Abstract
We demonstrate the suitability of a fast, green, easy-to-perform, and modified sample extraction procedure, i.e., dispersive liquid–liquid microextraction (DLLME) for the determination of efavirenz (EFV) in human plasma. Data acquisition was done by gas chromatography–mass spectrometry (GC–MS) in the selected ion monitoring (SIM) mode. The simplicity of the method lies in, among others, the avoidance of the use of large organic solvent volumes as mobile phases and non-volatile buffers that tend to block the plumbing in high-performance liquid chromatography (HPLC). Chromatographic and mass spectral parameters were optimized using bovine whole blood for matrix matching due to insufficient human plasma. Method validation was accomplished using the United States Food and Drug Administration (USFDA) 2018 guidelines. The calibration curve was linear with a dynamic range of 0.10–2.0 μg/mL and an R2 value of 0.9998. The within-run accuracy and precision were both less than 20% at the lower limit of quantification (LLOQ) spike level. The LLOQ was 0.027 μg/mL which compared well with some values but was also orders of magnitude better than others reported in the literature. The percent recovery was 91.5% at the LLOQ spike level. The DLLME technique was applied in human plasma samples from patients who were on treatment with EFV. The human plasma samples gave concentrations of EFV ranging between 0.14–1.00 μg/mL with three samples out of seven showing concentrations that fell within or close to the recommended therapeutic range.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Efavirenz (EFV) is still a widely used antiretroviral drug (ARV) in low- and middle-income countries (LMICs) [1, 2]. This is despite its deficiencies compared to dolutegravir (DTG) which is recommended by the World Health Organization (WHO), the United States of America (USA), and the European Union (EU). Until June 2018, EFV was part of the WHO preferred first-line combination antiretroviral therapy (cART) regimen for human immunodeficiency virus type 1 (HIV-1) [3]. DTG-based and low-dose EFV-based, i.e., EFV400, combinations are now considered as first-line treatments for HIV-1 in resource-limited settings [4]. EFV inhibits the reverse transcriptase enzyme causing inhibition of viral ribonucleic acid (RNA) and deoxyribonucleic acid (DNA)–dependent polymerase activities by disrupting the catalytic site [5, 6]. It is mainly prescribed as a combination single-tablet regimen with tenofovir disoproxil fumarate (TDF) and emtricitabine (EMT) brand named Atripla. It is a preferred non-nucleoside reverse transcriptase inhibitor (NNRTI) in many countries because it has low hepatotoxicity, does not require dose adjustment, and can also, like DTG, be used concomitantly with tuberculosis treatment [7,8,9,10]. EFV-based regimens have been the backbone of combination antiretroviral therapy for over 15 years, and this was mainly due to its antiretroviral efficacy and favorable pharmacokinetics [11]. These considerations make it imperative that accurate, sensitive, and easy-to-access analytical methods with potential for miniaturization in resource-limited LIMCs continue to be developed to monitor EFV in human plasma for clinical analysis.
Several methods have been reported in literature for the determination of EFV in human plasma. Solid-phase extraction (SPE) or liquid–liquid extraction (LLE) followed by high-performance liquid chromatography (HPLC) separation coupled to ultraviolet (UV) detection is common [12,13,14]. HPLC followed by UV detection has been reported in several analyses of EFV in human plasma [15,16,17,18] and dried blood spots [19] using traditional extraction methods, i.e., SPE, LLE, and protein precipitation (PPT). Reasonable limits of quantification (LOQs) have been reported using these methods. For instance, a LOQ of 0.10 μg/mL has been achieved in the determination of EFV in human plasma [9]. Despite HPLC coupled with UV detection meeting some of the required analytical performance characteristics such as good accuracy, precision, and low LOQs, this method suffers several shortcomings such as limited spectral identification that complicates the use of data systems to accurately quantify the separated components. In contrast to a separation system coupled to a mass spectrometer whose data system is two dimensional as opposed to one, the lack of specificity in HPLC alone exasperates the limitation to adequately identify analytes in a complex sample matrix such as human plasma.
Liquid chromatography–mass spectrometry (LC–MS) offers a better alternative to HPLC–UV for the determination of EFV in human plasma [20,21,22]. LC–MS/MS can identify the compound(s) of interest by both retention time and mass to charge (m/z) ratio of the precursor and product ions and therefore offers the advantage of greater analytical specificity [23]. LC-MS for the determination of EFV in human plasma [24,25,26] in hair [27] and in saliva [21] has been reported
Gas chromatography–mass spectrometry (GC–MS) is another technique that has been applied in the determination of EFV. GC–MS using electron ionization (EI) [28, 29] has been reported for the determination of EFV in human plasma in which SPE and LLE with derivatization were used respectively for sample preparation. The limitations of these sample preparation procedures, such as cost and use of large volumes of organic solvents in the case of SPE and a higher sample and extraction solvent volume in the case of LLE with derivatization, could be overcome by a simpler method such as DLLME requiring up to 100 μL of extraction solvent and lower sample volumes. This could be followed by GC–MS which is less cumbersome in contrast to HPLC and LC–MS where in a lot of instances, the latter methods’ use of large volumes of mobile phases and buffers that are not benign to the environment can be problematic.
