Abstract
Preventing clinical drug-induced liver injury (DILI) remains a major challenge, because DILI develops via multifactorial mechanisms. Immune and inflammatory reactions are considered important mechanisms of DILI; however, biomarkers from in vitro systems using immune cells have not been comprehensively studied. The aims of this study were (1) to identify promising biomarker genes for predicting DILI in an in vitro coculture model of peripheral blood mononuclear cells (PBMCs) with a human liver cell line, and (2) to evaluate these genes as predictors of DILI using a panel of drugs with different clinical DILI risk. Transcriptome-wide analysis of PBMCs cocultured with HepG2 or differentiated HepaRG cells that were treated with several drugs revealed an appropriate separation of DILI-positive and DILI-negative drugs, from which 12 putative biomarker genes were selected. To evaluate the predictive performance of these genes, PBMCs cocultured with HepG2 cells were exposed to 77 different drugs, and gene expression levels in PBMCs were determined. The MET proto-oncogene receptor tyrosine kinase (MET) showed the highest area under the receiver-operating characteristic curve (AUC) value of 0.81 among the 12 genes with a high sensitivity/specificity (85/66%). However, a stepwise logistic regression model using the 12 identified genes showed the highest AUC value of 0.94 with a high sensitivity/specificity (93/86%). Taken together, we established a coculture system using PBMCs and HepG2 cells and selected biomarkers that can predict DILI risk. The established model would be useful in detecting the DILI potential of compounds, in particular those that involve an immune mechanism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Drug-induced liver injury (DILI) is an adverse drug reaction that presents with hepatitis, cholestasis, jaundice, steatosis, nodular regenerative hyperplasia, or sinusoidal obstruction syndrome (Kleiner 2017). DILI is a frequent cause of acute liver failure, accounting for 20–40% of all cases of hepatic failure (Bell and Chalasani 2009; Björnsson 2014; Ostapowicz et al. 2002). While hepatotoxicity due to acetaminophen overdose is the most common, that is predictable and dose-related, liver injury associated with several other drugs appears to be of an idiosyncratic nature with no explicit dose–toxicity relationship, and is largely host-dependent (Funk and Roth 2017). Although idiosyncratic DILI is relatively rare with an incidence of 1 in 1000–100,000 treated patients (de Abajo et al. 2004), its occurrence has a significant impact on the pharmaceutical industry and regulatory agencies, because it can be a reason for discontinuation of clinical trials and postmarketing regulatory actions, including withdrawal from the market and labeling restrictions (Kaplowitz 2005). A survey reported that 462 drugs were withdrawn from the worldwide market between 1953 and 2013, and hepatotoxicity (81 drugs; 18%) was the most common adverse drug reaction that led to withdrawal, followed by immune-related reactions (79 drugs; 17%) (Onakpoya et al. 2016). For example, alatrofloxacin, bromfenac, troglitazone, and trovafloxacin were all approved by the U.S. Food and Drug Administration (USFDA) in 1997, but later withdrawn from the market because of severe liver toxicity or liver failure (Senior 2014). Therefore, it is one of the most important challenges to predict the toxicity of candidate compounds at the early nonclinical stage of drug development.
It is believed that DILI develops via multifactorial mechanisms involving both drug properties (dose, lipophilicity, reactive metabolites, mitochondrial liability, enzyme, and transporter inhibition) and host-specific factors (genetic variation, age, sex, microbiota, diet, and polypharmacy) (Weaver et al. 2020). Therefore, these risk factors can be applied to develop test platforms to evaluate the potential DILI risk of a compound. To date, researchers in the pharmaceutical industry, academia, and regulatory agencies have made great efforts in establishing DILI platforms for the detection of the toxic potential of compounds. Cell death, ATP, reactive oxygen species (ROS), mitochondrial membrane potential, bile salt export pump (BSEP) inhibition, and covalent binding in vitro have been examined using liver cell lines and primary hepatocytes. However, the problem is that these approaches largely identify the intrinsic toxicity of drugs, that are usually detected in experimental animals, and may not predict compounds at risk for idiosyncratic reactions.
Accumulating in vivo experimental evidence suggests that innate immunity and adaptive immunity and their interaction with drugs are important mechanisms of idiosyncratic DILI (Liguori et al. 2010; Song et al. 2019; Uetrecht 2019). These achievements suggest the possibility of in vitro platforms that incorporate immune factors for DILI risk assessment. A few such studies have been conducted by adapting coculture or media transfer methods using primary cells or cell lines of human hepatocytes and monocytic cells (Edling et al. 2009; Granitzny et al. 2017; Kim et al. 2017). We recently reported a cell-based assay in which human promyelocytic leukemia HL-60 cells were treated with conditioned media of HepG2 or HepaRG cells that were exposed to drugs (Oda et al. 2016). The study found that the expression levels of S100A9, IL-1B, and CXCL8 (IL-8) mRNA in HL-60 cells may be useful in predicting DILI potential to some extent. However, there is room for improvement to establish a more physiologically relevant assay.
In this study, we established an in vitro model by coculturing PBMCs as an immune cell source and hepatocellular carcinoma-derived HepG2 cells to predict the DILI potential of drugs. First, the transcriptome in PBMCs of the coculture that had been exposed to several DILI-positive and DILI-negative drugs was obtained to select candidate biomarker genes. Then, the identified genes were further evaluated for their predictivity of DILI using 77 drugs with different DILI potential.
Materials and methods
Test drugs
The supplier and the catalog number of the test drugs are provided in Table S1 and Table S2. A total of 10 and 77 drugs were examined by microarray and screening analyses, respectively. Information on the clinical DILI risk of the test drugs was collected from the USFDA-approved prescription drug labels cited in DailyMed (https://dailymed.nlm.nih.gov/dailymed/index.cfm) and the Liver Toxicity Knowledge Base Benchmark Dataset (LTKB-BD; https://www.fda.gov/ScienceResearch/BioinformaticsTools). USFDA-approved prescription drug labels classify DILI risk into the following four categories: boxed warning (BW), warnings and precautions (WP), adverse reactions (AR), or no mention (NM). Drug labeling balances the information of causality, incidence, and severity based on data from controlled trials, published literature reports, and spontaneous reports to AERS (adverse event reporting systems) (CDER 2006). LTKB-BD classifies DILI risks into four categories, namely, Most-DILI, Less-DILI, Ambiguous-DILI, and No-DILI concern, by considering the clinical severity of DILI and labeling approved by the USFDA (Chen et al. 2011). Specifically, a drug is classified into Most-DILI concern if the drug was withdrawn (WDN) from the market, had a BW-label regarding liver injury, or caused severe and moderate liver injury (fatal hepatotoxicity, acute liver failure, liver necrosis, jaundice, and hyperbilirubinemia) described in the WP section. A drug is classified into Less-DILI concern if it was associated with mild DILI as described in the WP and AR sections of the drug label. A No-DILI-concern drug had no mention regarding liver injury in its drug label. Among the drugs classified as Most-DILI and Less-DILI-concern previously, the drugs were recently reclassified into Ambiguous-DILI concern if the drugs were not verified as DILI cause (Chen et al. 2016). The classifications are shown as the “Drug label” and “LTKB annotation” in Table 1 and Tables S1–2. In this study, Most-DILI-concern drugs or withdrawn drugs were considered DILI-positive drugs, and Less-, No-, and Ambiguous-DILI-concern drugs were considered DILI-negative drugs (“DILI classification” in Table 1). Although rofecoxib was withdrawn from the market, due to cardiovascular events, it was considered a DILI-negative drug.
All drugs, except amikacin sulfate, chloroquine diphosphate, and valproate sodium, were dissolved in 100% dimethyl sulfoxide (DMSO, Sigma-Aldrich, St. Louis, MO) as a stock solution. Amikacin sulfate, chloroquine diphosphate, and valproate sodium were dissolved in distilled water.
