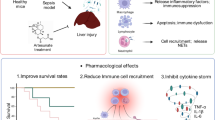
Abstract
The role of non-parenchymal cells (NPCs) in the early phase of acetaminophen (APAP)-induced liver injury (AILI) remains unclear. Therefore, single-cell sequencing (scRNA-seq) was performed to explore the heterogeneity and immune network of NPCs in the livers of mice with AILI. Mice were challenged with saline, 300 mg/kg APAP, or 750 mg/kg APAP (n = 3 for each group). After 3 h, the liver samples were collected, digested, and subjected to scRNA-seq. Immunohistochemistry and immunofluorescence were performed to confirm the expression of Makorin ring finger protein 1 (Mkrn1). We identified 14 distinct cell subtypes among the 120,599 cells. A variety of NPCs were involved, even in the early stages of AILI, indicating highly heterogeneous transcriptome dynamics. Cholangiocyte cluster 3, which had high deleted in malignant brain tumors 1 (Dmbt1) expression, was found to perform drug metabolism and detoxification functions. Liver sinusoidal endothelial cells exhibited fenestrae loss and angiogenesis. Macrophage cluster 1 displayed a M1 polarization phenotype, whereas cluster 3 tended to exhibit M2 polarization. Kupffer cells (KCs) exhibited pro-inflammatory effects due to the high expression of Cxcl2. qRT-PCR and western blotting verified that the LIFR-OSM axis might promote the activation of MAPK signaling pathway in RAW264.7 macrophages. Mkrn1 was highly expressed in the liver macrophages of AILI mice and AILI patients. Interaction patterns between macrophages/KCs and other NPCs were complex and diverse. NPCs were highly heterogeneous and were involved in the immune network during the early phase of AILI. In addition, we propose that Mkrn1 may serve as a potential biomarker of AILI.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Drug-induced liver injury (DILI) is a major cause of unexplained liver injury and a challenge in drug development (Andrade et al. 2019; Hoofnagle and Björnsson. 2019; Shen et al. 2019). Acetaminophen (N-acetyl-p-aminophenol, APAP) is widely used due to its antipyretic and analgesic effects (Ghanem et al. 2016; Yan et al. 2018). However, APAP overdose is currently the leading cause of DILI and acute liver failure (Li et al. 2022; Shen et al. 2019). At present, APAP-induced liver injury (AILI) is the most common and clinically relevant intrinsic DILI model. Hepatotoxicity induced by N-acetyl-p-benzoquinone imine (NAPQI), which is a reactive and toxic metabolite of APAP, is generally considered as the initiating factor of AILI (Stephens et al. 2014).
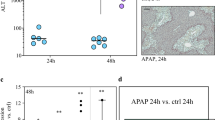
Non-parenchymal cells (NPCs) account for approximately 40% of the total cell population in the human liver, including liver sinusoidal endothelial cells (LSECs), Kupffer cells (KCs), hepatic stellate cells (HSCs), cholangiocytes, and diverse immune cells types (Alvaro et al. 2007; Friedman. 2008; Krenkel and Tacke. 2017; Li and Zeng. 2020; Poisson et al. 2017). In addition to hepatocytes, NPCs are also involved in the occurrence and development of AILI (Stephens et al. 2014). Recent evidence indicates that both innate and adaptive immune systems participate in the development of AILI (Yuan and Kaplowitz. 2013). Our previous study demonstrated that histopathological injury could be observed and alanine aminotransferase (ALT)/aspartate aminotransferase (AST) was significantly elevated during the very early phase of AILI after 3 h of APAP (300 mg/kg) challenge (Li et al. 2018). Simultaneously, levels of inflammatory factors including IL-6, IL-10, and TNF-α were also elevated, suggesting that immune cells may participate in the development of AILI, even during the early stage (Li et al. 2018). Moreover, our bulk RNA-seq results illustrated that the liver transcriptome was significantly altered after 3 h of APAP challenge and differential genes were enriched during the immune response (Li et al. 2018). This suggested that hepatocyte damage may activate an immune response by releasing damage-associated molecular patterns (DAMPs) during the early stages of AILI (Li et al. 2018). Further research has indicated that chemokine (C–C motif) receptor 2 (CCR2)-positive monocytes aggravate the early phase of AILI (Mossanen et al. 2016), and macrophages have also been implicated in the resolution of inflammation during AILI (Antoniades et al. 2012). Cellular interactions between antigen-presenting cells (APCs) and T-cells, APCs, cholangiocytes, HSCs, and hepatocytes are involved in the pathogenesis of DILI (Aithal and Grove. 2015). However, the global landscape of cells involved in the onset of AILI and the interaction among them remain unclear.
Single-cell RNA sequencing (scRNA-seq) is a powerful tool used to analyze specific cell populations and identifies cellular heterogeneity, cell state transitions, and intercellular communication (Hwang et al. 2018). Recently, scRNA-seq has been applied to map the human liver cell atlas and the landscape of liver cells in rodent models of non-alcoholic steatohepatitis (NASH), carbon tetrachloride- or bile duct ligation-induced acute liver injury (Aizarani et al. 2019; MacParland et al. 2018; Massalha et al. 2020; Ramachandran et al. 2019; Terkelsen et al. 2020; Xiong et al. 2019; Yang et al. 2021). A recent study published in 2022 performed spatial transcriptomics and scRNA-seq to reveal the spatiotemporal program of zonal liver regeneration following acute injury 24 h after APAP injection (300 mg/kg) in mice (Ben-Moshe et al. 2022). However, the single-cell transcriptional alterations of NPCs and immune network in an early-stage AILI mouse model are still unknown.
Using scRNA-seq analysis, we demonstrated an unbiased clustering of all liver cells, and focused on the changes in NPCs between healthy and AILI mice. We explored the transcriptional heterogeneity of NPCs and the immune microenvironment during the early phase (3 h) of AILI in mice for the first time. Our study may contribute to further exploration of the mechanisms of AILI and identification of new therapeutic targets.
Methods
AILI mouse model and liver collection
Male C57BL/6 mice, aged 6–8 weeks, were purchased from the Experimental Animal Center of Shanghai SLAC (Shanghai, China). The animals were housed in specific pathogen-free conditions at 24 ± 2 °C under a 12 h light–dark cycle, at a relative humidity of 50 ± 5%. Fresh acetaminophen (Sigma Aldrich, USA) solution was prepared by dissolving acetaminophen in saline in advance, which was warmed up to 55 °C and cooled down to 37 °C before use (Lei et al. 2022). All of the mice were fasted overnight for approximately 12 h before being administered an intraperitoneal (i.p.) injection of APAP at a dose of 750 mg/kg (n = 3), 300 mg/kg (n = 3), or warm phosphate-buffered saline (PBS, 37 °C) (n = 3). The mice were humanely euthanized after 3 h of treatment with APAP or saline. The intact liver tissue from each mouse was collected for subsequent digestion. All animal studies were conducted in accordance with the guidelines of the Institutional Animal Care and Use Committee of Renji Hospital, School of Medicine, Shanghai Jiao Tong University (No. RJ-2018–0930).