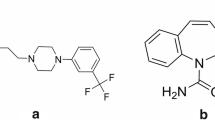
We report a method employing a modified non-traditional bioanalytical sample preparation method, i.e., DLLME followed by EI–GC–MS using the selected ion monitoring mode (SIM) data acquisition mode, for the determination of EFV in human plasma. DLLME employs microliter volumes of samples and extraction solvents, making it ideal for human fluid samples, and minimum extraction times compared to traditional bioanalytical sample preparation techniques, i.e., SPE, LLE, and PPT. DLLME also affords a cost-effective and environmentally benign, i.e., green technique, which can easily be automated in-house with commercially automated systems such as the Gerstel Multipurpose Sampler (MPS) robotic by Gerstel Inc. [30] already available on the market. Automated systems can improve the turnaround time for sample preparation during therapeutic drug monitoring (TDM) in clinical laboratories to furnish medical personnel with the required analytical data within a specified time. Figure 1 shows the structures of EFV and the internal standard used in this study, metronidazole (MNZ).
Materials and methods
Reagents and apparatus
The EFV standard was purchased from U.S. Pharmacopeia Convention Rockville (USA); metronidazole (MNZ), used as an internal standard, was purchased from Sigma-Aldrich (Seelze, Germany). HPLC-grade methanol (99.9% purity) was purchased from Sigma-Aldrich Co., St. Louis, MO (USA). HPLC-grade acetonitrile and 15-mL centrifuge tubes were purchased from Fisher Scientific, Waltham, MA (USA). Chloroform, HPLC grade, was purchased from Alpha Chemical, Mumbai (India). Formic acid (FA) of 85% purity was purchased from Afro World Impex, Gaborone (Botswana). Zinc sulfate was purchased from Rochelle Chemicals, Johannesburg (South Africa). Five thousand–microliter syringes were from Hamilton, Reno, NV (USA). Ultra-high-purity (99.999%) nitrogen and helium gas were purchase from Afrox Gaborone (Botswana). Deionized water was from the Milli Q Millipore water purification system from Millipore Sigma, Temecula, CA (USA). Human plasma samples were provided by the Botswana Harvard AIDS Institute Partnership for HIV Research and Education Center, Gaborone (Botswana). Fresh bovine blood was donated by a butchery in Ramotswa (Botswana).
GC–MS
All the parameters reported here were optimized in-house as described below. An Agilent 7890A GC system coupled to an Agilent 5975C single quadrupole mass spectrometer equipped with an Agilent G4513A auto-sampler module was used throughout the experiments. A HP 5MS capillary column of 30 m length, 0.25 mm internal diameter, and 0.25 μm film thickness (5%-phenyl-methyl-siloxane) was used throughout for separations. One microliter of standard/sample was injected in pulsed splitless mode. The injector, transfer line, and ion source temperatures were 260 °C, 280 °C, and 230 °C, respectively. Helium was used as a carrier gas at a constant flow rate of 3 mL/min. The oven temperature was initially held at 130 °C for 2 min and then ramped to 280 °C at a heating rate of 30 °C/min and held there for 2 min for a total run time of 9 min. Mass spectra were acquired using electron ionization (EI) in the full and selected ion monitoring (SIM) modes. The mass scan range was m/z 50–350. In the SIM mode, acquisition of spectra was based on three or more selected masses of interest, i.e., three or four ions with the highest intensity with the first most intense m/z ratio being the quantification ion, and the other less intense ions used for confirmation. The instrument was set to reject all other ions including background ions arising from the matrix. The ions selected for EFV were m/z ratios 180, 246, and 315 while for MNZ, m/z ratios 54, 81, 124, and 171. The positive chemical ionization (PCI) using methane as a reagent gas was done using standard Agilent PCI settings embedded in the tune system. The system uses EI tune parameters and only allows manually setting the reagent gas flow into the ionization chamber.
Preparation of standards
Individual stock standard solutions of EFV and MNZ (internal standard) were prepared separately at 1000 μg/mL. Each compound was dissolved in a 10-mL volumetric flask using methanol and stored in a freezer at a temperature of − 10 °C. Prior to analysis, serial dilution was made to prepare working standards of 5 μg/mL EFV and 1 μg/mL MNZ from the stock solutions using methanol. The working standard solutions were stored at 4 °C in vials and wrapped in aluminum foil to protect them from photo-degradation.
