Abstract
C-Reactive protein (CRP) is a biomarker of inflammatory responses and an index for assessing the risk of cardiovascular disease and estimating prognosis. In this study, we constructed a surface-enhanced Raman spectroscopy (SERS) biosensor composed of a multifunctional DNA three-way junction (DNA 3WJ), porous gold nanoplates (pAuNPs), and an Au-Te nanoworm structure for detection of CRP. The pAuNP and Au-Te nanostructures were synthesized by galvanic replacement reactions, and the morphology was confirmed by transmission electron microscopy, scanning electron microscopy, and dynamic light scattering (DLS). To generate the SERS signal, the Au-Te nanostructure was immobilized on an indium-tin oxide substrate, and the thiol-modified CRP aptamer was then self-assembled onto the modified substrate for CRP recognition. To amplify the SERS signal and identify the Raman tag, the multifunctional DNA 3WJ was conjugated with the pAuNPs, and each fragment of 3WJ was functionalized to biotin (pAuNP conjugation), methylene blue (Raman reporter), and CRP aptamer (target binding). The results were confirmed by gel electrophoresis. For conjugation between pAuNPs and DNA 3WJ, avidin was encapsulated in pAuNPs, and the conjugation structure was confirmed by DLS. The fabricated SERS biosensor showed detection limits of 2.23 pM in phosphate-buffered saline and 3.11 pM in diluted human serum. Overall, the proposed biosensor may have potential applications as a SERS biosensor platform.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Inflammation occurs acutely or chronically as the body’s biological immune response to stimulation. In particular, in cases of chronic inflammation, metabolic disease and chronic inflammation are closely related as a factor that causes cardiovascular disease [1, 2]. Cardiovascular disease can be diagnosed early if the level of inflammation is continuously monitored. Therefore, detection of inflammatory biomarkers is important in clinical practice.
C-Reactive protein (CRP), a plasma protein discovered in 1930, is synthesized by the liver in response to infection, inflammatory disease, and malignant tumors and is then secreted into the bloodstream, where it can act as a biological indicator of systemic inflammatory reactions [3, 4]. The American Heart Association (AHA) uses high-sensitivity C-reactive protein (hs-CRP) as an index to assess the risk of heart disease and estimate prognosis. The acceptable precision level is 0.3 μg/mL or less, and concentrations greater than this are thought to be associated with a high risk of cardiovascular disease [5].
General techniques for detecting such CRP include chemiluminescent immunoassay and enzyme-linked immunosorbent assay (ELISA) [6, 7]. However, ELISA technology is complicated and requires the use of antibodies, which increases the test cost. In order to overcome these shortcomings, researchers have attempted to develop novel approaches for sensitive detection of CRP. Methods for detection by electrochemistry [8, 9], fluorescence [10, 11], and surface plasmon resonance [12, 13] have also been studied. Among various detection methods, Raman spectroscopy, which is an easy approach for measurement and does not cause loss of sample, has attracted much attention. Raman spectroscopy is capable of multisignal labeling, and the use of surface-enhanced Raman scattering (SERS) using nanoparticles has the advantage of obtaining a signal sensitivity comparable to that of fluorescence [14, 15].
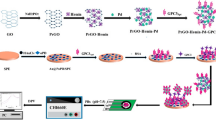
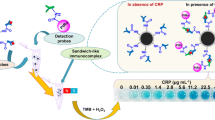
In this study, we developed a novel SERS biosensor consisting of multifunctional DNA three-way junction (DNA 3WJ)-conjugated porous gold nanoplates (pAuNPs) and an Au-Te nanoworm particle to detect CRP. To fabricate the SERS biosensor, we fabricated a substrate using Au-Te nanoworm particles on an indium-tin oxide (ITO) substrate used as the SERS platform to confirm the possibility of detecting protein signals [16]. Then, the thiol-modified CRP aptamer was anchored onto the Au-Te nanoworm by covalent bonding. After CRP was added onto the substrate, the second probe was added to amplify the SERS signal. Moreover, a multifunctional DNA 3WJ was designed to perform three functions within a single biomolecule [17,18,19]. Three pieces of single-stranded nucleic acids were modified to generate Raman signals, specifically binding to CRP and pAuNPs. The first single strand of DNA contained an aptamer sequence that specifically bound to CRP, and the second was labeled with methylene blue (MB) at the end of the DNA to generate a Raman signal. Finally, biotin was labeled for conjugation on the surface of nanoparticles, and a sandwich CRP detection probe was fabricated through assembly of these three strands. To further enhance the SERS signal, pAuNPs with a large surface area were synthesized, and avidin and MB were bound to the pAuNPs by electrostatic attraction (avidin/MB/pAuNPs). Next, biotin-labeled multifunctional 3WJ was conjugated as a secondary CRP detection probe. On the Au-Te nanoworm-modified ITO substrate, the CRP aptamer sequence that specifically bound to CRP was selected for the first capture of CRP, and the multifunctional DNA 3WJ/pAuNPs amplified the signal of MB on the surface. We were then able to detect the presence of CRP through the amplified Raman signal of MB. The overall schematic is shown in Fig. 1.
