Abstract
Metabolic flux analysis is based on the measurement of isotopologue ratios. In this work, a new GC-MS-based method was introduced enabling accurate determination of isotopologue distributions of sugar phosphates in cell extracts. A GC-TOFMS procedure was developed involving a two-step online derivatization (ethoximation followed by trimethylsilylation) offering high mass resolution, high mass accuracy and the potential of retrospective data analysis typical for TOFMS. The information loss due to fragmentation intrinsic for isotopologue analysis by electron ionization could be overcome by chemical ionization with methane. A thorough optimization regarding pressure of the reaction gas, emission current, electron energy and temperature of the ion source was carried out. For a substantial panel of sugar phosphates both of the glycolysis and the pentose phosphate pathway, sensitive determination of the protonated intact molecular ions together with low abundance fragment ions was successfully achieved. The developed method was evaluated for analysis of Pichia pastoris cell extracts. The measured isotopologue ratios were in the range of 55:1–2:1. The comparison of the experimental isotopologue fractions with the theoretical fractions was excellent, revealing a maximum bias of 4.6 % and an average bias of 1.4 %.

Isotopologue distribution of glucose-6-phosphate measured by GC-QTOFMS after derivatization
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
13C-based metabolic flux analysis (13C-MFA) is an emerging approach in biotechnology. In this specific application, isotope tracers, like 13C, are introduced into an organism, where the isotope label is distributed according to the tracer’s metabolic fate and will consequently lead to a characteristic pattern of labelled metabolites. Conducting these labelling experiments will not only allow you to unravel non-obvious metabolic pathways and their regulation, but has also become a key tool in metabolic engineering of microorganisms to prove successful genetic manipulations [1–3].
Evidently, the accurate measurement of isotopologue distributions by NMR or mass spectrometry forms the basis for such experiments, which can be carried out at different levels of complexity—either the isotopologue distribution of proteinogenic amino acids are addressed or in more sophisticated strategies isotopologue distributions of free metabolites have to be assessed. In these cases, the isotopologues of sugar phosphates are of utmost importance, since they represent the starting point of metabolization of the labelled substrate.
Gas chromatography employing capillary columns in combination with mass spectrometry is prominently used in the field of metabolomics due to its typically high separation efficiency and peak capacity. In combination with mass spectrometric detection systems, it represents a highly selective and sensitive methodology [4–7]. State-of-the-art GC-MS methods cover a wide range of analytes (e.g. organic acids, amino acids, sugars and sugar phosphates), which can be separated within a single chromatographic run after appropriate derivatization [8, 9].
Regarding the highly challenging analysis of sugar phosphates occurring in multiple isomers in the metabolic network, methods providing full coverage of all biologically important sugar phosphates in the pentose phosphate pathway are still missing. In the past, capillary electrophoresis (CE)-MS offered high separation efficiency, low sample consumption and selective detection covering a wide range of metabolites (as it is suitable for separation of cationic, anionic and neutral compounds), however, the limit of detection and limit of quantification posed severe limitations to the application of the method in quantitative metabolite profiling, thus also for the analysis of sugar phosphates [10–14]. The state-of-the-art LC-MS methods addressing quantification of sugar phosphates in biological samples [15, 16] involve mixed-mode, anion exchange, ion pairing or hydrophilic interaction liquid chromatography (HILIC). Moreover, reversed phase [17], porous graphitized carbon [18] and propylamine [19] chromatographic columns and specialized columns such as Nucleodex ß-OH [20, 21] were shown to be suitable for separation of at least a substantial number of the sugar phosphates. Han et al. introduced pre-column derivatization prior to separation on pentafluorophenyl core-shell particles with methylphosphonic acid as an efficient tailing-sweeping reagent to alleviate the typically poor retention on reversed phase chromatographic material [22]. Mixed-mode (HILIC-weak anion exchange) chromatography employing MS-friendly mobile phases combined with tandem mass spectrometry for phosphorylated compounds has been established by Hinterwirth et al. [23]. Ion pairing chromatography has been successfully carried out with different reagents and columns [24–27]. However, a major drawback of ion pairing chromatography is the irreversible contamination of mass spectrometry instrumentation by the ion pairing reagents. Ion exchange chromatography in combination with mass spectrometry has been applied successfully for profiling of ionic intermediates after eluent suppression. According to Wang et al., anion exchange chromatography in combination with mass spectrometry and with anion suppression covered a wider range of anionic metabolites than HILIC or ion pairing chromatography [28]. In this context, we have recently presented a method for strong anion exchange chromatographic separation of sugar phosphates with ICP-MS detection [29], which was used as the first chromatographic dimension for an LCxLC-ESI-MS/MS method circumventing the incompatibility of electrospray ionization with high buffer concentrations [30]. Although the method provided the separation of some important sugar phosphates, others had to be excluded due to instability at the high pH of the mobile phase.
