.org/10.1371/journal.pntd.0003473.
Talagrand-Reboul E, Westermann B, Raess MA, Schnell G, Cantero P, Barthel C, Ehret-Sabatier L, Jaulhac B, Boulanger N. Proteomic as an Exploratory Approach to Develop Vaccines Against Tick-Borne Diseases Using Lyme Borreliosis as a Test Case. Vaccines (Basel). 2020. https://doi.org/10.3390/vaccines8030463.
Magni R, Almofee R, Yusuf S, Mueller C, Vuong N, Almosuli M, Hoang MT, Meade K, Sethi I, Mohammed N, Araujo R, McDonald TK, Marcelli P, Espina V, Kim B, Garritsen A, Green C, Russo P, Zhou W, Vaisman I, Petricoin EF 3rd, Hoadley D, Molestina RE, McIntyre H, Liotta LA, Luchini A. Evaluation of pathogen specific urinary peptides in tick-borne illnesses. Sci Rep. 2020. https://doi.org/10.1038/s41598-020-75051-3.
Scott JD, Foley JE, Clark KL, et al. Established Population of Blacklegged Ticks with High Infection Prevalence for the Lyme Disease Bacterium, Borrelia burgdorferi Sensu Lato, on Corkscrew Island, Kenora District. Ontario Int J Med Sci. 2016;13(11):881–91. https://doi.org/10.7150/ijms.16922.
Johnson TL, Graham CB, Boegler KA, et al. Prevalence and Diversity of Tick-Borne Pathogens in Nymphal Ixodes scapularis (Acari: Ixodidae) in Eastern National Parks. J Med Entomol. 2017;54(3):742–51. https://doi.org/10.1093/jme/tjw213.
Olsthoorn F, Sprong H, Fonville M, et al. Occurrence of tick-borne pathogens in questing Ixodes ricinus ticks from Wester Ross, Northwest Scotland. Parasit Vectors. 2021;14(1):430. https://doi.org/10.1186/s13071-021-04946-5.
Dzieciatkowska M, Hill R, Hansen KC. GeLC-MS/MS analysis of complex protein mixtures. Methods Mol Biol. 2014;1156:53–66. https://doi.org/10.1007/978-1-4939-0685-7_4.
Paddock CD, Sumner JW, Comer JA, Zaki SR, Goldsmith CS, Goddard J, McLellan SL, Tamminga CL, Ohl CA. Rickettsia parkeri: a newly recognized cause of spotted fever rickettsiosis in the United States. Clin Infect Dis. 2004;38(6):805–11. https://doi.org/10.1086/381894.
Kristof MN, Allen PE, Yutzy LD, Thibodaux B, Paddock CD, Martinez JJ. Significant Growth by Rickettsia Species within Human Macrophage-Like Cells Is a Phenotype Correlated with the Ability to Cause Disease in Mammals. Pathogens. 2021;10(2):228. https://doi.org/10.3390/pathogens10020228.
Uchiyama T. Tropism and pathogenicity of rickettsiae. Front Microbiol. 2012. https://doi.org/10.3389/fmicb.2012.00230.
Godlewska R, Wiśniewska K, Pietras Z, Jagusztyn-Krynicka EK. Peptidoglycan-associated lipoprotein (Pal) of Gram-negative bacteria: function, structure, role in pathogenesis and potential application in immunoprophylaxis. FEMS Microbiol Lett. 2009;298(1):1–11. https://doi.org/10.1111/j.1574-6968.2009.01659.x.
Csicsay F, Flores-Ramirez G, Zuñiga-Navarrete F, Bartošová M, Fučíková A, Pajer P, Dresler J, Škultéty Ľ, Quevedo-Diaz M. Proteomic analysis of Rickettsia akari proposes a 44 kDa-OMP as a potential biomarker for Rickettsialpox diagnosis. BMC Microbiol. 2020;20(1):200. https://doi.org/10.1186/s12866-020-01877-6.