Optimization of the GC separation parameters
The following parameters were optimized using the one factor at a time (OFAT) strategy: (i) injection mode, (ii) injector temperature, (iii) carrier gas flow rate, and (iv) oven heating rate.
To select the injection mode, a cocktail of standards using the splitless and pulsed splitless modes of injection was injected. Peak areas were adduced from the chromatograms and the ratios of analyte to internal standard were compared. The highest peak area ratio was taken as the optimum.
The injector temperature was optimized by varying temperatures from 220 to 280 °C in increments of 20 °C. The optimum was the temperature that gave the highest peak ratio of the analyte to internal standard. In order to optimize the carrier gas flow rate, it was varied from 1 to 4 mL/min which is the maximum flow rate which the Agilent 5975C series GC-MS system can be operated at. This was done in order to obtain the optimum flow to attain better efficiency measured as the number of theoretical plates (N) and to obtain short retention times for higher sample throughput. Ultimately, a van Deemter plot was used to adduce the optimum flow rate.
In order to improve the quality of separation and optimize the analysis time, the optimum oven heating temperature was determined by varying the heating rate from 10 to 30 °C/min in increments of 5 °C/min. The optimum heating rate was taken as that which yielded the highest analyte-to-internal standard ratio.
Selection of the acquisition and ionization modes
To select an appropriate acquisition scan mode, the FS mode was compared to the SIM mode by determining the instrument limits of detection (IDLs) in the two scan modes. Further, the ionization modes, i.e., EI and PCI, were compared using detection and quantification limits as figures of merit and using the selected acquisition scan mode.
Samples for optimization of DLLME and analysis
Since the goal was to optimize a method to analyze a biological matrix which the USFDA 2018 [31] defines as “discrete material of biological origin that can be sampled and processed in a reproducible manner” such as “blood, serum, plasma, urine, feces, cerebrospinal fluid, saliva, sputum, and various discrete tissues,” we used a surrogate matrix, i.e., fresh bovine whole blood, which was consistent with the study matrix to optimize the DLLME procedure. This was because the human plasma samples were in very minute quantities (μL volumes), were rare and difficult to obtain [32], and thus could not be used for this purpose. Previously, the use of human blank sera for method validation and application in drug pharmacokinetics in rats has been reported [33, 34]. The bovine whole blood in this research was used for matrix matching the analytical standards and for in-house spiking to assess the efficiency of extraction. Seven de-identified archived plasma samples from adult non-pregnant HIV patients, on Atripla, from the Republic of Botswana, stored at the Botswana Harvard AIDS Institute Partnership (BHP) for HIV Research and Education Center in Gaborone, together with one control human blood plasma sample were donated by the BHP. Control samples were from subjects on any other antiretroviral treatment (not containing EFV). All the samples were stored frozen at a temperature of − 80 °C and only allowed to thaw prior to being used.
DLLME sample preparation
The DLLME procedure was adapted from [35] and modified as described below. Fifty microliters of bovine blood were pipetted into a 15-mL centrifuge tube and spiked with 30 μL of 5 μg/mL EFV dissolved in methanol. Five milliliters of 1 μg/mL MNZ (internal standard) dissolved in water was added to the centrifuge tube. The mixture was shaken for a few seconds to homogenize it. Ten microliters of 5% (v/w) ZnSO4 was then added to the centrifuge tube to precipitate the proteins instead of 4% w/v sulfosalicylic acid that was used by Heydari et al. [35]. This was one of the modifications to the procedure. From literature, ZnSO4 has shown 91% precipitation of protein in human plasma [36], and here, it also removed the red coloration from bovine blood compared to other deproteinization reagents that were tested. Clarification of the red color was the focus here, because deproteinization reagents have been proven not to improve the signal-to-noise (S/N) ratios of small-molecule peaks by GC–MS [37]. Moreover, methanol has also been reported to deproteinize human plasma, and thus, removal of one step further reduced sample pre-treatment time. The mixture was centrifuged for 2 min at 4500 rpm. The clear supernatant was transferred to another centrifuge tube and 1300 μL of a mixture of acetonitrile (dispersing solvent) containing 100 μL of chloroform (extraction solvent) was injected rapidly into the clear supernatant using a 5-mL glass syringe. A cloudy solution developed instantly, and the tube was shaken for a few seconds and centrifuged for 4 min at 6000 rpm to afford phase separation. After centrifugation, the chloroform settled at the bottom of the tube. The chloroform phase was drawn using a glass syringe into a GC vial and evaporated to dryness under nitrogen. The residue was reconstituted with 40 μL of methanol. This was another modification to the method; i.e., instead of reconstituting with 4.5 mL of phosphate buffer (20 mM, pH = 8) containing 2% w/v NaCl, methanol was used because it is compatible with GC–MS. One microliter of the methanol reconstituted extract was injected into the GC–MS system for analysis. After the analytical performance characteristics were evaluated and found to be satisfactory, the extraction procedure was applied to real human plasma samples. Here, to 50 μL of each of the human plasma samples was added 5 mL of 1 μg/mL MNZ in a 15-mL centrifuge tube, followed by the DLLME procedure used for extraction of bovine whole blood.