NPC single-cell suspensions and single-cell RNA sequencing
Fresh liver tissues were stored in sCelLiveTM Tissue Preservation Solution (Singleron Bio Co., Nanjing, China) and pre-cooled on ice immediately after being collected. Approximately 10 min later, the liver tissues were washed with Hank’s Balanced Salt Solution (HBSS) three times and then digested with 2 ml sCelLiveTM Tissue Dissociation Solution (Singleron) via the Singleron PythoN™ Automated Tissue Dissociation System (Singleron) at 37 °C for 15 min. The solution was then centrifuged at 500 × g for 5 min and suspended in PBS. Finally, the suspensions were stained with trypan blue (Sigma, United States), and cellular viability was greater than 90% microscopically. NPC single-cell suspensions were prepared at a concentration of 1 × 105 cells/ml in PBS. NPC single-cell suspensions were then loaded onto microfluidic devices, and scRNA-seq libraries were constructed using the GEXSCOPE® Single-Cell RNA Library Kit (Singleron Biotechnologies) according to the manufacturer’s instructions (Singleron GEXSCOPE®) (Dura et al. 2019). The individual libraries were diluted to 4 nM and pooled for sequencing. These libraries were then sequenced on an Illumina HiSeq X with 150 bp paired-end reads.
scRNA-seq quantifications and statistical analysis
Raw reads were processed using an internal pipeline to generate gene expression profiles. Briefly, the cell barcodes and Unique Molecular Identifiers (UMI) were extracted after filtering the read barcodes without poly T tails. Adapters and poly A tails were trimmed (fastp V1) before aligning read two to GRCh38 with ensemble version 92 gene annotation (fastp 2.5.3a and featureCounts 1.6.2) (Liao et al. 2014). Reads with the same cell barcode, UMI, and gene were grouped together to calculate the number of UMIs of the genes in each cell. The UMI count tables for each cellular barcode were used for further analyses. Cell type identification and clustering analysis were performed using the Seurat program (Butler et al. 2018; Satija et al. 2015), and further (http://satijalab.org/seurat/, R package, v.3.0.1) used to analyze the sequencing data. The UMI count tables were loaded into R using the read.table function. Next, a parameter resolution of 0.6 was set for the FindClusters function for clustering analyses. Differentially expressed genes (DEGs) between different samples or consecutive clusters were identified using the FindMarker function. Gene Ontology (GO) function and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were carried out on the gene set using the clusterProfiler software to explore biological functions or pathways that are significantly associated with the specifically expressed genes (Yu et al. 2012). Pseudotime trajectory analysis was performed using Monocle (Version 2.10.0) to identify differential gene expression and time-series analysis. The cell–cell interaction between cells was predicted based on known ligand-receptor pairs using Cellphone DB (Qiu et al. 2017) v2.1.0. CellCall v0.0.0.9000 (Zhang et al. 2021) was used to analyze the intercellular interaction based on the receptor-ligand interaction between two cell types/subtypes, and to infer the signaling pathways of internal regulation. The scRNA-seq data have been deposited into the Gene Expression Omnibus database under accession code GSE228305.
Human liver samples collection and process
Liver tissues from patients with AILI, herbal-induced liver injury (HILI), non-alcoholic fatty liver disease (NAFLD), chronic hepatitis B (CHB), liver cirrhosis, and primary biliary cholangitis (PBC) were collected to evaluate the expression of Dmbt1 or Mkrn1. Liver tissues were collected from healthy donors (used as the control groups) whose livers were subsequently used for liver transplantation. Paraffin embedding, slice sectioning, and immunohistochemistry or immunofluorescence were performed. The study of human samples was approved by the ethics committee of Renji Hospital, School of Medicine, Shanghai Jiao Tong University (No. RA-2019–175).
Statistical analysis
Data are expressed as mean ± standard error of the mean (SEM). Multiple group comparisons were performed using a one-way analysis of variance (ANOVA). P < 0.05 was considered as statistically significant. Calculations and graphs were prepared using the GraphPad Prism software (version 9.0; San Diego, CA, USA). For details regarding the materials and methods used, please refer to the Supplementary information.
Results
Single-cell atlas of liver NPCs in healthy and early-phase AILI mice
We successfully obtained liver NPCs from healthy and early-phase AILI mice. ScRNA-seq was performed as shown in Fig. 1a. After quality control analysis (Supplementary Fig. 1), approximately 120,599 cells were selected for further analysis. These cells were classified into 14 cell clusters: B-cells, T-cells, neutrophils, endothelial cells (ECs), epithelial progenitors, cholangiocytes, hepatocytes, plasmacytoid dendritic cells (pDCs), mononuclear phagocytes (Mps), fibroblasts, plasma cells, smooth muscle cells (SMCs), hepatic stellate cells (HepSCs), and erythrocytes (Fig. 1b). The Uniform Manifold Approximation and Projection (UMAP) plot revealed the distribution of cells between the control and AILI groups (Fig. 1c). The proportions of each cluster are shown in Fig. 1d. The violin plot displays the unique expression profile of the representative marker genes for each cluster (Fig. 1e).
Single-cell atlas of liver NPCs in healthy and early phase acetaminophen-induced liver injury (AILI) mice. a Study design, cell isolation, capture, RNA sequencing, and data analysis. The illustration was created with FigDraw.com. b Uniform Manifold Approximation and Projection (UMAP) plot of liver cell clusters based on 120,599 single-cell transcriptomes. A total of 14 different clusters were identified after unsupervised clustering. c The sample origin of cell clusters. d The distribution of cell clusters between the three groups. e Expression of representative marker genes for each cell type. Gene expression violin plots are shown in log-scale Unique Molecular Identifiers (UMI). f Gene expression of CYP450 isozymes in the different cell types. g Gene expression of Gst, Nat, Ugt, and Sult in different cell types
Considering that our model is a drug-injured model, we further analyzed the drug-metabolizing enzyme expression profiles of the selected cell clusters. Previous studies have reported that isoforms of the CYP2 and CYP3 families are highly expressed in mouse livers and are the main enzyme families for drug metabolism and detoxification (Nelson et al. 2004). Excess APAP led to increased CYP2E1 activity and decreased CYP1A2 and CYP3A4 expression in mouse livers (Xie et al. 2014). Here, we found that, in addition to Cyp2e1, the widely known APAP-metabolizing enzyme, and other CYP450 superfamily members, such as Cyp2d9, Cyp2f2, Cyp4a14, and Cyp4a10, were highly expressed in hepatocytes. More intriguingly, Cyp4b1 and Cyp26b1 were highly expressed in ECs, while Cyp26b1 was gradually decreased in ECs after APAP administration (Fig. 1f). Gsts were mainly expressed in cholangiocytes, hepatocytes, SMCs, and HepSCs, and we found that the expression of Gsts in cholangiocytes tended to increase after APAP treatment. Nat, Ugt, and Sult were mainly expressed in hepatocytes and cholangiocytes (Fig. 1g).