Hence, we dare to state that GC-MS remains unrivalled regarding separation power, selectivity and sensitivity in the case of sugar phosphates [31–35] and will prove to be the method of choice for the analysis of this metabolite panel.
In the context of 13C-MFA, two ionization modes, i.e. electron ionization (EI) and chemical ionization (CI) are applicable for the ionization of the alkyloximated/silylated derivatives of sugar phosphates. As a drawback, EI leads to nearly complete fragmentation of the molecular ion and does therefore not allow the isotopologue-selective measurement of labelled metabolites generated in biological tracer experiments. Moreover, high-intensity fragment ions (e.g. m/z 73, 147, 299 and 315) containing no carbon backbone of the sugar phosphate molecule are generated by, e.g. rearrangement of the silyl-group and do not deliver the required isotopologue information for 13C-MFA analysis. Contrarily, chemical ionization is regarded as a soft ionization process generating intact protonated molecule or [M-CH3]+ fragment ions, where a methylgroup of the trimethylsilyl is abstracted, and therefore allowing isotopologue-selective analysis. Different reagent gases for CI including methane, isobutane and ammonia have been investigated in the context of metabolomics [36]. The observed intensity of protonated molecular ions was at least ten times higher as compared to electron ionization and only a low degree of fragmentation and non-selective fragments were obtained. Accordingly, chemical ionization is to date predominantly used for identity confirmation of organic compounds [37, 38]. In the present work, a novel method employing capillary gas chromatography in combination with methane chemical ionization time-of-flight mass spectrometry (GC-CI-TOFMS) is presented for the separation of sugar phosphates and the determination of their isotopologue fractions in yeast cell extract samples. Important ionization parameters, i.e. emission current, electron energy, reagent gas pressure and source temperature were optimized. To the best of our knowledge, this is the first time that GC-CI-TOFMS was used for the determination of isotopologue fractions of sugar phosphates in cell extract samples from Pichia pastoris.
Materials and methods
Chemicals
2-Phosphoglyceric acid disodium salt hydrate (2PG), 3-phosphoglyceric acid disodium salt (3PG), 6-phosphogluconic acid barium salt (6PGA), dihydroxyacetone phosphate lithium salt (DHAP), erythrose-4-phosphate sodium salt (E4P), fructose-6-phosphate disodium salt hydrate (F6P), glucose-6-phosphate dilithium salt (G6P), glyceraldehyde-3-phosphate solution (3GAP), manose-6-phosphate disodium salt hydrate (M6P), mannitol-1-phosphate lithium salt (Mn1P), ribulose-5-phosphate sodium salt (Rb5P), ribose-5-phosphate disodium salt hydrate (R5P), xylulose-5-phosphate sodium salt (Xl5P) were purchased at Sigma Aldrich (Sigma Aldrich, Vienna, Austria) and sedoheptulose-7-phosphate lithium salt (S7P), was purchased from Carbosynth, UK. Stock standard solutions of all compounds were prepared by dissolving an appropriate amount of solid standard in LC-MS-grade water in amber vials. All single stocks were kept at −20 °C and were stable for a minimum of 3 weeks. The working standards were prepared daily in LC-MS grade water. Ethoxyamine hydrochloride was purchased from Sigma Aldrich, N-methyl-N-(trimethylsilyl)trifluoroacetamide (MSTFA) in with 1 % trimethylchlorosilane (TMCS) from Thermo Scientific (Waltham, MA, USA). Ethoxyamine solution was prepared daily by dissolving an appropriate amount of solid substances in water-free pyridine (Sigma Aldrich). LC-MS grade water was used for standard and sample preparation. A mixture of n-alkanes from C8 to C40 in hexane (Sigma Aldrich) was used for the calculation of modified retention indices.
Instrumentation
An Agilent 7200 GC-QTOFMS system (Agilent, Waldbronn, Germany) was used for the separation of sugar phosphates and analysis of their respective isotopologues. The system consisted of a G3440B Agilent 7890B network gas chromatograph equipped with a multimode injector and device backflush device and an Agilent G3851A orthogonal accelerator quadrupole time-of-flight mass spectrometer (QTOFMS) with removable ionization source (EI/CI ionization sources). A multipurpose sampler (MPSII, Gerstel, Germany) was used for automated two-step derivatization of the analytes and injection into the Agilent multimode injector. The sensitivity and accuracy of the mass spectrometer was checked before starting analysis with an internal calibrant solution containing perfluorotributylamine (PFTBA) and perfluoro-5,8-dimethyl-3,6,9-trioxydodecane (PFDTD) applying the mass calibration protocol of the system. The non-bleed septum and single tapered UltraInert Inlet Liners with O-ring pre-installed were changed after approximately hundred injections. The calibration of the mass axis of the TOFMS was programmed in each sequence after ten sample injections. Data acquisition and evaluation were carried out with MassHunter Acquisition and MassHunter Quantitative and Qualitative (Agilent Technologies, CA, USA), respectively.