Method validation
The method was partially validated using the USFDA [31] guidelines while referring to a discussion of bioanalytical method validation guidelines [29]. The analytical performance characteristics that were evaluated were the linearity and the instrument detection limit (IDL) that afforded estimation of the lower limit of quantification (LLOQ). Others were the method detection limits (MDLs), extraction efficiency expressed as percent recovery, the accuracy, the precision expressed as percent coefficient of variation (% CV), and matrix effects expressed in percent (% ME).
Instrument detection limit (IDL)
The IDL and the LLOQ were estimated at 3 and 10 times the signal-to-noise (S/N) ratio of the blank sample extract which was estimated at the y intercept of the calibration curve equivalent to the standard deviation of the line of regression of y on x (sy/x), respectively. The IDL was used as a guide for in-house spiking of real matrix blanks for recovery and precision calculations.
Method detection limit (MDL)
The MDL reflects a composite of instrumental noise and noise due to the matrix and is thus considered as the true LOD of a method. In this instance, a sample extract with the concentration of EFV close to the LLOQ was selected and run seven times on the GC–MS. Calculation of the MDL was done according to literature [38].
Accuracy and precision
To estimate the accuracy, three replicates were done at each spike level and this afforded calculation of the mean accuracy. Accuracy values were compared with the USFDA guidelines [31], where the accuracy should not exceed ± 15% of the nominal concentrations and should not exceed ± 20% at the LLOQ.
To evaluate precision, three replicate spiked samples were made per concentration at each spike level including at the LLOQ. The precision was expressed as the coefficient of variation (% CV). The precision values were compared with the USFDA guidelines which require that it should be ± 15% except at the LLOQ where it should not exceed ± 20%.
Matrix effects
The matrix effect expressed as % ME was calculated by comparing the peak area of the matrix matched standard with the solvent standard according to Matuszewski et al. (2003) [39]. This method has also been recommended as the best approach for evaluating matrix effects in the validation of bioanalytical chromatographic methods for the quantification of drugs in biological fluids [29]. Equation 1 was used in this case. A positive value indicates enhancement and a negative value indicates suppression of the analyte signal on the detector response.
where B is the average peak area of the analyte spiked into the extracted matrix and A is the average peak area of the analyte with the same concentration in a solution.
Results and discussion
Optimization of GC separation parameters
Figure 2(A) shows results of the optimization of the injection mode wherein the pulsed splitless mode was three times better than the splitless mode and (B) the van Deemter plot showing the optimum at 3 mL/min. In (C), the optimum oven heating rate was 30 °C, despite 10 °C showing similar results. Thirty degrees Celsius was selected because the IDL measured at this temperature was lower than at 10 °C. Separations were also faster at 30 °C without compromising peak symmetry. The optimum injector temperature of 280 °C is shown in (D). However, this injector temperature exhibited peak splitting which prompted the use of 260 °C as the optimum.
Selection of the acquisition and ionization modes
Figure 3(A) shows a comparison of the FS acquisition mode, IDLs using EI and positive chemical ionization (PCI) with methane as reagent gas, and (B) detection and quantification limits as well as nominal spike level performance of the SIM scan mode. (A) shows the FS mode where the analyte in PCI was not quantifiable but was in SIM. SIM yielded superior IDLs using EI compared to PCI. The performance of PCI using SIM was also inferior to EI as seen in (B). EI using SIM for data acquisition was ultimately taken as the best combination for further experiments.
Table 1 is a summary of the optimum values obtained after a series of experiments designed to optimize separation and mass spectrometry parameters.
Optimization of mass spectrometry parameters. (A) Comparison of the full-scan and selected ion monitoring acquisition modes, (B) comparison of EI and PCI limits of detection, quantification, and nominal spike level. FS = full scan; SIM = selected ion monitoring. All experiments were done in triplicate, i.e., n = 3
Applying the conditions tabulated gave the chromatogram in Fig. 4 showing the SIM acquisition chromatogram of a standard mixture of MNZ and EFV.
Figure 5 shows the FS mass spectra of MNZ and EFV. The mass spectra provided a basis for selecting the ions to use in the SIM mode based on their intensities.