Dmbt1+ cholangiocytes are involved in drug metabolism and detoxification processes
A total of 9,300 cholangiocytes were detected, and were further divided into four clusters (Fig. 2a). Figure 2b displays the unique expression profile of the representative marker genes for each cluster. Next, we analyzed the characteristics and heterogeneity of cholangiocytes after APAP challenge. As shown in Fig. 2c and Supplementary Fig. 2, the genes upregulated by APAP were associated with heat shock proteins, inflammation, adhesion, apoptosis, autophagy, and ubiquitination. Interestingly, we also observed the significantly high expression of genes related to the cell cycle, cell regeneration and proliferation, especially in cluster 3 (Fig. 2c).
Dmbt1+ cholangiocytes are involved in drug metabolism and detoxification processes. a The UMAP plot of 9,300 cholangiocytes was reclustered into four clusters: the sample origin of cholangiocytes and the distribution of cell clusters between the three groups. b Circle plots displaying the expression of the marker gene in four clusters. c Heat map representation of differentially expressed genes (DEGs) in four clusters between control and AILI groups. d The cholangiocyte trajectories of different clusters and different groups. e UMAP plot displaying Dmbt1 gene expression in cholangiocytes; trajectory distribution of Dmbt1 in cholangiocytes. f Heat map representation of gene expression in four clusters: Gene Ontology (GO) analysis of Dmbt1 + cholangiocytes. g Anti-Dmbt1 immunofluorescence staining of liver sections from control and AILI mice. h Anti-Dmbt1 immunofluorescence staining of liver sections from healthy and AILI patients. i Heatmap of genes that were significantly differentially expressed along the pseudotime axis
Next, we applied pseudotime methods to simulate the differentiation trajectory of the cholangiocytes. The results suggest that cholangiocyte cluster 1 may be at the starting point of cholangiocytes (Fig. 2d). The number of cells in cluster 3 decreased with increasing APAP doses (Fig. 2d). Among the genes upregulated in cluster 3 by APAP, Dmbt1, which is an important factor in liver regeneration and ductular cell differentiation, was the top gene (Fig. 2e). Therefore, we defined cluster 3 cholangiocytes as Dmbt1+ cholangiocytes. We found that Dmbt1+ cholangiocytes markedly expressed drug-metabolizing enzymes as well as genes exerting anti-inflammatory effects (Fig. 2f). KEGG analysis revealed that Dmbt1+ cholangiocytes were mainly involved in the response to xenobiotic stimuli, the glutathione metabolic process, the epoxygenase P450 pathway, and detoxification (Fig. 2f). Immunofluorescence confirmed that the expression of Dmbt1 in the mouse liver decreased after APAP treatment (Fig. 2g). Liver tissue sections from patients also indicated that the expression level of Dmbt1 was reduced in patients with AILI (Fig. 2h). DEGs along the pseudotime axis were clustered into four modules according to their expression patterns (Fig. 2i, Supplementary Fig. 2).
Overall, our results indicated that cholangiocytes play an important role in the early phases of AILI, and Dmbt1+ cholangiocytes are involved in drug metabolism and detoxification processes.
LSECs, fenestrae loss and pro-angiogenesis occur in early phase of AILI
In total, 8077 endothelial cells were detected, which were further divided into two subpopulations: LSECs (approximately 80%) and vascular endothelial cells (Fig. 3a). We found that LSECs expressed high levels of endothelial cell markers (Lyve1, Plpp3, Aadam23, Flt4, Gja4, and Kdr). In addition, genes related to antigen recognition, presentation, and endocytosis, such as Clec4g, Fcgr2b, Clec1b, and Dpp4 were also highly expressed in LSECs (Fig. 3b).
Liver sinusoidal endothelial cells (LSECs) experience fenestrae loss and pro-angiogenesis in early phase of AILI. a The UMAP plot of 8,077 endothelial cells (ECs) was reclustered into two clusters: the sample origin of ECs and the distribution of cell clusters between the three groups. b Canonical cell markers were used to label LSECs based on cell identity, as represented in the ECs UMAP. c Circle plots displaying the DEGs in LSECs between control and AILI groups. d Representative ligand-receptor signaling axes interactions between T-cells and LSECs. e Representative ligand-receptor signaling axes interactions between LSECs and T-cells. f Representative transmission electron microscopy images showing the fenestrae of LSECs in the control and AILI groups. Yellow arrows indicate fenestrae
As shown in Fig. 3c and Supplementary Fig. 3, among all the DEGs identified in the LSECs, genes involved in angiogenesis and vessel integrity, adhesion, actin cytoskeleton organization, hypoxia and oxidative stress, inflammation, and chemotaxis were the most significantly upregulated after APAP treatment. These genes were further upregulated in the 750 mg/kg APAP group when compared to the 300 mg/kg APAP group (Fig. 3c).
LSECs play an important role in immune tolerance by communicating with T-cells, thereby influencing pathological progression (Poisson et al. 2017). In the present study, we observed many pairs of such as TIGIT-Nectin2, ICAM1-ITGAL, CXCL10-CXCR3, and CXCL10-DPP4 between LSECs and T-cells were activated after APAP challenge (Fig. 3c, e). Transmission electron microscopy revealed that LSECs were morphologically regular with clear cell margins and a normal number of fenestrae (yellow arrows) under normal conditions (Fig. 3f). After treatment with 300 mg/kg APAP, LSECs were enlarged and widened with a reduced number of fenestrae (yellow arrows); after challenging with 750 mg/kg APAP, LSECs became swollen and deformed with blurred cell margins and even less or no fenestrae (Fig. 3f).
Overall, our results indicate that LSECs can be implicated in the pathogenesis of AILI through the loss of fenestrae and pro-angiogenesis.
NK cells exert cell-killing activity in early phase of AILI
A total of 40,316 T-cells were detected and further divided by marker annotation into six clusters including Treg, CD8Teff, natural killer (NK), NKT, Naïve T, and Gamma Delta T (GDT) cells. We found that the number of NK cells increased in a dose-dependent manner after APAP treatment (Fig. 4a). In addition, the heatmap of the top marker genes displayed a unique expression profile for each cluster (Fig. 4b).