Chemostat cultivation
The chemostat cultivations were performed in a 1.4-L benchtop bioreactor (DASGIP Parallel Bioreactor System, Germany) with a working volume of 400 mL. A 100-mL pre-culture medium (per litre: 10 g yeast extract, 20 g peptone, 10 g glycerol) was inoculated from the working cell bank (750 μl cryostock of P. pastoris CBS7435) and grown at 28 °C and 150 rpm overnight. This culture was used for inoculation of the bioreactor at an optical density (OD600) of 1.0. After a batch phase of approximately 24 h the feed and harvest for the continuous chemostat cultivation were started. The cells were grown under glucose-limited conditions with a dilution rate of 0.1 h−1 for at least seven residence times before taking the sample. Three independent chemostat cultivations were performed. Temperature, pH and dissolved oxygen were maintained at 25 °C, pH 5.0 (adjusted with 8 M KOH) and 20 % (by controlling the stirred speed and inlet gas flow), respectively. The batch medium contained per litre: 39.9 g glycerol, 1.8 g citric acid, 12.6 g (NH4)2HPO4, 0.022 g CaCl2·2H2O, 0.9 g KCl, 0.5 g MgSO4·7H2O, 2 mL Biotin (0.2 g L−1), 4.6 mL trace salts stock solution. The pH was set to 5.0 with 32 % (w/w) HCl. The chemostat medium contained per litre: 55 g glucose∙1H2O, 2.3 g citric acid, 21.75 g (NH4)2HPO4, 0.04 g CaCl2·2H2O, 2.5 g KCl, 1.0 g MgSO4·7H2O, 2 g Biotin (0.2 g L−1), and 2.43 g trace salts stock solution. The pH was set to 5.0 with 32 % (w/w) HCl. Trace salts stock solution contained per litre: 6.0 g CuSO4·5H2O, 0.08 g NaI, 3.0 g MnSO4∙H2O, 0.2 g Na2MoO4·2H2O, 0.02 g H3BO3, 0.5 g CoCl2, 20.0 g ZnCl2, 5.0 g FeSO4·7H2O, and 5.0 mL H2SO4 (95–98 % w/w).
Sampling and quenching
Approximately 50 mL of the fermentation broth was sampled into 200 mL of 60:40 methanol:water kept at approx. −30 °C for instantaneous quenching of the cellular metabolism. Subsequently, 2-mL aliquots of the quenched cell suspension (corresponding to approximately 10 mg cell dry weight) were filtered through cellulose acetate filters (0.45 μm, Sartorius BioLab Products) using two filtration units (Polycarbonate Filter Holder; Sartorius Lab Technologies Products) equipped with a vacuum pump in parallel. The cell pellet on the filter was washed once with cold quenching solvent before transfer to a 15-mL tube and stored at −80 °C until extraction. The exact biomass content of the sample aliquot was determined by drying five replicates of 2-mL quenched cell suspension to constant weight at 105 °C in pre-weighed glass tubes.
Extraction of sugar phosphates
The metabolite extraction procedure was adapted from Neubauer et al. [39]. In brief, 4 mL of pre-heated extraction solvent (75 % v/v ethanol) was poured onto the filter. After vortexing for approx. 20 s, the samples were incubated at 85 °C in a water bath for 3 min with intermediate mixing. After rapid cooling-down on dry ice, the samples were centrifuged at 4000×g for 10 min at −20 °C. The supernatant was decanted into pre-cooled 15-mL tubes. The ethanolic extracts were stored on at −80 °C.
Automated just in time online derivatization of sugar phosphates
Direct silylation of sugar phosphates leads to the occurrence of a large number of different chromatographic peaks caused by the presence of different isomers of each sugar phosphate, related to cyclic and open-chain structures as introduced by Harvey et al. [44]. In addition, the trimethylsilyl (TMS) derivative of ketose and aldose phosphates is unstable. By introducing an alkyloximation step prior to silylation, the carbonyl group is protected. For the protection of the ketogroups already during sample preparation [9], 10 μL of ethoxyamine hydrochloride in water-free pyridine c(EtOX) = 19 mg mL−1 was added to all sample aliquots and standard ethoximation before drying. The solutions were then dried to complete dryness in a vacuum centrifuge operating at low pressure (below 1 mbar). The two-step derivatization procedure was modified for the MPS2 based on the derivatization parameters published by Koek et al. [5]. For ethoximation, the dried sample was reconstituted in 18 μL ethoxyamine in water-free pyridine (18.7 mg mL−1) and incubated at 40 °C for 90 min Subsequently 42 μL MSTFA with 1 % TMS, was added and the samples were incubated at 40 °C for 50 min for silylation. The derivatized samples were put at 4 °C for approx. 4 min for sample cool down and where then injected by the MPS2.
