Abstract
Airborne bacteria are components of the atmospheric aerosol particles and can be responsible of allergic disease, regardless of their viability. In this paper, we report a method for the determination of total (viable and nonviable) bacterial content in airborne particles, using muramic and dipicolinic acids as biomarkers of bacteria and bacterial spores, respectively. The analytical procedure was optimized with bacteria and spores of Bacillus subtilis. After extraction and purification, the two biomarkers were analyzed by HPLC-ESI-MS/MS and their percentage was evaluated to be used as conversion factor. The present method for the determination of the total bacterial content was then applied to environmental samples, after a proper collection in an urban site. Thanks to the use of a low pressure impactor, capable of fractionating particles into the range of 0.03–10 μm, it was also possible to study the bacterial content in ultrafine, fine, and coarse particulate matter. The results from this study showed that muramic and dipicolinic acids can be determined together in one chromatographic run in reversed phase ion pair chromatography. Bacteria were more abundant than bacterial spores in the urban atmosphere, both showing a higher concentration in the coarse fraction of particles, although bacteria and bacterial spore amounts per unit mass of ultrafine particles were higher than in fine and coarse particles.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Primary biological aerosol particles (PBAPs)—in addition to pollen, plant, and animal debris, epithelial cells, viruses, and fungi—also include a minor content of bacteria and bacterial spores [1–3], with the Bacillus genus generally found to be the most abundant single genus represented at urban, forest, coastal, and rural locations [4, 5] and generally with more abundant Gram-positive bacteria accounting for about 80∼86% of the total [6].
Bacterial content in atmosphere can have an influence on climate change as a consequence of the ability of bacteria to act as cloud condensation nuclei; furthermore, more importantly, they are human, plant, and animal pathogens [7–9]. Airborne bacteria cells can be released into the atmosphere from water and soil surfaces, vegetation, and anthropogenic sources (wastewater treatment plants, sewage treatment, animal rendering, fermentation processes, agricultural activities, etc.) [10–13]; global transport of dust also affects the amount and composition of bacterial occurrence in atmosphere [14].
Bacteria are generally attached to the atmospheric particles [15] and can therefore be present in all size ranges of the aerosol. The size of an airborne bacterial particle obviously affects the degree of infectivity as a function of its penetration coefficient and deposition rate in the human respiratory tract.
Besides cells, bioaerosol also includes bacterial spores; sporulation by certain bacteria results in the formation of a metabolically dormant cell known as endospore, which is resistant to severe physical and chemical conditions.
In determining airborne bacterial content, differences in sampling and/or analysis methods cause discrepancies in the reported literature data [16].
Bacterial aerosol sampling can be performed either by culturable air sampling methods or by collecting particulate on membrane filters. In the case of culturable samples, only viable microorganisms are detected. When bioaerosol is collected on filters, both viable and nonviable bacteria can be detected, depending on the analysis method used. Since biological aerosol can cause adverse health effects regardless of viability [17], in order to determine the total content of bioaerosol in atmosphere, the analysis can be performed by molecular biology techniques or through the use of biomarkers [3, 18–22]. In the last case, after the extraction and analysis of suitable chemical markers from samples of particulate matter, their concentrations can be converted via conversion factors to specific bioaerosol [23].
Concerning bacteria, muramic acid can be considered a specific index of their presence in atmosphere [24]. This compound is a component of the peptidoglycan, the only cell wall polymer common to both Gram-negative and Gram-positive bacteria, with the Gram negatives containing relatively little peptidoglycan (less than 10% of the total cell wall), and Gram positives mainly composed of peptidoglycan (usually 30–70% of the total cell wall) [25]. Muramic acid is also a biologically active substance, influencing cellular immune response [25].
Dipicolinic acid (pyridine 2,6-dicarboxylic acid), identified as being extremely important in spore resistance and stability, is synthetized during bacteria sporulation and can be considered an index of bacterial spore presence in atmosphere [26, 27].
As dipicolinic and muramic acids are not found elsewhere in nature, their detection in environmental samples can therefore be used for an indirect estimate of both bacterial spores and bacteria, using conversion factors. Obviously, it is difficult to assign proper factors and an incorrect assignment can lead to significant errors.
Mielniczuk found an average percentage of muramic acid from airborne bacteria between 0.14 and 0.95% [28]. Black found a percentage in the range 0.06–1.04% of muramic acid in a number of bacteria, among which is Bacillus subtilis, with 0.806% of muramic acid. The quantitative analyses of various Gram-positive and Gram-negative bacteria resulted in average values of 0.486 and 0.289% muramic acid on a dry weight basis, respectively [29].