Natural killer (NK) cells exert cell-killing activity in early phase of AILI. a The UMAP plot of 40,316 T-cells was reclustered into six clusters: the sample origin of T-cells, and the distribution of cell clusters between the three groups. b Violin plot displaying representative marker gene expression for each cluster. c Circle plots displaying the DEGs in NK cells between control and AILI group. d GO analysis of NK cells between the control and 300 kg/mg AILI group, and between the control and 750 kg/mg AILI group. e KEGG analysis of NK cells between the control and 300 kg/mg AILI group, and between the control and 750 kg/mg AILI group
Next, we explored the transcriptional heterogeneity of each T-cell cluster in the AILI groups. We found that upregulated genes in NK cells were associated with cell killing activity (Klra8, Klra13-ps, and Klra4), hypoxia, oxidative stress (Rora, Mt1, Sesn1, Mt2, Txnip), inflammation, and chemotaxis (Litaf, Sqstm1, Cxcr4 and Cxcl2) (Fig. 4c). GO enrichment analysis revealed that NK cells were involved in cell killing, lysosomes, tumor necrosis factor production, T-cell-mediated cytotoxicity, and interferon-gamma production (Fig. 4d). KEGG enrichment analysis indicated that the inflammatory signaling pathway, necroptosis and NK cell-mediated cytotoxicity were primarily activated after APAP challenge (Fig. 4e). The upregulated DEGs and enrichment analysis of other T-cell clusters are shown in Supplementary Fig. 4. The results indicate that they are mainly involved in apoptosis, oxidative stress, inflammation, chemotaxis, T-cell proliferation, and activation (Supplementary Fig. 4).
Overall, we found that T-cells participate in the onset of AILI, which is mainly indicated by NK cells exerting cell-killing activity.
Macrophages differently polarized in the early phase of AILI
In our study, 11,432 Mps were detected, which were further clustered into eight groups (Supplementary Fig. 5). The proportion of macrophages was the highest in Mps, followed by monocytes and Kupffer Cells (Supplementary Fig. 5). A total of 2556 macrophages were further subdivided into three clusters. We found that upon treatment with 300 mg/kg and 750 mg/kg APAP, the percentage of cluster 1 and cluster 2 macrophages was significantly elevated, while that of cluster 3 macrophages was decreased (Fig. 5a).
Macrophages differently polarized in the early phase of AILI. a The UMAP plot of 2,556 macrophages was reclustered into three clusters: the sample origin of macrophages and the distribution of cell clusters between the three groups. b Violin plots displaying representative polarization gene expression in each cluster. c Volcano plots of DEGs upregulated in the 300 mg/kg AILI group in cluster 1 macrophages. d Volcano plots of DEGs upregulated in the 300 mg/kg AILI group in cluster 2 macrophages. e Volcano plots of DEGs upregulated in the 300 mg/kg AILI group in cluster 3 macrophages. f The GO and KEGG analysis of three clusters between control and AILI groups
Macrophages with different polarization statuses play different roles in the pathogenesis of liver disease (Krenkel and Tacke. 2017). We found that macrophages in cluster 1 were M1-polarized, with a high expression of Ccl2, Ms4a7, Cx3cr1, C3ar1, Trem2, Tlr2, Il1b, Ccrl2, Cxcl16 and Tlr7, while cluster 3 macrophages were M2 -polarized, with a high expression of Arg1, Cd5l, CD163, Serpina1b and Vsig4. Contrastingly, cluster 2 did not preferentially express M1 and M2 marker genes (Fig. 5b, Supplementary Fig. 5). Volcano plots display the DEGs identified in macrophage clusters 1, 2, and 3 from the 300 mg/kg APAP-injured liver and those from the saline-treated group (Fig. 5c, d, e). In cluster 3, M2 markers, such as Vsig4, Mrc1 and Cd163 were significantly downregulated (Fig. 5e). Therefore, in the early phase of AILI, more M1-like cluster 1 macrophages are present with high quantity and activation, and more M2-like cluster 3 macrophages are present with low quantity and dysfunction, which may play a role in the pathogenesis of liver injury. Functionally, clusters 1 and 2 were involved in inflammatory signaling pathways, chemotaxis, the pattern recognition receptor signaling pathway, the HIF-1 signaling pathway, and the nitric oxide metabolic process (Fig. 5f). For cluster 3 macrophages, pathways relevant to cytokine and chemokine production, tumor necrosis factor production, the nitric oxide metabolic process, tissue regeneration, and response to IL-4 occurred in the 750 mg/kg APAP-injured liver. Thus, it is suggested that cluster 3 may be involved in injury repair and tissue remodeling in APAP-induced liver failure (Fig. 5f).
Overall, we found that macrophages were differently polarized in the early phase of AILI. More M1-like cluster 1 macrophages tended to be activated, while simultaneously more M2-like cluster 3 macrophages presented dysfunction, which might be involved in the pathogenesis of AILI.
Kupffer cells express high levels of CXCL2 in early phase of AILI
A total of 2,653 Kupffer cells were identified. The cells were further divided into four groups. Cluster 3 demonstrated an APAP dose-dependent increase in cell number (Fig. 6a). The violin plot displays the unique expression profile for each cluster (Fig. 6b).
Kupffer cells express high levels of CXCL2 in early phase of AILI. a The UMAP plot of 2,653 Kupffer cells (KCs) was reclustered into four clusters: the sample origin of KCs and the distribution of cell clusters between the four groups. b Violin plot displaying representative marker gene expression for each cluster. c Heat map representation of DEGs in four clusters between the control and AILI groups. d The GO analysis of four clusters between control and 300 kg/mg AILI group. e The KC trajectories of different clusters and different groups. f Heat map representation of gene expression in the four clusters: GO analysis of cluster 3. g Trajectory distribution of CXCL2 in KCs. h Heatmap of genes that were significantly differentially expressed along the pseudotime axis
Genes involved in inflammation, endoplasmic reticulum stress, DNA damage, histone demethylase, autophagy, and ubiquitination were the most significantly upregulated DEGs after 300 mg/kg APAP treatment compared to the saline group. In addition, these genes were found to be further upregulated in the 750 mg/kg APAP group, especially in cluster 3 Kupffer cells (Fig. 6c). GO enrichment and KEGG pathway analyses revealed that upregulated DEGs in cluster 1, 2, and 3 Kupffer cells were mainly enriched for cell chemotaxis, cell adhesion, inflammatory pathway activation, and ubiquitin protein ligase binding. Interestingly, all Kupffer cell clusters upregulated ubiquitin protein ligase binding after APAP treatment (Fig. 6d, Supplementary Fig. 5).
Pseudotime analysis suggested that cluster 1 may be the starting point of Kupffer cells. The results indicated that the cell number in cluster 3 increased with an increasing APAP dose, indicating that APAP treatment contributed to the evolution and occurrence of cluster 3 (Fig. 6e). Cluster 3 Kupffer cells highly expressed Cxcl2, and Cxcl2 was the top feature gene in cluster 3 Kupffer cells (Fig. 6f). Cluster 3 Kupffer cells also highly expressed some key inflammatory factors and chemokines, such as Cxcl1, Il6, Ccl3, Ccrl2, Nlrp3, Nfkb2 and Tnf (Fig. 6f). We found that cluster 3 Kupffer cells were mainly involved in reactive oxygen species metabolic processes, cytokine production, and inflammatory responses (Fig. 6f). Furthermore, the expression level of Cxcl2 gradually increased with the evolution of Kupffer cells (Fig. 6g). DEGs along the pseudotime axis were clustered into four modules according to their expression patterns (Fig. 6h, Supplementary Fig. 5).
Overall, we found that Kupffer cells were significantly activated in the early stages of AILI through the secretion of high levels of inflammatory cytokines and chemokines, especially CXCL2.