Determination of retention indices
Because of the low sensitivity of alkane retention index compounds in CI, electron ionization was used to calculate modified retention indexes (gradient temperature retention indices). The retention index mixture was prepared by diluting the Sigma Aldrich retention index standard containing C8–C40 alkanes in 18 μL ethoxyamine in water-free pyridine and 42 μL of MSTFA with 1 % TMCS and injected into GC-EI-TOFMS. The modified retention indices were calculated as described elsewhere [40].
Analysis of sugar phosphates by GC-CI/EI-TOFMS
The sugar phosphate derivatives were separated on a nonpolar Optima 1MS Accent analytical column (Macherey-Nagel, Germany, 60 m × 0.25 mm i.d., 0.25 μm film thickness, 100 % dimethylpolysiloxane stationary phase), which was protected by a deactivated nonpolar guard column (5 m × 0.32 mm i.d., Agilent). Helium (5.0, Linde Austria) was used as carrier gas at a constant flow rate of 1.3 mL min−1. GC injection of 1.0 μL aliquots of sample solution was performed in hot split/splitless mode (splitless time 1 min, 250 °C). The following GC temperature gradient was used: 70 °C for 1 min, 20 °C min−1 to 190 °C, 5 °C min−1 to 225 °C, 3 °C min−1 to 260 °C and 20 °C min−1 to 310 °C (hold for 1 min). The solvent delay was set to 7 min. Back-flushing was employed for 3 min at 310 °C after each chromatographic run. A more detailed information and discussion regarding derivatization and separation of sugar phosphates on the nonpolar Optima 1MS Accent column is in preparation [41].
Chemical ionization was performed with methane (for GC-MS, 5.0, Linde Austria) as reagent gas. For internal standardization during optimization of the source for chemical ionization, meso-erythritol was chosen as internal standard. Optimum CI source parameters were 200 eV electron energy and 40 % methane gas flow (equal to 2.0 mL min−1) at 150 °C. The emission current was kept at 10 μA. The TOFMS was set to a mass acquisition range of 70–900 m/z. For electron impact ionization, the ion source temperature and emission current applied to the filament were set to 220 °C and 35 μA, respectively. Electron energy was kept at 70 eV. With both ionization modes, the TOFMS was operated in 2 GHz-EDR mode (2 GHz extended dynamic range) in order to extend the linear dynamic range.
Isotopologue distribution calculation
In order to evaluate the method regarding the trueness of the determined isotopologue fractions in real matrices, five independently prepared yeast cell extracts were analysed applying the conditions described above. Isotopologue fractions were calculation according to Eq. 1.
where, IF i is the measured isotopologue fraction i and n is the number of carbon atoms in the metabolite backbone. A i represents the peak areas of the respective isotopologues obtained by integration of the extracted ion chromatograms (±50 ppm mass extraction window). The trueness (bias) of the results was obtained by subtracting the natural isotopologue fractions (calculated via envipat open-source software [42]) from the experimental values.
Results and discussion
Optimization of chemical ionization source parameters
In this work, methane and ammonia were tested as reagent gases for ionization of sugar phosphates with CI. The spectra obtained with ammonia as reagent gas revealed less fragmentation of the derivatized sugar phosphates, whereas the intensities of the protonated molecules in the mass spectra were similar to those observed with methane (data not shown). However, due to safety considerations, methane was preferred for further optimization. Using methane, the three dominant reactive species formed by ionization of the reagent gas are CH5 + (m/z 17.0386), C2H5 + (m/z 29.0381) and C3H5 + (m/z 41.0393). Three crucial parameters of the CI source requiring optimization are the reaction gas pressure, the ion source temperature and the emission current applied to the CI filament [43]. For monitoring, the influence of reaction gas pressure on the ratio of C2H5 + and C3H5 +, the methane flow rate was stepwise increased from 10 to 50 % (corresponding to 0.5 to 2.5 ml min−1 methane gas flow) at a source temperature of 150 °C (see below). CH5 + could not be monitored, as the mass range is restricted to m/z 20 in the TOFMS software. The reagent gas pressure was optimized for positive chemical ionization (PCI) in a way that the abundance of the C3H5 + ion was 20 % higher than that of the C2H5 + ion. Otherwise, increased fragmentation would lead to low signal intensities of the protonated molecules due to increased fragmentation. As the proton affinities of C2H5 + and C3H5 + are higher than that of CH5 +, these ions give the highest possible yield of the protonated molecules as fragmentation is less favoured.
However, the application of the optimized reaction gas settings to the analysis of derivatized sugar phosphates revealed the need of further compound specific optimization. A mixture containing a 3PG, a tetrose phosphate (E4P), a pentose phosphate (R5P), hexose phosphate (G6P) and sedoheptulose 7 phosphate was used to find an optimum compromise regarding the settings of the source parameters mentioned above. For this purpose, the intensity ratios of the protonated molecular ion vs. the phosphorous containing base peak present in EI mass spectra (m/z 315.1022 (C9H20O4PSi+) or 299.0718 (C8H17O4PSi+), according to Harvey and Horning [44]) was calculated for the different source settings (see Fig. 1).