Calibration curve and IDL
The calibration curve of EFV in the SIM acquisition mode yielded a regression equation, i.e., y = 1.5611x + 0.6361, with a slope of 1.5611 ± 0.0101 and an intercept of 0.6361 ± 0.01175 (± standard deviations of the slope and intercept at p = 0.05) and was linear with an R2 value of 0.9998, i.e., indicating that 99.98% of the values fell within the straight line. The correlation coefficient (r) of the regression line was 0.9999 indicating a very strong relationship between the ratio of EFV to IS and the concentration of EFV. The linear range for the calibration curve was 0.10–2.0 μg/mL based on six calibrators with each calibrator run three (3) times (n = 3), which was adequate for the quantification of real samples. Preliminary experiments using HPLC (results not shown) indicated that the highest concentration to expect from the real samples was approximately 0.9 μg/mL. An IDL of 0.03 μg/mL was estimated from the curve which afforded the estimation of the LLOQ which was ~ 0.1 μg/mL. These values were used as guides for spiking the blank bovine whole blood for validation purposes.
MDL and LLOQ
The MDL (true LOD) was 0.0081 μg/mL, and from this value, the real LLOQ was estimated to be 0.027 μg/mL. Thus, the GC–MS detected concentrations of EFV at levels below its therapeutic range, i.e., 1–4 μg/mL [40].
Accuracy and precision
The accuracy at the LOD spike level was 34.3% (n = 3) as expected and at two times the limit of detection (LOD) spike level was 17.2% (n = 3) while at the LLOQ spike level, it was 8.1% (n = 3) and satisfied the USFDA guidelines, i.e., it was less than 20%. The precision expressed as % CV at the LOD spike level was 2.9% (n = 3) and at two times the LOD spike level was 2.0% while at the LLOQ spike level, it was 1.7% (n = 3) and was within the acceptable range, i.e., less than 15%.
Percentage recovery
The USFDA guidelines do not prescribe acceptable percent recoveries. Percentage recoveries were evaluated at the LOD spike level where it was 65.7% (n = 3), at a spike level of 2 times the LOD [29] where it was 82.8% (n = 3) and was 91.5% (n = 3) at the LLOQ.
Matrix effects
The matrix effects were estimated using a control human plasma sample spiked at 5 μg/mL according to Matuszewski et al. (2003) [39]. The % ME (Eq. 1) was + 4.6 indicating a S/N ratio enhancement. This implies that the selectivity of the method was also enhanced as a result of an improvement in at least one factor contributing to it. This was attributed to the SIM acquisition mode which scans selected ions unique to the analyte and rejects matrix ions. Figure 6 shows the SIM acquisition chromatograms of the control sample, the blank, and the real human sample HBP 1.
The internal standard, i.e., MNZ, and analyte, i.e., EFV peaks, in the control and the real sample extracts were obscure in the FS acquisition mode whereas the blank showed no peak for both the internal standard and the analyte. However, the selectivity of the method is demonstrated in Fig. 6 where MNZ and EFV were clearly quantifiable at 4.7 and 6.6 min in the sample extract from HBP1 and control sample extract, respectively, using the SIM acquisition mode.
Application of the method to the analysis of EFV in human plasma
Real human plasma samples collected from the BHP laboratory were analyzed in triplicate for EFV. Table 2 shows the results.
The proposed therapeutic range of efavirenz for treatment of HIV-1 in ARV-naive patients suitable to avoid virologic failure from literature is 1–4 μg/mL for efavirenz [40]. Three samples, i.e., HBP1, HBP4, and HBP6, were either within or close to the therapeutic range while others were not.
Comparison with literature values
This method was superior to some values reported in the literature employing GC-MS/SIM for the determination of EFV since the LLOQ in this study was lower as shown in Table 3. The table also shows that a relatively smaller sample volume of real human plasma was used with DLLME, i.e., 50 μL, compared to 200 μL in the other two methods employing GC–MS in the SIM mode of signal acquisition. This is an advantage considering the small human bio fluid sample volumes (μL) that are accessible for bioanalysis. Kankaanpää et al. [41] proposed that this problem could be overcome by either diluting the samples or validating at higher concentrations if the LLOQs were low and where the therapeutic ranges were significantly higher, i.e., 1–4 μg/mL, as in the case of EFV. This view is upheld in this study where the LLOQ was estimated at 0.027 μg/mL. It is proposed here that these suggestions could be considered when reviewing bioanalytical method validation guidelines in the future. The percent recovery obtained in this study at the LLOQ spike level was better or comparable to the literature values. Table 3 also shows literature quality control values for EFV obtained using DLLME, LLE, and SPE followed by HPLC–UV detection. The major differences can be seen in the volumes of samples that were employed using traditional bioanalytical sample preparation methods. Other validation parameters were either inferior or superior compared to those obtained in this study.
