Abstract
Exosomes are stable nanovesicles secreted by cells into the circulation. Their reported sizes differ substantially, which likely reflects the difference in the isolation techniques used, the cells that secreted them, and the methods used in their characterization. We analyzed the influence of the last factor on the measured sizes and shapes of hydrated and desiccated exosomes isolated from the serum of a pancreatic cancer patient and a healthy control. We found that hydrated exosomes are close-to-spherical nanoparticles with a hydrodynamic radius that is substantially larger than the geometric size. For desiccated exosomes, we found that the desiccated shape and sizing are influenced by the manner in which drying occurred. Isotropic desiccation in aerosol preserves the near-spherical shape of the exosomes, whereas drying on a surface likely distorts their shapes and influences the sizing results obtained by techniques that require surface fixation prior to analysis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cells actively secrete exosomes via an endocytic pathway [1–3]. Exosomes are found in the extracellular space and all body fluids, including blood, urine, and saliva. The molecular content of exosomes is derived from the cells that release them. The frequency of release depends on the cell environment, such as the level of dissolved oxygen and the pH [4, 5], and the cell type, with cancer cells known to release a larger number of exosomes. They carry small noncoding RNAs, although the number of microRNA (miRNA) molecules enveloped inside each exosome may not be high [6]. Nevertheless, some reports suggest that most miRNAs in the blood are contained within exosomes [7]. It is thought that by their fusing with recipient cells and releasing their RNA and other cargo [8–11], exosomes play a role in short-range and long-range intercellular signaling.
Exosomes can be differentiated from other circulating vesicles by the markers of the endosomal pathway and their small size, which distinguishes them from other extracellular vesicles. Although an important differentiator, the size of exosomes is often reported with substantial variability, as illustrated by a sample of published results summarized in Table 1.
Several factors likely contribute to the large range of reported sizes, including the source of exosomes—that is, the type of body fluid or cell line from which they were isolated. The exosome isolation method, which may include ultracentrifugation [12], solvent precipitation [13], size-exclusion chromatography [14], immunoaffinity isolation [15], microfluidic techniques [16], and ultrafiltration [17], is another factor in size variability [15, 18, 19].
The choice of characterization technique influences the measured size of exosomes as well. Although it is recognized that the precise determination of the size distribution of exosomes with any given technique is difficult [20–22], it is often underappreciated that different analytical methods estimate sizes on the basis of dissimilar physical principles, which produce identical results in only limited cases. The sample preparation steps used with different methods, such as sample verification, desiccation, or surface fixation, also affect sizing results. The geometric shape of exosomes influences the size measurements obtained with techniques that infer particle sizes from their mobility, such as nanoparticle tracking analysis (NTA), dynamic light scattering (DLS), and differential mobility analysis (DMA). The standard practice in NTA and DLS is to assume that the exosomes are spherical, contrary to numerous reports and reviews [23] suggesting that exosomes have a cup-shaped geometry. Therefore, to provide sizing consistency and accuracy by methods that rely on particle mobility, the shape of hydrated and desiccated exosomes becomes an important factor that must be examined.
In this article, we quantify the influence of different analytical techniques on the measurements of exosome sizes. By reanalyzing the same samples by different sizing techniques, we find significant variability in the results that is comparable with the variability seen in Table 1, where the difference between samples—by biofluids, cell lines, and the techniques used in their isolation—is also a contributing factor. We offer a model that explains this result, and suggest that the difference in exosome sizes measured by different techniques may itself be an important characteristic of a sample that depends on such biophysical properties of exosomes as their surface decoration, elasticity, and molecular content.
Materials and methods
Samples and exosome isolation
Serum samples from a 75-year-old female pancreatic cancer patient with high levels of the tumor marker carbohydrate antigen 19-9 (414 U/mL) and a healthy 42-year-old woman seen for routine cholesterol testing were used in this study. Both samples were obtained from ARUP Laboratories (Salt Lake City, UT, USA) and were de-identified according to the Institutional Review Board protocol. Exosomes were isolated from 1 mL serum using an ExoQuick kit (System Biosciences, Mountain View, CA, USA) following the manufacturer’s instructions. Briefly, serum was centrifuged at 3,000 g for 15 min to remove cells and cell debris. The supernatant was transferred to a sterile vessel, and 252 μL of ExoQuick solution was added. The mixture was refrigerated for 30 min and then centrifuged at 1,500 g for 30 min at room temperature. After centrifugation, the supernatant was discarded and the exosome pellet was saved. To spin down the residual ExoQuick solution, the pellet was centrifuged for another 5 min at 1,500 g, and the supernatant was removed without disturbing the pellet. The pellet was then resuspended in 200 μL of 2 mM ammonium acetate buffer. The buffer solution was prepared using ammonium acetate salt (Sigma-Aldrich, St Louis, MO, USA) and deionized water (Milli-Q filtration). The exosome size was then characterized by all the analytical techniques.
Analytical methods
Scanning electron microscopy
Prior to scanning electron microscopy (SEM) imaging, both samples were diluted 1:100 in deionized water. A glass slide was gently cleaned with nitrogen gas and placed on the specimen stage of the scanning electron microscope (FEI NanoNova 630 high-resolution scanning electron microscope). Five microliters of each sample was then placed on the glass slide and allowed to dry. The samples were imaged at 0.98 Torr using a low-vacuum secondary electron detector at magnifications in the × 35,000 to × 65,000 range. The acquired 1,024 pixel × 943 pixel images were analyzed using custom MATLAB software to determine the exosome size distribution. The grayscale SEM images were first converted into a binary form to define the boundary of each particle. Each exosome in each image was approximated by an ellipse that provided the best (in the least-squares sense) fit to its perimeter. The exosome diameter was calculated by geometrically averaging the lengths of the major axis and the minor axis (which are equal for spherical particles) of the fitted ellipse.