Dependence of sensitivity and degree of fragmentation of (a) 3PG, (b) R5P and (c) G6P on the emission current and methane pressure. The upper graphs show the measured intensity of the protonated molecules, whereas the lower graphs depict the intensity ratio of the protonated molecules vs. the derivatized phosphate ion (EC_5, EC_10 and EC_15 correspond to emission currents of 5, 10 and 15 μA)
As shown in Fig. 1, the intensity of the protonated molecular ions of 3PG, R5P and G6P revealed a maximum at an emission current of 10 μA at all selected methane gas flows. At 40 % methane, an emission current of 5 μA showed an improvement regarding sensitivity for high masses only. Independent of the methane pressure, extensive fragmentation was observed when applying higher emission currents on the filament. Considering the extent of fragmentation, it became evident that an emission current of 5 μA would be preferable, but as the precision of isotopologue ratio analysis is strongly depending on sensitivity (ion counting statistics), an emission current of 10 μA was selected as optimum value.
The collision probability between the analyte molecules and the reagent gas increases with gas pressure. As a consequence, the number of proton transfer reactions also increases, especially for high mass analytes. On the other hand, a high pressure of methane in the ion source will lead to lower transmission as observed by Schoots et al. [42]. Fragmentation was lower at lower flow rates, but with the drawback of low sensitivity. Over the investigated mass range, the highest sensitivity was obtained at a methane gas flow of 30 %. Since measurement precision was significantly higher at 40 % than at 30 % methane, we selected 40 % as optimum value.
Temperature and electron energy are additional source parameters affecting the degree of fragmentation and the intensity of protonated molecules. As fragmentation of the derivatized sugar phosphates increased with temperature, an ion source temperature of 150 °C was set as an optimum value. Even lower ion source temperatures were avoided in order to circumvent contamination of the ion source and the quadrupole analyser system. Optimization of electron energy was performed using the internal calibrant solution of the instrument. Changes of electron voltage affected the intensities of fragment ions and protonated molecule to the same degree. According to Gross [45], electron energy should not be lower than 200 eV. As higher voltages did not improve sensitivity, the electron voltage was set to this minimum value, which is also beneficial regarding filament life time. All remaining source parameters, e.g. repeller and ion focusing voltages were first optimized with the automated tuning procedure. Repeller voltage was then further optimized via manual tuning using the internal PFDTD calibrant solution following the suggestions of Abate et al. [37].
Methane chemical ionization mass spectra of derivatized sugar phosphates
In time-of-flight mass spectrometry, accurate mass measurement of protonated molecular ions enables identity confirmation of unknown compounds. Moreover, selectivity is significantly improved, as very narrow mass extraction windows can be used for generation of extracted ion chromatograms. The average mass accuracies obtained for five independent measurements for the investigated sugar phosphates are listed in Table 1. The TOFMS system employed within this work showed an excellent mass accuracy below 7 ppm (external mass calibration).
Representative mass spectra of R5P are depicted in Fig. 2a–c. With CI (2a, 2b), the protonated molecule [M + H]+ with m/z 634.2690 and [M-CH3]+ with m/z 618.2363 represented the most abundant ions. Additionally, an ion at m/z [M + 73.0486]+ was observed in the mass spectrum especially in the case of analytes with high molecular mass. This ion represents an adduct with the trimethysilylium ion which is formed in the source, as a consequence of the large excess of derivatization reagent [46]. Ions which are typically formed in EI (Fig. 2c), i.e. the phosphorus-based nonspecific fragments at m/z 315.1022 (C9H20O4PSi+) and at 299.0718 (C8H17O4PSi+) and unspecific fragments from trimethylsilylation, i.e. (CH3)3Si+ at m/z 73.0468 and (CH3)3SiOSi+(CH3)2 at m/z 147.0814 showed extremely low abundance in the CI spectra. Ions with masses higher than those of the protonated molecules are [M + C2H5]+ at m/z [M + 29.0386]+ and [M + C3H5]+ at m/z [M + 41.0386]+. However, the intensities of those signals were significantly lower than those of the protonated molecules.
Experimental mass spectra of ribose-5-phosphate obtained a in CI mode with 20 %, b in CI mode with 40 % of methane gas flow and c in EI mode at 70 eV. The m/z values given in the spectra represent the theoretical (exact) monoisotopic masses of the detected ions. The measured accurate mass values and are given in Table 1
Separation and detection of derivatized sugar phosphates via GC-CI-TOFMS
An aqueous sugar phosphate standard containing 25 μM of each compound was dried, derivatized and analysed according to a procedure developed by Troyer et al. [41] using the optimized conditions for GC-CI-TOFMS. As can be seen in Fig. 3, the chromatographic method provides excellent separation of all compounds of interest for at least one of the cis- and trans- isomers with the exception ofXl5P. The Xl5P isomers co-elute with R5P and Rb5P. However, as the ratio of the two derivatization products is constant, the area of Xl5P could be calculated as proposed by Cipollina et al. [8]. Additionally, the panel of substances amenable to GC methods is generally limited due to poor derivatization yields and instability of derivatives [9], hampering, e.g. the analysis of glucose-1-phosphate and fructose-1,6-biphosphate, which would be of biotechnological interest.