Godoacre reported that spores from bacteria of Bacillus species accumulate a substantial deposit (5–14% of dry weight) of dipicolinic acid [30]. Warth in 1971 showed a dipicolinic acid content in B. subtilis spores equal to 12% of the dry weight, whereas Hindle reported a percentage of dipicolinic in spores of B. subtilis equal to 9.8% [31, 32]. Paidhungat found that spores of Bacillus and Clostridium species normally contained ≥10% of their dry weight as dipicolinic acid [33].
Finally, as early as 1996, Popham reported that peptidoglycan is also present in bacterial spores, even if muramic acid is generally converted to muramic lactam [34].
The aim of this work was to develop an analytical method for the determination of airborne total bacteria (culturable and nonculturable) and spores. The optimization included the simultaneous extraction of the bacterial biomarkers, dipicolinic and muramic acids, from airborne particulate matter, and their analysis by high performance liquid chromatography–tandem mass spectrometry (HPLC–MS/MS). A proper methodology to extract, purify, and recover the acids was performed, utilizing cultivated species of Gram-positive B. subtilis bacteria and spores.
In order to convert marker values into biomass, we compared the load of biomarkers in cultivated species of B. subtilis found in the present work with literature data, being Bacillus the most common genus in the ambient air, and assessed the conversion factors that were found to be a good approximation.
Finally, we applied the optimized method for the biomarker determination to urban atmospheric particulate matter of different sizes, and through the conversion factors, we estimated the mass contribution of bacterial components and gave a quantitative interpretation of the data. Discussion of the method and a summary of the results are reported in this paper.
Experimental
Chemicals, reagents, and materials
All the chemical and chromatographic reagents were HPLC or analytical grade.
Ultra gradient methanol (MeOH), acetonitrile (CH3CN), and water (H2O) were obtained by ROMIL LDT (Cambridge, UK). Glacial acetic acid (CH3COOH), 30% ammonia (NH4OH) solution, and triethylamine (TEA) were purchased from Carlo Erba Reagents S.p.A. (Arese, Milan, Italy). A total of 30% suprapur hydrochloric acid (HCl) was obtained by Merck S.p.a. (Vimodrone, Milan, Italy), muramic and dipicolinic acids were supplied by Sigma-Aldrich S.r.l. (Milan, Italy), and LB Broth by Sigma-Aldrich (Milan, Italy).
Stock standard solutions of the two analytes were prepared by dissolving each compound in H2O/MeOH 75:25 (1 mg mL−1) and stored at −20 °C in amber vials. Working solutions were prepared by successive dilution of the stock standard solutions.
Pepsin, pancreatin, and lysozyme enzymes were purchased from Sigma-Aldrich S.r.l. (Milano, Italy). B. subtilis strain PB1831 and relative spores were grown as described by Schmalisch et al. [35] with a few modifications. Strata Phenyl (500 mg/ml), Strata SI-1 Silica (1000 mg/6 ml), Strata C18 cartridges (200 mg/3 mL) were obtained from Phenomenex (Castel Maggiore, BO, Italy). Discovery® C18 HPLC Column 5 μm particle size, L × I.D. 3 cm × 2.1 mm was purchased from Sigma Aldrich (Milano, Italy).
Polytetrafluoroethylene (PTFE) Membrane Disc Filters (diameter: 47 mm, porosity: 2.0 μm) Pall Corporation were from VWR International s.r.l., Milan, Italy, and PTFE filters (diameter: 25 mm, porosity: 0.45 μm, ALBET) were purchased from Exacta + Optech Labcenter SpA (San Prospero, MO, Italy). SRM1649a urban dust was supplied by NIST (Gaithersburg, MD, USA).
Instrumentation
Filters were weighed before and after the sampling using a microbalance (Sartorius ME5; weight uncertainty ±2 μg) after conditioning in climatic cabinet (Activa Climatic Cabinet, Aquaria Milan, Italy) for 24 h, at 20 °C, and 50% relative humidity. Extractions were carried out in ultrasonic bath (Sonica Ultrasonic Cleaner 3200 Serie ETH S3, Soltec, Milan, Italy). The extracts were evaporated by a Glas-Col SE 500 evaporator (Bioanalitica Strumenti srl, Italy) under a nitrogen stream. A vacuum manifold 12-Port model SPE (Alltech, Casalecchio di Reno, Bologna, Italy) was used to quickly elute the analytes from the solid-phase cartridges at a constant flow.
Sampling was realized by a HYDRA Dual Sampler (FAI Instruments, Fontenuova, Rome, Italy) and by a low-pressure impactor (DLPI, Dekati Italia, Albiate MB, Italy).