Electrospray DMA
DMA has been previously used to size biological particles, including cold viruses [24], lipoproteins [25], virus-like particles [26], and other macromolecular assemblies [27, 28]. This study is the first to use this technique to characterize exosome sizes. Patient and control samples were diluted 1:100 in ammonium acetate solution, and 0.5 mL of the preparation was used in the electrospray aerosol generator (model 3480; TSI, Shoreview, MN, USA) to confine individual exosomes inside charged droplets formed by atomizing the suspension in the Taylor cone formed at the end of 25 μm inner diameter capillary. Pressure-driven flow through the capillary was maintained by 26 kPa excess pressure across the capillary. A stream of gas consisting of air at 1.0 L/min and carbon dioxide at 0.2 L/min was used to entrain the charged droplets and carry them into a bipolar charge neutralizer, where they were bombarded by α-particles formed by radioactive decay of 210Po. The solvent and volatile ammonium acetate salt rapidly evaporated from the entrained droplets, whereas the bombardment by α-particles electrically neutralized most of the exosomes desiccated in the gas phase [29]. A fraction of the desiccated exosomes retained a single net positive or negative change (±1 electron charge), whereas particles with higher charges (±2, ±3, etc.) occur at an increasingly rare frequency [30]. For example, after the neutralization, 46.7 % of 74-nm particles carry an elementary charge of ±1 (26.2 % of particles will have an excess and 20.5 % will have a deficit of a single charge), 1.9 % and 3.2 % will have a charge of +2 and −2, respectively, and there will be a negligible number of higher-charged particles; the balance is formed by neutral particles [29].
After charge reduction in the neutralizer, the desiccated exosomes were carried by flowing gas into the differential mobility analyzer (TSI model 3080) operated with sheath flow of nitrogen at 10 L/min. Inside the differential mobility analyzer, a strong negative electrical potential deflects positively charged exosomes toward a collection slit (Fig. 5c). The exact deflection trajectory toward the collection slit depends on electrical and drag forces on the particles. The electrical force is constant for the vast majority of attracted exosomes because the neutralization process left them with the same charge of +1. Therefore, at a given potential only the drag force (which changes with the particle diameter) and the particle shape determine which particles pass through the collection slit and are counted one-by-one inside the condensation particle counter (TSI model 3025A). The result is expressed as the average number of particles analyzed per volume of inlet gas at a flow rate of 1.0 L/min over 20 s. By sweeping the bias potential from 1.7 to 2.5 kV and assuming that the desiccated exosomes are spherical particles, we characterized their size distribution in the range from 20 to 82 nm in 1-nm increments. Alternatively, by maintaining a constant potential, one can direct a narrow size fraction from the differential mobility analyzer into an electrostatic deposition chamber and they can be deposited on a substrate for subsequent analysis (Fig. 5c).
Cryo-transmission electron microscopy
The imaging procedure was described in detail previously [31]. Briefly, prior to cryo-transmission electron microscopy (TEM) imaging, the exosome samples were diluted 1:100 in deionized water. Approximately 3.5 μL of sample was placed on a holey carbon-coated copper grid. The unstained sample was vitrified with the aid of a robotic accessory (Vitrobot; FEI, Hillsboro, OR, USA) used to plunge-freeze the aqueous sample into liquid ethane maintained at the temperature of liquid nitrogen. Once vitrified, the samples were stored in liquid nitrogen. Prior to image acquisition, the stored samples were transferred to a cryoholder (model 626; Gatan, Pleasanton, CA, USA), which maintained their temperature at approximately − 180 °C during imaging. The 2,048 pixel × 2,048 pixel cryo-TEM images were obtained at 200 kV using a Tecnai F20 transmission electron microscope (FEI, Hillsboro, OR, USA) coupled to a FEI Eagle CCD camera and were sized using the same MATLAB image analysis code as used in SEM sizing.
Dynamic light scattering
The samples were diluted 1:100,000 in deionized water and were filtered through 0.2-μm syringe filters (Corning, Tewksbury, MA, USA). Prior to the measurements, 1 mL of the sample preparation was placed into a low-volume disposable sizing cuvette for analysis and given 5 min to reach 25 °C. The DLS measurements were performed with a Zetasizer Nano ZS (Malvern Instruments, Malvern, UK) at 173° which measures particles in the 0.3–10,000-nm size range. Water viscosity at 25 °C (0.8872 cP) and the refractive index of the solution of 1.33 were used to interpret the measurements. The refractive index for exosomes was set to 1.35. Samples were analyzed in three repeats, each consisting of 12 scattering measurements. The data obtained were processed using a general purpose model implemented in the Zetasizer software to obtain the size distribution, its mean, and the standard deviation.
Nanoparticle tracking analysis
NTA was performed using a NanoSight (Salisbury, UK) instrument (model LM10) by illuminating the sample with a 40-mW violet laser (405-nm wavelength), capturing the light scatted by exosomes with a high-sensitivity scientific CMOS camera (OrcaFlash2.8, Hamamatsu C11440), and analyzing the results using the software provided by the manufacture (NanoSight version 2.3). The minimal expected particle size, minimal track length, and blur size were set to Auto, gain was set to 1, brightness was set to 0, and the detection threshold set was to 10 Multi. The viscosity of deionized water depends on the temperature, and was adjusted automatically on the basis of the temperature measurements. The temperature of the cell was measured manually, and stayed at 20 °C with a maximum fluctuation of 0.1 °C throughout the nanoparticle tracking. The viscosity of water at these temperatures is nearly constant and equal to 1 cP. Prior to analysis, the exosome samples were diluted 1:1,000 in deionized water and allowed to equilibrate to room temperature (20 °C). Water used in the dilution was filtered using a Nanopure filtration system (Thermo Scientific, Waltham, MA, USA) and stored in a glass container prior to use. Samples were analyzed within 5 min of the initial dilution. By means of a 1-mL sterile syringe, each sample was injected into the test cell. Approximately 35 particles were observed in the field of view, and the typical concentration was approximately 5 × 108 particles per milliliter for each measurement. A 60-s video was recorded for each sample at 20 frames per second with 25.98-ms shutter speed and the camera gain set to 512 and was analyzed using the NTA software with the described settings. Each video consisted of more than 1,000 frames, and the total number of valid particle tracks for each 60-s measurement was approximately 1,200. The video data characterizing hydrodynamic mobility of particles in the field of view were analyzed with the NTA software, which reported the exosome size distribution, its mode, and the mean and the standard deviation.
Data analysis
Size-frequency measurements for a given sample obtained with different techniques were converted into the probability density functions (pdf) of particle sizes expressed as a histogram. The width of a data bin, h, in each histogram was calculated as \( h=3.5\sigma /{n}^{\frac{1}{3}} \), where n is the number of sized exosomes having diameters in the range characterized by the standard deviation σ. The hypothesis that the medians of two distributions are different was tested with 95 % confidence, assuming that the size measurements were normally distributed. The testing was found to be robust to the type of the assumed distribution. Specifically, the outcome of the test did not change when the assumption of normality was changed to the assumption that size measurements were log-normally distributed.