Kurtti TJ, Felsheim RF, Burkhardt NY, Oliver JD, Heu CC, Munderloh UG. Rickettsia buchneri sp. nov., a rickettsial endosymbiont of the blacklegged tick Ixodes scapularis. Int J Syst Evol Microbiol. 2015;65(Pt 3):965–70. https://doi.org/10.1099/ijs.0.000047.
Cull B, Burkhardt NY, Wang XR, Thorpe CJ, Oliver JD, Kurtti TJ, Munderloh UG. The Ixodes scapularis Symbiont Rickettsia buchneri Inhibits Growth of Pathogenic Rickettsiaceae in Tick Cells: Implications for Vector Competence. Front Vet Sci. 2022;8:748427. https://doi.org/10.3389/fvets.2021.748427.
Shapiro ED. Borrelia burgdorferi (Lyme disease). Pediatr Rev. 2014;35(12):500–9. https://doi.org/10.1542/pir.35-12-500.
Schwan TG, Piesman J. Temporal changes in outer surface proteins A and C of the lyme disease-associated spirochete, Borrelia burgdorferi, during the chain of infection in ticks and mice. J Clin Microbiol. 2000;38(1):382–8. https://doi.org/10.1128/JCM.38.1.382-388.2000.
Obonyo M, Munderloh UG, Fingerle V, Wilske B, Kurtti TJ. Borrelia burgdorferi in tick cell culture modulates expression of outer surface proteins A and C in response to temperature. J Clin Microbiol. 1999;37(7):2137–41. https://doi.org/10.1128/JCM.37.7.2137-2141.
Ramamoorthy R, Scholl-Meeker D. Borrelia burgdorferi proteins whose expression is similarly affected by culture temperature and pH. Infect Immun. 2001;69(4):2739–42. https://doi.org/10.1128/IAI.69.4.2739-2742.2001.
Xu Q, McShan K, Liang FT. Essential protective role attributed to the surface lipoproteins of Borrelia burgdorferi against innate defences. Mol Microbiol. 2008;69(1):15–29. https://doi.org/10.1111/j.1365-2958.2008.06264.x.
Wolf MJ, Watkins HR, Schwan WR. Ixodes scapularis: Vector to an Increasing Diversity of Human Pathogens in the Upper Midwest. WMJ. 2020;119(1):16–21.
Kumar D, Downs LP, Adegoke A, Machtinger E, Oggenfuss K, Ostfeld RS, Embers M, Karim S. An Exploratory Study on the Microbiome of Northern and Southern Populations of Ixodes scapularis Ticks Predicts Changes and Unique Bacterial Interactions. Pathogens. 2022;11(2):130. https://doi.org/10.3390/pathogens11020130.
Acknowledgements
The authors would like to thank Binghamton University School of Pharmacy for the opportunity to conduct this research, Dr. Tchilabalo Alayi for method developments and guidance, and Dr. Tracy Curtis for tick collection methods. This work was partially supported by the Binghamton University Organized Research Center Award to the Tick-borne Illness Center at Binghamton University. We also would like to thank Decker Foundation for supporting the purchase of the mass spectrometry instrument used in this study.
Funding
This work was partially supported by the Decker Foundation.
Author information
Authors and Affiliations
Contributions
HS collected the ticks, prepared the samples for analysis, generated the data, and wrote the first draft. EC helped with mass spectrometry analysis and data generation; RR helped with sample preparation and data generation; RS helped with PCR assay, data generation, and revising the manuscript; MS helped with DNA sequencing, data interpretation, and revising the manuscript; and YH helped with supervision, writing, and editing the manuscript.
Corresponding author
Ethics declarations
Ethics approval
NA
Consent to participate
NA
Consent for publication
NA
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
ABC Highlights: authored by Rising Stars and Top Experts.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Smith, H.R., Canessa, E.H., Roy, R. et al. A single tick screening for infectious pathogens using targeted mass spectrometry. Anal Bioanal Chem 414, 3791–3802 (2022). https://doi.org/10.1007/s00216-022-04054-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-022-04054-y