Isotopologue analysis and determination of isotopologue fractions
As sugar phosphates are important primary compounds of the central carbon metabolism both their selective chromatographic separation and isotopologue-selective detection for determination of isotopologue fractions in 13C tracer experiments is highly desirable. As a matter of fact, the accuracy, i.e. trueness and uncertainty, of isotopologue ratio analysis significantly affects all further data processing in the context of 13C metabolic flux analysis. Table 2 summarizes the isotopologue distributions determined for selected sugar phosphates in yeast cell extracts. Theoretical isotopologue fractions were calculated by isotope distribution calculator tool from envipat open-source software [42] and compared with the experimental values. As can be seen in Table 2, the experimental isotopologue fractions (IFexp) of all analysed sugar phosphates were in excellent agreement with the theoretical values (IFtheor). The average bias was 1.4 % with a maximum value of 4.6 %. The range of the accurately determined isotopologue ratios is directly reflecting the linearity of the method. According to our results, the method is capable to determine isotopologue ratios in the range of 55:1–2:1 (Table 2). The LODs given in Table 1 were calculated following the novel EURACHEM validation guideline and are representing the lower limit of the working range [47]. Repeatability of five independent replicates was below 1 % standard deviation in most cases. This result can be attributed to counting statistics and peak integration repeatability, but is, in contrast to quantitative analysis, independent of derivatization repeatability. Higher values can be explained by the low abundance of the respective metabolites in the yeast cell extracts.
Conclusion
The presented GC-CI-TOFMS method providing modified retention indices, high mass accuracy and accurate isotopologue distribution for sugar phosphates shows a high potential for 13C-based metabolic flux analysis. Compared to EI, the low degree of fragmentation and simpler mass spectra of CI are advantageous especially in isotopologue ratio measurement.
As a general drawback of derivatization, the addition of silicon and carbon atoms demands for correction of the isotopologue distribution before implementation in metabolic flux modelling. Current developments include the programming and validation of a software correcting for the contribution of the natural abundance of the isotopes entered via the derivatization procedure. For this purpose, an object-oriented software tool will be developed using the programming language Perl. It relies on data caching in the form of lookup tables. The use of lookup tables significantly reduces the execution time for the computation of the multinomial distributions and significantly improves the overall performance of the software. The data obtained after this correction step can be directly imported into dedicated software products for MFA such as OpenFLUX2 [48] or 13CFLUX2 [49].
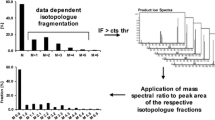
In order to obtain information on the position of the 13C labels, future developments will aim at isotopologue-selective fragmentation via GC-CI-QTOFMS.
References
Stephanopoulos G (1999) Metabolic fluxes and metabolic engineering. Metab Eng 1:1–11
Sauer U (2006), Metabolic networks in motion: 13C-based flux analysis. Mol Biosyst 2, doi:10.1038/msb4100109
Zamboni N, Fendt S, Ruehl M, Sauer U (2009) 13C-based metabolic flux analysis. Nat Protoc 4:878–892
Strelkov S, von Elstermann M, Schomburg D (2004) Comprehensive analysis of metabolite in Corynebacterium glutamicum by gas chromatography / mass spectrometry. Biol Chem 385:853–861
Koek MM, Muilwijk B, Van-Der-Werf MJ, Hankemeier T (2006) Microbial metabolomics with gas chromatography/mass spectrometry. Anal Chem 78:1272–1281
Lisec J, Schauer N, Kopla J, Willmitzer L, Fernie AR (2006) Gas chromatography mass spectrometry-based metabolite profiling in plants. Nat Protoc 1:387–396
Smart KF, Aggio RBM, Van Houtte JA, Villass-Boas SG (2010) Analytical platform for metabolome analysis of microbial cell using methyl chloroformate derivatization followed by gas chromatography mass spectrometry. Nat Protoc 5:1709–1729
Cipollina C, ten Pierick A, Canelas AB, Seifar RM, van Maris AJA, van Dam JC, Heijnen JJ (2009) A comprehensive method for the quantification of non-oxidative pentose phosphate pathway intermediates in Saccharomyces cerevisiae by GC-IDMS. J Chromatogr B 877:2231–3236