Results
Hydrodynamic sizing
We measured the size distribution of exosomes in solution by NTA and DLS. For spherical particles, both techniques estimate hydrodynamic diameters on the basis of the measurements of particle mobility in a liquid. The insert in Fig. 1b is a typical frame of an NTA video sequence of the particle motion captured by the CMOS sensor. The spots in the image are formed by scattered laser light collected by a × 20 optical objective. The temporal motion of each particle is used to estimate its diffusivity; the corresponding hydrodynamic size is found from the Stokes–Einstein equation. The results for individual particles are then summarized as the size distribution of the particle populations.
Figure 1a and b shows, respectively, the size distributions of exosomes (scaled to represent probability density functions) from a healthy woman control and a woman with pancreatic cancer measured by NTA. The mode for the control sample was 136 nm, whereas it was 121 nm for the sample from the pancreatic cancer patient, and the mean size (plus or minus the standard deviation) for the control sample was 182 ± 79 nm, and was 157 ± 72 nm for the pancreatic cancer sample. The exosome concentration in the control sample was 5.00 × 1011 particles per milliliter, but was higher (5.41 × 1011 particles per milliliter) in the patient sample.
The same samples analyzed by DLS gave similarly broad size distributions (Fig. 1c, d). The mode and the mean for the control sample were 91 nm and 119 ± 47 nm, whereas they were 92 nm and 130 ± 55 nm for the patient sample.
To rule out the possibility that osmolarity of the solution influences the results, two exosome samples (see the electronic supplementary material for details) were suspended in both deionized water and phosphate-buffered saline, and were sized by NTA. The measured size distributions (Fig. S1, Table S1) indicate that low osmolarity does not lead to substantial swelling of the exosomes.
Geometric sizing of hydrated exosomes by cryo-TEM
Figure 2a and b shows typical cryo-TEM images of exosomes in control and pancreatic cancer patient samples, respectively. The imaged particles are unstained, yet sufficient contrast was present to identify particle boundaries. The exosomes appear as close-to-circular projections, with mean eccentricities of 0.334 ± 0.084 and 0.290 ± 0.085 for control and patient exosomes, respectively. Computer analysis (illustrated in Fig. S2) shows that the geometric mean of the diameters of 106 exosomes derived from the control sample are in the range between 26 and 129 nm and those of 212 patient exosomes are in the 25-98 nm range. The size distributions for the two samples are shown in Fig. 2c. The average diameter plus or minus the standard deviation was found to be 71 ± 24 nm for the control sample and 55 ± 14 nm for the patient sample. Consistent with the NTA results, the concentration of patient exosomes was higher, and their tendency to cluster was notable.
Vitrified exosomes in cryo-transmission electron microscopy (TEM) images were isolated from the control (a) and pancreatic cancer patient (b) samples. The weblike features in a and b are the carbon-support film. Note that exosomes are found on the carbon support or between supports (inside the "holes"). Two clustered groups of exosomes can be seen in the lower-right quadrant in b. The exosome size distribution in c is based on computer analysis of 106 control and 212 patient exosomes. Exosomes at the highest resolution obtained are shown in d. Arrows shows faint, extra density surrounding some particles, which suggests the presence of macromolecules conjugated to their surface. pdf probability distribution function, Std standard deviation
Visual inspection indicated that algorithmic image analysis undercounted the number of exosomes present in cryo-TEM images. Manual sizing was therefore performed, the results of which are summarized and compared with those of algorithmic image analysis in Fig. S2. Although the results were generally consistent with computer sizing, several differences were revealed, as discussed in the electronic supplementary material. Briefly, a larger number of exosomes identified manually is related to the difficulty in algorithmically sizing clustered exosomes and exosomes that are either on or between carbon supports. Underfocusing in cryo-TEM images [32] leads to a coronal appearance of exosome membranes (for examples, see Fig. 2d). Individuals often ignore these coronas (Fig. S2b) and estimate sizes smaller than those identified by computer analysis (Fig. S2d). The computer analysis sizes are reported as a geometric mean of diameters along the major axis and the minor axis (Fig. S2e), but manual analysis was not robust in identifying the major axis and the minor axis, and they were reported for each particle as a single number.
Geometric sizing of desiccated exosomes
Scanning electron microscopy
A representative SEM image of desiccated exosomes is shown in Fig. 3a. In total, eight SEM images of the patient sample and 17 images of a control sample were used to size the exosomes. The histograms in Fig. 3b show the sizing result obtained from the analysis of 24,024 control and 12,298 patient exosomes. The mean (plus or minus the standard deviation) sizes were 52 ± 21 nm and 50 ± 18 nm for the control and patient samples, respectively.
a Typical scanning electron microscopy (SEM) image of desiccated exosomes. b The exosome size distribution is based on image analysis of 24,024 and 12,298 exosomes derived from control and patient samples, respectively. c The concentration with which the exosomes are deposited on the surface is nonuniform and resembles the coffee ring. We also observe that during the desiccation of the sample, the average size of deposited exosomes may change with spatial position. With reference to the four areas in c delineated by diagonal lines, we see that the largest size is observed in the top-left corner of the image and the size decreases toward the bottom-right corner. This size segregation is quantified in d. pdf probability distribution function
Image analysis of these desiccated samples revealed dependence of exosome sizes on their spatial location within the perimeter of the dried sample. This can be seen in Fig. 3c, which shows the particle size segregation, from larger to smaller, as we move diagonally away from the upper-left corner of the image. Quantification of this change in Fig. 3d shows a rather significant change in exosome sizes in the four areas of the image. In the top-left corner, the exosome sizes are in the range from 50 to 80 nm, and decrease to the predominant sizes of 20–50 nm as we move toward the bottom-right area of the image. Such size segregation of exosomes as a result of drying has not been reported previously. It is likely related to the coffee ring effect [33] known to occur when a suspension of particles dries on a surface. For the case of micrometer particles, the deposition pattern depends on capillary flow in the drying drop [33] and the roughness of the surface on which desiccation occurs [34]. Deposition also depends on several characteristics of the particles themselves—their size, shape [35], and surface activity—and the presence of other surface-active compounds in the solution [36]. These same factors likely influence the deposition of exosomes during surface desiccation of the sample.
The observed coffee ring phenomenon with size segregation suggests that, to avoid bias in characterizing the concentration and sizes of surface-deposited exosomes with techniques such as SEM, TEM, and atomic force microscopy (AFM), the entire area of the dried sample should be imaged and analyzed.