Materials and methods
Materials
ITO-glass (3 × 1 cm) was purchased from National Nanofab Center (10 Ω resistance; Daejeon, Korea). Recombinant human CRP was purchased from Sino Biological (Beijing, China). Triton-X solution, triethoxysilane, chloroauric acid, hydrazine monohydrate hydrate, poly (vinylpyrrolidone) (molecular weight [Mw] = 40 kDa), sodium tellurite, sodium hydroxide, ethylene glycol, avidin from egg white, MB, l-ascorbic acid, and citric acid were purchased from Sigma-Aldrich (St. Louis, MO, USA). Hydrogen tetrachloroaurate (III) hydrate was purchased from Kojima Chemicals Co. (Japan). Silver nitrate, hydrogen peroxide (30%), trisodium citrate dihydrate, and sodium borohydride were all purchased from Junsei (Japan). The sequences of the strands used as CRP probes were as follows: thiol-modified CRP capture aptamer, 5’-SH-GCCTGTAAGGTGGTCGGTGTGGCGAGTGTGTTAGGAGAGATTGC-3’ [20]; CRP aptamer-tagged DNA 3WJ-a, 5’-CGAAGGGGATTCGAGGGGTGATTGCGTGCTCCATTTGGTTTGCCATGTGTATGTGGG-3’ [21]; MB-tagged 3WJ-b, 5’-MB-CCCACATACTTTGTTGATCC-3’; biotin-tagged DNA 3WJ-c, 5’-biotin-GGATCAATCATGGCAA-3’.
Preparation of pAuNPs
pAuNPs were manufactured using the galvanic replacement reaction with a sacrificial Ag nanoplate template, as previously reported [22]. Briefly, 250 μL of 10 mM AgNO3, 1.5 mL of 3.5 mM PVP (Mw = 29 kDa) aqueous solution, 300 μL of 30 mM sodium citrate, and 24.75 mL deionized (DI) water were sequentially placed in 50-mL transparent glass vials. Then, 60 μL of 30% hydrogen peroxide and 250 μL of 100 mM NaBH4 solution were rapidly injected in sequence. After 3 h of incubation with magnetic stirring, 1 mL of 100 mM l-ascorbic acid and 330 μL of 75 mM sodium citrate were added to the as-grown Ag nanoseed. Then, 13 mL growth solution was added at a rate of 0.2 mL/s. The preparation of growth solution was accomplished by mixing of 2 mL of 10 mM AgNO3, 125 μL of 100 mM citric acid, and 17.875 mL DI water. Along with the addition of growth solution, the mixture color gradually changed from pale blue to deep blue. The synthesized Ag nanoplates were directly applied to galvanic replacement to obtain pAuNPs without any purification step.
As-synthesized Ag nanoplates were diluted five times with distilled water, and 20 v/v% of 1 mM AuCl4- aqueous stock solution was then rapidly injected under vigorous magnetic stirring (~500 rpm). After incubation for 2 h, the product was purified by centrifugation at 9000 rpm for 10 min and washed with DI water at least three times. Purified pAuNPs were redispersed at the initial volume of the Ag nanoplates, and the concentration was denoted as 1 volumetric equivalent (eq.).