Enhanced interactions between macrophages/Kupffer cells and other NPCs
To further investigate the mutual regulatory process between macrophages/Kupffer cells and other NPCs in AILI livers, we performed cell–cell communication analysis by mapping the ligand-receptor interactions. Overall, we observed extensive interactions between macrophages/Kupffer cells and other cells after APAP treatment (Supplementary Fig. 6).
We found that the co-stimulation signal, CD28-CD86, between Kupffer cells and T-cells was activated in AILI livers. In addition, ADORA2A-ENTPD1, ADORA2A-NAMPT, IL-1b-ADRB2, CRLF2-TSLPR, and HGF-CD44 were activated in Kupffer and T-cells. The interaction patterns of macrophages and Kupffer cells with other cells were similar (Fig. 7a). Using immunofluorescence validation in saline- and APAP-treated mice, we confirmed CD28-CD86 activation between T-cells and macrophages after APAP challenge (Fig. 7b).
Enhanced interactions between macrophages/KCs and other NPCs. a Representative ligand-receptor signaling axis interactions between macrophages/KCs and other NPCs. b Representative CD28-CD86 immunofluorescence staining images of healthy and AILI mice. White circles indicate double-stained cells. c Representative autocrine ligand-receptor signaling axis interactions in macrophages and KCs. d qRT-PCR confirmed that Osm, Lifr, Tnf, Il6, Cxcl2, Ccl2, Trem2, and Inos were upregulated in RAW264.7 cells treated with oncostatin M (OSM). Data are shown as mean ± SEM (*P < 0.05, **P < 0.01, vs. control). e Western blotting verified that mitogen-activated protein kinase (MAPK) signaling pathway-related protein expression was upregulated in RAW264.7 cells treated with OSM. f Graphical abstract displaying autocrine LIFR-OSM signaling pathway activation in macrophages. The illustration was created with BioRender.com
Furthermore, we found that ligand-receptor interactions were activated between macrophages and macrophages, and Kupffer cells and Kupffer cells, indicating that both paracrine and autocrine signaling play a significant role in AILI (Fig. 7c). We treated RAW264.7 macrophages with oncostatin M (OSM) to validate paracrine or autocrine signaling between macrophages through leukemia inhibitory factor receptor (LIFR)-OSM interactions. OSM is a member of the IL-6 family and is involved in tumor cell proliferation, inflammation and metabolism (Lantieri and Bachetti. 2022). The results indicated that OSM significantly induced Tnf, Il-6, Trem2, Inos, Ccl2, Ccl5, Cxcl2 and Cxcl1 gene expression and suppressed Il-10 and Arg1 gene expression transcriptionally in RAW264.7 cells (Fig. 7d, Supplementary Fig. 6). The possible mechanisms by which OSM drives the secretion of cytokines and chemokines by macrophages should be further explored. Using western blot analysis, we found that OSM significantly activated the mitogen-activated protein kinase (MAPK) signaling pathway by upregulating the protein expression of p-p38, p-ERK and p-SAPK/JNK (Fig. 7e). This suggests that OSM might directly induce macrophage inflammation activation and M1 polarization via the MAPK signaling pathway (Fig. 7f).
Overall, we observed a number of extensive interactions between macrophages/Kupffer cells and other cells at the onset of AILI. Furthermore, we found that macrophages are activated by LIFR-OSM autocrine signaling.
Mkrn1 is a potential macrophages-specific indicator of AILI
Next, we explored whether there is a potential diagnostic indicator in the early phase of AILI. Mps presented increased quantity and functional activation exclusively after APAP challenge. We found that 21 Mps marker genes were simultaneously upregulated by both 300 mg/kg APAP and 750 mg/kg APAP treatment (Fig. 8a). Among them, Mkrn1 was substantially and specifically expressed in Mps and was highly expressed in various subpopulations of Mps (Fig. 8b). Mkrn1 is an E3 ubiquitin ligase involved in metabolic diseases, malignancies, cell survival, and apoptosis through ubiquitination of substrate proteins (Kim et al. 2005). Immunohistochemical analysis demonstrated that protein expression of Mkrn1 in the liver was increased by APAP treatment in a dose-dependent manner (Fig. 8c). In addition, using immunofluorescence, we further confirmed that Mkrn1 expression was specific to macrophages in AILI mouse livers (Fig. 8d). Liver tissue sections from patients also validated that Mkrn1 was highly expressed in patients with AILI compared with the control patients (Fig. 8e, f, g). Furthermore, we found that Mkrn1 was barely expressed in other liver diseases such as herbal-induced liver injury (HILI), non-alcoholic fatty liver disease (NAFLD), chronic hepatitis B (CHB), liver cirrhosis, and primary biliary cholangitis (PBC) (Fig. 8h). These results suggest that Mkrn1 might be a macrophage-specific, highly expressed protein in the early phase of AILI and may serve as a potential indicator.
Makorin ring finger protein 1 (Mkrn1) is a potential macrophage-specific indicator of AILI. a Venn diagram displaying the APAP-induced upregulated genes in Mps marker genes. b Circle plot of Mkrn1 expression in main cell type; violin plot of Mkrn1 expression in the Mps. c Representative Mkrn1 immunohistochemistry staining images in healthy and AILI mice; red arrows indicate Mkrn1 stained cells. d Representative images of Mkrn1 and F4/80 double immunofluorescence staining in healthy and AILI mice; white circles indicate double-stained cells. e Representative images of Mkrn1 and CD68 double immunofluorescence staining in control and AILI patients; white arrows indicate double-stained cells. f Representative Mkrn1 immunohistochemistry staining images in control patients. g Representative Mkrn1 immunohistochemistry staining images in AILI patients; red arrows indicate Mkrn1 stained cells. h Representative Mkrn1 immunohistochemistry staining images in patients with herbal-induced liver injury (HILI), non-alcoholic fatty liver disease (NAFLD), chronic hepatitis B (CHB), liver cirrhosis, and primary biliary cholangitis (PBC)
Discussion
In the current study, we mapped a comprehensive single-cell atlas, demonstrating cell heterogeneity and immune networks during the early phase of AILI. The characteristics of cholangiocytes, LSECs, T-cells, macrophages, and Kupffer cells were identified. Furthermore, we found that Mkrn1 may serve as a macrophage-specific indicator of AILI.
Our study provides a new APAP metabolic profile at single-cell resolution. In addition to hepatocytes, NPCs may also be directly involved in the metabolism of APAP, especially ECs, cholangiocytes, SMCs, and HepSCs. At the onset of AILI, genes related to cell proliferation and regeneration in cholangiocytes, such as Spp1 and Sox9, were upregulated. Secreted phosphoprotein 1 (Spp1) encodes osteopontin (OPN), which is a glycoprotein involved in tumor cell survival, epidermal homeostasis, and keratinocyte differentiation (Wei et al. 2019). OPN has been identified as a novel serum biomarker of DILI in clinical practice (Church et al. 2019). Interestingly, we found that Dmbt1+ cholangiocytes drastically decreased after APAP treatment, which was validated by immunofluorescence staining. These cells are involved in detoxification and CYP450 drug metabolism. Therefore, it may be possible that Dmbt1+ cholangiocytes are the direct target cells of APAP overdose attack; however, the mechanism needs to be explored in further studies. In addition, we found a cluster of epithelial progenitors (highly expressed Tm4sf4, Epcam, and Krt19 as marker genes) in acute AILI. Interestingly, the epithelial progenitors tended to evolve toward a cholangiocyte-like phenotype rather than toward a hepatocyte-like phenotype upon APAP challenge, especially in the 750 mg/kg APAP-injured liver (Supplementary Fig. 7).