Differential mobility analysis
The electrospray DMA sizing results are shown in Fig. 4. Particles with diameters less than 20 nm were observed (Fig. S3), but were attributed to the contribution of macromolecules, such as globular proteins, lipids, lipoproteins, and agglomerates (dimers, higher-order oligomers, etc.) thereof [37]. For this reason, only particles larger than 20 nm were considered to be exosomes in the DMA results. This decision is supported by cryo-TEM and SEM results that show that very few exosomes smaller than 20 nm are present. Further justification is provided by TEM imaging of DMA-deposited exosomes (see the electronic supplementary material for the description of the experiment and Fig. S4 for a sample of acquired images). The image analysis summarized in Fig. S5 directly confirms that 20 nm is an appropriate lower limit for the diameters of exosomes desiccated in the aerosol. The smaller particles seen in Fig. S3 do not appear in Fig. S4 as distinct objects, which is consistent with biomolecular complexes of lower densities. The size distribution with a 20-nm cutoff seen in Fig. 4 is based on the condensation particle count of 5.50 × 106 particles from the patient sample and 6.22 × 106 particles derived from the control sample. The average diameter of the control and patient sample particles was 37 ± 12 nm and 40 ± 12 nm, respectively.
Discussion
Influence of the exosome isolation method
Exosome sizes overlap with those of other biological particles [38]. Specifically, the sizes of lipoprotein and protein agglomerates overlap with those of exosomes in the lower range of the size distribution, whereas larger extracellular microvesicles and cell debris interfere with exosome sizing at the high end of their size range. Therefore, the exosome isolation method has an influence on the population of the isolated particles and thus affects the sizing results.
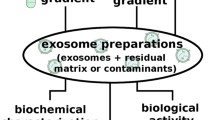
Exosome isolation is not a standardized procedure, and the benefits of different methods are hotly debated. Differential ultracentrifugation remains the most widely used approach, followed by gradient ultracentrifugation and precipitation techniques [39], such as ExoQuick used by us. Numerous studies have been conducted to reveal the influence of the isolation methods on the population of the isolated exosomes and the contamination of the isolated samples by soluble proteins, molecular complexes, and extracellular vesicles other than exosomes [15, 18, 19, 39–41]. For example, Van Deun et al. [39] used three different isolation method—ultracentrifugation, gradient centrifugation, and ExoQuick—to isolate exosomes produced by the MCF-7 breast cancer cell line transfected with a vector to stimulate exosomal production. They found that the isolation method used has an effect on proteomic and nucleic content of the sample, as well as the concentration and size distribution of isolated particles. They concluded that certain gradient fractions obtained with iodixanol-based gradient ultracentrifugation produce exosome samples most enriched in CD63 and other exosomal marker proteins, lowest contamination by extravesicular proteins, and a distinct messenger RNA profile. However, it is not clear if the same narrow-density fraction contains exosomes produced by different cell lines and whether the exosome-rich density band remains narrow in more complicated cases of heterogeneous exosomes secreted by many different types of cells and contained in biological fluids. A recent result shows that most individual exosomes do not carry biologically significant numbers of miRNAs [6], which suggests nonuniform distribution of nucleic material across the population and varying density between exosomes. This is further supported by Jeppesen et al. [41], who found different exosome subpopulations characterized by variable sedimentation characteristics. Reliance on protein biomarkers to verify the quality of exosome isolation has also been questioned because not all exosomes express marker proteins such as programmed cell death 6 interacting protein, tumor susceptibility gene 101 protein, CD9, and CD63 [42]. For exosomes secreted by RBL-2H3 cells, it was reported that only 47 %, 32 %, and 21 % of exosomes contain CD63, MHC class II, and CD81 markers, respectively [43]. As another example of variability in proteins biomarkers, Bobrie et al. [44] reported evidence of heterogeneous vesicle populations and different proportions of CD63, CD9, and milk fat globule–epidermal growth factor 8 protein (lactadherin) markers in exosomes isolated in sucrose gradients at “classic” densities of approximately 1.15 g/mL.
ExoQuick was compared with exosome isolation by ultracentrifugation in the study of Caradec et al. [40]. It was concluded that ExoQuick provides an efficient and reproducible method for exosome isolation for quantitative studies, whereas ultracentrifugation does not. They reported that for serum samples both methods isolate exosomes equivalently expressing several exosomal markers, but ultracentrifugation preparations were strongly contaminated by proteins (albumin and IgG). Rekker et al. [18] examined ultracentrifugation and ExoQuick isolation of exosomes for the purposes of miRNA profiling. They concluded that both methods are suitable for use with serum samples, but noted that the exosomal miRNA profile is slightly affected by the isolation method.
Although the influence of isolation techniques was not investigated in the current study, literature reports indicate that it is an important factor contributing to the reported size variability of exosomes. The source of the exosomes—that is, biological fluids or cell lines—was also not examined, but is likely a factor contributing to the variability seen in Table 1 and other studies.
Exosomes are spherical bioparticles
In the vitrified state, the properties of exosomes are similar to those observed in biological conditions. Cryo-TEM images are formed by electrons transmitted through a vitrified sample (Fig. 5a) and give two-dimensional projections of three-dimensional exosomes in their native hydrated state. Our results show that these projections are close to circular for both patient and control exosomes (mean eccentricities of 0.290 ± 0.085 and 0.334 ± 0.084, respectively). This finding is consistent with only a near-spherical shape of exosomes. It contradicts common claims that exosomes have a “cup shape” [23] (as illustrated in Fig. 5d) reported on the basis of electron microscopy imaging of desiccated samples [12, 45–47] and AFM of both hydrated and desiccated exosomes [48–50]. Our finding is supported by prior cryo-TEM results that report round morphology [51] of exosomes released by hepatocytes, spheroid shape [52] of human mesenchymal stem cell exosomes, and naturally spherical [53] exosomes secreted by prion-infected cells.