Peripheral blood mononuclear cells
Blood was collected from a healthy male donor aged 63 years into Vacutainer® CPT™ 8-mL tubes containing sodium citrate (Becton Dickinson Vacutainer Systems, Franklin Lakes, NJ) and centrifuged at 1800×g at room temperature for 20 min. The PBMC layer was transferred into a 50-mL tube and washed with 1 × PBS twice. Freshly isolated PBMCs were used for a coculture experiment with HepG2 or HepaRG cells for microarray analysis as described below. Informed consent for participation in this study was obtained from the donor, and all protocols were designed in accordance with the Declaration of Helsinki and approved by the ethics committee of Nagoya University School of Medicine.
Cryopreserved PBMCs were purchased from Cellular Technology Limited (Cleveland, OH). Cryopreserved PBMCs were thawed according to the manufacturer’s protocols. Briefly, cryopreserved PBMCs were thawed in RPMI 1640 medium (Nissui Pharmaceutical) supplemented with 2-mM L-glutamine and CTL Anti-Aggregate Wash™ solution (Cellular Technology Limited) and centrifuged at 330×g at room temperature for 20 min. The cells were then switched to CTL-Test™ medium (Cellular Technology Limited) supplemented with 2 mM L-glutamine and centrifuged at 330×g at room temperature for 20 min. The cryopreserved PBMCs were used for a coculture experiment with HepG2 for screening assays as described below. All cells were cultured in a humidified atmosphere of air containing 5% CO2 at 37 °C.
Liver cell line
The human hepatocellular carcinoma cell line HepG2 was purchased from the American Type Culture Collection (Manassas, VA). HepG2 cells were grown in Dulbecco’s modified Eagle’s medium (Nissui Pharmaceutical, Tokyo, Japan) containing 0.1-mM nonessential amino acids, 10% fetal bovine serum (FBS; Sigma-Aldrich), and 2-mM l-glutamine. The medium was renewed every 2 days.
Cryopreserved human hepatoma HepaRG® cells (HRP116), which are terminally differentiated hepatic cells, were purchased from BioPredic International (Rennes, France). The cryopreserved differentiated HepaRG cells were seeded on a 24-well collagen-coated plate at a density of 4.8 × 105 cells/well in Medium 670 (HepaRG Thawing/Plating/General purpose Medium; BioPredic International), and the media were shifted to Medium 620 (HepaRG Maintenance and Metabolism Medium; BioPredic International) 1 day after seeding. The cells were used 7 days after seeding for a coculture experiment with PBMCs.
Assessment of PBMC and HepG2 cell viability
Freshly isolated PBMCs (a healthy male donor aged 63 years) and commercial cryopreserved PBMCs (LP226; a healthy female donor, 55 years old, Cellular Technology Limited) were seeded onto a 96-well plate at 5.0 × 105 cells/well and treated with drugs at various concentrations corresponding to 100-, 50-, 25-, 10-, 5-, 2.5- or 1-fold ( ×) human plasma maximum concentration (Cmax). The final DMSO concentration was 0.4%. After a 24-h incubation period, Cell Counting Kit-8 (Dojindo molecular technologies, Kumamoto, Japan) assays for the freshly isolated PBMCs and CellTiter-Glo®2.0 (Promega, Madison, WI) assays for the commercial cryopreserved PBMCs were conducted to determine cell viability. The maximum drug concentration that yielded cell viability of more than 70% was used for the coculture experiments of microarray and screening as described below. In addition, a drug concentration producing a 50% decrease in cell viability (IC50) was determined from a plot of the concentration of the test drugs (X) versus the percentage (Y) of cell viabilities relative to the control by nonlinear regression analysis according to the following Hill equation with GraphPad Prism 8 software (Version 8.3.0, GraphPad Software, San Diego, CA):
IC50 values were expressed as the fold-over Cmax. For compounds for which IC50 values could not be calculated correctly due to low toxic effects to the cells, 100 × Cmax was assigned as their IC50 values.
HepG2 cells were seeded onto a 96-well plate at 2.5 × 104 cells/well and treated with drugs at a single concentration for each drug. The tested concentration (Table 1) was the maximum drug concentration that yielded PBMC viability of more than 70% determined above. The final DMSO concentration was 0.4%. After a 24-h incubation period, CellTiter-Glo®2.0 assays were conducted to determine cell viability.
Drug treatment for coculture and monoculture of PBMCs and HepG2 or HepaRG cells
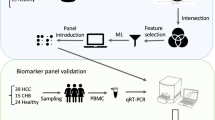
HepG2 (2.5 × 105 cells/well) and HepaRG (4.8 × 105 cells/well) cells were seeded onto a 24-well plate using 0.5 mL of the corresponding normal growth media as described above. After 1 day (HepG2) or 7 days (HepaRG), media were removed, and 0.75 mL of CTL-Test Medium containing test drugs was added. Immediately, Falcon® culture inserts (0.4-µm pore size; catalog number: 353095, Corning; Corning, NY) were placed in each well, and PBMCs were seeded into the inserts at 5.0 × 105 cells/well suspended in 0.125 mL of CTL-Test Medium. Then, 0.125 mL of CTL-Test Medium containing test drugs (at twofold of the final concentration) was added to the inserts. The drug concentration was set at a concentration for each drug based on PBMC viability. After 24 h of incubation (5% CO2 at 37 °C), PBMCs were collected and used for RNA extraction. These procedures are illustrated in Fig. 1.
Illustration of the research flow. Human PBMCs were cocultured with HepG2 or HepaRG cells using culture inserts. PBMCs were seeded into culture inserts and placed on 24-well plates containing HepG2 or HepaRG cells, and the tested drugs were added to both parts. For microarray analysis, the coculture was treated with 10 drugs, and RNA from PBMCs was extracted. For qPCR analysis, the coculture of PBMCs and HepG2 cells was treated with 77 drugs, and RNA from PBMCs was extracted
For microarray analysis, both HepG2 and HepaRG cells were used as liver cells, and freshly isolated PBMCs (a healthy male donor, 63 years old) were used as test cells. For these experiments, test drugs included amodiaquine, diclofenac sodium, ketoconazole, levofloxacin, tienilic acid, and trovafloxacin mesylate (DILI-positive) and chloroquine phosphate, ethacrinic acid, ketorolac tromethamine, and mebendazole (DILI-negative) as well as 0.4% DMSO as a control (Table S1).
For screening assays, only HepG2 cells were used as liver cells, commercial cryopreserved PBMCs (lot: LP226; a healthy female donor, 55 years old) were used, and 77 drugs were examined (Table S2) followed by qPCR analysis. To investigate donor-to-donor differences in predictive performance, cryopreserved PBMCs from another donor (lot: LP316; a healthy male donor, 25 years old, Cellular Technology Limited) were used, and 77 drugs were examined followed by the qPCR analysis.
For monoculture of PBMCs, commercial cryopreserved PBMCs (LP226; a healthy female donor, 55 years old) were cultured in the absence of liver cells using culture inserts as described above, and 77 drugs were examined followed by qPCR analysis at a concentration shown in Table S2.
Microarray analysis
For microarray analysis, total RNA was extracted from freshly isolated PBMCs, which were cocultured with HepG2 or HepaRG cells, using an RNeasy Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. The concentration and integrity of total RNA were determined with a 2100 Bio Analyzer (Agilent Technologies, Santa Clara, CA), and the RNA integrity number (RIN) was confirmed to be greater than 7, that is of sufficient quality for use in microarray experiments. Samples were analyzed by a microarray platform of Sure Print G3 Human 8 × 60 K ver.3.0 (Agilent Technology). The data were analyzed using Subio Platform ver. 1.21 (Subio, Amami, Japan). Raw signal intensities were normalized by a 75th percentile shift for either PBMCs/HepG2 or PBMCs/HepaRG groups, and fold changes in the drug-treated samples against each DMSO-treated group were calculated. Principal component analysis (PCA) of the expression levels of all genes was conducted using the “prcomp” function of R software version 3.6.0 (R Core Team 2019). Hierarchical clustering analysis of 12 predictor genes was performed by Euclidean distance and the “ward. D2” clustering method of the R software. A heatmap was generated using the “heatmap.2” function of the R package “gplots.” Microarray data were deposited in the Gene Expression Omnibus database (https://www.ncbi.nlm.nih.gov) under accession number GSE147866.