Dipicolinic and muramic acids were analyzed by liquid chromatography/negative ion electrospray ionization-tandem mass spectrometry (HPLC/ESI-MS/MS), with an Agilent 1290 series binary HPLC pump system, at a flow rate of 200 μL min−1, fitted with an autosampler (20 μL) Agilent G4226A, coupled to an Agilent Jet Stream 6460 triple quadrupole mass spectrometer with an IonSpray source (Agilent Technologies Italia S.p.A., Cernusco sul Naviglio MI, Italy).
Sampling
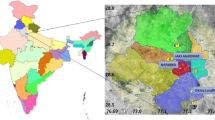
Sampling of atmospheric particulate matter was performed by a low-pressure impactor (DLPI), in the Botanic Garden of the University of Rome “Sapienza,” sited in a busy area of the city of Rome, Italy. The DLPI classifies airborne particles with aerodynamic diameter (Da) smaller than 10 μm (PM10) into 13 size fractions whose 50% cutoff diameters are 0.03, 0.06, 0.10, 0.17, 0.26, 0.40, 0.65, 1.0, 1.6, 2.5, 4.4, 6.8, and 10 μm. Therefore, the fraction of particles collected on impactor stages 1–3 (Dp < 0.1 μm) is referred to as the ultrafine fraction (UF), while the portions on impactor stages 4–8 (0.17 μm < Dp < 1.0 μm) and on impactor stages 9–13 (Dp > 1.0 μm) are referred to as the fine (F) and the coarse (C) fractions, respectively. Each fraction was collected on PTFE filters and weighed before and after the collection to obtain gravimetric size distribution of the particles. The operating flow rate was 10 L min−1, and the sampling duration was 13 consecutive days, with a final volume of sampled air equal to 188 m3. The amount of particulate matter on each filter was in the range 38–692 μg. The atmospheric concentrations of ultrafine, fine, and coarse PM were 1.0, 5.9, and 14.6 μg m−3. The sampling campaign was carried out in the spring of 2015 (March 26–April 8) at an average temperature of 17 °C.
In addition, a HYDRA Sampler, an automatic and sequential sampling system, equipped with 47 mm diameter PTFE membranes, was placed close to the first sampling device. Daily PM10 was collected on two different filters during two subsequent days within the sampling period of the DLPI (6 and 7 April). Each sampling duration was 24 h, and the operating flow rate was 2.3 m3 h−1, with a final volume of sampled air equal to 53.8 e 52.5 m3. The amount of particulate matter on the two filters was 631.7 and 914.7 μg, respectively.
Strain growth conditions
B. subtilis strain PB1831 was grown in LB broth at 37 °C for 16 h.
Spores were prepared as described by Schmalisch et al. [35]. B. subtilis was inoculated in 35 mL of Difco Sporulation Medium (DSM) and incubated at 37 °C on a rotary shaker (150 rpm) for 72 h. Spores were collected by centrifugation and washed twice with ice-cold distilled water. The obtained pellet was dried in an oven at 65 °C until constant weight.
Analytical procedure protocol
Details on method operation are provided in this paragraph. Blank filters, samples of B. subtilis and relative spores, blank filters spiked by standard solutions at different concentration levels, and sampled filters were processed as follows.
After the addition of 200 μL of water, the samples underwent a freezing (−20 °C) and thawing cycle for 7 times. Then 3 mL of 6 N HCl was added to the samples, successively placed in an ultrasonic bath at ambient temperature for half an hour, and hydrolyzed at 105 °C (2 h) under a gentle nitrogen gas flow.
The hydrolyzed solution (about 3 mL), neutralized by NH3, was evaporated to dryness, and dissolved in 100 μL of water/MeOH = 50:50, before loading onto a SPE Strata C18 cartridge. The cartridge was previously conditioned with 4 mL of MeOH and washed with 4 mL of H2O. Compounds loaded on the cartridge were eluted with 8 mL of MeOH/H2O 5:95 V/V. After evaporation to dryness, the extracted solution was redissolved in 500 μL of mobile phase and analyzed in HPLC/MS-MS in multiple reaction monitoring mode (MRM).
Standard calibration curves and matrix-matched calibration curves
Standard calibration curves were built by triplicate injections of 5 μL of multistandard solutions over a working range of 10–500 ng mL−1. These curves were used for the quantification of the analytes in bacteria and spores grown, as in “Strain growth conditions.”
Since matrix components in urban airborne particulate matter may result in response enhancement or depression, the most effective approach is the use of the standard addition method or matrix-matched calibration curves in order to quantify the environmental samples.