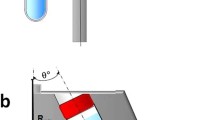
Size and shape of hydrated and desiccated exosomes and comparison of the methods we used to characterize them. a Cryo-TEM images give two-dimensional projections of the geometry of exosomes in their hydrated state. Close-to-circular projections indicate that exosomes are spherical particles. The diameter of the projections characterizes the size of the exosomes in the hydrated spherical state. b Nanoparticle tracking analysis (NTA) and dynamic light scattering (DLS) characterize the mobility of exosomes in solution and estimate their hydrodynamic sizes, which were found to be substantially larger than their geometric sizes. c Electrospray (ES) with charge reduction generates desiccated nanoparticles primarily charged to ±1 or 0. A uniform drying front during aerosol desiccation preserves an approximately spherical shape of the exosomes. DMA separates particles on the basis of their charge-to-size ratio. With predominantly identical +1 change of attracted particles, the separation is based on particle sizes. The condensation particle counter (CPC) counts particles one-by-one. Instead of directing particles for counting, one can deposit particles of a selected size on a substrate. d SEM is used to image exosomes desiccated on a surface. Surface desiccation proceeds with a nonuniform drying front, which likely leads to a shape distortion and the formation of previously observed cup-shaped particles
The cup-shaped morphology of exosomes was questioned before. For example, van der Pol et al. [54] noted that “identification of exosomes based on their cup-shaped morphology after negative staining and visualization by TEM seems questionable.” Our results confirm this suspicion. We furthermore suggest a mechanism (see Fig. 5) that explains this artifact by nonuniformity of capillary forces during surface desiccation, as discussed below.
The conclusion that the innate shape of exosomes is spherical should be taken into account when sizing is performed by techniques that produce shape-dependent results. For example, because particle mobility in a liquid or electrical field is shape dependent, sizing of exosomes by NTA, DLS, or DMA should account for their shape as being spherical, as we have done in this work. Furthermore, the shape distortion away from spherical in the sample preparation step—for example, owing to the electrostatic forces used to immobilize exosomes on a charged substrate prior to AFM characterization or surface desiccation prior to electron microscopy imaging—will likely affect the measured sizes and require correction for such distortion.
Exosome sizing is method specific
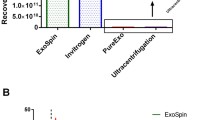
Of the three factors contributing to size variability—the difference in exosome sources, the difference in isolation techniques, and the difference in sizing methods—only the latter was present in this study. Nevertheless, as illustrated by the summary of the sizing results in Fig. 6 and Table 2, the difference in sizing techniques alone introduces variability consistent with what was observed when all three factors were present (Table 1). Specifically, the measured mode size obtained by NTA falls within the range reported by others [17, 55, 56]; the average sizes obtained by SEM and cryo-TEM imaging are also consistent with prior observations [17, 48, 56]. Sizing by DMA gives the average size of exosomes of approximately 40 nm, which is the smallest average size found with any technique.
Summary of sizing results for control and patient samples obtained by different methods. The box contains data between the first (q 1) and third (q 3) quantiles; a red line inside a box marks the second quantile (median). A notch on the box is used to establish if size medians are significantly different. Assuming a normal distribution, nonoverlapping notches imply that true medians are different with 95 % confidence. Comparison of medians is reasonably robust for other distributions and was tested to produce the same statistical conclusion under the assumption of log-normally distributed measurements of exosome sizes. Whiskers encompass data points larger than (2.5q 3 − 1.5q 1) and smaller than (2.5q 1 − 1.5q 3); all data points outside this range are marked with a plus sign +
The difference in the sizing results obtained by different techniques provides an important insight into biophysical properties of exosomes, the understanding of which requires a closer look at the physical principles used in exosome sizing.
Measurements of hydrodynamic sizes
NTA [17, 55] (which has emerged as the most widely used method for characterizing the size distribution and the concentration of exosomes) and DLS both estimate the hydrodynamic (liquid mobility) size of particles (Fig. 5b). The hydrodynamic size is obtained by first estimating particle diffusivity in the solution and then calculating the corresponding particle diameter to match the observations. NTA estimates diffusivity separately for each particle in the field of view of the instrument by measuring particle displacement over time. Unlike NTA, DLS [57]—another commonly used exosome sizing method—is an ensemble technique that estimates the hydrodynamic size distribution of the entire particle population by analyzing the temporal variation in the intensity of the scattered light measured at a fixed scattering angle.
If we disregard the contribution of the solvation layer formed around particles owing to changes in hydrogen bonding of water molecules at the particle–solvent interface [58, 59], the hydrodynamic and geometric sizes will be equal for smooth, hard, electrically neutral, spherical particles with zero surface charges. Several of these assumptions fail in the case of exosomes, which are elastic particles known to have a negative surface charge [44] and have a membrane with conjugated and transmembrane macromolecules. In an aqueous solution, any particle with a surface charge is surrounded by an electrical double layer, sometimes referred to as an ionic atmosphere [60]. As the particle moves, the ionic atmosphere moves with it, making an apparent particle’s size larger than its physical size. The thickness of the ionic atmosphere is approximately equal to the Debye length, which depends on the ionic strength and dielectric permittivity of the solution.
Surface-conjugated macromolecules also exert a strong influence on particle diffusivity. To complicate matters further, the configuration of such surface decoration (e.g., its lateral extension) may change with the concentration of surface ligands, the ionic strength of the solution, its pH, and the type of parent cells that secreted the exosomes [61].
Geometric sizing of exosomes
Desiccated samples
Too small for light microscopy, exosomes can be directly imaged by electron microscopy. Traditional electron microscopy (SEM and TEM) characterizes samples in a vacuum after exosomes have been desiccated on a surface. Exosomes desiccated in the gas phase may be characterized on the basis of their electrical mobility inside differential mobility analyzers. Drying, either on a surface or in the gas phase, introduces severe environmental transformation of biological samples. Figure 5c and d illustrates that a change in interfacial forces during desiccation likely alters the shape and size of exosomes measured by electron microscopy, AFM, and DMA. As discussed later, these changes depend on whether the desiccation occurred on the surface or in the gas phase.
Hydrated samples
Few options exist to characterize geometric sizes of hydrated exosomes, but of these cryo-TEM [62] is the gold standard. Prior to imaging, a thin sample of hydrated exosomes is pipetted on the TEM grid and blotted to remove excess liquid. The sample is then plunged into liquid ethane to impose the rapid cooling needed to obtain the vitrified (amorphous) form of ice, which has properties similar to those of liquid water [62]. In addition to the complexity of sample preparation, another notable disadvantage of cryo-TEM imaging is the relatively small number of exosomes seen in each acquired image. Consequently, the number of exosomes used to characterize the size distribution is small, and only the range of their sizes is often reported in the literature (Table 1).
AFM can be used to visualize the three-dimensional topography of hydrated biological particles. Prior to imaging in the solution, the particles must be immobilized on a substrate, by tethering them to a functionalized surface, by trapping them in a filter, or by electrostatic attraction to a charged surface. The negatively charged exosomal membrane makes electrostatic surface fixation particularly convenient. Note, however, that the same electrostatic forces that immobilize exosomes on the surface likely distort their spherical shape.