Vielhauer O, Zakhartsev M, Horn T, Takors R, Reuss M (2011) Simplified absolute metabolite quantification by gas chromatography-isotope dilution mass spectrometry on the basis of commercially source material. J Chromatogr B 879:3859–3870
Ramautar R, Mayboroda OA, Somsen GW, DeJong GJ (2011) CE-MS for metabolomics: developments and applications in the period 2008–2010. Electrophoresis 32:52–65
Ramautar R, Mayboroda OA, Somsen GW, De Jong GJ (2013) CE-MS for metabolomics: developments and applications in the period 2010–2012. Electrophoresis 34:86–98
Toya Y, Ishii N, Hirasawa T, Naba M, Hirai K, Sugawara K, Igarashi S, Shimizu K, Tomita M, Soga T (2007) Direct measurement of isotopomer of intracellular metabolites using capillary electrophoresis time of flight mass spectrometry for efficient metabolic flux analysis. J Chromatogr A 1159:134–141
Harada K, Ohyama Y, Tabushi T, Kobayashi A, Fukusaki A (2008) Quantitative analysis of anionic metabolites for Catharanthus roseus by capillary electrophoresis using sulfonated capillary coupled with electrospray ionization tandem mass spectrometry. J Biosci Bioeng 105:249–260
Sato S, Yanagisawa S (2010) Capillary electrophoresis electrospray ionization mass spectrometry using fused silica capillaries to profile anionic metabolites. Metabolomics 6:529–540
Fang Z-Z, Gonzalez FJ (2014) LC-MS-based metabolomics: an update. Arch Toxicol 88:1491–1502
Siegel D, Permentier H, Reijngoud D-J, Bischoff R (2014) Chemical and technical challenges in the analysis of central carbon metabolites by liquid-chromatography mass spectrometry. J Chromatogr B 966:21–33
Sun T, Wetzel SJ, Johnson ME, Surlow BA, Patton-Vogt J (2012) Development and validation of a hydrophilic interaction liquid chromatography-tandem mass spectrometry method for quantification of lipids-related extracellular metabolites in Saccharomyces cerevisiae. J Chromatogr B 897:1–9
Antonio C, Larson T, Gilday A, Graham I, Bergstroem E, Thomas-Oates J (2007) Quantification of sugar and sugar phosphates in Arabidopsis thaliana tissues using porous graphitic carbon liquid chromatography-electrospray ionization mass spectrometry. J Chromatogr A 1172:170–178
Lu V, Bennett BD, Rabinowitz JD (2008) Analytical strategies for LC-MS-based targeted metabolomics. J Chromatogr B 871:236–242
Buchholz R, Takors R, Wandrey C (2001) Quantification of intracellular metabolites in Escherichia coli K12 using liquid chromatography-electrospray ionization tandem mass spectrometry techniques. Anal Biochem 295:129–137
Vizan P, Alcarraz-Vizan G, Diaz-Moralli S, Rodriguez-Prados JC, Zanuy M, Centelles JJ, Jauregui O, Cascante M (2007) Quantification of intracellular phosphorylated carbohydrates in HT29 human colon adenocarcinoma cell line using liquid chromatography-electrospray ionization tandem mass spectrometry. Anal Chem 79:5000–5005
Han J, Tschernutter V, Yang J, Eckle T, Borchers CH (2013) Analysis of selected sugar and sugar phosphates in mouse heart tissue by reductive amination and liquid chromatography electrospray ionization mass spectrometry. Anal Chem 85:5965–5973
Hinterwirth H, Lämmerhofer M, Preinerstorfer B, Gargano A, Reischl R, Bicker W, Trapp O, Brecker L, Lindner W (2010) Selectivity issues in targeted metabolomics: separation of phosphorylated carbohydrate isomers by mixed mode hydrophilic interaction/weak anion exchange chromatography. J Sep Sci 33:3273–3282
Huck JHJ, Struys EA, Verhoeven NM, Jakobs C, Van Der Knaap MS (2003) Profiling of pentose phosphates pathway intermediates in blood spots by tandem mass spectrometry: Application to transaldolase deficiency. Clin Chem 49:1375–1380
Michopoulos F, Whalley N, Theodoridis G, Wilson ID, Dunkley TPJ, Critchlow SE (2014) Targeted profiling of polar intracellular metabolites using ion-pair-high performance liquid chromatography and -ultra high performance liquid chromatography coupled to tandem mass spectrometry: application to serum, urine and tissue extracts. J Chromatogr A 1349:60–68
Luo B, Groenke K, Takors R, Wandrey C, Oldiges M (2007) Simultaneous determination of multiple intracellular metabolites in glycolysis, pentose phosphate pathway and tricarboxylic acid cycle by liquid chromatography-mass spectrometry. J Chromatogr A 1147:153–164
Buescher JM, Moco S, Sauer U, Zamboni N (2010) Ultrahigh performance liquid chromatography-tandem mass spectrometry method for fast and robust quantification of anionic and aromatic metabolites. Anal Chem 82:4403–4412