In the present work, the standard addition method was applied to PM10 collected by the HYDRA. The two sampled filters were submitted to the analytical procedure of “Analytical procedure protocol.” After the SPE cleanup, the solutions were divided in six aliquots and five of them were added with standard solution of dipicolinic and muramic acids at growing concentrations in the range 10–400 ng mL−1. Curves were built, reporting a linear plot of the analyte peak area (ordinate) against standard concentrations (abscissa), and the endogenous acids were determined by the x-intercept (on the negative x-axis). Regarding size segregated particles collected on 13 different filters with DLPI, in low amount mostly for the smaller sizes, we used the matrix-matched calibration curves built with the standard addition method on environmental samples collected by the HYDRA and shifted to the axis origin for subtracting the endogenous acid concentrations in PM10 [36].
Results and discussion
LC/MS/MS analysis
The choice of appropriate chromatographic conditions was particularly challenging because of the nature of the two molecules. Reversed phase columns with increasing polarity, hydrophilic reversed phase columns, and an anion exchange column were tested with different mobile phases.
Because of the wide pKa range of the two acids and pH ranges ensuring the stability of the columns, none of the columns tested proved completely suitable for the two compounds. A good compromise was achieved with the C18 column, working in ion pair chromatography (IPC).
Thus, reversed-phase HPLC separation of muramic and dipicolinic acids was performed on a C18 Discovery column in isocratic mode. One microliter was injected, the flow rate was 200 μL min−1, and the column oven was held at 40 °C. Mobile phase was 10% of MeOH/90% H2O containing triethylamine 5 mM, as ion pair agent, and glacial acetic acid 5 mM adjusted to pH 7.0.
The compounds co-eluted with a retention time (1.4 min) of about twice K’ value; the total run time was 3 min. The LC/MS/MS in MRM mode allowed us to obtain high specificity and sensitivity, even if the two compounds did not chromatographically separate.
To achieve the highest sensitivity and find the specific MS–MS transitions, the instrument parameters were previously optimized by continuous pump infusion of standard solutions of the analytes, at 10 ng μL−1, in MeOH/H2O 10:90, at a flow rate of 10 μL min−1, directly into the mass spectrometer, both in full and product ion scan. Experiments were carried out under positive and negative polarity with different additives to improve ionization efficiency. The best results were obtained, operating in negative ion mode. The compounds showed an abundant [M–H]− ion on Q1, which was chosen as precursor ion for MS–MS analysis. The MS spectra, the MS/MS fragmentations, and the extracted ion chromatograms (XIC) in MRM mode for both the compounds are provided in Figs. 1a–c and 2a–c.
(a) Negative ESI-Q1 and (b) MS-MS spectra of muramic acid; infusion 10 μL min−1 at concentration 10 ng μL−1; precursor ion m/z 250, parameters as in Tables 1 and 2. (c) Extracted ion currents (XICs) HPLC-MS/MS in MRM mode of a solution of muramic acid at concentration of 500 ng mL−1. The XIC refers to the most intense chosen transition (see Table 2)
(a) Negative ESI-Q1 and (b) MS-MS spectra of dipicolinic acid; infusion 10 μL min−1 at concentration 10 ng μL−1; precursor ion m/z 166, parameters as in Tables 1 and 2. (c) Extracted ion currents (XICs) HPLC-MS/MS in MRM mode of a solution of dipicolinic acid at concentration of 500 ng mL−1. The XICs refer to the most intense chosen transitions (see Table 2)
Table 1 shows the electrical parameters and Table 2 reports precursor ions (Q1) and MS–MS fragments for each compound with the transitions chosen for the quantification. The mass spectrum of muramic acid (Fig. 1a) shows the expected pseudomolecular ion at m/z 250. The side chain of lactic acid (lactate: C3O3H5 −) gives rise to the major fragment at m/z 89 [29]. As for dipicolinic acid (Fig. 1b), the m/z 122 and the m/z 78 fragments are due to the loss of one or two formic acid molecules from the pseudomolecular ion (m/z 166).
Extraction, cleanup, and recovery
Different attempts and preliminary experiments were carried out to extract and purify muramic acid from B. subtilis bacteria and dipicolinic acid from their spores, before extending the analytical procedure to PM environmental samples. The optimized method requires that the samples are added with 200 μL of water and subjected to a freezing (−20 °C) and thawing cycle to break the cellular membrane. Acidic hydrolysis is necessary to obtain free muramic acid from the peptidolglycan polymer. Enzymatic hydrolysis with pepsin, and in alternative with lysozyme, followed by pancreatin, was tried, but was unsuccessful. Acidic hydrolysis, previously tested directly on dipicolinic and muramic acid standard, proved to have no influence on the analyte stability.