Hydrodynamic size of exosomes is substantially larger than their geometric size
In view of the differences discussed, it is expected that the hydrodynamic size of exosomes will always be larger than their geometric size. The degree to which the measured hydrodynamic size exceeds the geometric size is, however, surprising. With reference to Fig. 6 and Table 2, we see that the mean hydrodynamic diameter measured by NTA and DLS is in the range between 119 and 182 nm for both the patient sample and the control sample, whereas the average geometric sizes of hydrated and desiccated exosomes are in the 37–71-nm range. Under the conditions of our experiments, the formation of the electrical double layer alone cannot explain this very large difference. The factor most likely responsible for the substantially larger hydrodynamic size of the exosomes observed by us is their membrane-conjugated macromolecular decoration, which may include transmembrane and tethered proteins, lipids, and saccharides, as well as functional and self-assembled surface-active compounds present in biological fluids and the cell growth medium. We observed faint excess density around some exosomes (Fig. 2d, arrows) that suggests the presence of such molecular decoration on the surface of these particles. The interaction of surface molecules and complexes with a biofluid or buffer impedes the mobility of exosomes, which increases their hydrodynamic size to the extent consistent with our observations. A similarly large increase in the hydrodynamic size due to surface decoration was observed after PEGylation of gold nanoparticles [63], which increased their hydrodynamic diameters to 75 ± 33 nm and 122 ± 50 nm from the initial geometric diameters of 38 and 89 nm, respectively.
Effect of aerosol and surface desiccation on exosome shapes and sizes
DMA characterizes exosome sizes after they have been rapidly desiccated while suspended in a gas. The drying front in an airborne exosome progresses isotropically in the exosome’s radial direction, as illustrated in Fig. 5c. This balances the capillary forces during desiccation and minimizes shape distortion of desiccated exosomes away from their hydrated spherical shape. To confirm that isotropic drying largely preserves the innate spherical shape of hydrated exosomes, the aerosol-desiccated exosomes were imaged by conventional TEM (see the electronic supplementary material for details). Although some exosomes showed signs of shape distortion after experiencing electrospray atomization, aerosol desiccation, and surface impact with the deposition surface, Fig. S4 indicates that they maintain close-to-circular two-dimensional TEM projections (mean eccentricity of 0.420 ± 0.175), as would be expected if a close-to-spherical shape was maintained.
Surface desiccation, unlike aerosol drying, proceeds with a nonisotropic front. This leads to unbalanced capillary forces that likely distort the shape of desiccated exosomes to produce the often observed cup shape of surface-desiccated exosomes, as conceptualized in Fig. 5d. Two symptoms of shape distortion were observed. First, as Table 2 shows, the eccentricity of surface-desiccated control and patient exosomes increased to 0.493 ± 0.313 and 0.540 ± 0.260, respectively from 0.334 ± 0.084 and 0.290 ± 0.085, respectively, for hydrated exosomes observed by cryo-TEM. The second observation is that when the spherical shape is distorted by desiccation on the surface, the two-dimensional projection of SEM-imaged exosomes would be expected to be larger than the diameter of spherical aerosol-desiccated exosomes. This is exactly what we see in Figs. 6 and 7, which show with 95 % statistical confidence that the diameters of aerosol-desiccated exosomes, measured by DMA, are smaller than those of surface-desiccated exosomes, measured by SEM.
Size comparison of exosomes desiccated in aerosol and on the surface for control (top panel) and patient samples. Smaller DMA sizes reflect a near-spherical shape of exosomes uniformly desiccated in the gas phase. Surface drying prior to SEM imaging creates a nonuniform drying front, which causes shape distortion (Fig. 5d), and a larger diameter of the area occupied by surface desiccated exosomes. pdf probability distribution function
Conclusions
We examined the shape and size of serum exosomes in their hydrated state and after desiccation. We found that sizing results for the same sample change significantly when different analytical techniques are used to size the exosomes. The size variability between different methods is significant and consistent with the variability in sizes reported in the literature for cases when the types of cells that secreted the exosomes, the body fluids from which they were isolated, and the exosome isolation methods were also contributing factors. This conclusion places further emphasis on the need to standardize the size quantification techniques and improve our understanding of the biophysical properties of exosomes responsible for the difference in the sizing results obtained with different methods.
Our results indicate that the innate shape of hydrated exosomes is spherical, which implies that the commonly reported “cup shape” of exosomes is an artifact of the sample preparation steps, such as desiccation and surface fixation. Although there is a broad agreement that the sample preparation and fixation steps may be responsible for the commonly observed “cup-shaped” morphology, our report appears to be the first that provides a mechanistic explanation for this artifact, and quantifies its influence on the reported exosome sizes.
We report a large difference in hydrodynamic and geometric sizes of exosomes and attribute it to the presence of membrane-conjugated macromolecules that impede their hydrodynamic mobility. The difference between the geometric and hydrodynamic sizes may therefore be useful in analyzing the conjugation of macromolecules to the surface of exosomes.
The method dependence of exosome sizing extends beyond the differences in the hydrodynamic and hydrated geometric sizes. For desiccated samples, we find that exosome sizing is influenced by the manner in which the drying occurs. We found that isotropic drying during aerosol desiccation preserves the near-spherical shape of the exosomes, whereas drying on a surface likely distorts their shapes and influences the sizing results obtained by electron microscopy, AFM, and other techniques that require fixation on the surface for analysis.