RNA extraction and real-time reverse transcription (RT)-PCR
For qPCR analysis of RNA samples from screening assays, total RNA was extracted from the PBMCs using RNAiso Plus (Takara Bio, Kusatsu, Japan) according to the manufacturer’s instructions. Five hundred nanograms of total RNA were reverse transcribed to cDNA using a ReverTra Ace® qPCR RT kit (Toyobo, Osaka, Japan) according to the manufacturer’s instructions. Quantitative real-time RT-PCR was performed with total volumes of 10 μL containing 0.5 μL of cDNA, 5 μL of 2 × TB Green® Premix Ex Taq (Tli RNaseH Plus, Takara Bio, Kusatsu, Japan) solution, 0.2 μL of 50-fold ROX reference dye, and 4 pmol each of the forward and reverse primers. Sequences of the primers that were commercially synthesized (Hokkaido System Sciences, Sapporo, Japan) are shown in Table S3. Real-time PCR was performed using an Mx3000P instrument (Agilent Technologies) with an initial denaturation at 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s and 60 °C for 30 s. Each assay included 1:10 serial dilution points (104–109-fold) of PCR amplicons of each gene, and a calibration curve was constructed by plotting the PCR threshold cycle (Ct) value against the log10 (template dilution value). The relative mRNA expression levels in experimental samples were calculated from the standard curve and normalized to GAPDH mRNA levels.
Statistical analysis
A comparison of the two groups was performed with an unpaired Student’s t test. Correlation analyses were performed using Pearson’s test.
Logistic regression analyses were performed for each biomarker individually and in combination with simple logistic model and multiple logistic model, respectively, to evaluate the discriminatory ability of biomarkers between DILI-positive and DILI-negative drugs. In the evaluation of each biomarker, we used the simple logistic regression model with the mRNA expression level \({{x}_{g}}\) of each biomarker \(g\) and the probability of DILI-positive drug \({P}_{g}\) for each biomarker \(g\):
In case of evaluating the combination of biomarkers, the multiple logistic regression model was used with the probability of DILI-positive drug \(P\):
The best combination of biomarkers was selected by minimizing the Akaike information criterion (AIC) through a backward stepwise selection. For each logistic regression model, receiver-operating characteristic (ROC) curves were drawn by plotting the true positive fraction (sensitivity) as a function of the false-positive fraction (1-specificity) for all possible thresholds. The optimal cut-off values were determined according to the maximum value of Youden’s index [J]\(([J] = \mathrm{ Sensitivity} + \mathrm{Specificity} -1\)). The overall discriminatory performance was estimated based on the area under the ROC curve (AUC). Sensitivity, specificity, and accuracy were calculated using the following formulas:
where TP, FN, FP, and TN denote the numbers of true positives, false negatives, false positives, and true negatives, respectively. Statistical analyses were performed using JMP Pro 14 Software (Version 14. 2. 0, SAS Institute, Cary, NC) or GraphPad Prism 8 software. In all tests, a p value < 0.05 was considered statistically significant.
Results
Selection of biomarkers for predicting DILI potential by microarray analysis
For gene expression analysis, we considered that treatment with the tested drugs should not cause a significant reduction in PBMC viability. Freshly isolated PBMCs (a healthy male donor aged 63 years) alone were exposed to ten drugs, including amodiaquine (AMQ), chloroquine (CLQ), diclofenac (DCF), ethaclinic acid (ETA), ketoconazole (KTZ), ketorolac (KTL), levofloxacin (LVF), mebendazole (MBZ), tienilic acid (TNA), and trovafloxacin (TVF), at concentrations corresponding to 100, 50, 25, 10, 5, 2.5, or 1 × Cmax of each drug, and viability was determined after 24 h using the Cell Counting Kit-8, which measures intracellular dehydrogenase activities (Table S4). Since cell death-dependent decrease in intracellular RNA content hinders a successful transcriptome-wide expression analysis, we chose a maximum concentration for each drug up to 100 × Cmax at which viability was over 70% of the 0.4% DMSO vehicle-treated control. The selected concentrations are shown in bold in Table S4. Since TNA at 25 × Cmax was insoluble in media, 10 × Cmax was selected.
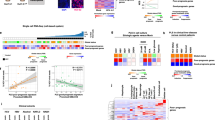
There is increasing evidence that idiosyncratic DILI is in part immune-mediated (Andrade et al. 2019; Fontana 2014) and that drugs can initiate immune reactions by activating innate immune reactions via the release of DAMPs from hepatocytes (Holt and Ju 2006). Therefore, an in vitro model that allows communication between immune cells and hepatic parenchymal cells may mimic the in vivo circumstance. In this study, we established an in vitro coculture system of PBMCs together with the liver-derived cell lines HepaRG and HepG2 using culture inserts (Fig. 1). For microarray analysis, freshly isolated PBMCs (a healthy male donor aged 63 years) were used. The coculture system (both insert and transwell sides) was treated with ten separate drugs mentioned earlier at the selected concentration, and transcriptome-wide expression analysis in PBMCs was performed. PCA of all detected genes showed an overall discrimination between DILI-positive and DILI-negative drugs, although the separation was not perfect (Fig. 2a). This separation was attributable to PC2, which accounts for 15.6% of the variance. To uncover the genes contributing to the PC, factor loading was calculated for each gene. Factor loadings range from − 1 to 1, and values close to − 1 and 1 indicate a strong association. Here, we selected genes with factor loadings below the 1st quartile (25th percentile) or more than the 4th quartile (75th percentile), some of which were further subjected to validation analysis between microarray and RT-qPCR data (not shown). The following 12 genes showing a significant correlation between microarray and RT-qPCR were selected as biomarkers (Fig. 2b): bone morphogenetic protein 6 (BMP6), C-X-C motif chemokine ligand 8 (CXCL8), epiregulin (EREG), interleukin 1 alpha (IL-1A), interleukin 1 beta (IL-1B), interleukin 24 (IL-24), MET proto-oncogene, receptor tyrosine kinase (MET), phosphotyrosine interaction domain containing 1 (PID1), prostaglandin-endoperoxide synthase 2 (PTGS2), solute carrier family 7 member 11 (SLC7A11, also known as xCT), secretory leukocyte peptidase inhibitor (SLPI), and TNF alpha-induced protein 6 (TNFAIP6). Hierarchical clustering of the 12 genes indicated that DILI-positive and DILI-negative drugs were mostly clustered separately regardless of whether the cocultured cells were HepG2 or HepaRG cells (Fig. 2b). Thus, HepG2 cells were used as a counterpart of PBMCs in the subsequent study.
Principal component analysis and hierarchical clustering of gene expression. Microarray analysis was performed using RNA from PBMCs that had been cocultured with HepG2 or HepaRG cells exposed to ten drugs as well as 0.4% DMSO vehicle-treated control (n = 1). a Principal component analysis of transcriptome-wide analysis. b Hierarchical clustering of selected genes. DILI-negative and DILI-positive drugs are shown in blue and red, respectively. AMQ amodiaquine, CLQ chloroquine, DCF diclofenac, ETA ethaclinic acid, KTL ketorolac, KTZ ketoconazole, LVF levofloxacin, MBZ mebendazole, TNA tienilic acid, TVF trovafloxacin
Evaluation of viabilities of PBMC and HepG2 cells as biomarkers for discriminating between DILI-negative and DILI-positive drugs
To set an optimal drug concentration used for coculture of PBMCs and HepG2 cells for the screening assay, commercial cryopreserved PBMCs (LP226; a healthy female donor, 55 years old) alone were exposed to 77 test drugs at concentrations corresponding to 100, 50, 25, 10, 5, 2.5, or 1 × Cmax of each drug, and viability was determined after 24 h using CellTiter-Glo®2.0 reagent, which measures the intracellular ATP content (Table S5). A maximum concentration for each drug up to 100 × Cmax at which viability was over 70% of the 0.4% DMSO-treated control was selected for coculture of PBMCs and HepG2 cells. The selected concentrations are shown in Table S5. In addition, the IC50 values expressed as the fold above Cmax of each drug were calculated (Table S5). When IC50 values could not be calculated due to low toxicity up to 100 × Cmax, they were set at 100 × Cmax. Next, we calculated the predictive performance of the IC50 value for DILI. For this analysis, “No-DILI” and “Less-DILI-concern” drugs annotated at LTKB-BD were considered to be DILI-negative drugs, while “Most-DILI-concern” drugs at LTKB-BD were considered DILI-positive drugs. “Most-DILI-concern” drugs (DILI-positive) exhibited significantly lower IC50 values in comparison to “No-DILI” and “Less-DILI-concern” drugs (DILI-negative drugs) (Fig. 3a, top and Table 1), suggesting that DILI-positive drugs were cytotoxic to some extent. Logistic regression analysis followed by ROC analysis yielded an AUC of 0.70 with 52%/86% (sensitivity/specificity) (Fig. 3a, bottom; Table 2), suggesting that cytotoxicity can flag approximately half of DILI-positive drugs.