After acidic hydrolysis and neutralization, a cleanup step is fundamental to reduce ion suppression and eliminate interferences before HPLC-MS-MS analysis. Three different SPE cartridges (Strata Si, Strata C18, Strata Phenyl), loaded with standard solutions, were compared to choose the most efficient one. After conditioning with 4 mL of MeOH and washing with 4 mL of H2O, the analytes were loaded on the cartridge and eluted with different mixtures of MeOH and/or CH3CN–H2O. Strata Si cartridge showed recoveries equal to 50 and 33% for dipicolinic and muramic acids, respectively. Strata Phenyl cartridge showed a recovery of 92% of muramic acid but of 45% of dipicolinic acid if eluted with 75% of MeOH in H2O. By eluting with 25% of MeOH in H2O, the percentages were reversed. A Strata C18 provided the best recoveries (95.4% for dipicolinic and 94.6 for muramic acids) with H2O/MeOH 95:5 as eluting solvent. The loss due to the evaporation step ranged between 1 and 5%.
Total recovery was calculated on blank filters added with equal amount of SRM 1649a and spiked with different amount of acids (0.05, 0.10, 0.25, and 0.50 μg ml−1) before the whole procedure, comparing the results with blank filters added with equal amount of SRM 1649a and spiked with the same amount of acids at the end of the analytical procedure. The recovery was estimated in the range 67.5–80.5% for muramic acid and 66.5–83.7% for dipicolinic acid.
Quantitative analysis, linearity, limits of detection, and reproducibility
Standard calibration curves (equation: y = 14.40x + 6.71 and R 2 = 0.997 for dipicolinic acid; equation: y = 1.45x + 26.79 and R 2 = 0.998 for muramic acid), used for the quantification of muramic and dipicolinic acids on spiked filters and in B. subtilis bacteria and spores, were compared with the results of the matrix-matched calibration curve (equation: y = 2.28x and R 2 = 0.996 for dipicolinic acid; equation: y = 0.05x and R 2 = 0.995 for muramic acid), used for PM analysis.
A good linearity was obtained for both the acids over a range of 10–500 and 10–400 ng mL−1 for the standard and the matrix-matched calibration curves, respectively. The latter exhibits much lower slopes, that is, a lower sensitivity, and indicates a strong matrix effect. Thus, the use of the matrix-matched calibration curves, obtained from fortified samples submitted to the full procedure, showed to be necessary and ensured the reliability of the method.
Instrumental detection limit (LOD), defined as the smallest concentration of analyte that has a signal significantly larger than the signal from a suitable blank [30], was calculated based on the relationship Cx(LOD) ≥ (μb + 3σ b)/a, where μb is the average of the blank responses, σ b is the standard deviation of responses of blank, and a is the slope of the calibration curve [37]. LOD values were 0.04 ng mL−1 for muramic and. 0.2 ng mL−1 for dipicolinic acids.
Intra-day instrumental reproducibility values were obtained by comparing 10 replicate injections of standard solutions at three different concentrations. Inter-day values were obtained by the average of 10 replicate injections over three consecutive days. The results were expressed as relative standard deviation (RSD). The intra-day precision ranged from 0.3 to 7%. The inter-day precision varied from 2 to 11%.
Amount of muramic and dipicolinic acids in B. subtilis bacteria and spores
Three aliquots of B. subtilis bacteria, about 3 mg each, were subjected to the extraction and cleanup procedure as in “Analytical procedure protocol” and analyzed in HPLC/MS-MS. Each solution, suitably diluted, was injected three times, and the quantitative results were obtained from the standard calibration curve.
The results showed a weight to weight percentage (w/w%) of muramic acid in B. subtilis bacteria samples equal to 0.4 ± 0.05%.
Because few data are available on genera of bacteria mainly found in air samples and on muramic acid content, we started from the earlier mentioned assumptions that Bacillus generally was found to be the most abundant single genus represented at urban locations, where generally Gram-positive bacteria occur in larger amounts, accounting for about 80∼86% of the total, and that muramic acid in a number of Gram-positive bacteria was on average 0.486% on a dry weight basis. Hence, we came to the conclusion that our coefficient (muramic acid in B. subtilis bacteria equal to 0.4%) was a good approximation to convert muramic acid concentration found in environmental samples into concentrations of total airborne bacteria, although the use of this approximation is a weakness of this study.
Three samples of about 3 mg of B. subtilis spores were also treated and analyzed after suitable dilution so as to fall within the standard calibration curve range. The w/w% of dipicolinic acid in bacterial spore samples was 9.6 ± 1.0% in agreement with literature data [31, 32].
Similarly to bacteria, also spores vary considerably in their content of dipicolinic acid, depending on the species [31], nevertheless, our result falls within the range 5–15%, as reported by Hindle [32], to convert the amount of acid found in environmental samples into airborne spores. Therefore, we assessed that the conversion factor 9.6 was a good approximation to measure bacterial spores in atmosphere from dipicolinic acid.
As expected, B. subtilis spores showed the presence of a very small amount of muramic acid (about 0.15% of the weight of the spores), as to be neglected.