We revealed that Klf2 and Klf4 were highly upregulated in LSECs in the livers of AILI mice. Klf2 and Klf4 are zinc finger transcription factors that inhibit vascular endothelial growth factor (VEGF)-induced angiogenesis (Black et al. 2001). Several studies have reported that Klf2 and Klf4 exhibit anti-inflammatory and anti-vascular remodeling effects in cirrhotic portal hypertension and atherosclerosis (Marrone et al. 2015; Niu et al. 2019). Thus, further enhancement of Klf2 or Klf4 transcription may contribute to LSEC dysfunction. The contraction of fenestrae is regulated by the calcium-calmodulin-actin system (Yokomori et al. 2003). In the present study, we found that DEGs were enriched for calcium transport and homeostasis in LSECs. Polymerized actin microfilaments induced by increased calcium levels may lead to fenestrae contraction and capillarization in LSECs. Furthermore, we found that NK cells might play the main role in cell killing in the early stage of AILI, which is consistent with a previous report (Liu et al. 2004). In addition, T-cells may further activate other cells by producing diverse cytokines, such as IL-17, IL-6, TNF-α, and IFN-γ, which participate in the pathogenesis of AILI.
Liver macrophages play a dual role in AILI (Yang et al. 2022). It was reported that pre-ablation of rat KCs demonstrated a protective effect in AILI (Holt et al. 2010). However, macrophages are also necessary for tissue repair and angiogenesis after AILI (Starkey Lewis et al. 2020). Here, for the first time, we used single-cell sequencing to profile liver macrophages, Kupffer cells, and their sub-clusters in the early stages of AILI. Macrophages were characterized by the high expression of Pf4 (Supplementary Fig. 5). Pf4/Cxcl4 is involved in heparin neutralization, chemotaxis, and angiogenesis inhibition (Warkentin. 2022). In our study, macrophage cluster 1 displayed an M1 polarization phenotype, whereas cluster 3 tended to exhibit M2 polarization. Macrophage clusters 1 and 2 may be involved in the initiation of innate immunity through cytokine release, chemotaxis, and inflammatory pathway activation in acute AILI. KCs are highly effective in expressing Cxcl2, which may further induce the entry of peripheral monocytes and neutrophils into the liver. Enrichment analysis revealed that ubiquitin protein ligase binding is involved in the pathogenesis of AILI in KCs.
A distinct advantage of single-cell analysis is its unprecedented accuracy in delineating the cellular origins of secreted ligands and their potential target cells. Inflammatory interaction pairs such as the TGFB3-TGFBR3 and IL-1B-ADRB2 signaling axis were observed between macrophages/KCs and other NPCs. Furthermore, we found that KCs could communicate with CD8Teff and NK cells through CCL24-CCR2, which may exacerbate liver injury. We also demonstrated that the autocrine LIFR-OSM signaling axis promoted M1 polarization through the activation of the MAPK signaling pathway in macrophages.
Mkrn1 is an E3 ubiquitin ligase that catalyzes the covalent attachment of ubiquitin moieties to substrate proteins (Kim et al. 2005). In general, Mkrn1 keeps cells alive by suppressing p53/TP53 under normal conditions but stimulates apoptosis by repressing Cyclin-dependent kinase inhibitor 1A (CDKN1A) under stress conditions (Lee et al. 2009). Studies have found that Mkrn1 affects glucose metabolism and adipose tissue by mediating the ubiquitination of AMP-activated protein kinase (Lee et al. 2018). We found that Mkrn1 was highly expressed in the liver macrophages of AILI mice and patients with AILI, suggesting that Mkrn1 may be a potential macrophage-specific indicator of AILI. Interestingly, compared to other liver disease conditions, Mkrn1 seems to be specifically expressed in the liver macrophages of AILI. However, the mechanism underlying the high expression of Mkrn1 and its exact role in AILI require further systematic experimental validation.
A limitation of this study is that all of the data were generated from an AILI mouse model, as fresh human AILI samples are very difficult to obtain. In addition, due to the limitations of current technology and the specificity of the acute liver injury model, it is difficult to obtain sufficient amounts of hepatocytes and NPCs at the same time. Thus, we were unable to analyze the interaction between hepatocytes and NPCs at the onset of AILI.
In conclusion, this is the first report to describe the transcriptional heterogeneity of NPCs and the immune environment during the early phase of AILI. By identifying multiple cell subtypes, highlighting a catalog of altered genes and pathways, and cell–cell interactions, we have provided a comprehensive atlas of the NPCs and immune network in the livers of AILI mice. Furthermore, we demonstrated that Mkrn1 may serve as a macrophage-specific biomarker of AILI. We believe that this study contributes to the further understanding of AILI pathogenesis and provides a rich resource for identifying biomarkers from NPCs in AILI.
Data availability
Data and materials used and/or analyzed during the current study are available from the corresponding authors on reasonable request.