Preparation of Au-Te nanoworms
Au-Te nanoworms were prepared by galvanic replacement of a sacrificial Te nanorod template, as previously reported [23]. To a 50-mL transparent glass vial, we added 1 g PVP (Mw = 40 kDa), 92.2 mg sodium tellurite, and 0.5 mg NaOH and fully dissolved these components in 40 mL ethylene glycol as the solvent. Next, 1.3 mL hydrazine hydrate was injected for initiation of Te nanorod formation and incubated for 3 h at 70°C in a water bath under gentle magnetic stirring (~300 rpm). Synthesized Te nanorods were purified by dialysis (30-kDa cutoff) in a DI water reservoir to eliminate unreacted ions and reductants.
Next, 250 μL of 10 mM AuCl4- aqueous stock solution was rapidly injected into 2 mL purified Te nanorods, and the mixture was incubated for 2 h at room temperature. After the reaction, Au-Te was purified by centrifugation at 8000 rpm for 10 min and washed with DI water at least three times to ensure the removal of dissolved ions. Finally, manufactured Au-Te was redispersed at the initial volume of the Te nanorods, and the concentration was denoted as 1 volumetric equivalent (eq.).
Characterization
Nanoparticles were characterized by energy-filtering transmission electron microscopy (TEM; LIBRA 120; Carl Zeiss, Germany) and field-emission scanning electron microscopy (SEM; SUPRA 55VP; Carl Zeiss). Dynamic light scattering (DLS) and zeta-potential were measured using a Zetasizer Nano ZS (Malvern, UK). The ultraviolet-visible (UV-Vis) spectra of nanoparticles were obtained using a Lambda 465 UV-Vis spectrophotometer (PerkinElmer, USA).
Preparation of multifunctional DNA 3WJ-conjugated pAuNPs
To prepare CRP Apt/biotin/MB-tagged DNA 3WJ, we used a protocol described in a previous study [18]. The three single strands were diluted in TMS buffer (50 mM Tris-HCl, 10 mM MgCl2, 100 mM NaCl) at the same molar ratio, heated to 80°C for 5 min, slowly cooled to 4°C, and stored in a refrigerator. The assembled DNA 3WJ was identified on an 8% native TBM polyacrylamide gel. For conjugation of the multifunctional DNA 3WJ to pAuNPs with signal amplification function, pAuNPs were reacted with avidin to bind the surface of pAuNPs via electrostatic attraction. Then, 0.1 mg/mL avidin (20 μL) and 100 mM HEPES buffer (100 μL) were added to pAuNPs (4 eq.) solution, and the solution was incubated at 4°C for 24 h. Subsequently, 5 μL of 1 μM MB was added, the solution was incubated at room temperature for 2 h and centrifuged at 4000 rpm for 10 min, the supernatant was discarded, and the pellet was washed with 10 mM HEPES solution. The washing process was carried out three times. Next, CRP Apt/biotin/MB-tagged DNA 3WJ solution in 10 μM TMS (10 μL) was added to a total volume of 100 μL, and samples were incubated at room temperature for 2 h. Finally, samples were washed three times. The CRP Apt/biotin/MB-tagged DNA 3WJ-conjugated avidin/MB-encapsulated pAuNPs were purified with an Amicon Ultra centrifugal filter (MWCO: 50k; Millipore, USA) [24]. The prepared SERS tag could capture CRP and generate Raman signals simultaneously.
Fabrication of the SERS biosensor
To fabricate the SERS biosensor, an ITO substrate immobilized with Au-Te was used [16]. All DNA was heat-treated at 80°C for 5 min before use, slowly cooled to 4°C, washed, and dried with DIW and nitrogen gas at each step. Thiol-modified CRP capture aptamer (1 μM, 30 μL) was immobilized on the prepared Au-Te substrate and incubated for 24 h at room temperature. Thereafter, 30 μL CRP solution was placed on the substrate and reacted at room temperature for 3 h, and 30 μL of the finally prepared SERS tag solution was placed on the substrate, followed by incubation for 3 h. Measurements were performed using a SENTERRA confocal Raman spectroscope (Bruker Optics, USA) with a 785-nm diode laser, 10-mW power, and exposure time of 5 s for all experiments. Results were obtained from 10 different points for 10 independent samples.