Bacteria and bacterial spores in atmospheric particulate matter
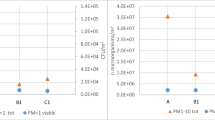
Starting from the extrapolation on x-axis of the curves built with the standard addition method (see “Standard calibration curves and matrix-matched calibration curves”), muramic and dipicolinic acid concentrations (6 and 7 April) were calculated in PM10. Dipicolinic acid found on the filters was 74.4 and 49.8 ng and muramic acid was 58.5 and 52.8 ng (Table 3).
Filters from the DLPI were first assembled into three fractions: UF, F, and C (see “Sampling”), then subjected to the procedure of “Analytical procedure protocol.” Quantitative analysis was performed by using the matrix matched calibration curves (“Standard calibration curves and matrix-matched calibration curves”), and the amounts of acids found on each set of filters were 48.1, 64.1, and 208.3 ng for dipicolinic acid and 22.1, 42.5, and 108.7 for muramic acid in ultrafine, fine, and coarse fractions of PM (Table 3).
Taking into account that dipicolinic acid is 9.6% of spores and muramic acid is 0.4% of bacteria, Table 4 reports spore and bacteria atmospheric concentrations.
The results from the two different samplers agree reasonably despite referring to different time periods, and one sampler collects PM for 13 consecutive days, suggesting no chemical degradation or artifacts occurred due to the protracted sampling time. In contrast to the fungi and fungal spore results [3], bacteria are more abundant than bacterial spores in the urban atmosphere.
Relative to the size distribution, as expected [15, 38], bacteria concentrations found in the coarse fraction of particulate matter are 2.5 and 5 times higher than concentrations in fine and ultrafine particles, respectively. Similarly for bacterial spores, their concentration found in coarse particles is 3 and 4 times higher than spores in fine and ultrafine particles (Table 3).
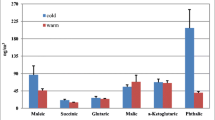
However, since coarse particles are 6 and 15 times higher than fine and ultrafine particulate matter, bacteria and bacterial spore amounts per unit mass of PM (mg g−1) in the three size fractions (UF, F, and C) show that contribution of both bacteria and spores to the ultrafine particles was higher than to the fine and coarse fractions (see Fig. 3). Fine and coarse fractions showed a similar percentage contribution. This leads to the hypothesis that the bacteria are an important and hitherto underestimated fraction of atmospheric aerosols of nanometric dimensions.
Ultrafine particles are known to contain a higher content of organic carbon (OC) than the other fractions [20, 39], and the present results demonstrate that ultrafine OC is partially due to bacteria. In addition, inhaled ultrafine particles can penetrate deeply into the cells and blood vessels of the lung. Therefore, although the percentage of ultrafine particles was only 5%, ultrafine particles demonstrate to have a significant influence on human infections.
Assuming an average weight of 1 × 10−12 g for each bacterial cell [40], the number of bacteria found in atmosphere for cubic meter is also shown in Table 3. In the urban PM10, at spring, at an average temperature of 17 °C, we found about 250,000 cells m−3. Although very few data are present in literature reporting the number of bacterial cells found in atmosphere, as almost all the scientific community measures the CFU, we could compare our results with few other sampling sites. Bauer in Austrian Alps found 1.2 × 104 bacteria m−3 [15]. Harryson found an average concentration of 17,504 cells m−3 in the urban atmosphere of Birmingham (GB) [5], while Tong found 2000 to ten million cells per cubic meter [41] in the mid-Willamette River Valley (Oregon, USA). It demonstrates a wide range of bacteria variability due to the sampling site.
Conclusion
Since the total bacterial concentration is responsible of allergic disease and the only culturable determination might be insufficient for a right investigation, we determined the total bacterial component of bioaerosol in outdoor environment through muramic and dipicolinic acids as biomarkers of bacteria and relative spores, respectively.
A reliable method for their determination was optimized on bacteria and spores of B. subtilis and then applied to environmental samples. Such study also allowed us to determine the w/w percentages of the two acids in bacteria and bacterial spores of B. subtilis, respectively; those percentages were then used as conversion factors, although such an approximation is a weakness of the present approach.
Bacteria and bacterial spores were studied in PM10 sampled in two spring days in an urban atmosphere. These results were also compared with those obtained from a DLPI sampling in a parallel period, showing a good agreement.
DLPI sampling also allowed to study the size distribution of bacteria and spores in UF, F, and C fractions of PM10. Although the higher concentration of bacteria is in the coarse particulate fraction, ultrafine particles have the highest bacteria percentage contribution, equal to 3%.