Effects of drugs (n = 77) on PBMC and HepG2 cell viability and their discriminatory abilities for DILI. a PBMCs (lot: LP226) in monoculture were treated with various concentrations (100, 50, 25, 10, 5, 2.5, 1, 0.5, or 0.25 × Cmax) of DILI-positive (n = 27) and DILI-negative drugs (n = 50), and cell viabilities were determined based on cellular ATP levels (n = 3 or 4 biological replicates). Top. IC50 values of test drugs for the inhibition of PBMC viability. IC50 values are expressed as the fold change ( ×) of the human plasma maximum concentration (Cmax) and are shown as dots. The red horizontal line represents the mean of all drugs. Bottom. ROC curve for the discrimination of DILI-positive drugs from DILI-negative drugs using the IC50 values. b HepG2 cells in monoculture were treated with a single concentration of DILI-positive (n = 27) and DILI-negative drugs (n = 50), and cell viabilities were determined based on cellular ATP levels (n = 3 or 4 biological replicates). The tested concentration shown in Table 1 was a maximum concentration for each drug up to 100 × Cmax at which PBMC viability was over 70% of the 0.4% DMSO vehicle-treated control. Top. HepG2 cell viability expressed as the percentage of the DMSO vehicle-treated control is shown as dots. Bottom. ROC curve for the discrimination of DILI-positive drugs from DILI-negative drugs using the values of HepG2 cell viability. CI confidence interval
To evaluate the cytotoxicity of drugs to HepG2 cells, the selected single concentration for each drug was treated to HepG2 cells in a monoculture condition (Table 1). DILI-positive drugs significantly reduced viability (p = 0.0212) of HepG2 cells compared to those treated with DILI-negative drugs (Fig. 3b, top; Table 1). The predictive performance of HepG2 cell viability for DILI resulted in an AUC of 0.69 with 67%/68% (sensitivity/specificity) (Fig. 3b, bottom and Table 2), which was almost equal to that of PBMC viability (AUC = 0.70) (Fig. 3a, bottom).
Evaluation of gene expression levels as biomarkers for discriminating between DILI-negative and DILI-positive drugs
We next compared the coculture system of cryopreserved PBMCs with HepG2 cells to the PBMC monoculture system in terms of the inducibility of selected genes to evaluate the utility of the coculture system (Fig. 4). The PBMC (lot: LP226)/HepG2 coculture system and PBMC (lot: LP226) monoculture were treated with DILI-positive drugs, including diclofenac, etodolac, flucloxacillin, flutamide, nevirapine, nimesulide, troglitazone, trovafloxacin, and zafirlukast, at each concentration that yielded more than 70% cell viability (Table S5), and mRNA expression in PBMCs was determined. Treatment with most DILI-positive drugs highly induced BMP6, CXCL8, and MET mRNA expression levels in the PBMC/HepG2 coculture system compared to the PBMC monoculture (Fig. 4). These results indicate that the PBMC/HepG2 coculture system has a higher reactivity than the PBMC monoculture system, suggesting that marker gene expression in the PBMC/HepG2 coculture system could be applied to DILI assessment. Therefore, we next investigated the utility of the selected biomarker genes for predicting the DILI potential of 77 randomly selected marketed or withdrawn drugs with different DILI risks.
Effects of coculture with HepG2 cells on the induction of gene expression in in vitro culture of PBMCs exposed to DILI-positive drugs. Human PBMCs (lot: LP226) were cultured with (PBMC + HepG2 coculture) or without (PBMC monoculture) HepG2 cells using the culture insert and exposed to DILI-positive drugs (diclofenac, etodolac, flucloxacillin, flutamide, nevirapine, nimesulide, troglitazone, trovafloxacin, and zafirlukast) at the concentrations shown in Table 1. BMP6, CXCL8, and MET mRNA expression levels were determined by qPCR and normalized to that of GAPDH. Data are the mean ± SD (n = 3 biological replicates). *p < 0.05 and **p < 0.01 by two-tailed unpaired Student’s t test
Cryopreserved PBMCs (lot: LP226) were cocultured with HepG2 cells exposed to the test drugs at the selected drug concentration that yields more than 70% cell viability, and mRNA expression was determined by qPCR. The mRNA expression levels of BMP6, MET, and PID1 were significantly higher in the cocultured PBMCs treated with DILI-positive drugs than in those treated with DILI-negative drugs (Fig. 5a). DILI-positive drugs increased the expression levels of CXCL8, EREG, IL-24, and SLPI mRNA, although they were not statistically significant. Then, logistic regression analyses were conducted for each gene to evaluate how accurately these genes discriminate DILI-positive drugs versus DILI-negative drugs (Table S6). The ROC analysis revealed that the AUC values were the highest for MET (AUC: 0.81), followed by BMP6 (0.76), IL-24 (0.73), CXCL8 (0.72), and EREG (0.67), with significance (Fig. 5b). The AUC values of the four genes (MET, BMP6, IL-24, and CXCL8) were higher than that of the IC50 of PBMC viability (Fig. 3b). MET and BMP6 showed sensitivity/specificity of 85%/66% and 78%/70%, respectively (Table 2). Thus, marker gene expression levels could be of utility for the prediction of DILI potential.
Effects of drugs (n = 77) on the mRNA expression of biomarkers and their discriminatory ability for DILI in PBMC and HepG2 cell coculture. Human PBMCs (lot: 226) were cultured with HepG2 cells and exposed to DILI-positive (n = 27) and DILI-negative drugs (n = 50) at the concentrations shown in Table 1. a The mRNA expression levels in the PBMCs were normalized to that of GAPDH and are shown as relative to 0.4% DMSO vehicle-treated control. Each dot represents the mean of each drug (n = 2 biological replicates), and the red horizontal line represents the mean of all drugs. P values determined by two-tailed unpaired Student’s t test are shown. b ROC curves for the discrimination of DILI-positive drugs from DILI-negative drugs using the mRNA expression levels. CI confidence interval. c MET gene expression levels in PBMCs cocultured with HepG2 cells exposed to drugs. Pharmacological categories are shown above the columns. Each column represents the mean of each drug (n = 2). Blue: DILI-negative drug; red: DILI-positive drug; FN false negative, FP false positive (color figure online)
We next investigated whether the marker gene levels reflect the DILI potential of drugs within the same pharmacological class. For this purpose, the MET expression levels were compared among drugs within the same pharmacological categories, because its AUC was the highest (Fig. 5c). Using the cut-off value which maximizes Youden’s index, eight NSAIDs were successfully predicted as DILI-positive or DILI-negative by MET expression, as were antibacterials except the DILI-positive drug levofloxacin, which was falsely predicted as DILI-negative. However, the antidiabetic thiazolidinedione pioglitazone and dopaminergic entacapone were cases of false-positive prediction. Thus, the single parameter analysis of marker gene expression levels in the PBMC/HepG2 coculture system showed superiority to PBMC viability.