Abbreviations
- NPCs:
-
Non-parenchymal cells
- APAP:
-
N-acetyl-p-aminophenol, acetaminophen
- AILI:
-
Acetaminophen-induced liver injury
- scRNA-seq:
-
Single-cell RNA sequencing
- Mkrn1:
-
Makorin ring finger protein 1
- Dmbt1:
-
Deleted in malignant brain tumor 1
- KCs:
-
Kupffer cells
- DILI:
-
Drug-induced liver injury
- NAPQI:
-
N-acetyl-p-benzoquinone imine
- LSECs:
-
Liver sinusoidal endothelial cells
- HSCs:
-
Hepatic stellate cells
- ALT:
-
Alanine aminotransferase
- AST:
-
Aspartate aminotransferase
- DAMPs:
-
Damage-associated molecular patterns
- CCR2:
-
Chemokine (C–C motif) receptor 2
- APCs:
-
Antigen-presenting cells
- NASH:
-
Non-alcoholic steatohepatitis
- UMI:
-
Unique molecular identifiers
- DEGs:
-
Differentially expressed genes
- GO:
-
Gene ontology
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- HILI:
-
Herbal-induced liver injury
- NAFLD:
-
Non-alcoholic fatty liver disease
- CHB:
-
Chronic hepatitis B
- PBC:
-
Primary biliary cholangitis
- OSM:
-
Oncostatin M
- TUNEL:
-
Terminal dUTP nick‑end labeling
- ECs:
-
Endothelial cells
- pDCs:
-
Plasmacytoid dendritic cells
- Mps:
-
Mononuclear phagocytes
- SMCs:
-
Smooth muscle cells
- HepSCs:
-
Hepatic stellate cells
- UMAP:
-
Uniform manifold approximation and projection
- LIFR:
-
Leukemia inhibitory factor receptor
- MAPK:
-
Mitogen-activated protein kinase
- Spp1:
-
Secreted phosphoprotein 1
- OPN:
-
Osteopontin; VEGF, vascular endothelial growth factor
- CDKN1A:
-
Cyclin-dependent kinase inhibitor 1A
- LDH:
-
Lactate dehydrogenase
- Epcs:
-
Epithelial progenitors
- GSVA:
-
Gene set variation analysis
- ALP:
-
Alkaline phosphatase
- GGT:
-
γ-Glutamyl transferase
- TBIL:
-
Total bilirubin
References
Aithal GP, Grove JI (2015) Genome-wide association studies in drug-induced liver injury: step change in understanding the pathogenesis. Semin Liver Dis 35(4):421–431. https://doi.org/10.1055/s-0035-1567829
Aizarani N, Saviano A, Sagar et al (2019) A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature 572(7768):199–204. https://doi.org/10.1038/s41586-019-1373-2
Alvaro D, Mancino MG, Glaser S et al (2007) Proliferating cholangiocytes: a neuroendocrine compartment in the diseased liver. Gastroenterology 132(1):415–431. https://doi.org/10.1053/j.gastro.2006.07.023
Andrade RJ, Chalasani N, Björnsson ES et al (2019) Drug-induced liver injury. Nat Rev Dis Primers 5(1):58. https://doi.org/10.1038/s41572-019-0105-0
Antoniades CG, Quaglia A, Taams LS et al (2012) Source and characterization of hepatic macrophages in acetaminophen-induced acute liver failure in humans. Hepatology 56(2):735–746. https://doi.org/10.1002/hep.25657
Ben-Moshe S, Veg T, Manco R et al (2022) The spatiotemporal program of zonal liver regeneration following acute injury. Cell Stem Cell 29(6):973–989. https://doi.org/10.1016/j.stem.2022.04.008
Black AR, Black JD, Azizkhan-Clifford J (2001) Sp1 and krüppel-like factor family of transcription factors in cell growth regulation and cancer. J Cell Physiol 188(2):143–160. https://doi.org/10.1002/jcp.1111
Butler A, Hoffman P, Smibert P, Papalexi E, Satija R (2018) Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol 36(5):411–420. https://doi.org/10.1038/nbt.4096
Church RJ, Kullak-Ublick GA, Aubrecht J et al (2019) Candidate biomarkers for the diagnosis and prognosis of drug-induced liver injury: an international collaborative effort. Hepatology 69(2):760–773. https://doi.org/10.1002/hep.29802
Dura B, Choi J-Y, Zhang K et al (2019) scFTD-seq: freeze-thaw lysis based, portable approach toward highly distributed single-cell 3’ mRNA profiling. Nucleic Acids Res 47(3):e16. https://doi.org/10.1093/nar/gky1173
Friedman SL (2008) Hepatic stellate cells: protean, multifunctional, and enigmatic cells of the liver. Physiol Rev 88(1):125–172. https://doi.org/10.1152/physrev.00013.2007
Ghanem CI, Pérez MJ, Manautou JE, Mottino AD (2016) Acetaminophen from liver to brain: new insights into drug pharmacological action and toxicity. Pharmacol Res 109:119–131. https://doi.org/10.1016/j.phrs.2016.02.020
Holt MP, Yin H, Ju C (2010) Exacerbation of acetaminophen-induced disturbances of liver sinusoidal endothelial cells in the absence of Kupffer cells in mice. Toxicol Lett 194(1–2):34–41. https://doi.org/10.1016/j.toxlet.2010.01.020
Hoofnagle JH, Björnsson ES (2019) Drug-induced liver injury - types and phenotypes. N Engl J Med 381(3):264–273. https://doi.org/10.1056/NEJMra1816149
Hwang B, Lee JH, Bang D (2018) Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp Mol Med 50(8):1–4. https://doi.org/10.1038/s12276-018-0071-8
Kim JH, Park S-M, Kang MR et al (2005) Ubiquitin ligase MKRN1 modulates telomere length homeostasis through a proteolysis of hTERT. Genes Dev 19(7):776–781. https://doi.org/10.1101/gad.1289405
Krenkel O, Tacke F (2017) Liver macrophages in tissue homeostasis and disease. Nat Rev Immunol 17(5):306–321. https://doi.org/10.1038/nri.2017.11
Lantieri F, Bachetti T (2022) OSM/OSMR and interleukin 6 family cytokines in physiological and pathological condition. Int J Mol Sci 23(19):11096. https://doi.org/10.3390/ijms231911096
Lee E-W, Lee M-S, Camus S et al (2009) Differential regulation of p53 and p21 by MKRN1 E3 ligase controls cell cycle arrest and apoptosis. EMBO J 28(14):2100–2113. https://doi.org/10.1038/emboj.2009.164
Lee M-S, Han H-J, Han SY et al (2018) Loss of the E3 ubiquitin ligase MKRN1 represses diet-induced metabolic syndrome through AMPK activation. Nat Commun 9(1):3404. https://doi.org/10.1038/s41467-018-05721-4
Lei X, Xu Q, Li C et al (2022) Egr1 confers protection against acetaminophen-induced hepatotoxicity via transcriptional upregulating of Acaa2. Int J Biol Sci 18(9):3800–3817. https://doi.org/10.7150/ijbs.71781
Li L, Zeng Z (2020) Live imaging of innate and adaptive immune responses in the liver. Front Immunol 11:564768. https://doi.org/10.3389/fimmu.2020.564768
Li C, Ming Y, Hong W et al (2018) Comparison of hepatic transcriptome profiling between acute liver injury and acute liver failure induced by acetaminophen in mice. Toxicol Lett 283:69–76. https://doi.org/10.1016/j.toxlet.2017.11.020
Li X, Tang J, Mao Y (2022) Incidence and risk factors of drug-induced liver injury. Liver Int 42(9):1999–2014. https://doi.org/10.1111/liv.15262
Liao Y, Smyth GK, Shi W (2014) FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30(7):923–930. https://doi.org/10.1093/bioinformatics/btt656
Liu Z-X, Govindarajan S, Kaplowitz N (2004) Innate immune system plays a critical role in determining the progression and severity of acetaminophen hepatotoxicity. Gastroenterology 127(6):1760–1774. https://doi.org/10.1053/j.gastro.2004.08.053