Results and discussion
Characterization of pAuNPs and Au-Te nanoworms
The synthesis of pAuNPs and Au-Te nanoworms was performed using the galvanic replacement reaction for sacrificial Ag nanoplates and the Te nanorod template, respectively [22, 23]. The size and morphology of pAuNPs and Au-Te nanoworms were characterized by TEM and SEM. According to the obtained images, platelet two-dimensional pAuNPs with a rough surface morphology and partial porosity were clearly identified from pAuNPs (Fig. 2a). For Au-Te nanoworms, twisted one-dimensional worm-like irregular nanostructures were successfully manufactured (Fig. 2b). Both distinguished nanostructures originated from the characteristics of sacrificial nanotemplates.
DLS measurement of pAuNPs and Au-Te nanoworms exhibited average hydrodynamic diameters of 128.8 and 101.5 nm, respectively (Fig. 2c). Because of the rough surface morphology and one- and two-dimensional characteristics, Stoke-Einstein equation-mediated size calculation of DLS provided slightly different size dimensions identified from TEM and SEM images. However, the polydispersity indices of pAuNPs (0.165) and Au-Te nanoworms (0.353) indicated the degree of reliability based on their irregular morphology (Fig. 2d). The negative ζ-potential values of pAuNPs (−23.9 mV) and Au-Te nanoworms (−18.3 mV) showed similarities derived from the environment of the exposed metallic gold surface of nanoparticles.
Confirmation of the assembly of the multifunctional bioprobe
To confirm the binding affinity of the CRP aptamer, gel electrophoresis (8% native TBE-polyacrylamide gel electrophoresis [PAGE]) was performed. Figure 3a shows the expected two-dimensional CRP aptamer structure used in this study, and Fig. 3b shows the expected two-dimensional assembled multifunctional DNA 3WJ structure used in this study. The two images were obtained from UNAfold. Figure 3c shows the TBE-PAGE results of the selective binding of the CRP aptamer to CRP. In a comparison of the CRP capture probe (lane 2) and the CRP capture probe reacted with CRP (lane 3), band migration was confirmed in lane 3. As a result, the size of the band appeared to change owing to changes in the three-dimensional structure as DNA bound to CRP. In addition, a control experiment with myoglobin was conducted to confirm that myoglobin, another plasma protein, did not bind to other proteins. Notably, CRP capture probe plus myoglobin (lane 4) showed the same band shape as CRP capture probe alone (lane 2), indicating that myoglobin did not bind to proteins other than CRP.
(a) Schematic image of expected 2D structure of CRP aptamer. (b) Schematic image of expected multi-functional DNA 3WJ structure from 3WJ-a of CRP aptamer for detection (red line), 3WJ-b for signal reporter (blue line), and biotin modified 3WJ-c for conjugation (purple line). (c) 8% TBE PAGE gel result of CRP aptamer, CRP and myoglobin, (d) TBM PAGE gel result of multi-functional DNA 3WJ, CRP and myoglobin. Results of investigation of pAuNP SERS tag (e) diameter of fabricated each particle, (f) zeta potential of each fabricated particle surface
Furthermore, the assembly results of the multifunctional DNA 3WJ and binding tests between multifunctional DNA 3WJ and CRP were confirmed by 8% native TBM-PAGE (Fig. 3d). The Apt/MB/Bio-tagged DNA 3WJ band (lane 5) was heavier than each DNA 3WJ band. These findings confirmed that the assembly of multifunctional DNA 3WJ was successful. Additionally, we compared the binding of the assembled DNA 3WJ and CRP. The shape of the band changed owing to the slight increase in the height of the band; thus, the DNA became heavier through binding with CRP, and the three-dimensional structure changed. In addition, for the control, a band position on the same line as DNA 3WJ was observed, confirming selective binding only to CRP. As a result, we confirmed that the DNA probes selectively bound only to CRP, and the assembly of DNA 3WJ was also successful in further experiments.