Future works in this area should include an accurate study on factors to convert muramic and dipicolinic acids into bacteria and spores. The percentages should be calculated for each bacterium genera, and on a case-to-case basis, any available database on the main bacteria content in atmosphere should be considered.
References
Matthias-Maser S, Brinkmann J, Schneider W. The size distribution of marine atmospheric aerosol with regard to primary biological aerosol particles over the South Atlantic Ocean. Atmos Environ. 1999;33:3569–75.
Matthias-Maser S, Jaenicke R. The size distribution of primary biological aerosol particles in the multiphase atmosphere. Aerobiologia. 2000;16:207–10.
Di Filippo P, Pomata D, Riccardi C, Buiarelli F, Perrino C. Fungal contribution to size-segregated aerosol measured through biomarkers. Atmos Environ. 2013;64:132–40.
Shaffer BT, Lighthart B. Survey of culturable airborne bacteria at four diverse locations in Oregon: urban, rural, forest, and coastal. Microbial Ecol. 1997;34:167–77.
Harrison RM, Jones AM, Biggins PDE, Pomeroy N, Cox CS, Kidd SP, et al. Climate factors influencing bacterial count in background air samples. Int J Biometeorol. 2005;49:167–78.
Fang Z, Ouyang Z, Zheng H, Wang X, Hu L. Culturable airborne bacteria in outdoor environments in Beijing, China. Microbial Ecol. 2007;54(3):487–96.
Lindemann J, Constantinidou HA, Barchet WR, Upper CD. Plants as sources of airborne bacteria, including ice nucleation-active bacteria. Appl Environ Microbiol. 1982;44(5):1059–63.
Bauer H, Giebl H, Hitzenberger R, Kasper-Giebl A, Reischl G, Zibuschka F, et al. Airborne bacteria as cloud condensation nuclei. J Geophys Res-Atmos. 2003;108(D21):AAC2.1–5.
Georgakopoulos DG, Després V, Fröhlich-Nowoisky J, Psenner R, Ariya PA, Pósfai M, Ahern HE, Moffett BF, Hill TCJ. Microbiology and atmospheric processes: biological, physical and chemical characterization of aerosol particles. Biogeosciences Discussions, European Geosciences Union. 2008;5,2;1469–1510. <hal-00297993 > .
Hejkal TW, Larock PA, Winchester JW. Water-to-air fractionation of bacteria. Appl Environ Microbiol. 1980;39(2):335–8.
Lindemann J, Upper CD. Aerial dispersal of epiphytic bacteria over bean plants. Appl Environ Microbiol. 1985;50(5):1229–32.
Sawyer B, Rao KC, O’Brien P, Elenbogen G, Zenz DR, Lue-Hing C. Changes in bacterial aerosols with height above aeration tanks. J Environ Eng. 1996;122:368–73.
Jaenicke R. Abundance of cellular material and proteins in the atmosphere. Science. 2005;308(5718):73.
Brodie EL, De Santis TZ, Moberg Parker JP, Zubietta IX, Piceno YM, Andersen GL. Urban aerosols harbor diverse and dynamic bacterial populations. Proc Natl Acad Sci. 2007;104(1):299–304.
Bauer H, Kasper-Giebl A, Löflund M, Giebl H, Hitzenberger R, Zibuschka F, et al. The contribution of bacteria and fungal spores to the organic carbon content of cloud water, precipitation and aerosols. Atmos Res. 2002;64:109–19.
Després VR, Huffman JA, Burrows SM, Hoose C, Safatov AS, Buryak G, et al. Primary biological aerosol particles in the atmosphere: a review. Tellus B. 2012;64:15598.
Wady L, Shehabi A, Szponar B, Pehrson C, Sheng Y, Larsson L. Heterogeneity in microbial exposure in schools in Sweden, Poland and Jordan revealed by analysis of chemical markers. J Expo Anal Environ Epidemiol. 2004;14:293–9.
Buiarelli F, Canepari S, Di Filippo P, Perrino C, Pomata D, Riccardi C, et al. Extraction and analysis of fungal spore biomarkers in atmospheric bioaerosol by HPLC-MS-MS and GC-MS. Talanta. 2013;105:142–51.
Buiarelli F, Gallo V, Di Filippo P, Pomata D, Riccardi C. Development of a method for the analysis of underivatized amino acids by liquid chromatography/tandem mass spectrometry: application on Standard Reference Material 1649a (urban dust). Talanta. 2013;115:966–72.
Di Filippo P, Pomata D, Riccardi C, Buiarelli F, Gallo V, Quaranta A. Free and combined amino acids in size-segregated atmospheric aerosol samples. Atmos Environ. 2014;98:178–89.
Blais-Lecours P, Perrott P, Duchaine C. Non-culturable bioaerosols in indoor settings: impact on health and molecular approaches for detection. Atmos Environ. 2015;110:45–53.