Conclusion
A DLLME extraction procedure employing a GC–MS in the selected ion monitoring (SIM) method for the determination of EFV in human plasma has been developed. This is the first method employing DLLME and GC–MS–SIM reported in the literature. The procedure consumed extremely low volumes of samples and extraction solvents, i.e., in microliters, and was very simple and rapid to perform. These advantages in conjunction with the use of a very selective separation and detection system simplified this method to produce excellent limits of detection and quantification. For resource-poor countries, DLLME could be a method of choice for sample preparation. Using the USFDA guidelines, the method was partially validated and found suitable for use in clinical analysis because the therapeutic range for EFV was well above the LLOQ.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Neogi U, Häggblom A, Singh AK, Rogers LC, Rao SD, Amogne W, Schülter E, Zazzi M, Arnold E, Sarafianos SG, Sönnerborg A. Factors influencing the efficacy of rilpivirine in HIV-1 subtype C in low- and middle-income countries. J Antimicrob Chemother. 2016;71(2):367–71. https://doi.org/10.1093/jac/dkv359.
Hemelaar J, Gouws E, Ghys PD, et al. Global trends in molecular epidemiology of HIV-1 during 2000–2007. AIDS. 2011;25:679–89. https://doi.org/10.1097/QAD.0b013e328342ff93.
The NAMSAL ANRS 12313 Study Group. Dolutegravir-based or low-doseefavirenz–based regimen for the treatment of HIV-1. N Engl J Med. 2019;381:816–26. https://doi.org/10.1056/NEJMoa1904340.
World Health Organization. (WHO) (2019) Update of recommendations on first- and second-line antiretroviral regimens. Geneva, Switzerland: World Health Organization; (WHO/CDS/HIV/19.15). Licence: CC BY-NC-SA 3.0 IGO. Last Accessed 29th September 2020.
Hartman TL, Buckheit RW. The continuing evolution of HIV-1 therapy: identification and development of novel antiretroviral agents targeting viral and cellular targets. Mol Biol Int J. 2012;401965. https://doi.org/10.1155/2012/401965.
Hoen E, Berger J, Calmy A, Moon S. Driving a decade of change: HIV/AIDS, patents and access to medicines for all. J Intern AIDS Soc. 2011;14:15–27. https://doi.org/10.1186/1758-2652-14-15.
Arendt G, De-Nocker D, Von-Giesen HJ, Nolting T. Neuropsychiatric side effects of efavirenz therapy. Expert Opin Drug Saf. 2007;6:147–54. https://doi.org/10.1517/14740338.6.2.147.
Evans-Jones JG, Cottle LE, Back DJ, Gibbons S, Beeching NJ, Carey PB, Khoo SH. Recognition of risk for clinically significant drug interactions among HIV-infected patients receiving antiretroviral therapy. J Clin Infect Dis. 2010;50:1419–21. https://doi.org/10.1086/652149.
Gallant JE, DeJesus E, Arribas JR, Pozniak AL, Gazzard B, Campo RE, Lu B, McColl D, Chuck S, Enejosa J, Toole JJ, Cheng AK. Tenofovir DF, emtricitabine, and efavirenz vs. zidovudine, lamivudine and efavirenz for HIV. New Engl J Med. 2006;354:251–60. https://doi.org/10.1056/NEJMoa051871.
Russo G, Paganotti GM, Soeria-Atmadja S, Haverkamp M, Ramogola-Masire D, Vullo V, Gustafsson LL. Pharmacogenetics of non-nucleoside reverse transcriptase inhibitors (NNRTIs) in resource-limited settings: influence on antiretroviral therapy response and concomitant anti-tubercular, antimalarial and contraceptive treatments. Infect Genet Evol. 2016;37:192–207. https://doi.org/10.1016/j.meegid.2015.11.014.
Apostolova N, Funes HA, Blas-Garcia A, Galindo MJ, Alvarez A, Esplugues JV. Efavirenz and the CNS: what we already know and questions that need to be answered. J Antimicrob Chemother. 2015;70:2693–708. https://doi.org/10.1093/jac/dkv183.
Donnerer J, Kronawetter M, Kapper A, Haas I, Kessler HH. Therapeutic drug monitoring of the HIV / AIDS drugs abacavir, zidovudine, efavirenz, nevirapine, indinavir, lopinavir, and nelfinavir. Aust J Pharm. 2003;69:197–204. https://doi.org/10.1159/000073664.
Langmann P, Zilly M, Weissbrich B, Desch S, Väth T, Klinker H. Therapeutic drug monitoring of Indinavir in HIV-infected patients undergoing HAART. J Infect Dis. 2002;30:13–6. https://doi.org/10.1007/s15010-001-1111-0.