To confirm the conjugation of multifunctional DNA 3WJ with pAuNPs, the particle size and zeta potential were measured. Figure 3e shows the results of measuring the particle diameter; we found that the particle diameter size changed as avidin and DNA 3WJ bound to pAuNPs. Figure 3f shows the results of zeta potential measurements for the particle surface at each stage. Synthesized pAuNPs had a negative charge on the surface and therefore spontaneously bound to avidin with an isoelectric point of 10. As avidin bound to the surface, the charge of the entire particle became positive. Subsequently, it was possible to confirm that the surface of the particle was negatively charged again through binding with DNA 3WJ. As a result, we confirmed that pAuNPs and avidin bound well at each step and that DNA 3WJ was evenly distributed over the particles.
Investigation of the feasibility of the fabricated SERS biosensor
The fabricated biosensor was confirmed by SERS (Fig. 4). After placement of the capture probe on the Au-Te substrate, a blue signal was observed. By contrast, a green signal was observed when applying the combination of the capture probe and CRP, and a red line with peaks at 447, 496, 765, 1388, and 1614 cm-1 was observed when combining the SERS tag. This seemed to be caused by MB in the SERS tag produced as a peak region characteristic of MB [25]. The capture probe and CRP were not labeled, and the characteristic peak changes could not be confirmed. However, because the peak of the SERS tag specifically binding to CRP was confirmed, the capture probe, CRP, and SERS tag were thought to all be immobilized on the substrate. Thus, the manufactured biosensor could effectively detect CRP.
SERS biosensor performance
Raman spectra were obtained at various concentrations of CRP, and the detection limit was estimated by comparing the intensity at 447 cm-1 (Fig. 5a). The test was performed in a buffer of 100 nM to 1 pM and in serum. The intensity at the 447 cm-1 gradually decreased as the CRP concentration decreased. A linear fitting, i.e., y = 1397.9 ln(x) + 38126, was observed (Fig. 5b), and the correlation coefficient (R2) was 0.9654. The detection limit was calculated as 3 (standard deviation/slope of the linear calibration curve), and the detection limit in the buffer was 2.23 pM. In addition, the Raman spectrum was measured in human serum. CRP was serially diluted from 100 nM to 1 pM in 10% serum. The detection result is shown in Fig. 5c, and Fig. 5d shows the results of the linear fitting, i.e., y = 1922.8ln[x] + 52271 (correlation coefficient [R2] = 0.8913). The limit of detection was determined to be 3.11 pM. The results of the detection limit were compared with those in a previous study (Table 1). Figure 6 shows the selectivity test of the CRP SERS biosensor. Hemoglobin, interferon-γ, myoglobin, and troponin, which are all present in human serum, were used as controls. The results confirmed that the manufactured biosensor had high selectivity for CRP and could be detected in human serum.
Evaluation of CRP SERS biosensor performance: (a) Raman spectrum in different concentrations of CRP (PBS buffer), (b) calibration curve in different concentrations of CRP in buffer; linear range from 100 nM to 1 pM, (c) Raman spectrum in different concentrations of CRP (serum), (d) calibration curve in different concentrations of CRP in serum; linear range from 100 nM to 1 pM
Conclusion
In this study, we fabricated a SERS biosensor composed of multifunctional DNA 3WJ/pAuNP conjugates and Au-Te nanoworm substrate for CRP detection. The prepared Au-Te nanoworm structure showed usefulness as a CRP biosensor when combined with a thiol-tagged CRP aptamer. Through the versatility of DNA 3WJ, DNA 3WJ can easily exert different functionalities, including target recognition, signal reporter, and conjugation with nanoparticles. Additionally, pAuNPs can be easily combined with MB and avidin, and through this modification, it is possible to easily create a SERS tag that can detect CRP and generate Raman signals. The multifunctional DNA 3WJ/pAuNP conjugates, capable of detecting CRP with Raman signal amplification, provided a useful SERS tag. As the concentration of CRP decreased, the Raman signal intensity of the SERS tag also gradually decreased, enabling the detection limit of the fabricated biosensor to be estimated. We therefore identified detection limits of 2.23 pM in buffer and 3.11 pM in human serum. Based on these results, the proposed biosensor shows potential applications as a SERS biosensor platform. Thus, the combination of DNA 3WJ and nanoparticles may enable the development of various pathogen detection SERS platforms.