The combination of biomarker genes better predicts DILI potential
To investigate whether the combination of marker gene expression levels affects prediction, multiple logistic analysis followed by ROC analysis was performed. By the stepwise procedure, we found that a combination of BMP6, EREG, IL-1A, MET, PID1, PTGS2, SLC7A11, SLPI, and TNFAIP6 (Table S7) provided the highest AUC value of 0.94 (Fig. 6a, top). The estimated probability of being a DILI-positive drug was calculated and shown with the cut-off value, which maximizes Youden’s index (Fig. 6a, bottom). As a result, the sensitivity and specificity were calculated as 93% and 86%, respectively, with an accuracy of 88% (Fig. 6a, bottom; Table 3). Two out of 27 DILI-positive drugs were mispredicted as false negatives, including didanosine and tolcapone, while 7 out of 50 DILI-negative drugs were mispredicted as false positives, including atropine, chloroquine, deferoxamine, isoproterenol, mebendazole, pamabrom, and primaquine (Fig. 6a, bottom).
Development of a multiple logistic regression model of gene expression levels for detecting DILI-positive drugs. Top. ROC curves for the discrimination of DILI-positive drugs (n = 27) from DILI-negative drugs (n = 50) using a multiple logistic regression model of mRNA expression levels of nine genes (BMP6, EREG, IL-1A, MET, PID1, PTGS2, SLC7A11, SLPI, and TNFAIP6) in a PBMC (lot: LP226) + HepG2 coculture (n = 2 biological replicates), b PBMC (lot: LP316) + HepG2 coculture (n = 1), and c PBMC (lot: LP226) monoculture (n = 1). Bottom. The probability of being DILI-positive for each test drug (n = 77) was calculated from the logistic model for each culture. The red horizontal line represents the cut-off probability. CI confidence interval
Evaluation of donor-to-donor differences and comparison of PBMC monoculture with coculture
To investigate donor-to-donor differences in the predictive performance of gene expression levels, another donor-derived PBMC (lot: LP316) were cocultured with HepG2 and the 77 drugs were treated. mRNA expression levels of the nine genes used for the multiple logistic regression analysis were measured in PBMCs (Supplementary Fig. S1). In addition to BMP6, MET, and PID1, the mRNA expression levels of EREG were highly induced by DILI-positive drugs in this lot (Supplementary Fig. S1A and Fig. 5a). AUC values of BMP6, MET, PID1, and SLPI mRNA levels were significant in the coculture, among which BMP6 and MET were common to both PBMC donors (Supplementary Fig. S1B and Fig. 5b). Furthermore, multiple logistic regression analysis of the nine genes resulted in an AUC of 0.91 (Fig. 6b) and sensitivity/specificity of 74%/100% (Table 3) in PBMC (LP316) + HepG2 coculture, which were almost equal to those of PBMC (LP226) + HepG2 coculture (AUC = 0.94, Fig. 6a; sensitivity/specificity 93%/86%, Table 3). Therefore, although there were some donor-to-donor differences in the predictive performance of each gene, the integrated analysis suggests that predictive performance was almost not affected by the donor difference.
Finally, we compared the predictive performance of mRNA expression levels of the nine genes between PBMC monoculture and PBMC/HepG2 coculture. For this purpose, PBMC (lot: LP226) was cultured on culture inserts and 77 drugs were treated in the absence of liver cells. The mRNA expression levels in PBMC monoculture were not different between DILI-positive drugs and DILI-negative drugs except SLC7A11 (Supplementary Fig. S2A), and AUCs of each gene ranges from 0.53 to 0.77 (Supplementary Fig. S2B). These values were lower than those of PBMC + HepG2 coculture (AUC range: 0.51 to 0.81) (Fig. 5b). Furthermore, multiple logistic regression analysis of the nine genes resulted in an AUC of 0.83 (Fig. 6c) and sensitivity/specificity of 74%/86% in PBMC monoculture (Table 3), which were lower than those of PBMC + HepG2 coculture (AUC = 0.94; sensitivity/specificity 93%/86%, Fig. 6a). These results demonstrate that PBMC + HepG2 coculture has a better predictive performance for DILI than PBMC monoculture, supporting the utility of PBMC + HepG2 coculture.
Discussion
Most cell-based assays for the assessment of DILI reported so far utilize cytotoxicity in hepatic parenchymal cells as readouts (Gómez-Lechón et al. 2016). However, their predictabilities for DILI are not so high. It is important to note that clinical features of immune-mediated or hypersensitivity drug reactions can be observed in one-quarter of patients (Andrade et al. 2019). Thus, biological responses in immune cells could be applied to the risk assessment of DILI. However, there is a paucity of such studies that attempt to evaluate clinical DILI risks comprehensively by immune cell responses in vitro. In this study, we explored and evaluated marker genes in human PBMCs cocultured with a liver-derived HepG2 cell line by transcriptome-wide analysis and evaluated their predictability for DILI.
We previously reported a cell-based assay in which the human promyelocytic leukemia cell line HL-60 was incubated with conditioned media of HepG2 or HepaRG cells that had been treated with test drugs. In that study, immune- and inflammatory-related genes in HL-60 cells, including MCP-1, S100A9, IL-1B, CXCL8, and TNFA, were tested as biomarkers (Oda et al. 2016), because these genes were upregulated in the liver of mice treated with DILI-positive drugs compared to mice treated with DILI-negative drugs (Yano et al. 2014). Screening of 79 pharmaceuticals revealed that CXCL8 in HL-60 cells treated with conditioned media of HepaRG cells that had been treated with test drugs provided acceptable discriminatory performance for DILI with an AUC of 0.75. However, the previous study (Oda et al. 2016) subjectively selected biomarker genes from in vivo mouse studies, which motivated us to identify more suitable genes for DILI assessment in this study focusing on human cell-based assays that included an immune component (PBMCs). Therefore, this study implemented transcriptome-wide gene expression analysis to objectively select candidate biomarker genes. In addition to CXCL8 and IL-1B, which are the only common biomarkers between this and previous studies (Oda et al. 2016), we found several promising biomarkers.
We found that each of the mRNA expression levels of MET, BMP6, and IL-24 in PBMCs provided superior predictability (AUC 0.73–0.81) of DILI compared to CXCL8 (IL-8) and IL-1B (Fig. 5b). In particular, the MET level is promising, because it showed the highest AUC among all single parameters evaluated with a sensitivity of 85% and a specificity of 66% (Table 2). Furthermore, MET expression levels could differentiate some DILI-positive drugs from DILI-negative drugs within the same pharmaceutical categories (Fig. 5c). Thus, evaluation of DILI potential by MET levels in PBMCs may aid in selecting compounds at the lead optimization or early stage of drug development. Most in vitro assays for evaluating DILI potential established so far utilize cell viability, superoxide production, and mitochondrial membrane integrity in hepatic parenchymal cells as readouts (O'Brien et al. 2006; Wink et al. 2014; Xu et al. 2008). These conventional markers have no satisfactory predictive power (cell viability: 10% sensitivity and 92% specificity; glutathione depletion: 19% sensitivity and 85% specificity) in HepG2 cells when assayed at a single concentration (O’Brien et al. 2006). Recent studies, however, utilized long-term spheroid cultures of primary human hepatocytes for the evaluation of hepatotoxic liabilities of drugs (Proctor et al. 2017; Vorrink et al. 2018). They reported improved sensitivities/specificities of 59%/80% in 110 drugs (Proctor et al. 2017) and 69%/100% in 123 drugs at 100 × Cmax concentration (Vorrink et al. 2018), suggesting that hepatocytes retaining physiological cellular phenotypes can be translated into an improved prediction. However, it should be noted that the overall predictive performance of these studies was determined by a single donor of hepatocytes. More recently, with a support vector machine-based classifier using human whole blood Cmax and EC10 of cytotoxicity to sandwich cultured primary human hepatocytes (corresponding to the drug concentration where the sigmoidal curve reaches the value of 90%), sensitivity and specificity were 88% and 100%, respectively, although the number of tested drugs was only 28 (Albrecht et al. 2019). Although the number of tested drugs and exposure concentrations as well as target cells (monoculture of hepatocytes versus coculture of PBMCs and HepG2 cells) are different between the two studies and our study, the predictability of DILI by MET expression levels is comparable to those of the previous studies. Taken together, single gene expression, such as MET, in PBMCs can be an alternative in vitro tool for detecting DILI potential.