Rezk NL, Tidwell RR, Kashuba ADM. High-performance liquid chromatography assay for the quantification of HIV protease inhibitors and non-nucleoside reverse transcriptase inhibitors in human plasma. J Chromatogr B. 2004;805:241–7. https://doi.org/10.1016/j.jchromb.2004.03.002.
Sarasa-Nacenta M, Lopez-pua Y, Lopez-Cortes LF, Mallolas J, Gatell JM, Carne X. Determination of efavirenz in human plasma by high-performance liquid chromatography with ultraviolet detection. J Chromatogr B. 2001;763:53–9. https://doi.org/10.1016/s0378-4347(01)00357-7.
Dailly E, Raffi F, Jolliet P. Determination of atazanavir and other antiretroviral drugs (indinavir, amprenavir, nelfinavir and its active metabolite M8, saquinavir, ritonavir, lopinavir, nevirapine and efavirenz) plasma levels by high performance liquid chromatography with UV detection. J Chromatogr B. 2004;813:353–8. https://doi.org/10.1016/j.jchromb.2004.10.005.
Ramachandran G, Kumar AKH, Swaminathan S, Venkatesan P, Kumaraswami V, Greenblatt DJ. Simple and rapid liquid chromatography method for determination of efavirenz in plasma. J Chromatogr. 2006;835(2006):131–5. https://doi.org/10.1016/j.jchromb.2006.03.014.
Fox D, Connor RO, Mallon P, Mcmahon G. Simultaneous determination of efavirenz, rifampicin and its metabolite desacetyl rifampicin levels in human plasma. J Pharmaceut Biomed Anal. 2011;56:785–91. https://doi.org/10.1016/j.jpba.2011.07.041.
Hoffman JT, Rossi SS, Espina-Quinto R, Letendre S, Capparelli EV. Determination of efavirenz in human dried blood spots by reversed-phase high-performance liquid chromatography with UV detection. J Ther Drug Monit. 2013;35:203–8. https://doi.org/10.1097/FTD.0b013e31827fb72b.
Ren C, Fan-Havard P, Schlabritz-Loutsevitch N, Ling Y, Chan KK, Liu Z. A sensitive and specific liquid chromatography/tandem mass spectrometry method for quantification of nevirapine and its five metabolites and their pharmacokinetics in baboons. J Biomed Chromatogr. 2009;24:717–26. https://doi.org/10.1002/bmc.1353.
Theron A, Cromarty D, Rheeders M, Viljoen M. Determination of salivary efavirenz by liquid chromatography coupled with tandem mass spectrometry. J Chromatogr B. 2010;878:2886–90. https://doi.org/10.1016/j.jchromb.2010.08.051.
Zhang L, Sun J, Chen J, Jia XF, Shen F, Lu HZ. An LC–MS/MS method for simultaneous quantification of seven anti-HIV medicines in plasma of HIV-infected patients. Pharm Anal Acta. 2010;1:102. https://doi.org/10.4172/2153-2435.1000102.
Soldin SJ, Soukhov N, Janici N, Jonklaas J, Soldin OP. The measurement of free thyroxine by isotope dilution tandem mass spectrometry. Clin Chimica Acta. 2005;358:113–8. https://doi.org/10.1016/j.cccn.2005.02.010.
Sichilongo K, Mwando E, Sepako E, Massele A. Comparison of efficiencies of selected sample extraction techniques for the analysis of selected antiretroviral drugs in human plasma using LC-MS. J Pharmacol Toxicol Methods. 2018;89:1–8. https://doi.org/10.1016/j.vascn.2017.10.001.
Srivastava P, Moorthy GS, Gross R, Barrett JS. A sensitive and selective liquid chromatography/tandem mass spectrometry method for quantitative analysis of efavirenz in human plasma. PLoS One. 2013;8:1–9. https://doi.org/10.1371/journal.pone.0063305.
Rouzes A, Karine B, Xuere F, Djabarouti S, Pellegrin I, Pellegrin JL, Coupet AC, Augagneur, Budzinski H, Saux MC, Breilh D. Simultaneous determination of the antiretroviral agents: amprenavir , lopinavir, ritonavir, saquinavir and efavirenz in human peripheral blood mononuclear cells by high-performance liquid chromatography – mass spectrometry. J Chromatogr B. 2004;813:209–16. https://doi.org/10.1016/j.jchromb.2004.09.041.
Huang Y, Gandhi M, Greenblatt RM, Gee W, Lin EST, Messenkoff N. Sensitive analysis of anti-HIV drugs, efavirenz, lopinavir and ritonavir, in human hair by liquid chromatography coupled with tandem mass spectrometry. Rapid Commun Mass Spectrom. 2008;22:3401–9. https://doi.org/10.1002/rcm.3750.