Perrino C, Marcovecchio F. A new method for assessing the contribution of primary biological atmospheric particles to the mass concentration of the atmospheric aerosol. Environ Int. 2016;87:108–15.
Pietrogrande MC, Bacco D, Chiereghin S. GC/MS analysis of water-soluble organics in atmospheric aerosol: optimization of a solvent extraction procedure for simultaneous analysis of carboxylic acids and sugars. Anal Bioanal Chem. 2013;405:1095–104.
Fox A, Wright L, Fox K. Gas chromatography-tandem mass spectrometry for trace detection of muramic acid, a peptidoglycan chemical marker, in organic dust. J Microbiol Methods. 1995;22:11–26.
Schleifer KH, Kandler O. Peptidoglycan. Types of bacterial cell walls and their taxonomic implications. Bacteriol Rev. 1972;36(4):407–77.
Janssen FW, Lund AJ, Anderson LE. Colorimetric assay for dipicolinic acid in bacterial spores. Science. 1958;127(3288):26–7.
Taylor KML, Lin W. Hybrid silica nanoparticles for luminescent spore detection. J Mater Chem. 2009;19:6418–22.
Mielniczuk Z, Mielniczuk E, Larsson L. Determination of muramic acid in organic dust by gaschromatography-mass spectrometry. J Chromatogr B. 1995;670:167–72.
Black GE, Fox A, Fox K, Snyder AP, Smith PBW. Electrospray tandem mass spectrometry for analysis of native muramic acid in whole bacterial cell hydrolysates. Anal Chem. 1994;66(23):4171–6.
Goodacre R, Shann B, Gilbert RJ, Timmins ÉM, McGovern AC, Alsberg BK, et al. Detection of the dipicolinic acid biomarker in Bacillus spores using Curie-point pyrolysis mass spectrometry and Fourier transform infrared spectroscopy. Anal Chem. 2000;72:119–27.
Warth AD, Strominger JL. Structure of the peptidoglycan from vegetative cell walls of Bacillus subtilis. Biochemistry. 1971;10(24):4349–58.
Hindle AA, Hall EAH. Dipicolinic acid (DPA) assay revisited and appraised for spore detection. Analyst. 1999;124:1599–604.
Paidhungat M, Setlow B, Setlow ADP. Characterization of spores of Bacillus subtilis which lack dipicolinic acid. J Bacteriol. 2000;182(19):5505–12.
Popham DL, Helin J, Costello CE, Setlow P. Muramic lactam in peptidoglycan of Bacillus subtilis spores is required for spore outgrowth but not for spore dehydration or heat resistance. PNAS - Proc Natl Acad Sci – Microbiol – USA. 1996;93:15405–10.
Schmalisch M, Maiques E, Nikolov L, Camp AH, Chevreux B, Muffler A, et al. Small genes under sporulation control in the Bacillus subtilis genome. J Bacteriol. 2010;192(20):5402–12.
Pomata D, Di Filippo P, Riccardi C, Buiarelli F, Gallo V. Determination of non-certified levoglucosan, sugar polyols and ergosterol in NIST Standard Reference Material 1649a. Atmos Environ. 2014;84:332–8.
MacDougall D, Crummett WB. Guidelines for data acquisition and data quality evaluation in environmental chemistry. Anal Chem. 1980;52(14):2242–9.
Alexis NE, Lay JC, Zeman K, Bennett WE, Peden DB, Soukup JM, et al. Biological material on inhaled coarse fraction particulate matter activates airway phagocytes in vivo in healthy volunteers. J Allergy Clin Immunol. 2006;117:6.
Ramgolam K, Favez O, Cachier H, Gaudichet A, Marano F, Martinon L, et al. Size-partitioning of an urban aerosol to identify particle determinants involved in the proinflammatory response induced in airway epithelial cells. Part Fibre Toxicol. 2009;6(10):1–12.
Davis B, Dulbecco R, Eisen H, Ginsberg H. Bacterial physiology: microbiology, 2nd ed., Maryland: Harper and Row. 1973;96–97.
Tong Y, Lighthart B. Diurnal distribution of total and culturable atmospheric bacteria at a rural site. Aerosol Sci Technol. 1999;30:246–54.
Acknowledgments
This work was supported by MIUR 2015 – Project C26A15R5ZS and by INAIL/Ricerca Discrezionale/2016-2018/P3O3/UO2
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Di Filippo, P., Pomata, D., Riccardi, C. et al. Muramic and dipicolinic acids in atmospheric particulate matter as biomarkers of bacteria and bacterial spores. Anal Bioanal Chem 409, 1657–1666 (2017). https://doi.org/10.1007/s00216-016-0111-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-0111-y