Wang Christison TT, Misuno K, Lopez L, Huhmer AF, Huang Y, Hu S (2014) Metabolomic profiling of anionic metabolites in head and neck cancer cells by capillary ion chromatography with orbitrap mass spectrometry. Anal Chem 86:5116–5124
Chu DB, Klavins K, Koellensperger G, Hann S (2014) Speciation analysis of sugar phosphates via anion exchange chromatography combined with inductively coupled plasma dynamic reaction cell mass spectrometry-optimization for the analysis of yeast cell extracts. J Anal At Spectrom 29:915–925
Klavins K, Chu DB, Koellensperger G, Hann S (2014) Fully automated on-line two-dimensional liquid chromatography in combination with ESI MS/MS detection for quantification of sugar phosphates in yeast cell extracts. Analyst 139:1512–1520
Werner E, Heiliera JF, Ducruix C, Ezan E, Junot C, Tabet JC (2008) Mass spectrometry for the identification of the discriminating signals from metabolomics: current status and future trends. J Chromatogr B 871:143–163
Crutchfield CA, Lu W, Melamud E, Rabinowitz JD (2010) Mass spectrometry-based metabolomics of yeast, Chapter 16. In: Abelson J, Simon M, Verdine G, Pyle A (ed) Method in enzymology, 1rd edn. Elsevier
Pasikanti KK, Ho PC, Chan ECY (2008) Gas chromatography/mass spectrometry in metabolic profiling of biological fluids. J Chromatogr B 871:202–211
Ramautar R, Somsen GW, de Jong GJ (2009) CE-MS in metabolomics. Electrophoresis 30:276–291
Wittmann C (2007) Fluxome analysis using GC-MS. Microb Cell Factories 6:6. doi:10.1186/1475-2859-6-6
Warren CR (2013) Use of chemical ionization for GC-MS metabolite profiling. Metabolomics 9:110–120
Abate S, Ahn YG, Kind T, Cataldl TRI, Fiehn O (2010) Determination of elemental compositions by gas chromatography/time-of-flight mass spectrometry using chemical and electron ionization. Rapid Commun Mass Spectrom 24:1172–1180
Portoles T, Pitarch E, Lopez FJ, Hernandez F, Niessen WMA (2011) Use of soft and hard ionization techniques for elucidation of unknown compounds by gas chromatography/time of flight mass spectrometry. Rapid Commun Mass Spectrom 25:1589–1599
Neubauer S, Haberhauer-Troyer C, Klavins K, Russmayer H, Steiger MG, Gasser B, Sauer M, Mattanovich D, Hann S, Koellensperger G (2012) U13C cell extract of Pichia pastoris—a powerful tool for evaluation of sample preparation in metabolomics. J Sep Sci 35:3091–3105
VanDenDool H, Kratz PD (1963) A generalization of the retention index system including linear temperature programmed gas–liquid partition chromatography. J Chromatogr 11:463–471
Troyer C, Koellensperger G, Hann S. GC-EI-MS/MS with just in time online derivatization for accurate quantification of primary metabolites in biotechnological samples, in preparation
http://www.envipat.eawag.ch/index.php, accessed 20 August 2014
Schoots AC, Leclercq PA (1979) Chemical ionization mass spectrometry of trimethylsilylated carbohydrates and organic acids retained in uremic serum. Biomed Mass Spectrom 6:502–507
Harvey DJ, Horning MG (1973) Characterization of the trimethylsilyl derivatives of sugar phosphates and related compounds by gas chromatography and gas chromatography mass spectrometry. J Chromatogr 76:51–62
Gross JH (2011) Chapter 7 in Mass spectrometry, 2nd ed, Springer
Harrison AG (1992) Chemical ionization mass spectrometry, 2nd ed, CRC
Magnusson B, Örnemark U (eds) (2014) Eurachem Guide: the fitness for purpose of analytical methods – a laboratory guide to method validation and related topics (2nd ed.), ISBN 978-91-87461-59-0. Available from http://www.eurachem.org
Quek L-E, Wittmann C, Nielsen LK, Krömer JO (2009) OpenFLUX: efficient modelling software for 13 C-based metabolic flux analysis. Microb Cell Factories 8:25. doi:10.1186/1475-2859-8-25
Weitzel M, Noeh K, Dalman T, Niedenführ S, Stute B, Wiechert W (2013) 13CFLUX2—high-performance software suite for 13C-metabolic flux analysis. Bioinformatics 29(1):143–145
Acknowledgments
Dinh Binh Chu gratefully acknowledges Erasmus Mundus Action 2 (www.eurasia2.cz) for financial support. EQ BOKU Vienna is acknowledged for providing mass spectrometric instrumentation. This work has been supported by the Austrian BMWFJ, BMVIT, SFG, Standortagentur Tirol and ZIT through the Austrian FFG-COMET-Funding Program.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chu, D.B., Troyer, C., Mairinger, T. et al. Isotopologue analysis of sugar phosphates in yeast cell extracts by gas chromatography chemical ionization time-of-flight mass spectrometry. Anal Bioanal Chem 407, 2865–2875 (2015). https://doi.org/10.1007/s00216-015-8521-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-015-8521-9