Recently, the DILI potential of drugs has been evaluated by a hybrid evaluation system that includes several biological parameters (cytotoxicity, mitochondrial dysfunction, and bile salt export pump inhibition) and physicochemical properties (dose, lipophilicity, ionization state, and carbon bond saturation) (Aleo et al. 2020). These approaches take into consideration the nature of idiosyncratic DILI, which develops by multifactorial mechanisms (Kaplowitz 2005; Stephens et al. 2014). In this study, using the nine identified biomarker genes, a multiple logistic model of BMP6, EREG, IL-1A, MET, PID1, PTGS2, SLC7A11, SLPI, and TNFAIP6 expression levels was modeled (Table S7). This model provided the best prediction in distinguishing DILI-positive drugs from DILI-negative drugs, with the highest AUC value of 0.94 (Fig. 6a) with a sensitivity of 93% and a specificity of 86% (Table 3), which was higher than the 85% sensitivity of MET. This trend was also observed in the hierarchical clustering analysis, where the consideration of several genes rather than a single gene appears to discriminate between DILI-positive and DILI-negative drugs (Fig. 2b). Thus, the use of a combination of several biomarker genes is expected to improve the predictive performance of DILI potential. Two drugs, didanosine and tolcapone, were consistently predicted to be false negatives in PBMC + HepG2 cell coculture irrespective of PBMC donor (Fig. 6a, b). Tolcapone is reported to form reactive metabolites that are trapped by glutathione, whereas the structural analog entacapone, a non-DILI drug, is not (Dragovic et al. 2016; Smith et al. 2003; Stepan et al. 2011). Thus, it should be emphasized that DILI potential should be evaluated by considering multiple factors, including the formation of reactive metabolites, cytotoxicity, drug plasma/tissue concentration, and immune-related responses.
Of particular interest in this study was that the extent of induction of BMP6, CXCL8, and MET mRNA by DILI-positive drugs (diclofenac, etodolac, flutamide, nevirapine, troglitazone, and zafirlukast) was largely higher in the coculture of PBMCs with HepG2 cells than in the PBMC monoculture (Fig. 4). This suggests that humoral factors secreted or released from drug-exposed HepG2 cells may affect the inducibility of these genes in PBMCs. It is known that stressed hepatocytes have the potential to release DAMPs such as high mobility group box-1 (HMGB1) and heat shock proteins (HSPs) to activate the immune system through Toll-like receptors (TLRs) and the receptor for advanced glycation end products (RAGEs) (Holt and Ju 2006; Kono and Rock 2008). This study did not identify which DAMPs are released from HepG2 cells and which immune cells receive the stimuli and are activated. Other working groups also suggested that hepatocytes treated with DILI drugs (such as amodiaquine, diclofenac, ketoconazole, nevirapine, tolcapone, and troglitazone) release DAMPs to activate immune cells (Granitzny et al. 2017; Kato and Uetrecht 2017; Mak et al. 2018). This immune cell activation in vitro by DILI drugs would be linked with DILI, because some drugs discussed here are likely to cause liver injury via an immune-mediated mechanism as follows: flucloxacillin-induced liver injury is associated with the human leukocyte antigen (HLA)*B5701 allele (Daly et al. 2009; Wuillemin et al. 2014); nevirapine-induced skin rash is associated with the HLA-B3505 allele (Chantarangsu et al. 2009); flutamide-induced liver injury is associated with Th2 cytokines in wild-type mice (Higuchi et al. 2012); diclofenac (Deng et al. 2006) and trovafloxacin (Shaw et al. 2009) cause liver injury in rats or mice pretreated with lipopolysaccharide; and troglitazone and tolcapone cause liver injury only in PD1 knockout mice under anti-CTLA4 antibody treatment (Mak et al. 2018).
Although we found several genes that can predict a drug’s potential to cause DILI, their association with the mechanism of DILI remains unknown. One such gene, MET, also known as hepatocyte growth factor receptor (HGFR), is a proto-oncogenic receptor tyrosine kinase that recognizes the endogenous ligand hepatocyte growth factor/scatter factor (HGF) (Trusolino et al. 2010). MET is expressed in monocytes, B cells, and natural killer (NK) cells but not in peripheral blood CD8+ or CD4+ T cells (Molnarfi et al. 2012), although the function of HGF-MET signaling in these cells remains unclear. Another study, however, demonstrated that MET is expressed in memory T cells in lymph nodes and that HGF–MET signaling facilitates T-cell recruitment to heart tissue to contribute to inflammation (Komarowska et al. 2015). BMP6 is a member of the TGF-β superfamily of BMPs and exhibits a broad spectrum of biological functions, including cell growth and differentiation (Miyazono et al. 2010). BMPs, including BMP6, are known to produce and activate naïve CD4+ T cells in response to T-cell receptor stimulation (Martinez et al. 2015). Although the pathophysiological roles of MET and BMP6 in DILI are unknown, the significant upregulation of MET and BMP6 mRNA in PBMCs by DILI-positive drugs may reflect the activation of monocytes, B cells, NK cells, or T cells. Because of limited information on the association between these genes and DILI, further study is required.
Idiosyncratic DILI is thought to be associated with multifactorial mechanisms, including patient-associated factors (i.e., sex, genetic variation, and infectious status) as well as the compound potential to cause DILI. In fact, certain alleles of HLA are associated with the increased incidence of DILI caused by specific drugs (Usui and Naisbitt 2017). However, such an association can be identified only after the drug reaches the market under the current circumstance. Thus, predictive models for DILI are urgently required. With respect to in vitro models utilizing immune cells for the risk assessment of DILI, Kegel et al. established an in vitro culture of human Kupffer cells treated with conditioned media of hepatocytes that were exposed to diclofenac and acetaminophen (Kegel et al. 2015). They found a compound- and donor-dependent increase in proinflammatory cytokines produced by Kupffer cells. However, this system seems to be unsuitable for screening large numbers of compounds due to the limited availability of primary human liver cells. In addition, Kupffer cells alone may not be sufficient to capture potential immune activation induced by drugs. In this regard, our coculture model using PBMCs and HepG2 cells may be useful in that cells are readily available and PBMCs are comprised of several immune cell types, including subsets of lymphocytes (T cells, B cells, and NK cells), monocytes, and dendritic cells.
There are several limitations in this study. First, most of the DILI-positive drugs selected for our microarray experiments have been demonstrated to have immunoallergic features, including amodiaquine, diclofenac, levofloxacin, (LiverTox: https://www.ncbi.nlm.nih.gov/books/NBK547852/), tienilic acid (Lecoeur et al. 1996), and trovafloxacin (Ball et al. 1999). This would constitute a bias with respect to biomarker selection as many other DILI-positive drugs not having an immune component in their mechanisms may be excluded. Second, although we considered Less-DILI-concern drugs as DILI-negative in this study, other database reported some DILI concern for them. For example, acetylsalicylic acid and trazodone, which are classified as Less-DILI concern in LTKB-BD, are reported to cause clinically apparent liver injury in the LiverTox database. In addition, although rofecoxib is not assigned in LTKB-BD and, hence, is considered as DILI-negative in this study, it can cause clinically apparent, symptomatic drug-induced liver injury with jaundice in rare instance according to the LiverTox database. Third, although we used 77 drugs, the validity of this assay needs to be confirmed using another set of drugs. Finally, we used only two donors for investigating donor-to-donor differences in the predictability of DILI, because we focused on the identification and evaluation of biomarker genes with a panel of drugs. PBMC composition, and preexisting inflammatory status are different between donors (Ter Horst et al. 2016); therefore, further study is warranted to elucidate the effects of individual differences in PBMCs on the predictability of DILI. However, we believe that information regarding the evaluation of DILI potential by gene expression provided from this study is important, even though the data were obtained using cells from one donor.