References
Ridker PM, Hennekens CH, Buring JE, Rifai N. C-reactive protein and other markers of inflammation in the prediction of cardiovascular disease in women. N Engl J Med. 2000;342:836–43.
Timpson NJ, Lawlor DA, Harbord RM, Gaunt TR, Day INM, Palmer LJ, et al. C-reactive protein and its role in metabolic syndrome: mendelian randomisation study. Lancet. 2005;366:1954–9.
Tillett WS, Francis T. Serological reactions in pneumonia with a non-protein somatic fraction of pneumococcus. J Exp Med. 1930;52:561–71.
Danesh J, Wheeler JG, Hirschfield GM, Eda S, Eiriksdottir G, Rumley A, et al. C-reactive protein and other circulating markers of inflammation in the prediction of coronary heart disease. N Engl J Med. 2004;350:1387–97.
Myers GL, Rifai N, Tracy RP, Roberts WL, Alexander RW, Biasucci LM, et al. CDC/AHA workshop on markers of inflammation and cardiovascular disease: application to clinical and public health practice: report from the laboratory science discussion group. Circ J. 2004;110:e545–9.
Parra MD, Tuomola M, Cabezas-Herrera J, Cerón JJ. Use of a time-resolved immunofluorometric assay for determination of canine C-reactive protein concentrations in whole blood. Am J Vet Res. 2005;66:62–6.
Clarke JL, Anderson JL, Carlquist JF, Roberts RF, Horne BD, Bair TL, et al. Comparison of differing C-reactive protein assay methods and their impact on cardiovascular risk assessment. Am J Cardiol. 2005;95:155–8.
Bryan T, Luo X, Bueno PR, Davis JJ. An optimised electrochemical biosensor for the label-free detection of C-reactive protein in blood. Biosens Bioelectron. 2013;39:94–8.
Wang J, Guo J, Zhang J, Zhang W, Zhang Y. RNA aptamer-based electrochemical aptasensor for C-reactive protein detection using functionalized silica microspheres as immunoprobes. Biosens Bioelectron. 2017;95:100–5.
Cai Y, Kang K, Liu Y, Wang Y, He X. Development of a lateral flow immunoassay of C-reactive protein detection based on red fluorescent nanoparticles. Anal Biochem. 2018;556:129–35.
Zhu X, Duan D, Publicover NG. Magnetic bead based assay for C-reactive protein using quantum-dot fluorescence labeling and immunoaffinity separation. Analyst. 2010;135:381–9.
Vance SA, Sandros MG. Zeptomole detection of C-reactive protein in serum by a nanoparticle amplified surface plasmon resonance imaging aptasensor. Sci Rep. 2014;4:5129.
Hu WP, Hsu HY, Chiou A, Tseng KY, Lin HY, Chang GL, et al. Immunodetection of pentamer and modified C-reactive protein using surface plasmon resonance biosensing. Biosens Bioelectron. 2006;21:1631–7.
Stoerzinger KA, Lin JY, Odom TW. Nanoparticle SERS substrates with 3D Raman-active volumes. Chem Sci. 2011;2:1435–9.
Lee T, Mohammadniaei M, Zhang H, Yoon J, Choi HK, Guo S, et al. Single functionalized pRNA/gold nanoparticle for ultrasensitive microRNA detection using electrochemical surface-enhanced Raman spectroscopy. Adv Sci. 2020;7:1902477.
Kim SM, Lee T, Gil YG, Kim GH, Park C, Jang H, et al. Fabrication of bioprobe self-assembled on Au–Te nanoworm structure for SERS biosensor. Materials. 2020;13:3234.
Lee T, Park SY, Jang H, Kim GH, Lee Y, Park C, et al. Fabrication of electrochemical biosensor consisted of multi-functional DNA structure/porous au nanoparticle for avian influenza virus (H5N1) in chicken serum. Mater Sci Eng Res Rep. 2019;99:511–9.
Park SY, Kim J, Yim G, Jang H, Lee Y, Kim SM, et al. Fabrication of electrochemical biosensor composed of multi-functional DNA/rhodium nanoplate heterolayer for thyroxine detection in clinical sample. Colloids Surf B. 2020;195:111240.