References
Thery C, Zitvogel L, Amigorena S (2002) Exosomes: composition, biogenesis and function. Nat Rev Immunol 2:569–579
Trams EG, Lauter CJ, Salem N Jr, Heine U (1981) Exfoliation of membrane ecto-enzymes in the form of micro-vesicles. Biochim Biophys Acta 645:63–70
Palma J et al (2012) MicroRNAs are exported from malignant cells in customized particles. Nucleic Acids Res 40:9125–9138
King HW, Michael MZ, Gleadle JM (2012) Hypoxic enhancement of exosome release by breast cancer cells. BMC Cancer 12:421
Kucharzewska P, Belting M (2013) Emerging roles of extracellular vesicles in the adaptive response of tumour cells to microenvironmental stress. J Extracell Vesicles. doi:10.3402/jev.v2i0.20304
Chevillet JR et al (2014) Quantitative and stoichiometric analysis of the microRNA content of exosomes. Proc Natl Acad Sci U S A 111:14888–14893
Gallo A, Tandon M, Alevizos I, Illei GG (2012) The majority of microRNAs detectable in serum and saliva is concentrated in exosomes. PLoS ONE 7:e30679
Pucci F, Pittet MJ (2013) Molecular pathways: tumor-derived microvesicles and their interactions with immune cells in vivo. Clin Cancer Res 19:2598–2604
Ge R, Tan E, Sharghi-Namini S, Asada HH (2012) Exosomes in cancer microenvironment and beyond: have we overlooked these extracellular messengers? Cancer Microenviron 5:323–332
Zhang Y et al (2010) Secreted monocytic miR-150 enhances targeted endothelial cell migration. Mol Cell 39:133–144
Regev-Rudzki N et al (2013) Cell-cell communication between malaria-infected red blood cells via exosome-like vesicles. Cell 153:1120–1133
Théry C, Amigorena S, Raposo G, Clayton A (2006) Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr Protoc Cell Biol 30:3.22.1–3.22.29
Taylor DD, Zacharias W, Gercel-Taylor C (2011) Exosome isolation for proteomic analyses and RNA profiling. Methods Mol Biol 728:235–246
Witwer KW et al (2013) Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J Extracell Vesicles. doi:10.3402/jev.v2i0.20360
Tauro BJ et al (2012) Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods 56:293–304
Chen C et al (2010) Microfluidic isolation and transcriptome analysis of serum microvesicles. Lab Chip 10:505–511
Vlassov AV, Magdaleno S, Setterquist R, Conrad R (2012) Exosomes: current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Biochim Biophys Acta 1820:940–948
Rekker K et al (2014) Comparison of serum exosome isolation methods for microRNA profiling. Clin Biochem 47:135–138
Alvarez ML, Khosroheidari M, Kanchi Ravi R, DiStefano JK (2012) Comparison of protein, microRNA, and mRNA yields using different methods of urinary exosome isolation for the discovery of kidney disease biomarkers. Kidney Int 82:1024–1032
György B et al (2011) Membrane vesicles, current state-of-the-art: emerging role of extracellular vesicles. Cell Mol Life Sci 68:2667–2688
van der Pol E, Coumans F, Varga Z, Krumrey M, Nieuwland R (2013) Innovation in detection of microparticles and exosomes. J Thromb Haemost 11(Suppl 1):36–45
van der Pol E et al (2010) Optical and non-optical methods for detection and characterization of microparticles and exosomes. J Thromb Haemost 8:2596–2607
Thery C, Ostrowski M, Segura E (2009) Membrane vesicles as conveyors of immune responses. Nat Rev Immunol 9:581–593
Bacher G et al (2001) Charge-reduced nano electrospray ionization combined with differential mobility analysis of peptides, proteins, glycoproteins, noncovalent protein complexes and viruses. J Mass Spectrom 36:1038–1052
Caulfield MP et al (2008) Direct determination of lipoprotein particle sizes and concentrations by ion mobility analysis. Clin Chem 54:1307–1316
Guha S, Pease LF III, Brorson KA, Tarlov MJ, Zachariah MR (2011) Evaluation of electrospray differential mobility analysis for virus particle analysis: potential applications for biomanufacturing. J Virol Methods 178:201–208
Guha S, Li M, Tarlov MJ, Zachariah MR (2012) Electrospray-differential mobility analysis of bionanoparticles. Trends Biotechnol 30:291–300
Pease LF et al (2010) Packing and size determination of colloidal nanoclusters. Langmuir 26:11384–11390
Wiedensohler A (1988) An approximation of the bipolar charge distribution for particles in the submicron size range. J Aerosol Sci 19:387–389
Flagan RC (2008) Differential mobility analysis of aerosols: a tutorial. KONA Powder Part J 26:254–268
Lattin JR, Belnap DM, Pitt WG (2012) Formation of eLiposomes as a drug delivery vehicle. Colloids Surf B 89:93–100
Belnap DM, Grochulski WD, Olson NH, Baker TS (1993) Use of radial density plots to calibrate image magnification for frozen-hydrated specimens. Ultramicroscopy 48:347–358
Deegan RD et al (1997) Capillary flow as the cause of ring stains from dried liquid drops. Nature 389:827–829
Nguyen TAH, Hampton MA, Nguyen AV (2013) Evaporation of nanoparticle droplets on smooth hydrophobic surfaces: the inner coffee ring deposits. J Phys Chem C 117:4707–4716
Yunker PJ, Still T, Lohr MA, Yodh AG (2011) Suppression of the coffee-ring effect by shape-dependent capillary interactions. Nature 476:308–311
Still T, Yunker PJ, Yodh AG (2012) Surfactant-induced Marangoni eddies alter the coffee-rings of evaporating colloidal drops. Langmuir 28:4984–4988
Kaufman SL, Skogen JW, Dorman FD, Zarrin F, Lewis KC (1996) Macromolecule analysis based on electrophoretic mobility in air: globular proteins. Anal Chem 68:1895–1904
György B et al (2011) Detection and isolation of cell-derived microparticles are compromised by protein complexes resulting from shared biophysical parameters. Blood 117:e39–e48
Van Deun J et al (2014) The impact of disparate isolation methods for extracellular vesicles on downstream RNA profiling. J Extracell Vesicles. doi:10.3402/jev.v3.24858
Caradec J et al (2014) Reproducibility and efficiency of serum-derived exosome extraction methods. Clin Biochem 47:1286–1292
Jeppesen DK et al (2014) Comparative analysis of discrete exosome fractions obtained by differential centrifugation. J Extracell Vesicles. doi:10.3402/jev.v3.25011
Mathivanan S, Simpson RJ (2009) ExoCarta: a compendium of exosomal proteins and RNA. Proteomics 9:4997–5000
Laulagnier K et al (2005) Characterization of exosome subpopulations from RBL-2H3 cells using fluorescent lipids. Blood Cells Mol Dis 35:116–121
Bobrie A, Colombo M, Krumeich S, Raposo G, Théry C (2012) Diverse subpopulations of vesicles secreted by different intracellular mechanisms are present in exosome preparations obtained by differential ultracentrifugation. J Extracell Vesicles. doi:10.3402/jev.v1i0.18397