In conclusion, we developed an in vitro coculture model using PBMCs with HepG2 cells to discriminate DILI-positive drugs from DILI-negative drugs based on gene expression signatures. The MET mRNA level provided good sensitivity and specificity for predicting DILI, but a combination of biomarker genes showed superior predictivity. However, because the number of DILI-positive drugs in the dataset was limited to 27, additional studies using a large number of compounds are needed to confirm the usefulness of these coculture systems. Nevertheless, to the best of our knowledge, this study is the first to identify biomarkers from transcriptome-wide analysis of PBMCs cocultured with liver cells for predicting the DILI potential of drugs. The established coculture model could be useful in detecting the DILI potential of compounds, in particular those that are immune-mediated.
References
Albrecht W, Kappenberg F, Brecklinghaus T et al (2019) Prediction of human drug-induced liver injury (DILI) in relation to oral doses and blood concentrations. Arch Toxicol 93(6):1609–1637. https://doi.org/10.1007/s00204-019-02492-9
Aleo MD, Shah F, Allen S et al (2020) Moving beyond binary predictions of human drug-induced liver injury (DILI) toward contrasting relative risk potential. Chem Res Toxicol 33(1):223–238. https://doi.org/10.1021/acs.chemrestox.9b00262
Andrade RJ, Chalasani N, Bjornsson ES et al (2019) Drug-induced liver injury. Nat Rev Dis Primers 5(1):58. https://doi.org/10.1038/s41572-019-0105-0
Ball P, Mandell L, Niki Y, Tillotson G (1999) Comparative tolerability of the newer fluoroquinolone antibacterials. Drug Saf 21(5):407–421. https://doi.org/10.2165/00002018-199921050-00005
Bell LN, Chalasani N (2009) Epidemiology of idiosyncratic drug-induced liver injury. Semin Liver Dis 29(4):337–347. https://doi.org/10.1055/s-0029-1240002
Björnsson ES (2014) Epidemiology and risk factors for idiosyncratic drug-induced liver injury. Semin Liver Dis 34(2):115–122. https://doi.org/10.1055/s-0034-1375953
CDER (2006) Warnings and precautions, contraindications, and boxed warning sections of labeling for human prescription drug and biological products—content and format. https://www.fda.gov/downloads/Drugs/GuidanceComplianceRegulatoryInformation/Guidances/ucm075096.pdf. Accessed 31 Mar 2020
Chantarangsu S, Mushiroda T, Mahasirimongkol S et al (2009) HLA-B*3505 allele is a strong predictor for nevirapine-induced skin adverse drug reactions in HIV-infected Thai patients. Pharmacogenet Genomics 19(2):139–146. https://doi.org/10.1097/FPC.0b013e32831d0faf
Chen M, Vijay V, Shi Q, Liu Z, Fang H, Tong W (2011) FDA-approved drug labeling for the study of drug-induced liver injury. Drug Discov Today 16(15–16):697–703. https://doi.org/10.1016/j.drudis.2011.05.007
Chen M, Suzuki A, Thakkar S, Yu K, Hu C, Tong W (2016) DILIrank: the largest reference drug list ranked by the risk for developing drug-induced liver injury in humans. Drug Discov Today 21(4):648–653. https://doi.org/10.1016/j.drudis.2016.02.015
Daly AK, Donaldson PT, Bhatnagar P et al (2009) HLA-B*5701 genotype is a major determinant of drug-induced liver injury due to flucloxacillin. Nat Genet 41(7):816–819. https://doi.org/10.1038/ng.379
de Abajo FJ, Montero D, Madurga M, Garcia Rodriguez LA (2004) Acute and clinically relevant drug-induced liver injury: a population based case-control study. Br J Clin Pharmacol 58(1):71–80. https://doi.org/10.1111/j.1365-2125.2004.02133.x
Deng X, Stachlewitz RF, Liguori MJ et al (2006) Modest inflammation enhances diclofenac hepatotoxicity in rats: role of neutrophils and bacterial translocation. J Pharmacol Exp Ther 319(3):1191–1199. https://doi.org/10.1124/jpet.106.110247
Dragovic S, Vermeulen NP, Gerets HH et al (2016) Evidence-based selection of training compounds for use in the mechanism-based integrated prediction of drug-induced liver injury in man. Arch Toxicol 90(12):2979–3003. https://doi.org/10.1007/s00204-016-1845-1
Edling Y, Sivertsson LK, Butura A, Ingelman-Sundberg M, Ek M (2009) Increased sensitivity for troglitazone-induced cytotoxicity using a human in vitro co-culture model. Toxicol Vitro 23(7):1387–1395. https://doi.org/10.1016/j.tiv.2009.07.026
Fontana RJ (2014) Pathogenesis of idiosyncratic drug-induced liver injury and clinical perspectives. Gastroenterology 146(4):914–928. https://doi.org/10.1053/j.gastro.2013.12.032
Funk C, Roth A (2017) Current limitations and future opportunities for prediction of DILI from in vitro. Arch Toxicol 91(1):131–142. https://doi.org/10.1007/s00204-016-1874-9
Gómez-Lechón MJ, Tolosa L, Donato MT (2016) Metabolic activation and drug-induced liver injury: in vitro approaches for the safety risk assessment of new drugs. J Appl Toxicol 36(6):752–768. https://doi.org/10.1002/jat.3277
Granitzny A, Knebel J, Muller M et al (2017) Evaluation of a human in vitro hepatocyte-NPC co-culture model for the prediction of idiosyncratic drug-induced liver injury: a pilot study. Toxicol Rep 4:89–103. https://doi.org/10.1016/j.toxrep.2017.02.001
Higuchi S, Kobayashi M, Yano A et al (2012) Involvement of Th2 cytokines in the mouse model of flutamide-induced acute liver injury. J Appl Toxicol 32(10):815–822. https://doi.org/10.1002/jat.1706
Holt MP, Ju C (2006) Mechanisms of drug-induced liver injury. AAPS J 8(1):E48–54. https://doi.org/10.1208/aapsj080106
Kaplowitz N (2005) Idiosyncratic drug hepatotoxicity. Nat Rev Drug Discov 4(6):489–499. https://doi.org/10.1038/nrd1750
Kato R, Uetrecht J (2017) Supernatant from hepatocyte cultures with drugs that cause idiosyncratic liver injury activates macrophage inflammasomes. Chem Res Toxicol 30(6):1327–1332. https://doi.org/10.1021/acs.chemrestox.7b00065
Kegel V, Pfeiffer E, Burkhardt B et al (2015) Subtoxic concentrations of hepatotoxic drugs lead to Kupffer cell activation in a human in vitro liver model: an approach to study DILI. Mediators Inflamm 2015:640631. https://doi.org/10.1155/2015/640631
Kim DE, Jang MJ, Kim YR et al (2017) Prediction of drug-induced immune-mediated hepatotoxicity using hepatocyte-like cells derived from human embryonic stem cells. Toxicology 387:1–9. https://doi.org/10.1016/j.tox.2017.06.005
Kleiner DE (2017) Drug-induced liver injury: the hepatic pathologist’s approach. Gastroenterol Clin North Am 46(2):273–296. https://doi.org/10.1016/j.gtc.2017.01.004
Komarowska I, Coe D, Wang G et al (2015) Hepatocyte growth factor receptor c-Met instructs T cell cardiotropism and promotes T cell migration to the heart via autocrine chemokine release. Immunity 42(6):1087–1099. https://doi.org/10.1016/j.immuni.2015.05.014
Kono H, Rock KL (2008) How dying cells alert the immune system to danger. Nat Rev Immunol 8(4):279–289. https://doi.org/10.1038/nri2215
Lecoeur S, Andre C, Beaune PH (1996) Tienilic acid-induced autoimmune hepatitis: anti-liver and-kidney microsomal type 2 autoantibodies recognize a three-site conformational epitope on cytochrome P4502C9. Mol Pharmacol 50(2):326–333