MacParland SA, Liu JC, Ma X-Z et al (2018) Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat Commun 9(1):4383. https://doi.org/10.1038/s41467-018-06318-7
Marrone G, Maeso-Díaz R, García-Cardena G et al (2015) KLF2 exerts antifibrotic and vasoprotective effects in cirrhotic rat livers: behind the molecular mechanisms of statins. Gut 64(9):1434–1443. https://doi.org/10.1136/gutjnl-2014-308338
Massalha H, Bahar Halpern K, Abu-Gazala S et al (2020) A single cell atlas of the human liver tumor microenvironment. Mol Syst Biol 16(12):9682. https://doi.org/10.15252/msb.20209682
Mossanen JC, Krenkel O, Ergen C et al (2016) Chemokine (C-C motif) receptor 2-positive monocytes aggravate the early phase of acetaminophen-induced acute liver injury. Hepatology 64(5):1667–1682. https://doi.org/10.1002/hep.28682
Nelson DR, Zeldin DC, Hoffman SMG et al (2004) Comparison of cytochrome P450 (CYP) genes from the mouse and human genomes, including nomenclature recommendations for genes, pseudogenes and alternative-splice variants. Pharmacogenetics 14(1):1–18. https://doi.org/10.1097/00008571-200401000-00001
Niu N, Xu S, Xu Y, Little PJ, Jin Z-G (2019) Targeting mechanosensitive transcription factors in atherosclerosis. Trends Pharmacol Sci 40(4):253–266. https://doi.org/10.1016/j.tips.2019.02.004
Poisson J, Lemoinne S, Boulanger C et al (2017) Liver sinusoidal endothelial cells: physiology and role in liver diseases. J Hepatol 66(1):212–227. https://doi.org/10.1016/j.jhep.2016.07.009
Qiu X, Hill A, Packer J et al (2017) Single-cell mRNA quantification and differential analysis with census. Nat Methods 14(3):309–315. https://doi.org/10.1038/nmeth.4150
Ramachandran P, Dobie R, Wilson-Kanamori JR et al (2019) Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575(7783):512–518. https://doi.org/10.1038/s41586-019-1631-3
Satija R, Farrell JA, Gennert D, Schier AF, Regev A (2015) Spatial reconstruction of single-cell gene expression data. Nat Biotechnol 33(5):495–502. https://doi.org/10.1038/nbt.3192
Shen T, Liu Y, Shang J et al (2019) Incidence and etiology of drug-induced liver injury in Mainland China. Gastroenterology. https://doi.org/10.1053/j.gastro.2019.02.002
Starkey Lewis P, Campana L, Aleksieva N et al (2020) Alternatively activated macrophages promote resolution of necrosis following acute liver injury. J Hepatol 73(2):349–360. https://doi.org/10.1016/j.jhep.2020.02.031
Stephens C, Andrade RJ, Lucena MI (2014) Mechanisms of drug-induced liver injury. Curr Opin Allergy Clin Immunol 14(4):286–292. https://doi.org/10.1097/ACI.0000000000000070
Terkelsen MK, Bendixen SM, Hansen D et al (2020) Transcriptional dynamics of hepatic sinusoid-associated cells after liver injury. Hepatology 72(6):2119–2133. https://doi.org/10.1002/hep.31215
Warkentin TE (2022) Platelet-activating anti-PF4 disorders: an overview. Semin Hematol 59(2):59–71. https://doi.org/10.1053/j.seminhematol.2022.02.005
Wei J, Marisetty A, Schrand B et al (2019) Osteopontin mediates glioblastoma-associated macrophage infiltration and is a potential therapeutic target. J Clin Invest 129(1):137–149. https://doi.org/10.1172/JCI121266
Xie Y, McGill MR, Dorko K et al (2014) Mechanisms of acetaminophen-induced cell death in primary human hepatocytes. Toxicol Appl Pharmacol 279(3):266–274. https://doi.org/10.1016/j.taap.2014.05.010
Xiong X, Kuang H, Ansari S et al (2019) Landscape of intercellular crosstalk in healthy and NASH liver revealed by single-cell secretome gene analysis. Mol Cell. https://doi.org/10.1016/j.molcel.2019.07.028
Yan M, Huo Y, Yin S, Hu H (2018) Mechanisms of acetaminophen-induced liver injury and its implications for therapeutic interventions. Redox Biol 17:274–283. https://doi.org/10.1016/j.redox.2018.04.019
Yang W, He H, Wang T et al (2021) Single-cell transcriptomic analysis reveals a hepatic stellate cell-activation roadmap and myofibroblast origin during liver fibrosis in mice. Hepatology 74(5):2774–2790. https://doi.org/10.1002/hep.31987
Yang T, Wang H, Wang X, Li J, Jiang L (2022) The dual role of innate immune response in acetaminophen-induced liver injury. Biology (Basel) 11(7):1057. https://doi.org/10.3390/biology11071057
Yokomori H, Oda M, Ogi M et al (2003) Endothelin-1 suppresses plasma membrane Ca++-ATPase, concomitant with contraction of hepatic sinusoidal endothelial fenestrae. Am J Pathol 162(2):557–566. https://doi.org/10.1016/S0002-9440(10)63849-7
Yu G, Wang L-G, Han Y, He Q-Y (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16(5):284–287. https://doi.org/10.1089/omi.2011.0118
Yuan L, Kaplowitz N (2013) Mechanisms of drug-induced liver injury. Clin Liver Dis 17(4):507–518. https://doi.org/10.1016/j.cld.2013.07.002
Zhang Y, Liu T, Hu X et al (2021) Cell Call: integrating paired ligand-receptor and transcription factor activities for cell-cell communication. Nucleic Acids Res 49(15):8520–8534. https://doi.org/10.1093/nar/gkab638
Acknowledgements
This work was supported by the National Natural Science Foundation of China (NSFC 81970513, NSFC 82270619, and NSFC 32171175), National Key R&D Program of China (2022YFC3502101), Clinical Research Innovation and Development Fund of Renji Hospital, School of Medicine, Shanghai Jiao Tong University (PY120-05).
Funding
This work was supported by the National Natural Science Foundation of China (NSFC 81970513, NSFC 82270619, and NSFC 32171175), the National Key R&D Program of China (2022YFC3502101), and the Clinical Research Innovation and Development Fund of Renji Hospital, School of Medicine, Shanghai Jiao Tong University (PY120-05).
Author information
Authors and Affiliations
Contributions
YXL, YZ, JL, and BXL performed the experiment and wrote and drafted the manuscript. HXL, YJ, TYZ, and FYZ retrieved the data and performed the statistical analysis. MXK, FX, WZ, and YXC performed the bioinformatics analysis. TJT, BXL, and MYM acquired funding. BXL and MYM revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
All animal experiments were approved by the Animal Ethics Committee of Shanghai Jiao Tong University Renji Hospital. Animal research was conducted in accordance with international guidelines. The clinical research was approved by the Ethics Committee of Shanghai Jiaotong University Renji Hospital in accordance with the ethical guidelines of the 1975 Declaration of Helsinki.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, X., Zhi, Y., Li, J. et al. Single-cell RNA sequencing to reveal non-parenchymal cell heterogeneity and immune network of acetaminophen-induced liver injury in mice. Arch Toxicol 97, 1979–1995 (2023). https://doi.org/10.1007/s00204-023-03513-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-023-03513-4