Lemmer P, Schneider S, Schuman M, Omes C, Arendt V, Tayari JC, Fundira L, Wennig R. Determination of nevirapine and efavirenz in plasma using GC/MS in selected ion monitoring mode. J Ther Drug Monit. 2002;27:521–5. https://doi.org/10.1097/01.ftd.0000164613.91555.2e.
González O, Alonso RM. Validation of bioanalytical chromatographic methods for the quantification of drugs in biological fluids. Handbook of Analytical Separations. 2020;7:115–34.
Ridgway K (2019). https://www.anatune.co.uk/wp-content/uploads/2019/04/AS224-Automated-solvent-extraction-LLME-and-DLLME-of-Flavours-from-a-selection-of-fruit-beverages.pdf last accessed 3rd august 2020.
USFDA (2018). Bioanalytical method validation guidance for industry, https://www.fda.gov/media/70858/download last accessed 7th august 2020.
Azadeh M, Gorovits B, Kamerud J, MacMannis S, Safavi A, Sailstad J, Sondag P. Calibration curves in quantitative ligand binding assays: recommendations and best practices for preparation, design, and editing of calibration curves. AAPS J. 2018;20:22. https://doi.org/10.1208/s12248-017-0159-430.
Cheng CL, Chou CH, Yoa-Pu Huc O. Determination of lamotrigine in small volumes of plasma by high-performance liquid chromatography. J Chromatogr B. 2005;817:199–206. https://doi.org/10.1016/j.jchromb.2004.12.004.
Cheng CL, Chou CH, Yoa-Pu Huc O. Determination of delavirdine in very small volumes of plasma by high-performance liquid chromatography with fluorescence detection. J Chromatogr B. 2002;769:297–303. https://doi.org/10.1016/S1570-0232(02)00015-6.
Heydari R, Rashidipour M, Naleini N. Determination of efavirenz in plasma by dispersive liquid-liquid micro-extraction coupled to high-performance liquid chromatography. Current Anal Chem. 2014;10:280–7. https://doi.org/10.2174/15734110113099990003.
Polson C, Sarkar P, Incledon B, Raguvaran V, Grant R. Optimization of protein precipitation based upon effectiveness of protein removal and ionization effect in liquid chromatography– tandem mass spectrometry. J Chromatogr B. 2003;785:263–75. https://doi.org/10.1016/S1570-0232(02)00914-5.
Toribio-Delgado AF, Robles-Gil MC, Maynar-Mariño M, Olcina-Camacho G, Maynar-Mariño JI. Is deproteinization necessary in the determination of human plasmatic steroids by GC/IT-MS/MS analysis? J Applied Bioanal. 2015;1:10–8. https://doi.org/10.17145/jab.15.003(ISSN 2405-710X).
Wisconsin Department of Natural Resources Laboratory Certification Program (1996) Analytical detection limit guidance & laboratory guide for determining method detection limits. https://dnr.wi.gov/regulations/labcert/documents/guidance/-LODguide.pdf Last Accessed 8th March 2021.
Matuszewski BK, Constanzer ML, Chavez-Eng CM. Strategies for the assessment of matrix effect in quantitative bioanalytical methods based on HPLC - MS / MS. Intern J Anal Chem. 2003;75:3019–30. https://doi.org/10.1021/ac020361s.
Bednasz CJ, Venuto CS, Ma Q, Daar ES, Sax PE, Fischl MA, Collier AC, Smith KY, Tierney C, Yang Y, Wilding GE, Morse GD. Efavirenz therapeutic range in HIV-1 treatment-naive participants. Ther Drug Monit. 2017;39:596–603. https://doi.org/10.1097/FTD.0000000000000443.
Kankaanpää A, Turtiainen S, Meririnne E, Ariniemi K, Ristola M, Kuoppasalmi K. Determination of the non-nucleoside reverse transcriptase inhibitor efavirenz in human plasma by gas chromatography mass spectrometry. Ther Drug Mont. 2007;29:513–4.
Acknowledgements
The authors thank the University of Botswana and the Ministry of Health and Wellness for granting ethical approval.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
Ethical approval to conduct this study was obtained from the University of Botswana Office of Research and Development (Reference Number RES/IRB/CHMS/014) and Human Research Development Division (HRDD) (Reference Number HPDME: 13/18/1).
Consent to participate
Informed consent was obtained from all participants before enrolment in the study.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Masenga, W., Paganotti, G.M., Seatla, K. et al. A fast-screening dispersive liquid–liquid microextraction–gas chromatography–mass spectrometry method applied to the determination of efavirenz in human plasma samples. Anal Bioanal Chem 413, 6401–6412 (2021). https://doi.org/10.1007/s00216-021-03604-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-021-03604-0