Yellon DM, Davidson SM (2014) Exosomes: nanoparticles involved in cardioprotection? Circ Res 114:325–332
Kobayashi M et al (2014) Ovarian cancer cell invasiveness is associated with discordant exosomal sequestration of Let-7 miRNA and miR-200. J Transl Med 12:4
Petersen KE et al (2014) A review of exosome separation techniques and characterization of B16-F10 mouse melanoma exosomes with AF4-UV-MALS-DLS-TEM. Anal Bioanal Chem 406:7855–7866
Sharma S, Gillespie BM, Palanisamy V, Gimzewski JK (2011) Quantitative nanostructural and single-molecule force spectroscopy biomolecular analysis of human-saliva-derived exosomes. Langmuir 27:14394–14400
Sharma S et al (2010) Structural-mechanical characterization of nanoparticle exosomes in human saliva, using correlative AFM, FESEM, and force spectroscopy. ACS Nano 4:1921–1926
Sharma S, Das K, Woo J, Gimzewski JK (2014) Nanofilaments on glioblastoma exosomes revealed by peak force microscopy. J R Soc Interface 11:20131150
Conde-Vancells J et al (2008) Characterization and comprehensive proteome profiling of exosomes secreted by hepatocytes. J Proteome Res 7:5157–5166
Zhou Y et al (2013) Exosomes released by human umbilical cord mesenchymal stem cells protect against cisplatin-induced renal oxidative stress and apoptosis in vivo and in vitro. Stem Cell Res Ther 4:34
Coleman BM, Hanssen E, Lawson VA, Hill AF (2012) Prion-infected cells regulate the release of exosomes with distinct ultrastructural features. FASEB J 26:4160–4173
van der Pol E, Böing AN, Harrison P, Sturk A, Nieuwland R (2012) Classification, functions, and clinical relevance of extracellular vesicles. Pharmacol Rev 64:676–705
Momen-Heravi F et al (2012) Impact of biofluid viscosity on size and sedimentation efficiency of the isolated microvesicles. Front Physiol 3:162
Sokolova V et al (2011) Characterisation of exosomes derived from human cells by nanoparticle tracking analysis and scanning electron microscopy. Colloids Surf B 87:146–150
Tscharnuter WT (2006) Photon correlation spectroscopy in particle sizing. In Encyclopedia of analytical chemistry. Wiley, Hoboken. doi:10.1002/9780470027318.a1512
Besseling NAM (1997) Theory of hydration forces between surfaces. Langmuir 13:2113–2122
He L, Hu Y, Wang M, Yin Y (2012) Determination of solvation layer thickness by a magnetophotonic approach. ACS Nano 6:4196–4202
Tathireddy P, Choi Y-H, Skliar M (2008) Particle AC electrokinetics in planar interdigitated microelectrode geometry. J Electrostat 66:609–619
Iyer S, Gaikwad RM, Subba-Rao V, Woodworth CD, Sokolov I (2009) Atomic force microscopy detects differences in the surface brush of normal and cancerous cells. Nat Nanotechnol 4:389–393
Frank J (2002) Single-particle imaging of macromolecules by cryo-electron microscopy. Annu Rev Biophys Biomol Struct 31:303–319
Oh E et al (2011) Cellular uptake and fate of PEGylated gold nanoparticles is dependent on both cell-penetration peptides and particle size. ACS Nano 5:6434–6448
Lavialle F et al (2009) Nanovesicles released by Dictyostelium cells: a potential carrier for drug delivery. Int J Pharm 380:206–215
Varga Z et al (2014) Towards traceable size determination of extracellular vesicles. J Extracell Vesicles. doi:10.3402/jev.v3.23298
Hardij J et al (2013) Characterisation of tissue factor-bearing extracellular vesicles with AFM: comparison of air-tapping-mode AFM and liquid peak force AFM. J Extracell Vesicles. doi:10.3402/jev.v2i0.21045
Dragovic RA et al (2011) Sizing and phenotyping of cellular vesicles using nanoparticle tracking analysis. Nanomedicine 7:780–788
Momen-Heravi F et al (2012) Alternative methods for characterization of extracellular vesicles. Front Physiol 3:354
Hood JL, Pan H, Lanza GM, Wickline SA (2009) Paracrine induction of endothelium by tumor exosomes. Lab Investig 89:1317–1328
Atay S, Gercel-Taylor C, Kesimer M, Taylor DD (2011) Morphologic and proteomic characterization of exosomes released by cultured extravillous trophoblast cells. Exp Cell Res 317:1192–1202
Tian T, Wang Y, Wang H, Zhu Z, Xiao Z (2010) Visualizing of the cellular uptake and intracellular trafficking of exosomes by live-cell microscopy. J Cell Biochem 111:488–496
Tatischeff I, Larquet E, Falcón-Pérez JM, Turpin P-Y, Kruglik SG (2012) Fast characterisation of cell-derived extracellular vesicles by nanoparticles tracking analysis, cryo-electron microscopy, and Raman tweezers microspectroscopy. J Extracell Vesicles. doi:10.3402/jev.v1i0.19179
Wahlgren J, Karlson TDL, Glader P, Telemo, Valadi H (2012) Activated human T cells secrete exosomes that participate in IL-2 mediated immune response signaling. PLoS ONE 7:e49723
Ng YH et al (2013) Endometrial exosomes/microvesicles in the uterine microenvironment: a new paradigm for embryo-endometrial cross talk at implantation. PLoS ONE 8:e58502
Acknowledgments
The authors are indebted to Brian F. Woodfield of Brigham Young University (Department of Chemistry) for providing access to the NTA instrument. The authors acknowledge financial support from the National Science Foundation (award number IGERT-0903715) and the University of Utah (Department of Chemical Engineering Seed Grant and the Graduate Research Fellowship award).
Author contributions
M.S. and P.S.B. conceived the study, R.R. isolated exosomes, V.S.C. and R.R. performed NTA measurements, DLS measurements were performed by V.S.C. and M.S, D.M.B performed cryo-TEM imaging, SEM imaging was performed by Y.J. and V.S.C., electrospray DMA measurements were performed by Y.H.T., V.S.C., L.F.P., and M.S., V.S.C. and M.S. analyzed the experimental results; A.E.B. wrote the MATLAB code to analyze the imaged exosomes, M.S. performed statistical analysis, and V.S.C. and K.J.B. performed manual sizing of the imaged exosomes. The manuscript was written by V.S.C. and M.S., and was edited by all authors.
Conflict of interest
The authors declare that they have no competing financial interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 551 kb)
Rights and permissions
About this article
Cite this article
Chernyshev, V.S., Rachamadugu, R., Tseng, Y.H. et al. Size and shape characterization of hydrated and desiccated exosomes. Anal Bioanal Chem 407, 3285–3301 (2015). https://doi.org/10.1007/s00216-015-8535-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-015-8535-3