Lee T, Lee Y, Park SY, Hong K, Kim Y, Park C, et al. Fabrication of electrochemical biosensor composed of multi-functional DNA structure/Au nanospike on micro-gap/PCB system for detecting troponin I in human serum. Colloids Surf B. 2019;175:343–50.
Jarczewska M, Rębiś J, Górski Ł, Malinowska E. Development of DNA aptamer-based sensor for electrochemical detection of C-reactive protein. Talanta. 2018;189:45–54.
Wu B, Jiang R, Wang Q, Huang J, Yang X, Wang K, et al. Detection of C-reactive protein using nanoparticle-enhanced surface plasmon resonance using an aptamer-antibody sandwich assay. ChemComm. 2016;52:3568–71.
Kang S, Kang K, Huh H, Kim H, Chang SJ, Park TJ, et al. Reducing agent-assisted excessive galvanic replacement mediated seed-mediated synthesis of porous gold nanoplates and highly efficient gene-thermo cancer therapy. ACS Appl Mater Interfaces. 2017;9:35268–78.
Kang S, Gil YG, Kim YJ, Kim YK, Min DH, Jang H. Environmentally Friendly Synthesis of Au–Te-clustered nanoworms via galvanic replacement for wavelength-selective combination cancer therapy. ACS Appl Mater Interfaces. 2020;12:5511–9.
Lee T, Kim SU, Min J, Choi JW. Multilevel biomemory device consisting of recombinant azurin/cytochrome c. Adv Mater. 2010;22:510–4.
Zhao Y, Li X, Du Y, Chen G, Qu Y, Jiang J, et al. Strong light–matter interactions in sub-nanometer gaps defined by monolayer graphene: toward highly sensitive SERS substrates. Nanoscale. 2014;6:11112–20.
António M, Ferreira R, Vitorino R, Daniel-da-Silva AL. A simple aptamer-based colorimetric assay for rapid detection of C-reactive protein using gold nanoparticles. Talanta. 2020;214:120868.
Kim BB, Im WJ, Byun JY, Kim HM, Kim MG, Shin YB. Label-free CRP detection using optical biosensor with one-step immobilization of antibody on nitrocellulose membrane. Sensors Actuators B Chem. 2014;190:243–8.
Choi YH, Ko H, Lee GY, Chang SY, Chang YW, Kang MJ, et al. Development of a sensitive SPR biosensor for C-reactive protein (CRP) using plasma-treated parylene-N film. Sensors Actuators B Chem. 2015;207:133–8.
Russell SM, de la Rica R. Paper transducers to detect plasmon variations in colorimetric nanoparticle biosensors. Sensors Actuators B Chem. 2018;270:327–32.
Nooney R, Rebello V, Keegan G, O'Connell CL, Byrne D, McDonagh C. Highly sensitive detection of C-reactive protein using a novel dissolution approach in a dye-doped silica nanoparticle-based fluorescence immunoassay. Anal Methods. 2017;9:994–1003.
Lee WB, Chen YH, Lin HI, Shiesh SC, Lee GB. An integrated microfluidic system for fast, automatic detection of C-reactive protein. Sensors Actuators B Chem. 2011;157:710–21.
Funding
This work was supported by a National Research Foundation of Korea (NRF) grant funded by the Korean government (MSIT) (No. 2021R1C1C1005583) and by the Ministry of Trade, Industry, and Energy (Grant. 10062995) and by Korea Ministry of Environment (MOE) as Program (2020003030001) and by was supported by BioNano Health-Guard Research Center funded by the Ministry of Science and ICT(MSIT) of Korea as Global Frontier Project (Grant number H-GUARD_2018M3A6B2057261 and by the Excellent researcher support project of Kwangwoon University in 2021
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Published in the topical collection Point-of-Care Testing with guest editors Oliver Hayden, Peter B. Luppa, and Junhong Min.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kim, S.M., Kim, J., Yim, G. et al. Fabrication of a surface-enhanced Raman spectroscopy-based analytical method consisting of multifunctional DNA three-way junction-conjugated porous gold nanoparticles and Au-Te nanoworm for C-reactive protein detection. Anal Bioanal Chem 414, 3197–3204 (2022). https://doi.org/10.1007/s00216-021-03559-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-021-03559-2