Liguori MJ, Ditewig AC, Maddox JF et al (2010) Comparison of TNFα to lipopolysaccharide as an inflammagen to characterize the idiosyncratic hepatotoxicity potential of drugs: Trovafloxacin as an example. Int J Mol Sci 11(11):4697–4714. https://doi.org/10.3390/ijms11114697
Mak A, Kato R, Weston K, Hayes A, Uetrecht J (2018) An impaired immune tolerance animal model distinguishes the potential of troglitazone/pioglitazone and tolcapone/entacapone to cause IDILI. Toxicol Sci 161(2):412–420. https://doi.org/10.1093/toxsci/kfx219
Martinez VG, Sacedon R, Hidalgo L et al (2015) The BMP pathway participates in human naive CD4+ T cell activation and homeostasis. PLoS ONE 10(6):e0131453. https://doi.org/10.1371/journal.pone.0131453
Miyazono K, Kamiya Y, Morikawa M (2010) Bone morphogenetic protein receptors and signal transduction. J Biochem 147(1):35–51. https://doi.org/10.1093/jb/mvp148
Molnarfi N, Benkhoucha M, Bjarnadottir K, Juillard C, Lalive PH (2012) Interferon-β induces hepatocyte growth factor in monocytes of multiple sclerosis patients. PLoS ONE 7(11):e49882. https://doi.org/10.1371/journal.pone.0049882
O'Brien PJ, Irwin W, Diaz D et al (2006) High concordance of drug-induced human hepatotoxicity with in vitro cytotoxicity measured in a novel cell-based model using high content screening. Arch Toxicol 80(9):580–604. https://doi.org/10.1007/s00204-006-0091-3
Oda S, Matsuo K, Nakajima A, Yokoi T (2016) A novel cell-based assay for the evaluation of immune- and inflammatory-related gene expression as biomarkers for the risk assessment of drug-induced liver injury. Toxicol Lett 241:60–70. https://doi.org/10.1016/j.toxlet.2015.10.029
Onakpoya IJ, Heneghan CJ, Aronson JK (2016) Post-marketing withdrawal of 462 medicinal products because of adverse drug reactions: a systematic review of the world literature. BMC Med 14:10. https://doi.org/10.1186/s12916-016-0553-2
Ostapowicz G, Fontana RJ, Schiodt FV et al (2002) Results of a prospective study of acute liver failure at 17 tertiary care centers in the United States. Ann Intern Med 137(12):947–954. https://doi.org/10.7326/0003-4819-137-12-200212170-00007
Proctor WR, Foster AJ, Vogt J et al (2017) Utility of spherical human liver microtissues for prediction of clinical drug-induced liver injury. Arch Toxicol 91(8):2849–2863. https://doi.org/10.1007/s00204-017-2002-1
R Core Team (2019) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/
Senior JR (2014) Evolution of the Food and Drug Administration approach to liver safety assessment for new drugs: current status and challenges. Drug Saf 37(Suppl 1):S9–17. https://doi.org/10.1007/s40264-014-0182-7
Shaw PJ, Ganey PE, Roth RA (2009) Trovafloxacin enhances the inflammatory response to a Gram-negative or a Gram-positive bacterial stimulus, resulting in neutrophil-dependent liver injury in mice. J Pharmacol Exp Ther 330(1):72–78. https://doi.org/10.1124/jpet.109.151068
Smith KS, Smith PL, Heady TN, Trugman JM, Harman WD, Macdonald TL (2003) In vitro metabolism of tolcapone to reactive intermediates: relevance to tolcapone liver toxicity. Chem Res Toxicol 16(2):123–128. https://doi.org/10.1021/tx025569n
Song B, Aoki S, Liu C, Ito K (2019) A toll-like receptor 9 agonist sensitizes mice to mitochondrial dysfunction-induced hepatic apoptosis via the Fas/FasL pathway. Arch Toxicol 93(6):1573–1584. https://doi.org/10.1007/s00204-019-02454-1
Stepan AF, Walker DP, Bauman J et al (2011) Structural alert/reactive metabolite concept as applied in medicinal chemistry to mitigate the risk of idiosyncratic drug toxicity: a perspective based on the critical examination of trends in the top 200 drugs marketed in the United States. Chem Res Toxicol 24(9):1345–1410. https://doi.org/10.1021/tx200168d
Stephens C, Andrade RJ, Lucena MI (2014) Mechanisms of drug-induced liver injury. Curr Opin Allergy Clin Immunol 14(4):286–292. https://doi.org/10.1097/ACI.0000000000000070
Ter Horst R, Jaeger M, Smeekens SP et al (2016) Host and environmental factors influencing individual human cytokine responses. Cell 167(4):1111–1124. https://doi.org/10.1016/j.cell.2016.10.018
Trusolino L, Bertotti A, Comoglio PM (2010) MET signalling: principles and functions in development, organ regeneration and cancer. Nat Rev Mol Cell Biol 11(12):834–848. https://doi.org/10.1038/nrm3012
Uetrecht J (2019) Mechanistic studies of idiosyncratic DILI: clinical implications. Front Pharmacol 10:837. https://doi.org/10.3389/fphar.2019.00837
Usui T, Naisbitt DJ (2017) Human leukocyte antigen and idiosyncratic adverse drug reactions. Drug Metab Pharmacokinet 32(1):21–30. https://doi.org/10.1016/j.dmpk.2016.11.003
Vorrink SU, Zhou Y, Ingelman-Sundberg M, Lauschke VM (2018) Prediction of drug-induced hepatotoxicity using long-term stable primary hepatic 3D spheroid cultures in chemically defined conditions. Toxicol Sci 163(2):655–665. https://doi.org/10.1093/toxsci/kfy058
Weaver RJ, Blomme EA, Chadwick AE et al (2020) Managing the challenge of drug-induced liver injury: a roadmap for the development and deployment of preclinical predictive models. Nat Rev Drug Discov 19(2):131–148. https://doi.org/10.1038/s41573-019-0048-x
Wink S, Hiemstra S, Huppelschoten S et al (2014) Quantitative high content imaging of cellular adaptive stress response pathways in toxicity for chemical safety assessment. Chem Res Toxicol 27(3):338–355. https://doi.org/10.1021/tx4004038
Wuillemin N, Terracciano L, Beltraminelli H et al (2014) T cells infiltrate the liver and kill hepatocytes in HLA-B*57:01-associated floxacillin-induced liver injury. Am J Pathol 184(6):1677–1682. https://doi.org/10.1016/j.ajpath.2014.02.018
Xu JJ, Henstock PV, Dunn MC, Smith AR, Chabot JR, de Graaf D (2008) Cellular imaging predictions of clinical drug-induced liver injury. Toxicol Sci 105(1):97–105. https://doi.org/10.1093/toxsci/kfn109
Yano A, Oda S, Fukami T, Nakajima M, Yokoi T (2014) Development of a cell-based assay system considering drug metabolism and immune- and inflammatory-related factors for the risk assessment of drug-induced liver injury. Toxicol Lett 228(1):13–24. https://doi.org/10.1016/j.toxlet.2014.04.005
Acknowledgements
The authors wish to thank Dr. Tomoya Shimokata and Yuka Murasaki from Department of Clinical Oncology and Chemotherapy, and Dr. Kosuke Yoshida from Department of Drug Safety Sciences/Department of Obstetrics and Gynecology Nagoya University Graduate School of Medicine for taking blood.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to disclose.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the 1964 Helsinki Declaration and approved by the ethics committee of Nagoya University School of Medicine (No. 2016-0094).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Oda, S., Uchida, Y., Aleo, M.D. et al. An in vitro coculture system of human peripheral blood mononuclear cells with hepatocellular carcinoma-derived cells for predicting drug-induced liver injury. Arch Toxicol 95, 149–168 (2021). https://doi.org/10.1007/s00204-020-02882-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-020-02882-4